Abstract
In this review, we present a critical analysis of the current status of wild Lactuca L. germplasm in relation to its utility for lettuce breeding. We discuss wild Lactuca germplasm in ex situ collections from the perspectives of taxonomy, biogeography, biology and ecology, gene pools, field exploration and acquisition, descriptor development, characterization and evaluation, and enhancement. Future research and other activities related to wild Lactuca germplasm and their continued exploitation in lettuce breeding are considered.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genetic resources of wild Lactuca species as conserved in the world’s genebanks are an integral part of our global plant heritage, and they play important role in modern lettuce breeding (Lebeda et al. 2007c; Mikel 2007; Maggioni et al. 2008; Mou 2008). Considerable progress in both fundamental research on Lactuca germplasm and its practical applications has been achieved during last 25 years (Lebeda et al. 2007c). The most important remaining gaps, problems and sources of confusion related to the effective use of these key resources are highlighted in this paper along with a recent progress report.
Taxonomy of Lactuca L.
Taxonomic and phylogenetic studies clearly place the genus Lactuca L. in the tribe Lactuceae, subfamily Cichorioideae, of the Compositae (Asteraceae) (Funk et al. 2005), one of the largest plant families. A careful review of published literature confirmed the existence of about 100 wild Lactuca spp., with the highest number of autochthonous species and species richness in Asia (51 species) and Africa (43 species) (Lebeda et al. 2004b). The most recent molecular data on phylogenetic relationships among Lactuca species (Koopman et al. 1998) confirmed, with some modifications, a previously elaborated broader generic concept (summarized by Lebeda et al. 2007c). However, formal classification, including subgeneric divisions (Lebeda et al. 2007c), need critical reconsideration and further elaboration.
In addition, serious taxonomic discrepancies can be found in the main world collections of Lactuca L. germplasm. Doležalová et al. (2004) studied 49 accessions of 24 Lactuca species received from the main world genebanks. It was found, after taxonomic review, that 35% of accessions were wrongly taxonomically described and were redetermined (on the genus, species and subspecific level). Therefore, good knowledge of classical taxonomy combined with the comparative study of original herbarium specimens must be considered as a most important step for the efficient management and utilization of Lactuca genetic resources (Lebeda et al. 1999), and correct interpretation of experimental data (Lebeda et al. 2002).
Geographic distribution and hot-spots of diversity
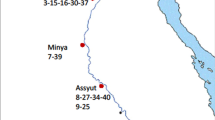
There is increasing interest in the potential value of genes from wild species in crop improvement (Gass and Frese 1999). For lettuce, there are two crucial issues related to the utilization of genes from wild species: loss of genetic diversity in situ and limited access to wild Lactuca in current ex-situ germplasm collections (Lebeda et al. 2004a, 2007c). To overcome these challenges, genebanks should focus on rapidly acquiring lettuce progenitors and wild relatives from the probable center of origin of lettuce and from those areas with the highest genetic diversity of Lactuca species (Lebeda et al. 2004c). High levels of diversity of Lactuca species found in the Mediterranean basin and southwestern Asia indicate that those regions should be seriously considered as hot-spots for lettuce conservation (Beharav et al. 2008a, b; Kitner et al. 2008; Lebeda et al. 2001b, c, 2008d). Future ecogeographic studies should also focus on central and southern Africa, central Asia, and North America to determine if other hot-spots exist and to develop collecting strategies accordingly (Lebeda et al. 2007c) (Fig. 1).
Biology and ecology
The genus Lactuca includes annual, biennial and perennial herbs, and rarely shrubs, with abundant latex. Sections Phoenixopus, Mulgedium, Lactucopsis, Tuberosae, Micranthae and Sororiae (see Table 1) are mostly biennial or perennial (Lebeda and Astley 1999). The division of section Lactuca into two subsections, Lactuca and Cyanicae, is based on the life cycle of their members (Feráková 1977). Subsection Lactuca comprises annual, winter annual or biennial herbs; perennial species belong to subsection Cyanicae. The autochthonous North American species are mostly biennial; however, at least one perennial species, L. tatarica subsp. pulchella (syn. L. oblongifolia), is also reported (McGregor et al. 1986). The African species are annual or perennial herbs or sub-shrubs, rarely scandent (Lebeda et al. 2004b).
The genus Lactuca comprises species with various ecological requirements occupying diverse habitats. The species of lettuce’s genepool (those of the breeders’ main interest), L. serriola, L. saligna and L. virosa, are weedy and occur on waste places and ruderal habitats—mainly along roads, highways and ditches (Lebeda et al. 2001b, c, 2004b, 2007a) (Fig. 2). Most species, i.e. L. perennis, L. viminea, L. graeca, and L. tenerrima, are calciphilous plants found in limestone and dolomite areas, often on rocky slopes. Endemic, lianalike species are found in rain forests of East Africa. Comprehensive surveys regarding the biology and ecology of European Lactuca species were conducted by Feráková (1977) and Lebeda et al. (2004b), who summarized available information on about 100 species from current world literature. However, basic data about the biology and ecology of most species, especially those of African and Asian origin, are still unavailable.
Gene pools and genetic diversity
Human effort in the process of domestication probably involved selection of wild relatives for leaves that had a reduction in leaf spines, latex content and bitter flavor, and for plants with delayed bolting. Domestication has also led to a shortening of internodes, bunching of leaves, increased seed size and non-shattering (Fig. 3), and changes in photoperiodism (enabling cultivation under various daylengths). These processes have been accompanied by bottlenecks that restricted the genetic diversity available in the primary gene pool.
In general, the primary gene pool of cultivated lettuce comprises the numerous cultivars and landraces of L. sativa and its wild ancestor, L. serriola. The wild serriola-like species from southwestern Asia (i.e., L. aculeata, L. altaica, L. azerbaijanica, L. georgica, and L. scarioloides) and the African species, L. dregeana, all display similar levels of interfertility with the crop and belong to the primary gene pool as well (Lebeda et al. 2007c). Although L. saligna and L. virosa have been intensively studied by both evolutionary biologists and plant breeders, their categorization to the secondary or tertiary gene pools has remained an open question (Fig. 4). A view rather different from that outlined above was proposed by Koopman et al. (1998), who suggested that section Lactuca subsection Lactuca comprises the primary and secondary gene pools, while sections Phaenixopus, Mulgedium and Lactucopsis make up the tertiary gene pool. However, the categorization of many Lactuca species into gene pools is still unclear and needs additional attention.
