Abstract
Transgressive segregation is a common phenomenon in plant species. In this study, transgressive segregation for kilo-grain weight (KGW) was observed in a recombinant inbred line (RIL) population derived from the cross between an indica variety, Teqing, and a wide compatible japonica variety, 02428, in three environments. A genetic linkage map with 154 single sequence repeat markers (SSR) was developed. Effects on KGW of quantitative trait loci (QTLs), digenetic epistasis, and their environmental interaction (QE) were determined using a mixed linear model approach. 13 QTLs with additive effects and 8 digenetic interactions involving 16 loci were identified. Eight QTLs were involved in interactions. Two QTLs and one epistasis showed QE. 30.0 and 14.0% of variation was explained by the additive effects and epistasis, respectively, which were much greater than the 4.4% of variation explained by QE. According to the model used, predicted KGW values of extreme phenotypes in the RIL population were very close to their observed values. This indicated that complementary action of additive QTLs and epistasis can adequately account for the important genetic bases of transgressive segregation for KGW in the rice RIL population.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Many important agronomic traits, such as yield and its components, are complex traits, which are controlled by multiple quantitative trait loci (QTLs) and modified by the internal and external environment (Xing et al. 2002; Li et al. 2003; Liu et al. 2008; Wan et al. 2008). Their phenotypes of a hybrid segregating population often present an approximately normal distribution. However, the phenotypes of individuals in either or both sides of the distribution usually are extreme or novel compared to those of either parental line. The generation of these extreme or novel phenotypes in a population is referred to as transgressive segregation (Grant 1975; Devicente and Tanksley 1993).
Transgressive segregation has been the rule rather than the exception in studies of quantitative traits because in most studies of quantitative genetics at least one transgressive trait and a considerable portion of traits examined were transgressive (Devicente and Tanksley 1993; Rieseberg and Ellstrand 1993; Rieseberg et al. 1999). A part of the transgressive progenies generated by hybridization is fixed by nature selection, and have different ecologic niches from their parents. Then they survive in different environments, ultimately leading to speciation. Thus, transgressive segregation, combined with nature selection, is one of the important mechanisms for evolutionary transition (Rieseberg et al. 2003; Albertson and Kocher 2005; Bell and Travis 2005).
To explain the phenomena of transgressive segregation, several hypotheses have been proposed, including an increased mutation rate, the exposure of recessive alleles (Rick and Smith 1953), overdominance (Devicente and Tanksley 1993), breakdown of linkage (Hagiwara et al. 2006; Winn et al. 2009), complementary gene action (Devicente and Tanksley 1993; Xu et al. 1998), and epistasis (Brem and Kruglyak 2005). The rate of mutation would be increased in a hybrid population, which may be caused by activation of transposable elements such as P elements. However, the frequency of mutation was unable to reach the high frequency of transgression (Engels 1983; Barton and Hewitt 1985). In addition, the same mutation did not occur in one more repeat experiment. Overdominance exists only in the F2 and back-crossed (BC) populations and is not inherited, whereas transgressive segregation also exists in doubled haploid (DH) lines, recombinant inbred lines (RILs) in addition to F2 and BC populations, and can be inherited. Thus, overdominance cannot explain transgressive segregation in RIL and DH populations (Devicente and Tanksley 1993; Xu et al. 1998). In fact, breakdown of linkage responsible for transgressive segregation was through complementary gene action because by recombination, the linkage of QTLs for the same trait was broken and then the recombinants for extreme phenotypes appeared (Hagiwara et al. 2006; Winn et al. 2009). Complementary genes appear to be common for most traits, and complementary gene action has been confirmed as one of the genetic bases of transgressive segregation in previous studies (Devicente and Tanksley 1993; Xu et al. 1998; Rieseberg et al. 2003). However, transgressive segregation was sometimes observed for traits that appear to lack complementary alleles in the parental lines (Monforte et al. 1997; Kim and Rieseberg 1999). These observations could be taken as evidences for non-additive gene action. As a result of the difficulty in assessing the contributions of epistasis, epistasis was just assumed to be another major reason for transgression (Rieseberg et al. 1999). Wang et al. (2008) reported that epistasis was responsible for transgressive segregation in resistance to the root-knot nematode in cotton. In their study, a major gene, rkn1, on chromosome 11 for resistance to Meloidogyne incognita in cv. Acala NemX was identified; RKN2 was located in the vicinity of rkn1 on chromosome 11, RKN2 did not confer resistance in Pima S-7, but when combined with rkn1, high levels of resistance were produced in the F1 and segregating F2, F3, and BC1F1 populations. Consequently, RKN2 was called a transgressive segregation factor. A pair of epistatic interactions between LD1.1 and LD1.5 have been fine-mapped for seeds or root development (or both) in Arabidopsis (Bikard et al. 2009). All combinations of alleles at LD1.5 and its interactor LD1.1 result in phenotypically normal plants, except for the combination of Col at LD1.1/Heterozygous at LD1.5, which shows a reduced primary root length (weak-root phenotype), and another combination (Col at LD1.1/Cvi at LD1.5), which shows embryo lethality at an early stage in the silique. These genes were identified as a pair of histidinol-phosphate aminotransferase genes coding for a protein (HPA) that catalyzes an important step in the biosynthetic pathway leading to histidine. Both studies were focused only on a pair of interactions responding to transgressive segregation of a qualitative trait. However, interactions between unlinked genes at the genome-wide level to explain transgressive segregation of quantitative traits still require work.