Germplasm collections—recent status and problems
Collections, their structure and gaps
Data describing wild Lactuca germplasm collections in Europe and around the world were summarized by Lebeda and Boukema (2001) and Lebeda et al. (2007c), and information concerning the exploitation of these wild relatives in commercial lettuce breeding has been summarized by Lebeda et al. (2007c) and Mou (2008). From these reports, it is clear that there are only few important collections in Europe (ca. 5) and the USA (ca. 3). In geographic centres of high species richness and diversity there are no significant germplasm collections with local accessions. Analysis of the International Lactuca Database (ILDB) showed that over 90% of wild collections are represented by only three species, L. serriola, L. saligna, and L. virosa, mostly of European origin. The autochthonous species from other continents (Asia, Africa, and the Americas), which form ca. 83% of known Lactuca species richness (Lebeda et al. 2004b), are represented in collections by only about 3% of the accessions (Lebeda et al. 2004a). Recently, Pandey et al. (2008) collected 373 species of wild crop relatives representing 120 genera and 48 families in the Indian gene centre; however, they made no mention of lettuce nor of its wild relatives. This example illustrates the underrepresentation of wild Lactuca species in recent collecting activities, which is a crucial feature needing attention for the future development of these collections (Lebeda et al. 2007c; Beharav et al. 2008b).
Taxonomic status of accessions and duplicates
The correct use of botanical nomenclature and, more importantly, the accurate taxonomic identification of genebank accessions are core tasks for the effective management and utilization of plant genetic resources. Insufficient or incorrect passport data, including taxonomic identification, complicate the evaluation of accessions (van Hintum and Boukema 1999; Lebeda et al. 2007c; Rajicic and Dehmer 2008), and make it more difficult to preserve genetic integrity, reduce collection redundancy, and interpret research findings.
Basic errors in the taxonomic status of accessions as reported by genebanks have been found repeatedly. When evaluating a set of 49 accessions of 24 wild Lactuca species for morphological characters, chromosome number, relative DNA content and isozyme polymorphisms, 17 accessions were reclassified and/or their taxonomic status criticized (Doležalová et al. 2004). Within a set of 95 accessions provided by gene banks in the Czech Republic, Germany, Netherlands, UK and the USA, nominally representing 12 species (L. aculeata, L. altaica, L. dentata, L. dregeana, L. indica, L. livida, L. perennis, L. quercina, L. saligna, L. serriola, L. tatarica and L. virosa), a morphological assessment confirmed the taxonomic identities of only 50 accessions; 31 accessions were re-determined (Lebeda et al. 2007d). Examples are given in Fig. 5. The remaining 14 accessions represented mixtures of L. serriola forms, mixtures of different Lactuca species, and interspecific hybrids (Doležalová et al. 2007a) (Fig. 6).
Taxonomic status of a set of 95 Lactuca species accessions representing 12 species received from main world gene banks (RICP, IPK, GNG, HRI, WG, LET). 1) plants of 31 morphologically uniform accessions, their taxonomic status re-determined; 2) 14 accessions represented by mixtures of L. serriola forms, different Lactuca species or interspecific hybrids; 3) taxonomic status of 15 accessions of L. serriola completed by determination of a lower taxonomic unit (f. serriola, f. integrifolia); 4) plants of 35 morphologically uniform accessions, their declared taxonomic status confirmed
An understanding of accession redundancy/duplication within and among genebanks is another important aspect of efficient plant genetic resource management (Spooner et al. 2005). Comparison of passport data from four large Lactuca collections (CGN, WRPIS, IPK and HRI) showed that 60% of 95 accessions are duplicated at least once among these collections (van Hintum and Boukema 1999). A morphological assessment of the abovementioned set of 95 Lactuca species accessions identified 34 duplicate groups on the basis of passport data, and showed that 69 accessions can be considered as morphological duplicates (Doležalová et al. 2007a; Lebeda et al. 2007d).
Field studies and collection activities
An increasing interest in determining the geographic distribution of wild Lactuca populations and sampling those populations in natural habitats resulted in the initiation of collection expeditions coordinated by the Department of Botany, Palacký University in Olomouc (Czech Republic) beginning in the early 1990s. From 1995 to 2008, expeditions were conducted in 14 European countries (Austria, Croatia, the Czech Republic, France, Hungary, Germany, Greece, Italy, Lithuania, The Netherlands, Slovakia, Slovenia, Spain, Sweden, Switzerland, the United Kingdom), 15 states of the USA (Arizona, California, Colorado, Idaho, Iowa, Nevada, North Carolina, Minnesota, Montana, Oregon, South Dakota, Utah, Washington, Wisconsin, Wyoming), and Canada. Field studies and germplasm collections were also made in Turkey, Israel, Jordan, Kazakhstan, South Korea, New Zealand and South Africa (Fig. 7). Collectively, these efforts have resulted in the collection of nearly 1,300 seed samples of 12 wild Lactuca species (Křístková and Lebeda 1999; Doležalová et al. 2001; Křístková et al. 2001; Lebeda et al. 2001b, c, 2007e; Beharav et al. 2008b; Doležalová et al. 2008).
Expeditions to collect L. serriola germplasm in four European countries were conducted within the framework of the EU-funded project “GENE-MINE” in 2001 (Lebeda et al. 2007a). The seed material (800 accessions from 50 locations) was used for regeneration, inclusion in national genebanks in the respective countries, and research purposes in follow-up studies (e.g., Lebeda and Petrželová 2004a; Lebeda et al. 2008a).
In cooperation with the Institute of Evolution (University of Haifa, Israel), expeditions focusing on the wild species, L. saligna, were conducted in 2004–2007 in Israel (Beharav et al. 2008a, b), following international standards for germplasm acquisition (Guarino et al. 1995) designed in a manner to avoid the collection of duplicates (van Hintum and Boukema 1999; Lebeda et al. 2004a).
Descriptor development
Precise descriptions of genetic resources serve as tool for their correct taxonomic determination and help define both interspecific and intraspecific variation (Lebeda et al. 2007b). A basic international descriptor list has been described for the genetic resources of L. sativa and of L. serriola and related species from the primary gene pool by a representatives of European genebanks within the activities of the European Cooperative Programme (ECP/GR), Working Group of Leafy Vegetables (Lebeda and Boukema 2005; Maggioni et al. 2008). In addition, an international list of the most important morphological characters of wild Lactuca species was created through the EU-funded project “GENE-MINE” (Doležalová et al. 2003a).
There have also been national descriptor lists published for the characterization of major Lactuca collections, including those from the Centre for Genetic Resources (CGN, Wageningen, The Netherlands) (Boukema et al. 1990), the Western Regional Plant Introduction Station (Pullman, Washington, USA) (McGuire et al. 1993), and the National Programme of Conservation and Utilization of Plant Genetic Resources of the Czech Republic for both cultivated lettuce (Lactuca sativa L.) (Křístková et al. 2008) and wild species (Doležalová et al. 2002a).
As descriptor lists for cultivated lettuce are developed or revised, serious consideration should be given to the inclusion of phenotypic traits of the wild Lactuca species being used to breed new lettuce cultivars. Because of these breeding efforts, during last decade new lettuce cultivars have been released with characteristics that do not conform to earlier groups of morphotypes.
Characterization and evaluation of wild Lactuca germplasm
Morphology
Morphological assessment of accessions can be performed during their regeneration by genebanks (Doležalová et al. 2002a, 2003a; Křístková et al. 2008), but this can have serious limitations, as the expression of morphological traits under controlled, regeneration conditions may differ significantly from expression under typical field conditions. Detailed studies of morphological variation have been performed for collections of L. serriola and L. saligna. Fifty L. serriola populations collected in four European countries (Czech Republic, Germany, The Netherlands, United Kingdom) (Lebeda et al. 2007a) were cultivated in a greenhouse under controlled conditions. Assessment included 26 quantitative and qualitative characters of stems (e.g., stem length), rosette and cauline leaves (e.g., depth of incisions) (Fig. 8), inflorescences and flowers (e.g., anthocyanin coloration on bracts) (summarized in Lebeda et al. 2007b), and fruits (e.g., length and width of achene body, length of achene beak and number of ribs) (Doležalová et al. 2007b) (Fig. 9). A similar morphological assessment was performed for about 70 populations of L. saligna from Czech Republic, France, Italy, Portugal, Israel, Jordan and Turkey (Křístková et al. 2007a; Beharav et al. 2008a, b). These studies revealed considerable (and previously unremarked) phenotypic variation, which must be seriously considered in future research and characterization activities with wild Lactuca germplasm, especially in relation to genotyping.
Phenology
The genus Lactuca is extremely variable in terms of phenology and plant development. Among developmental characteristics, substantial differences in time of anthesis were recorded among a geographically diverse set of accessions of L. serriola (Doležalová et al. 2005; Lebeda et al. 2007b, c). Substantial differences in developmental stages (beginning of bolting and flowering) were recorded among 89 L. serriola samples from different ecogeographic regions in Europe when grown under common conditions in a greenhouse. Developmental stages of plants, as influenced through selective processes under the original eco-geographic conditions where they evolved (Lebeda et al. 2001b, c), can persist when plants are cultivated under common environmental conditions and may be fixed genetically (Křístková et al. 2007b).
Karyology and DNA content
Wild Lactuca species can be divided into three main groups, according to their base chromosome number (Feráková 1977). The first group is relatively small and contains perennial species of Europe and the Himalayas with haploid chromosome number, n = 8. The haploid chromosome number, n = 9, characterizes the majority of European and Mediterranean species, as well as species from the Middle East, Africa and India. The third group, containing of North American species distributed from Canada to Florida, is marked by a haploid chromosome number of n = 17. It is of amphidiploid origin and is somewhat geographically and genetically isolated. Our understanding of genus remains incomplete, because the chromosome numbers of numerous Lactuca species are not known (Lebeda and Astley 1999) or may differ from reported data, as was reported by Doležalová et al. (2002b) for certain North American species.
To date, analyses of variation in nuclear DNA content have been performed on only a limited number of Lactuca species (i.e., L. sativa, L. serriola, L. saligna, and L. virosa) (Bennett and Leitch 1995; Koopman and De Jong 1996; Koopman 1999, 2000). Flow cytometry was tested for its reliability as a tool to distinguish among Lactuca species (Koopman 1999, 2000). Doležalová et al. (2002b) analyzed 50 accessions of 25 Lactuca species, along with Mycelis muralis, for chromosome number and relative DNA-content variation. Later, Koopman (2002) showed that five Lactuca species (L. viminea, L. virosa, L. serriola, L. sativa and L. sibirica) have significant intraspecific variation in DNA content, but concluded that only the variation within L. virosa seemed to have evolutionary significance. More recent studies have focused on intraspecific differences in DNA content in L. serriola germplasm originating from 12 European countries (Lebeda et al. 2004c, 2007c).
Karyotype analysis and relative DNA content were used to help characterize L. sativa, L. serriola, L. saligna and L. virosa and describe their evolutionary relationships (Koopman and De Jong 1996). Matoba et al. (2007) described detailed karyotype analyses of lettuce and allied species. These analyses revealed a dissimilarity between L. virosa and the remaining species. The simultaneous FISH (Fluorescence in situ hybridization) of 5S and 18S rDNAs revealed that both rDNA loci of L. sativa, L. serriola and L. saligna were identical; however, those of L. virosa differed from the other species, supporting a closer relationship between L. sativa/L. serriola and L. saligna than with L. virosa.
Protein and molecular diversity
The status of characterization of Lactuca species germplasm by protein and molecular markers has been recently summarized by Dziechciarková et al. (2004a) and Lebeda et al. (2007c). Various methods and approaches have been applied for this purpose; however, only a relatively limited number of wild Lactuca species and accessions have been analysed (Table 2) (e.g., Jansen et al. 2006). More extensive studies covering a broader geographic range and a larger number of populations are needed to describe relationships between ecogeographical conditions and corresponding genetic polymorphisms. Studies of this sort for L. saligna and L. serriola are now underway (Dziechciarková et al. 2004b; Kitner et al. 2008; Kuang et al. 2006, 2008; Lebeda et al. 2008c).
Biochemical diversity
The body of research on the evaluation of Lactuca germplasm also includes work on the detection and characterization of secondary phytochemicals, such as sesquiterpene lactones, phenolics, glucosides, and flavonoids, of pharmacological importance (Rees and Harborne 1984; Kisiel and Barszcz 1998; Kisiel and Zielinska 2000) (Fig. 10). This aspect has probably been underestimated, but we see increasing potential for the exploitation of at least some wild Lactuca germplasm in medicinal and pharmacological applications (Chen et al. 2007; Kim et al. 2007). Recently, phytochemical analyses have also been applied to the clarification of taxonomical relationships among various Asteraceae (Bohm and Stuessy 2001) including Lactuca species (Michalska et al. 2008).