With the release of the rice genome sequence (International Rice Genome Sequencing Project [IRGSP] 2005), plenty of molecular markers were developed and greatly facilitated determination of the genetic bases of transgressive segregation. The RIL population developed by single-seed descendants from a cross between two different varieties is suitable for determining the genetic bases of complex traits because RILs are homozygous for most of their loci and can be grown in different environments with several replicates (Young 1996; Loudet et al. 2002). This kind of genetic constitution is suitable for creating a simple model for epistasis analysis. Besides environment, only additive main effects and additive-by-additive epistatic effects are included in the model. Based on the mixed-model based composite interval method (MCIM), a computer software program (QTLMapper version 1.6) was developed, which was suitable for interval mapping of QTLs with additive, additive × additive epistasis, and their environment interactions (Wang et al. 1999).
Kilo-grain weight (KGW) was frequently reported with the highest heritability among three yield components (Rabiei et al. 2004). Dissection of its genetic bases was promising because comparatively reliable phenotypic data were obtained. In this study, an RIL population was used to uncover the mechanism of transgressive segregation of KGW, whose phenotypic data were collected from three different environments. Our study aimed to elucidate the genetic basis of KGW at the digenetic level and the genetic mechanism of transgressive segregation of KGW.
Materials and methods
Experimental population and phenotypic measurements
The population consisted of 177 RILs derived by single-seed descendent from a cross between an indica variety, Teqing (TQ), and a wide compatible japonica variety, 02428. The F7, F8, and F9 generations of the RILs, as well as two parents, were grown in the Experimental Station of Huazhong Agricultural University, Wuhan (30°N, 114°E), China, in the summer of 2005 and 2006, and Lingshui (18°N, 110°E), Hainan province, China, in the spring of 2007, respectively. Field trials were carried out following the randomized complete block design with two replications within each year. Ten seedlings at approximately 25 days old for each RIL were transplanted into the main field with a distance of 16.5 cm between plants within a row and 26.4 cm between rows. Eight plants in the middle of the row of each entry were harvested individually and measured for KGW. KGW was evaluated as the total filled grains weight divided by the number of grains from each plant multiplied by 1000, and the averaged trait measurements over the two replications within each year were used as the KGW of each line for analysis.
DNA extraction and molecular makers assay
Fresh leaves from one plant of each RIL were collected for DNA extraction according to the cetyl-trimethyl ammonium bromide (CTAB) method (Murray and Thompson 1980) with minor modifications. In total, 445 simple sequence repeat (SSR) markers covering all 12 chromosomes were screened for the polymorphism between the parents. The markers entered with three numbers were designed according to Temnykh et al. (2001, 2000), and those with four numbers according to the rice genome sequences of the Monsanto Company (McCouch et al. 2002). The SSR assay was carried out essentially as described by Wu and Tanksley (1993).
Data analysis
A molecular marker linkage map was constructed using MAPMAKER/EXP (version 3.0) (Lander et al. 1987). The linkage groups and the orders of markers on maps were determined by using the ‘group’ and ‘order’ commands of the Mapmaker program. The recombination values were converted to genetic distances in centimorgans (cM) using the Kosambi function (Kosambi 1944).
A method of mixed linear model based composite interval mapping (MCIM) was used to discover QTLs and to estimate additive effects, epistatic effects and QTLs × environments (QE) interaction effects by QTLMapper version 1.6 (Wang et al. 1999). For testing likelihood ratio (LR), we set P ≤ 0.005 as the threshold value, which was equal to LR ≥ 7.9. When we did two-dimensional scan using stepwise regression for pair-wise interaction markers and if a LR value at the peak of the scanning intervals was above the threshold, the testing point with that LR was thought to being of putative QTLs. The significant levels of additive effects and epistatic effects were confirmed by t-test and QE interaction effects were further tested by Bayesian test. The relative contributions were calculated as the proportion of variance caused by additive QTLs, epistatic QTLs and QE interactions in the total phenotypic variance. According to the model, a positive additive-additive (AA) value implied that the two-locus genotypes being the same as parents 02428 or Teqing took the positive effects, whereas the two-locus genotypes of recombination between parents took the negative effects and vice versa. For ease of description, the significant main effect QTLs were termed as “qKGWx,” in which x indicates the chromosome number and epistatic QTLs were designated as “iKGWy,” where y means the coding number of interactions.
Results
Transgressive segregation in the population
TQ expressed larger KGW compared to parent 02428 in all three environments. The performance of the population and two parents in 2007 (short-day condition) was better than that in 2005 and 2006 (long-day condition), in which there was an equivalent performance for KGW. The RIL population in 2007 showed the largest KGW variation, ranging from 17.1 to 35.5 g. In all the three environments, RILs with the largest grains had doubled KGW compared to the lowest one. Extreme RILs to either parent were observed in all 3 years. It was assumed that the RIL population had the same distribution as those of either parent line. The χ2 tests were used to compare the expected numbers of RILs in a population, whose KGWs exceeded either parent at levels of 2-fold the standard deviation (SD), with the actual number of RILs that exceeded those thresholds. The number of lines whose KGWs were less than the mean minus 2 SDs of the lower value parent 02428 significantly exceeded the expected number (P < 0.01) in three environments. Accordingly, the number of lines, whose KGWs were larger than the mean plus 2 SDs of the high value parent TQ significantly exceeded the expected number (P < 0.01). These results suggested that transgressive segregation occurred in two directions in the RIL population (Fig. 1). At least 10 and 43 lines showed extreme KGWs in small parent and large parent directions in the three environments (Table 1).
Genotyping and the linkage map
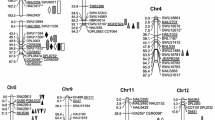
A total of 445 SSR markers were used for screening polymorphisms between the parents TQ and 02428; 222 markers (49%) showed polymorphisms between two parents, which indicated that the parents were diverse. Finally, 154 evenly distributed polymorphic markers were selected for genotyping the RIL population. These markers were assigned into 13 linkage groups, in which chromosome 3 was cut into two linkage groups. The total genetic map covered 1576 cM, with the average distance between adjacent markers 10.3 cM.
QTL and QE interactions resolved by two-locus analyses
Thirteen QTLs were identified for KGW in this RIL population (Table 2). Of these, 3 QTLs (qKGW3a, qKGw3b, and qKGw3c) were located on chromosome 3, two on each of chromosomes 4 (qKGW4a and qKGW4b), 6 (qKGW6a and qKGW6b), and 12 (qKGW12a and qKGW12b), and one on each of chromosomes 1 (qKGW1), 5 (qKGW5), 9 (qKGW9), and 10 (qKGW10). The additive effects of the QTLs ranged from 0.2 to 0.7 g and explained 0.7–4.7% of the phenotypic variance. Of these, TQ alleles at eight QTLs (qKWG3b, qKWG4b, qKWG5, qKWG6, qKWG8, qKWG9, qKWG10, and qKWG12) increased KGW, whereas 02428 alleles increased KGW at other QTLs.
Eight digenetic interactions were detected with significant epistatic effects on KGW, involving 16 loci dispersed on chromosomes 1, 4, 5, 8, 9, 10, and 12. One interaction (iKGW8) occurred between two loci with significant additive effects. Six interactions (iKGW1, iKGW3, iKGW4, iKGW5, iKGW6, and iKGW7) each involved a major additive-effect QTL and a locus without additive effect at the single locus level. Interestingly, the remaining one (iKGW2) occurred between two loci without significant additive effects. The effects of additive-additive interaction ranged from 0.2 to 0.6 g and explained 0.4–3.6% of the phenotypic variance. Parental genotypes for six interactions (iKGW1, iKGW2, iKGW3, iKGW4, iKGW6, and iKGW7) had positive effects on KGW. For the remaining two interactions (iKGW5 and iKGW8), recombinant genotypes had positive effects on KGW.