A dendrogram showing clustering (Tree clustering method) of Lactuca species in relationship to the content of different sesquiterpene lactones in leaves [elaborated by authors on the basis of data in Michalska et al. (2008)]
Resistance to diseases and pests
Recent advancement in research and breeding of lettuce for resistance to diseases and pests has been summarized elsewhere (Lebeda et al. 2007c; Mou 2008). Many sources of resistance to pathogens and pests have been found and described in wild Lactuca species (Table 3). Traditionally, Bremia lactucae has been considered the most important pathogen causing disease in cultivated lettuce. Limited availability of durable sources of resistance to Bremia has stimulated interest among breeders in new sources from wild Lactuca species (Lebeda et al. 2002, 2007c). Numerous reports (Lebeda and Zinkernagel 2003b; Beharav et al. 2006; Petrželová et al. 2007; Lebeda et al. 2008a) have demonstrated that wild Lactuca germplasm, especially of L. saligna and L. serriola, has enormous potential. More intensive exploitation of these new sources of resistance is primarily based on the increasing number of wild, characterized Lactuca accessions from various ecogeographical areas (Lebeda et al. 2004a, b, 2007c, 2008a, b, c). Lactuca saligna is currently considered to be the most important source of highly efficient resistance, which is expected to be nonhost specific (Lebeda et al. 2002; Lebeda and Zinkernagel 2003b; Petrželová et al. 2007). However, our understanding of both the mechanism (Lebeda and Reinink 1994; Lebeda and Pink 1998; Lebeda et al. 2001b, c, 2002, 2006, 2008b; Sedlářová et al. 2007a) and genetics (Jeuken and Lindhout 2002, 2004; Jeuken et al. 2001, 2008; Zhang 2008) of this resistance is still incomplete.
Conclusions and future prospects
Despite enormous progress in research on wild Lactuca germplasm, this review has demonstrated many important gaps in our understanding. The following list of topics can be considered as key challenges for future research and exploitation in lettuce breeding:
-
1.
Complex taxonomic and phylogenetic relationships within the genus;
-
2.
Detailed floristic, biogeographic and ecologic delimitation of the distributions of known Lactuca spp.;
-
3.
Clarification of the structure of Lactuca gene pools;
-
4.
Reconsideration of germplasm collection structure from the viewpoint of diversity, quality, and quantity;
-
5.
Collecting and exploration missions, especially to areas of high species richness and diversity (e.g., South Africa and Asia);
-
6.
Enlargement of activities focused on complex characterization and evaluation with importance for the management of wild Lactuca genebank collections and their efficient utilization in lettuce breeding;
-
7.
Broad international cooperation among diverse institutions, including Bioversity International.
References
Beharav A, Lewinsohn D, Lebeda A, Nevo E (2006) New wild Lactuca genetic resources with resistance against Bremia lactucae. Genet Resour Crop Evol 53:467–474. doi:10.1007/s10722-004-1932-7
Beharav A, Ben-David R, Doležalová I, Lebeda A (2008a) Eco-geographical distribution of Lactuca saligna natural populations in Israel. Isr J Plant Sci 56: (in press)
Beharav A, Lebeda A, Doležalová I, Ben-David R (2008b) Collecting of genetic diversity in natural populations of wild plant species: case study on Lactuca saligna and L. aculeata in Israel. In: Prohens J, Badenes ML (eds) Modern variety breeding for present and future needs. Editorial Universidad Politécnica de Valencia, Valencia, Spain, pp 75–76 (Abstract)
Bennett MD, Leitch IJ (1995) Nuclear DNA amounts in angiosperms. Ann Bot (Lond) 76:113–176. doi:10.1006/anbo.1995.1085
Bohm BA, Stuessy TF (2001) Flavonoids of the sunflower family (Asteraceae). Springer, Wien and New York
Bonnier FJM, Reinink K, Groenwold R (1992) New sources of major gene resistance in Lactuca to Bremia lactucae. Euphytica 61:203–211. doi:10.1007/BF00039659
Bos L, Huijberts N (1990) Screening for resistance to big-vein disease of lettuce (Lactuca sativa). Crop Prot 9:446–452. doi:10.1016/0261-2194(90)90135-T
Boukema IW, Hazekamp T, van Hintum TJL (1990) The CGN collections reviews, the CGN lettuce collection. Centre for Genetic Resources, Wageningen
Chen YH, Chen HY, Hsu CL, Yen GC (2007) Induction of apoptosis by the Lactuca indica L. in human leukemia cell line and its active components. J Agric Food Chem 55:1743–1749. doi:10.1021/jf063118t
Cole RA, Sutherland RA, Riggall WE (1991) The use of polyacrylamide gradient gel electrophoresis to identify variation in isozymes as markers for Lactuca species and resistance to the lettuce root aphid Pemphigus bursarius. Euphytica 56:237–242. doi:10.1007/BF00042370
Crute IR (1990) Resistance to Bremia lactucae (downy mildew) in British populations of Lactuca serriola (prickly lettuce). In: Burdon JJ, Leather SR (eds) Pests, pathogens and plant communities. Blackwell Scientific Publications, Oxford, pp 203–217
Crute IR (1992a) Downy mildew of lettuce. In: Chaube HS, Kumar J, Mukhopadhyay AN, Singh US (eds) Plant diseases of international importance, vol II. Diseases of vegetable and oil seed crops. Prentice Hall, Englewood Cliffs, pp 165–185
Crute IR (1992b) From breeding to cloning (and back again?): a case study with lettuce downy mildew. Annu Rev Phytopathol 30:485–506. doi:10.1146/annurev.py.30.090192.002413
Crute IR (1992c) The role of resistance breeding in the integrated control of downy mildew (Bremia lactucae) in protected lettuce. Euphytica 63:95–102. doi:10.1007/BF00023915
Doležalová I, Lebeda A, Křístková E (2001) Prickly lettuce (Lactuca serriola L.) germplasm collecting and distribution study in Slovenia and Sweden. Plant Genet Resour Newsl 128:41–44
Doležalová I, Křístková E, Lebeda A, Vinter V (2002a) Description of morphological characters of wild Lactuca L. spp. genetic resources (English-Czech version). Hortic Sci Prague 29:56–83