Two QTLs (qKGW3a and qKGW3b) showed significant QE, and their QE effects were −0.4 and +0.5 g and explained 1.2 and 1.5% of the variation, respectively. However, the QE effects were only detected in the short-day environment (2007), indicating that alleles on qKGW 3a and qKGW3b were sensitive to the photoperiod. The short-day condition was more favorable than the long-day condition for expression of the 02428 allele of qKGW3a, resulting in an increased KGW. In contrast, the short-day condition was favorable for expression of the TQ allele of qKGW3b, resulting in an increased KGW.
Only one epistatic QTL (iKGW1) was detected to significantly interact with environment. Its epistatic effect on KGW was 0.4 g and it explained 1.3% of the variation. The short-day condition was more favorable than the long-day condition for parent genotypes for iKGW1.
Overall, main-effect QTLs accounted for 30.0% of the phenotype variation; epistatic QTLs and QEs explained 14.0 and 4.4% of the variation, respectively.
Prediction of KGW for extreme lines
For the trait KGW, transgressive segregation was observed in the RIL population in all three environments (Table 1). For example, in the first environment (2005), at least 8.5% of RILs had higher KGW values than those of larger grain parent TQ, and at least 24.3% of RILs possessed lower values than those of the smaller grain parent 02428 (Table 1). Of these lines, seven with the largest KGW and seven with the lowest KGW were removed to explain transgressive segregation (Table 3).
According to the genetic model used in this study, the genetic effects of RILs were dissected into additive effects (A), additive-additive epistatic effects (AA), and QE interaction effects. The expectation of KGW could be expressed as KGWex = μ + (∑Ai +∑Aj + ∑AiAj) + (∑AiE + ∑AjE + ∑AiAjE), where KGWex was the expected KGW phenotypic value of each RIL, μ was the mean value of the population in all three environments, and ∑A (∑Ai and ∑Aj) or and ∑AiAj were summations of all additive effects (Ai or Aj) and all additive-additive epistatic effects (AiAj), respectively. ∑AiE, ∑AjE, and ∑AiAjE were the summation of Ai, Aj or AiAj by E interaction effects, respectively. Because three QE interactions were detected, but had no effects in 2005, then Q by E was not included in Table 3. For each RIL, the genotype of each main QTL or epistatic QTL was determined by the genotype of their flanking markers. Thus, the effects of main QTLs, epistatic QTLs, and QE interactions were estimated for each line. According to the model, the expected values of seven lines with the largest observed KGW were estimated. Their expected values ranged from 24.3 to 26.3 g, and their observed values ranged from 24.0 to 25.8 g, whereas for seven lines with the least observed KGW, the expected values ranged from 14.7 to 17.7 g and their observed values ranged from 13.8 to 16.7 g. The expectations were close to the experimental values, but there were still some differences probably derived from experimental error, which was impossible to estimate in the model. However, the differences between expected and observed values should display in a normal distribution. The pairwise differences of 14 lines ranged from −1.0 to 1.1 g, of which the mean (m) was 0.08 g and the SD was 0.74; they fell into the range from −1.4 (m − 1.96 SD) to 1.5 g (m + 1.96 SD), which equated to a 95% confidence interval in theory. That meant the model was suitable for predicting KGW for these lines.
Discussion
QTLs and epistasis explained the genetic bases of transgressive segregation of KGW
Several explanations account for the observations of transgression in a hybrid population (Rieseberg et al. 1999). These include: (1) an increased mutation rate in hybrids; (2) the exposure of rare recessive alleles that are normally heterozygous in the parental lines; (3) restricted developmental stability; (4) chromosome number variation; (5) overdominance (nonadditivity of allelic effects within a locus); (6) epistasis (nonadditivity of allelic effects between loci); and (7) the complementary action of additive alleles that are dispersed between the parental lines. In this study, because the first five situations did not exist in the RIL population, transgression could be explained only by the latter two hypotheses. The complementary action of additive alleles assumed that parental lines often harbored sets of alleles with opposite effects within lines, and most or all of the alleles with positive or negative effects in different loci were assembled into the same progenies by recombination, which would result in transgression in hybrids. This was a popular general explanation for transgressive segregation and illustrated in many previous studies (Devicente and Tanksley 1993; Xu et al. 1998; Rieseberg et al. 2003). In this study, 13 QTLs at the level of a single locus were identified for KGW; TQ alleles of six QTLs increased KGW but decreased KGW at seven QTLs. This also confirmed that the complementary action of additive alleles was an important genetic factor for transgression. However, the complementary action of additive alleles alone could not adequately explain transgression. For example, although line 174 had approximately the same number of positive alleles as the negative alleles (7+/6−), it still outperformed the higher parent TQ. This could be accounted for by digenetic interactions because line 174 possessed many more positive interactions than negative ones (7+/1−). Finally, its net genetic effect was positive. In contrast, line 164 had only three more negative alleles than positive alleles (5+/8−), but it outperformed the lower parent 02428 because it possessed many more negative digenetic interactions (1+/7−) and, consequently, its net genetic effect was negative, which resulted in a lower KGW than the lower KGW parent. Actually, those seven lines with the largest KGW possessed either many more positive alleles, many more positive digenetic interactions, or more positive alleles and interactions than the negative counterparts and vice versa (Table 3). Hence, the complementary action of additive alleles and epistasis could well explain transgressive segregation.
Epistasis played an important role in the genetic bases of KGW
13 additive QTLs identified in this study could explain 30.0% of the KGW variation. Seven QTLs were previously identified in other primary populations, including qKGW1, qKGW3b, qKGW3c, qKGW5, qKGW6, qKGW9, and qKGW12b (Lu et al. 1997; Xing et al. 2002; Thomson et al. 2003; Ge et al. 2005). Eight digenetic interactions were detected and explained 14% of the variation. In detail, eight QTLs of additive effects were involved in six digenetic interactions. Of these, two pairs of QTLs interacted each other. The other four QTLs separately interacted with the locus that had no significant additive effect at the single-locus level. Two interactions occurred between two loci that had no significant additive effects at the single-locus level. Only in five QTLs were their additive effects not statistically affected by other genetic backgrounds. These results agreed with the previous findings that some epistatic QTLs did not have main effects at the single-locus level. In contrast, a large proportion of the main-effect QTLs were involved in the epistatic interactions (Li et al. 1997; Xing et al. 2002). This indicated epistasis played an important role in controlling KGW.
The biologic nature of epistasis interaction detected statistically for KGW is unknown. Currently, the pathway involved in KGW expression is still unclear. Cloning and characterizing KGW QTLs would be helpful to understand the molecular mechanism underlying epistasis. Currently, only four QTLs for KGW have been cloned including GW2, GS3, GW5 (or qSW5) and GIF1. GW2 is located on chromosome 2 coded a RING-type E3 ubiqutin ligase (Song et al. 2007). GW5 (or qSW5) encoded a novel nuclear protein of 144 amino acids (Shomura et al. 2008; Weng et al. 2008). Both GW2 and GW5 negatively regulated grain weight and functioned in the pathway of ubiquitin–proteasome; they may work together to target degradation of certain substrates that promote cell division and grain growth as indicated by the interaction between GW2 and GW5 (or qSW5) that probably occurred. With the progress being made in metabolic studies, the biologic meaning of statistical epistasis may be gradually uncovered.
Exploitation of transgression segregation in rice breeding
For complicated traits in rice, such as yield-related traits, seedling vigor, phosphorus deficiency tolerance, and so on, transgressive segregation is frequently present in populations of not only low generation (F2) but also high generation (RIL) derived from the cross between intra-subspecies (Li et al. 1995; Redona and Mackill 1996a; Ni et al. 1998; Xu et al. 1998; Matsubara et al. 2008), between inter-subspecies (Redona and Mackill 1996b; Xiao et al. 1996; Xue et al. 1997), or between cultivar and wild rice (Xiao et al. 1998; Brondani et al. 2002; Haq et al. 2008). More attention should be paid to exploit transgressive segregation in rice breeding. The positive transgression such as heterosis is being pursued in agricultural production. To get a high-yield or high-quality variety, usually multiple superior QTLs are pyramided into a planting variety by recombination and marker-assisted selection (Lande and Thompson 1990; Ribaut and Hoisington 1998; Xu and Crouch 2008). The complementary action of additive alleles is critical to this method. However, in this study, epistasis was another important factor for transgression. In addition, epistasis has been reported to play an important role not only in the genetic bases of heterosis (Yu et al. 1997), but also in hybrid breakdown (Yamamoto et al. 2007; Jiang et al. 2008). Epistasis should be considered when a breeding program is designed. For example, the expected KGW of plants pyramiding all detected positive QTLs was 27.7 g. If the plant also pyramids all positive interactions, its KGW rose to 30.9 g. Obviously, reasonable application of positive interactions could cause a considerable effect on a target trait.