Doležalová I, Lebeda A, Janeček J, Číhalíková J, Křístková E, Vránová O (2002b) Variation in chromosome numbers and nuclear DNA contents in genetic resources of Lactuca L. species (Asteraceae). Genet Resour Crop Evol 49:383–395
Doležalová I, Křístková E, Lebeda A, Vinter V, Astley D, Boukema IW (2003a) Basic morphological descriptors for genetic resources of wild Lactuca spp. Plant Genet Resour Newsl 134:1–9
Doležalová I, Lebeda A, Dziechciarková M, Křístková E, Astley D, van de Wiel CCM (2003b) Relationships among morphological characters, isozymes polymorphism and DNA variability—the impact on Lactuca germplasm taxonomy. Czech J Genet Plant Breed 39:59–67
Doležalová I, Lebeda A, Tiefenbachová I, Křístková E (2004) Taxonomic reconsideration of some Lactuca spp. germplasm maintained in world genebank collections. Acta Hortic 634:193–201
Doležalová I, Lebeda A, Křístková E, Novotná A (2005) Morphological variation of Lactuca serriola populations from some European countries. In: Abstracts, XVII International Botanical Congress, Vienna, 17–23 July 2005, p 548
Doležalová I, Lebeda A, Křístková E, Novotná A (2007a) Relevance of morphologic assessment of wild Lactuca spp. germplasm for their taxonomic determination. Bulletin of Botanical Gardens, Museums & Collections, Polish Botanical Society. Botanical Garden—Center for Biological Diversity Conservation of the Polish Academy of Science, Warsaw, 16A: 22
Doležalová I, Lebeda A, Novotná A, Kršková M (2007b) Variation in fruit morphology of Lactuca serriola populations from Czech Republic, Germany, Netherlands and United Kingdom. In: Eucarpia Leafy Vegetables 2007 Conference Abstracts, Warwick HRI, Wellesbourne, 18–20 April 2007, p 8
Doležalová I, Lebeda A, Vondráková D (2008) Current status of the Lactuca working collection of Palacký University in Olomouc, Czech Republic. In: Maggioni L, Lebeda A, Boukema I, Lipman E (eds) Report of a working group on leafy vegetables. First meeting, 13–14 October 2005, Olomouc, Czech Republic. Bioversity International, Rome, pp 27–33
Dziechciarková M, Lebeda A, Doležalová I, Astley D (2004a) Characterization of Lactuca spp. germplasm by protein and molecular markers—a review. Plant Soil Environ 50:47–58
Dziechciarková M, Lebeda A, Doležalová I, Křístková E (2004b) Isozyme variation in European Lactuca serriola germplasm. In: Vollmann J, Grausgruber H, Ruckenbauer P (eds) Genetic variation for plant breeding. Proceedings of the 17th Eucarpia general congress, 8–11 September 2004, Tulln, Austria. BOKU-University of Natural Resources and Applied Life Sciences, Vienna, pp 103–107
Feráková V (1977) The genus Lactuca L. in Europe. Univerzita Komenského, Bratislava
Funk VA, Bayer RJ, Keeley S, Chan R, Watson L, Gemeinholzer B, Schilling E, Panero JL, Baldwin BG, Garcia-Jacas N, Susanna A, Jansen RK (2005) Everywhere but Antarctica: using a supertree to understand the diversity and distribution of the Compositae. Biol Skr 55:343–374
Gass T, Frese L (eds) (1999) Implementation of the global plan of action in Europe—conservation and sustainable utilization of plant genetic resources for food and agriculture. Proc of the European Symp, 1998, Braunschweig (Germany). Int Plant Gen Res Ins (IPGRI), Rome, Italy
Grube RC, Hayes R, Mou B, McCreight JD (2005a) Lettuce breeding. California lettuce research board annual report, 2004–2005. California Lettuce Research Board, Salinas
Grube RC, Wintermantel WM, Hand P, Aburomia R, Pink DAC, Ryder EJ (2005b) Genetic analysis and mapping of resistance to lettuce dieback: a soilborne disease caused by tombusviruses. Theor Appl Genet 110:259–268. doi:10.1007/s00122-004-1825-3
Guarino L, Rao RV, Reid R (eds) (1995) Collecting plant genetic diversity. Technical guidelines. CAB International, Wallingford
Gustafsson I (1989) Potential sources of resistance to lettuce downy mildew (Bremia lactucae) in different Lactuca species. Euphytica 40:227–232
Hayes RJ, Ryder EJ (2007) Introgression of novel alleles for partial resistance to big vein disease from Lactuca virosa into cultivated lettuce. HortScience 42:35–39
Hayes RJ, Ryder E, Robinson B (2004) Introgression of big vein tolerance from Lactuca virosa L. into cultivated lettuce (Lactuca sativa L.). HortScience 39:881
Hayes RJ, Ryder EJ, Wintermantel WM (2008) Genetic variation for big-vein symptom expression and resistance to Mirafiori lettuce big-vein virus in Lactuca virosa L., a wild relative of cultivated lettuce. Euphytica 164:493–500. doi:10.1007/s10681-008-9738-x
Hill M, Witsenboer H, Zabeau M, Vos P, Kesseli R, Michelmore R (1996) PCR-based fingerprinting using AFLPs as a tool for studying genetic relationships in Lactuca spp. Theor Appl Genet 93:1202–1210. doi:10.1007/BF00223451
Hooftman DAP, Nieuwenhuis BPS, Posthuma KI, Oostermeijer JGB, den Nijs HJCM (2007) Introgression potential of downy mildew resistance from lettuce to Lactuca serriola and its relevance for plant fitness. Basic Appl Ecol 8:135–146. doi:10.1016/j.baae.2006.03.008
Hu J, Ochoa OE, Truco MJ, Vick BA (2005) Application of the TRAP technique to lettuce (Lactuca sativa L.) genotyping. Euphytica 144:225–235. doi:10.1007/s10681-005-6431-1
Jansen J, Verbakel H, Peleman J, van Hintum TJL (2006) A note on the measurement of genetic diversity within genebank accessions of lettuce (Lactuca sativa L.) using AFLP markers. Theor Appl Genet 112:554–561. doi:10.1007/s00122-005-0162-5
Jeuken M, Lindhout P (2002) Lactuca saligna, a non-host for lettuce downy mildew (Bremia lactucae), harbors a new race-specific Dm gene and three QTLs for resistance. Theor Appl Genet 105:384–391. doi:10.1007/s00122-002-0943-z
Jeuken MJW, Lindhout P (2004) The development of lettuce backcross inbred lines (BILs) for exploitation of the Lactuca saligna (wild lettuce) germplasm. Theor Appl Genet 109:394–401. doi:10.1007/s00122-004-1643-7