References
Albertson RC, Kocher TD (2005) Genetic architecture sets limits on transgressive segregation in hybrid cichlid fishes. Evolution 59:686–690
Barton NH, Hewitt GM (1985) Analysis of hybrid zones. Ann Rev Ecol Syst 16:113–148
Bell MA, Travis MP (2005) Hybridization, transgressive segregation, genetic covariation, and adaptive radiation. Trends Ecol Evol 20:358–361
Bikard D, Patel D, Metté CL, Giorgi V, Camilleri C, Bennett MJ, Loudet O (2009) Divergent evolution of duplicate genes leads to genetic incompatibilities within A. thaliana. Science 323:623–626
Brem RB, Kruglyak L (2005) The landscape of genetic complexity across 5, 700 gene expression traits in yeast. Proc Natl Acad Sci USA 102:1572–1577
Brondani C, Rangel PNN, Brondanil RPV, Ferreira ME (2002) QTL mapping and introgression of yield-related traits from Oryza glumaepatula to cultivated rice (Oryza sativa) using microsatellite markers. Theor Appl Genet 104:1192–1203
Devicente MC, Tanksley SD (1993) QTL analysis of transgressive segregation in an interspecific tomato cross. Genetics 134:585–596
Engels WR (1983) The P family of transposable elements in Drosophila. Annu Rev Genet 17:315–344
Ge XJ, Xing YZ, Xu CG, He YQ (2005) QTL analysis of cooked rice grain elongation, volume expansion, and water absorption using a recombinant inbred population. Plant Breeding 124:121–126
Grant V (1975) Genetics of flowering plants. Columbia University Press, New York
Hagiwara WE, Onishi K, Takamure I, Sano Y (2006) Transgressive segregation due to linked QTLs for grain characteristics of rice. Euphytica 150:27–35
Haq TU, Aakhtarl J, Gorham J, Steele KA, Khalid M (2008) Genetic mapping of QTLs, controlling shoot fresh and dry weight under salt stress in rice (Oryza sativa L.) cross between Co39 × Moroberekan. Pak J Bot 40:2369–2381
International Rice Genome Sequencing Project (2005) The map-based sequence of the rice genome. Nature 436:793–800
Jiang WZ, Chu SH, Piao RH, Chin JH, Jin YM, Lee JH, Qiao YL, Han LZ, Piao ZZ, Koh HJ (2008) Fine mapping and candidate gene analysis of hwh1 and hwh2, a set of complementary genes controlling hybrid breakdown in rice. Theor Appl Genet 116:1117–1127
Kim SC, Rieseberg LH (1999) Genetic architecture of species differences in annual sunflowers: implications for adaptive trait introgression. Genetics 153:965–977
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugenics 12:172–175
Lande R, Thompson R (1990) Efficiency of marker-assisted selection in the improvement of quantitative traits. Genetics 124:743–756
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Li ZK, Pinson SRM, Stansel JW, Park WD (1995) Identification of quantitative trait loci (QTLs) for heading date and plant height in cultivated rice (Oryza sativa L.). Theor Appl Genet 91:374–381
Li ZK, Pinso SRM, Park WD, Paterson AH, Stansel JW (1997) Epistasis for three grain yield components in rice (Oryza sativa L.). Genetics 145:453–465
Li ZK, Yu SB, Lafitte HR, Huang N, Courtois B, Hittalmani S, Vijayakumar CHMl, Liu GF, Wang GC, Shashidhar HE, Zhuang JY, Zheng KL, Singh VP, Sidhu JS, Srivantaneeyakul S, Khush GS (2003) QTL × environment interactions in rice. I. Heading date and plant height. Theor Appl Genet 108:141–153
Liu GF, Zhang ZM, Zhu HT, Zhao FM, Ding XH, Zeng RZ, Li WT, Zhang GQ (2008) Detection of QTLs with additive effects and additive-by-environment interaction effects on panicle number in rice (Oryza Sativa L.) with single-segment substitution lines. Theor Appl Genet 116:923–931