Jeuken M, van Wijk R, Peleman J, Lindhout P (2001) An integrated interspecific AFLP map of lettuce (Lactuca) based on two L. sativa × L. saligna F-2 populations. Theor Appl Genet 103:638–647. doi:10.1007/s001220100657
Jeuken MJW, Pelgrom K, Stam P, Lindhout P (2008) Efficient QTL detection for nonhost resistence in wild lettuce: backcross inbred lines versus F2 population. Theor Appl Genet 116:845–857. doi:10.1007/s00122-008-0718-2
Kesseli RV, Michelmore RW (1986) Genetic variation and phylogenies detected from isozyme markers in species of Lactuca. J Hered 77:324–331
Kesseli RV, Ochoa O, Michelmore RW (1991) Variation at RFLP loci in Lactuca spp. and origin of cultivated lettuce (L. sativa). Genome 34:430–436
Kim KH, Kim YH, Lee KR (2007) Isolation of quinic acid derivatives and flavonoids from the aerial parts of Lactuca indica L. and their hepatoprotective activity in vitro. Bioorg Med Chem Lett 17:6739–6743. doi:10.1016/j.bmcl.2007.10.046
Kisiel W, Barszcz B (1998) A germacrolide glucoside from Lactuca tatarica. Phytochemistry 48:205–206. doi:10.1016/S0031-9422(97)01106-0
Kisiel W, Zielinska K (2000) Sesquiterpenoids and phenolics from Lactuca perennis. Fitoterapia 71:86–87. doi:10.1016/S0367-326X(99)00112-4
Kitner M, Lebeda A, Doležalová I, Maras M, Křístková E, Nevo E, Pavlíček T, Meglic V, Beharav A (2008) AFLP analysis of Lactuca saligna germplasm collections from four European and three Middle Eastern countries. Isr J Plant Sci 56: (in press)
Koopman WJM (1999) Plant systematics as useful tool for plant breeders, examples from lettuce. In: Lebeda A, Křístková E (eds) Eucarpia leafy vegetables ′99. Proceedings of the Eucarpia meeting on leafy vegetables genetics and breeding, Palacký University in Olomouc, Olomouc, pp 95–105
Koopman WJM (2000) Identifying lettuce species (Lactuca subs. Lactuca, Asteraceae). A practical application of flow cytometry. Euphytica 116:151–159. doi:10.1023/A:1004086503349
Koopman WJM (2002) Zooming in on the lettuce genome: species relationships in Lactuca s.l. inferred from chromosomal and molecular characters. Ph.D. dissertation, Wageningen University, Wageningen
Koopman WJM, De Jong HJ (1996) A numerical analysis of karyotypes and DNA amounts in lettuce cultivars and species (Lactuca subsp. Lactuca, Compositae). Acta Bot Neerl 45:211–222
Koopman WJM, Guetta E, Van de Wiel CCM, Vosman B, Van den Berg RG (1998) Phylogenetic relationships among Lactuca (Asteraceae) species and related genera based on ITS-1 DNA sequences. Am J Bot 85:1517–1530. doi:10.2307/2446479
Koopman WJM, Zevenbergen MJ, van den Berg RG (2001) Species relationships in Lactuca s.l. (Lactuceae, Asteraceae) inferred from AFLP fingerprints. Am J Bot 88:1881–1887. doi:10.2307/3558364
Křístková E, Lebeda A (1999) Collection of Lactuca spp. genetic resources in the Czech Republic. In: Lebeda A, Křístková E (eds) Eucarpia leafy vegetables ′99. Proceedings of the Eucarpia meeting on leafy vegetables genetics and breeding, Palacký University in Olomouc, Olomouc, pp 109–116
Křístková E, Lebeda A, Doležalová I (2001) Collecting and evaluating of Lactuca serriola germplasm in Europe. In: Swiecicki W, Naganowska B, Wolko B (eds) Broad variation and precise characterization—limitation for the future. Eucarpia, Section Genetic Resources, Poznań, pp 49–52
Křístková E, Lebeda A, Doležalová I (2007a) Phenotypic variability of Lactuca saligna germplasm collected in Italy and France. In: Eucarpia leafy vegetables 2007, Conference Abstracts, University of Warwick, Warwick HRI, UK, Poster Presentations, 18–20 April 2007, p 15
Křístková E, Lebeda A, Doležalová I, Vinter V, Křístková A (2007b) Variation in developmental stages of Lactuca serriola L. (prickly lettuce) germplasm from different European countries. In: Eucarpia leafy vegetables 2007, Conference Abstracts, University of Warwick, Warwick HRI, UK, Poster Presentations, 18–20 April 2007, p. 16
Křístková E, Doležalová I, Lebeda A, Vinter V, Novotná A (2008) Description of morphological characters of lettuce (Lactuca sativa L.) genetic resources. Hortic Sci Prague 38:113–129
Kuang H, Ochoa OE, Nevo E, Michelmore RW (2006) The disease resistance gene Dm3 is infrequent in natural populations of Lactuca serriola due to deletions and frequent gene conversions at the RGC2locus. Plant J 47:38–48. doi:10.1111/j.1365-313X.2006.02755.x
Kuang H, van Eck HJ, Sicard D, Michelmore R, Nevo E (2008) Evolution and genetic population structure of prickly lettuce (Lactuca serriola) and its RGC2 resistance gene cluster. Genetics 178:1547–1558. doi:10.1534/genetics.107.080796
Lebeda A (1985a) Occurrence of natural infection of powdery mildew (Erysiphe cichoracearum) by the genus Lactuca in Czechoslovakia. Acta Phytopath Acad Sci Hung 20:149–162
Lebeda A (1985b) Differences in resistance of wild Lactuca species to natural infection of lettuce powdery mildew (Erysiphe cichoracearum). Euphytica 34:521–523. doi:10.1007/BF00022949
Lebeda A (1986) Specificity of interactions between wild Lactuca spp. and Bremia lactucae isolates from Lactuca serriola. J Phytopathol 117:54–64. doi:10.1111/j.1439-0434.1986.tb04360.x
Lebeda A (1989) Response of lettuce cultivars carrying the resistance gene Dm11 to isolates of Bremia lactucae from Lactuca serriola. Plant Breed 102:311–316. doi:10.1111/j.1439-0523.1989.tb01261.x
Lebeda A (1990) The location of sources of field resistance to Bremia lactucae in wild Lactuca species. Plant Breed 105:75–77. doi:10.1111/j.1439-0523.1990.tb00455.x
Lebeda A (1994) Evaluation of wild Lactuca species for resistance of natural infection of powdery mildew (Erysiphe cichoracearum). Genet Resour Crop Evol 41:55–57. doi:10.1007/BF00051424
Lebeda A (1999) Powdery mildew on lettuce and wild Lactuca species. The First International Powdery Mildew Conference, August 29–September 2, 1999, Avignon (France); Abstracts, pp 16–17
Lebeda A (2002) Occurrence and variation in virulance of Bremia lactucae in natural populations of Lactuca serriola. In: Spencer-Phillips PTN, Gisi U, Lebeda A (eds) Advances in downy mildew research. Kluwer, Dordrecht, pp 179–183
Lebeda A, Astley D (1999) World genetic resources of Lactuca spp., their taxonomy and biodiversity. In: Lebeda A, Křístková E (eds) Eucarpia leafy vegetables ′99. Proceedings of the Eucarpia meeting on leafy vegetables genetics and breeding. Palacký University in Olomouc, Olomouc, pp 81–94
Lebeda A, Boukema IW (1991) Further investigation of the specificity of interactions between wild Lactuca spp. and Bremia lactucae isolates from Lactuca serriola. J Phytopathol 133:57–64. doi:10.1111/j.1439-0434.1991.tb00137.x
Lebeda A, Boukema IW (2001) Leafy vegetables genetic resources. In: Maggioni L, Spellman O (eds) Report of a network coordinating group on vegetables; ad hoc meeting, 26–27 May 2000, Vila Real, Portugal. International Plant Genetic Resources Institute, Rome, pp 48–57
Lebeda A, Boukema IW (2005) Ad hoc meeting on leafy vegetables. In: Thomas G, Astley D, Boukema IW, Daunay MC, Del Greco A, Diez MJ, van Dooijweert W, Keller J, Kotlińska T, Lebeda A, Lipman E, Maggioni L, Rosa E (eds) Report of a vegetable network. Joint Meeting with an ad hoc group of leafy vegetables, 22–24 May 2003, Skierniewice, Poland. International Plant Genetic Resources Institute, Rome, pp 82–94
Lebeda A, Buczkowski J (1986) Occurrence of Erysiphe cichoracearum perithecia on wild Lactuca species. J Phytopathol 115:21–28. doi:10.1111/j.1439-0434.1986.tb00857.x
Lebeda A, Jendrůlek T (1989) Application of multivariate analysis for characterizing the relationships between wild Lactuca spp. and Bremia lactucae. Acta Phytopathol Entomol Hung 24:317–331
Lebeda A, Mieslerová B (2003) Lettuce powdery mildew—an unknown disease of lettuce. In: van Hintum ThJL, Lebeda A, Pink DA, Schut JW (eds) Eucarpia leafy vegetables 2003. Proceedings of the Eucarpia meeting on leafy vegetables genetics and breeding, Noordwijkerhout, Centre for Genetics Resources, Wageningen, p 164
Lebeda A, Petrželová I (2001) Occurrence and characterization of race-specific resistance to Bremia lactucae in wild Lactuca spp. In: Swiecicki W, Naganowska B, Wolko B (eds) Broad variation and precise characterization—limitation for the future. Eucarpia, Section Genetic Resources, Prodruk, Poznan, pp 232–233
Lebeda A, Petrželová I (2004a) Occurrence of race-specific resistance to Bremia lactucae in Lactuca serriola germplasm originating from four European countries. In: Vollmann J, Grausgruber H, Ruckenbauer P (eds) Genetic variation for plant breeding. Proceedings of the 17th EUCARPIA General Congress, 8–11 September 2004, Tulln, Austria. BOKU-University of Natural Resources and Applied Life Sciences, Vienna, pp 113–116
Lebeda A, Petrželová I (2004b) Variation and distribution of virulence phenotypes of Bremia lactucae in natural populations of Lactuca serriola. Plant Pathol 53:316–324. doi:10.1111/j.0032-0862.2004.01003.x
Lebeda A, Petrželová I (2005) Comparison of resistance to Bremia lactucae in populations of Lactuca serriola occurring in Central Europe (Czech Republic) and the British Isles (England, UK). In: Bullitta S (ed) Plant genetic resources of geographical and “other” islands (Conservation, evaluation and use for plant breeding). Book of Abstracts. XVII Eucarpia Genetic Resources Section Meeting, 30 March–2 April 2005, CNR-ISPAAM, sezione Sassari Publisher, Sassari, p 14
Lebeda A, Petrželová I (2007) Race-specific resistance to Bremia lactucae in European populations of Lactuca serriola. In: Eucarpia leafy vegetables 2007. Conference Abstracts, University of Warwick, Warwick HRI, UK, Oral Presentations, 18–20 April 2007, p 11
Lebeda A, Pink DAC (1998) Histological aspects of the response of wild Lactuca spp. and their hybrids, with L. sativa to lettuce downy mildew (Bremia lactucae). Plant Pathol 47:723–736
Lebeda A, Reinink K (1994) Histological characterization of resistance in Lactuca saligna to lettuce downy mildew (Bremia lactucae). Physiol Mol Plant Pathol 44:125–139. doi:10.1016/S0885-5765(05)80106-7
Lebeda A, Zinkernagel V (2003a) Evolution and distribution of virulence in the German population of Bremia lactucae. Plant Pathol 52:41–51. doi:10.1046/j.1365-3059.2003.00802.x
Lebeda A, Zinkernagel V (2003b) Characterization of new highly virulent German isolates of Bremia lactucae and efficiency of resistance in wild Lactuca spp. germplasm. J Phytopathol 151:274–282
Lebeda A, Doležalová I, Křístková E, Janeček J, Vinter V, Vránová O, Doležal K, Tarkowski P, Petrželová P, Trávníček B, Novotný R, Janeček J (1999) Complex research of taxonomy and ecobiology of wild Lactuca spp. genetic resources. In: Lebeda A, Křístková E (eds) Eucarpia leafy vegetables ′99. Proceedings of the Eucarpia meeting on leafy vegetables genetics and breeding, Palacký University in Olomouc, Olomouc, pp 117–131
Lebeda A, Doležalová I, Křístková E, Janeček J, Vinter V, Vránová O, Doležal K, Tarkowski P, Petrželová P, Trávníček B, Novotný R (2001a) Biodiversity of genetic resources of wild Lactuca spp. In: Święcicki W, Naganowska B, Wolko B (eds) Broad variation and precise characterization—limitation for the future. Prodruk, Poznań, pp 53–56
Lebeda A, Doležalová I, Křístková E, Mieslerová B (2001b) Biodiversity and ecogeography of wild Lactuca spp. in some European countries. Genet Resour Crop Evol 48:153–164. doi:10.1023/A:1011265614395
Lebeda A, Pink DAC, Mieslerová B (2001c) Host-parasite specificity and defense variability in the Lactuca spp.—Bremia lactucae pathosystem. J Plant Pathol 83:25–35
Lebeda A, Pink DAC, Astley D (2002) Aspects of the interactions between wild Lactuca spp. and related genera and lettuce downy mildew (Bremia lactucae). In: Spencer-Phillips PTN, Gisi U, Lebeda A (eds) Advances in downy mildew research. Kluwer, Dordrecht, pp 85–117