Loudet FO, Chaillou S, Camilleri C, Bouchez D, Daniel-Vedele F (2002) Bay-0 × Shahdara recombinant inbred line population: a powerful tool for the genetic dissection of complex traits in Arabidopsis. Theor Appl Genet 104:1173–1184
Lu CF, Shen LH, Tan ZB, Xu YB, He P, Chen Y, Zhu LH (1997) Comparative mapping of QTLs for agronomic traits of rice across environments by using a doubled-haploid population. Theor Appl Genet 94:145–150
Matsubara K, Kono I, Hori K, Nonoue Y, Ono N, Shomura A, Mizubayashi T, Yamamoto S, Yamanouchi U, Shirasawa K, Nishio T, Yano M (2008) Novel QTLs for photoperiodic flowering revealed by using reciprocal backcross inbred lines from crosses between japonica rice cultivars. Theor Appl Genet 117:935–945
McCouch SR, Teytelman L, Xu YB, Lobos KB, Clare K, Walto MN, Fu BY, Maghirang R, Li ZK, Xing YZ, Zhang QF, Kono I, Yano M, Fjellstrom R, DeClerck G, Schneider D, Cartinhour S, Ware D, Stein L (2002) Development and mapping of 2240 new SSR markers for rice (Oryza sativa L.). DNA Res 9:199–207
Monforte AJ, Asins MJ, Carbonell EA (1997) Salt tolerance in Lycopersicon species V. Does genetic variability at quantitative trait loci affect their analysis? Theor Appl Genet 95:284–293
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4326
Ni JJ, Wu P, Senadhira D, Huang N (1998) Mapping QTLs for phosphorus deficiency tolerance in rice (Oryza sativa L.). Theor Appl Genet 97:1361–1369
Rabiei BM, Valizadeh M, Ghareyazie B, Moghaddam M (2004) Evaluation of selection indices for improving rice grain shape. Field Crops Res 8:359–367
Redona ED, Mackill DJ (1996a) Molecular mapping of quantitative trait loci in japonica rice. Genome 39:395–403
Redona ED, Mackill DJ (1996b) Mapping quantitative trait loci for seedling vigor in rice using RFLPs. Theor Appl Genet 92:395–402
Ribaut JM, Hoisington D (1998) Marker-assisted selection: new tools and strategies. Trends Plant Sci 3:236–239
Rick CM, Smith PG (1953) Novel variation in tomato species hybrids. Am Nat 87:359–375
Rieseberg LH, Ellstrand NC (1993) What can morphological and molecular markers tell us about plant hybridization? Crit Rev Plant Sci 12:213–241
Rieseberg LH, Arche MA, Wayne RK (1999) Transgressive segregation, adaptation and speciation. Heredity 83:363–372
Rieseberg LH, Raymond O, Rosenthal DM, Lai Z, Livingstone K, Nakazato T, Durphy J, Schwarzbach AE, Donovan LA, Lexer C (2003) Major ecological transitions in wild sunflowers facilitated by hybridization. Science 301:1121–1126
Shomura A, Izawa T, Ebana K, Ebitani T, Kanegae H, Konishi S, Yano M (2008) Deletion in a gene associated with grain size increased yields during rice domestication. Nat Genet 40:1023–1028
Song XJ, Huang W, Shi M, Zhu MZ, Lin HX (2007) A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat Genet 39:623–630
Temnykh S, Park WD, Ayres N, Cartinhour S, Hauck N, Lipovich L, Cho YG, Ishii T, McCouch SR (2000) Mapping and genome organization of microsatellite sequences in rice (Oryza sativa L.). Theor Appl Genet 100:697–712
Temnykh S, DeClerck G, Lukashova A, Lipovich L, Cartinhour S, McCouch SR (2001) Computational and experimental analysis of microsatellites in rice (Oryza sativa L.): frequency, length variation, transposon associations, and genetic marker potential. Genome Res 11:1441–1452
Thomson MJ, Tai TH, McClung AM, Lai XH, Hinga ME, Lobos KB, Xu Y, Martinez CP, McCouch SR (2003) Mapping quantitative trait loci for yield yield components and morphological traits in an advanced backcross population between Oryza rufipogon and the Oryza sativa cultivar Jefferson. Theor Appl Genet 107:479–493
Wan XY, Weng JF, Zhai HQ, Wang JK, Lei CL, Liu XL, Guo T, Jiang L, Su N, Wan JM (2008) Quantitative trait loci (QTL) analysis for rice grain width and fine mapping of an identified QTL allele gw-5 in a recombination hotspot region on chromosome 5. Genetics 179:2239–2252
Wang DL, Zhu J, Li ZK, Paterson AH (1999) User manual for QTLMapper version 1.0—a computer software for mapping quantitative trait loci (QTLs) with main effects, epistatic effects and QTL × environment interactions. http://ibi.zju.edu.cn/software/qtlmapper/QTLMaperManual.PDF
Wang CL, Ulloa M, Roberts PA (2008) A transgressive segregation factor (RKN2) in Gossypium barbadense for nematode resistance clusters with gene rkn1 in G. hirsutum. Mol Genet Genomics 279:41–52
Weng JF, Gu SH, Wan XY, Gao H, Guo T, Su N, Lei CL, Zhang X, Cheng ZJ, Guo XP, Wang JL, Jiang L, Zhai HQ, Wan JM (2008) Isolation and initial characterization of GW5, a major QTL associated with rice grain width and weight. Cell Res 18:1199–1209
Winn JA, Mason RE, Robbins AL, Rooney WL, Hay DB (2009) QTL mapping of a high protein digestibility trait in Sorghum bicolor. Int J Plant Genomics 2009, 471853:6. doi:10.1155/2009/47185
Wu KS, Tanksley SD (1993) Abundance, polymorphism and genetic mapping of microsatellites in rice. Mol Gen Genet 241:225–235
Xiao JH, Li JM, Yuan LP, Tanksley SD (1996) Identification of QTLs affecting traits of agronomic importance in a recombinant inbred population derived from a subspecific rice cross. Theor Appl Genet 92:230–244
Xiao JH, Li JM, Grandillo S, Ahn SN, Yuan LP, Tanksley SD, McCoucha SR (1998) Identification of trait-improving quantitative trait loci alleles from a wild rice relative, Oryza rufipogon. Genetics 150:899–909
Xing YZ, Tan YF, Hua JP, Sun XL, Xu CG, Zhang QF (2002) Characterization of the main effects, epistatic effects and their environmental interactions of QTLs on the genetic basis of yield traits in rice. Theor Appl Genet 105:248–257
Xu YB, Crouch JH (2008) Marker-assisted selection in plant breeding: from publications to practice. Crop Sci 48:391–407
Xu YB, McCouch SR, Shen ZT (1998) Transgressive segregation of tiller angle in rice caused by complementary gene action. Crop Sci 38:12–19
Xue Y, Jiang L, Su N, Wang JK, Deng P, Ma JF, Zhai HQ, Wan JM (1997) The genetic basic and Wne-mapping of a stable quantitative-trait loci for aluminium tolerance in rice. Planta 227:255–262
Yamamoto E, Takashi T, Morinaka Y, Lin SY, Kitano H, Matsuoka M, Ashikari M (2007) Interaction of two recessive genes, hbd2 and hbd3, induces hybrid breakdown in rice. Theor Appl Genet 115:187–194
Young ND (1996) QTL mapping and quantitative disease resistance in plants. Annu Rev Phytopath 34:479–501
Yu SB, Li JX, Xu CG, Tan YF, Gao YJ, Li XH, Zhang QF, Saghai Maroof MA (1997) Importance of epistasis as the genetic basis of heterosis in an elite rice hybrid. Proc Natl Acad Sci USA 94:9226–9231
Acknowledgments
This work was supported in part by Grants from the National Program on the Development of Basic Research (2007CB109001) and the National Natural Science Foundation of China (30971749, 30830064).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mao, D., Liu, T., Xu, C. et al. Epistasis and complementary gene action adequately account for the genetic bases of transgressive segregation of kilo-grain weight in rice. Euphytica 180, 261–271 (2011). https://doi.org/10.1007/s10681-011-0395-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-011-0395-0