Lebeda A, Doležalová I, Astley D (2004a) Representation of wild Lactuca spp. (Asteraceae, Lactuceae) in world genebank collections. Genet Resour Crop Evol 51:167–174. doi:10.1023/B:GRES.0000020860.66075.f7
Lebeda A, Doležalová I, Feráková V, Astley D (2004b) Geographical distribution of wild Lactuca spp. (Asteraceae, Lactuceae). Bot Rev 70:328–356. doi:10.1663/0006-8101(2004)070[0328:GDOWLS]2.0.CO;2
Lebeda A, Doležalová I, Janeček J, Gasmanová N (2004c) Differences in relative DNA content of Lactuca serriola germplasm collected in Europe. In: Summaries and Program, 17th International lettuce and leafy vegetable conference, 28–31 August 2004, Sandman Hotel, Montreal-Longueuil. Agriculture and Agri-Food Canada, Montreal, pp 29–30
Lebeda A, Sedlářová M, Lynn J, Pink DAC (2006) Phenotypic and histological expression of different genetic backgrounds in interactions between lettuce, wild Lactuca spp., L. sativa × L. serriola hybrids and Bremia lactucae. Eur J Plant Pathol 115:431–441. doi:10.1007/s10658-006-9034-3
Lebeda A, Doležalová I, Křístková E, Dehmer KJ, Astley D, Van de Wiel CCM, Van Treuren R (2007a) Acquisition and ecological characterization of Lactuca serriola L. germplasm collected in the Czech Republic, Germany, the Netherlands and United Kingdom. Genet Resour Crop Evol 54:555–562. doi:10.1007/s10722-006-0012-6
Lebeda A, Doležalová I, Křístková E, Novotná A (2007b) Comparative study of variation of some morphological characteristics of Lactuca serriola germplasm collected in Central Europe (Czech Republic) and the British Isles (England, UK). (Abstract). In: Bullitta S (ed) Plant genetic resources of geographical and “other” islands (conservation, evaluation and use for plant breeding). Proceedings of the XVII Eucarpia genetic resources section meeting; 30 March–2 April 2005, Castelsardo, Italy. CNR-ISPAAM, sezione Sassari Publisher, Sassari, pp 95–96
Lebeda A, Ryder EJ, Grube R, Doležalová I, Křístková E (2007c) Lettuce (Asteraceae; Lactuca spp.), Chapter 9. In: Singh R (ed) Genetic resources, chromosome engineering, and crop improvement series, vol 3—vegetable crops. CRC Press, Boca Raton, pp 377–472
Lebeda A, Doležalová I, Křístková E, Novotná A (2007d) Taxonomic determination of plant genetic resources—impact and consequences: case study of Lactuca spp. In: Hauptvogel P, Benediková D, Hauptvogel R (eds) Plant genetic resources and their exploitation in the plant breeding for food and agriculture. Book of Abstracts. 18th EUCARPIA genetic resources section meeting, May 23–26, 2007, Piešťany, Slovak Republic. PNprint, s.r.o., Piešťany, Slovak Republic, pp 39–40
Lebeda A, Doležalová I, Křístková E, Mieslerová B, Kitner M, Navrátilová B, Duchoslav M, Havránek P, Vondráková D (2007e) Germplasm collections of crop wild relatives—research, study and use on the Department of Botany, Palacký University in Olomouc (Czech Republic). In: Hauptvogel P, Benediková D, Hauptvogel R (eds) Plant genetic resources and their exploitation in the plant breeding for food and agriculture. Book of Abstracts. 18th EUCARPIA genetic resources section meeting, May 23–26, 2007, Piešťany, Slovak Republic. PNprint, s.r.o., Piešťany, Slovak Republic, pp 94–95
Lebeda A, Petrželová I, Maryška Z (2008a) Structure and variation in the wild-plant pathosystem: Lactuca serriola–Bremia lactucae. Eur J Plant Pathol 122:127–146. doi:10.1007/s10658-008-9291-4
Lebeda A, Sedlářová M, Petřivalský M, Prokopová J (2008b) Diversity of defence mechanisms in plant-oomycete interactions: a case study of Lactuca spp. and Bremia lactucae. Eur J Plant Pathol 122:71–89. doi:10.1007/s10658-008-9292-3
Lebeda A, Doležalová I, Dziechciarková M, Kitner M, Křístková E, Lindhout P (2008c) Genetic polymorphism of European populations of Lactuca serriola. Genet Resour Crop Evol (submitted)
Lebeda A, Doležalová I, Křístková E, Kitner M, Petrželová I, Mieslerová B, Novotná A (2008d) Wild Lactuca germplasm for lettuce breeding: recent status, gaps and challenges. In: Prohens J, Badenes MJ (eds) Modern variety breeding for present and future needs. Proceedings of the 18th EUCARPIA General Congress, September 9–12, 2008, Editorial Universidad Politécnica de Valencia, Valencia, pp 49–60
Maggioni L, Lebeda A, Boukema I, Lipman E (eds) (2008) Report of a working group on leafy vegetables. First meeting, 13–14 October 2005, Olomouc, Czech Republic. Bioversity International, Rome
Maisonneuve B (2003) Lactuca virosa, a source of disease resistance genes for lettuce breeding: results and difficulties for gene introgression. In: van Hintum TJL, Lebeda A, Pink DAC, Schut JW (eds) Eucarpia leafy vegetables ‘03. CGN Wageningen, The Netherlands, pp 31–35
Maisonneuve B, Chovelon V, Lot H (1991) Inheritance of resistance to beet western yellows virus in Lactuca virosa L. HortScience 26:1543–1545
Maisonneuve B, Chupeau MC, Bellec Y, Chupeau Y (1995) Sexual and somatic hybridization in the genus Lactuca. Euphytica 85:281–285. doi:10.1007/BF00023957
Maisonneuve B, Bellec Y, Souche S, Lot H (1999) New resistance against downy mildew and lettuce mosaic potyvirus in wild Lactuca spp. In: Lebeda A, Křístková E (eds) Eucarpia leafy vegetables ′99. Proceedings of the Eucarpia meeting of leafy vegetables genetics and breeding, Palacký University in Olomouc, Olomouc, pp 191–197
Matoba H, Mizutani T, Nagano K, Hoshi Y, Uchiyama H (2007) Chromosomal study of lettuce and its allied species (Lactuca spp., Asteraceae) by means of karyotype analysis and fluorescence in situ hybridization. Hereditas 144:235–243. doi:10.1111/j.2007.0018-0661.02012x
McCreight JD (1987) Resistance in wild lettuce to lettuce infectious yellows. HortScience 22:640–642
McGregor RL, Barkley TM, Brooks RE, Schofield EK (eds) (1986) Flora of the Great Plains. University Press of Kansas, Lawrence
McGuire PE, Ryder EJ, Michelmore RW, Clark RL, Antle R, Emery G, Hannan RW, Kesseli RV, Kurtz EA, Ochoa O, Rubatzky VE, Waycott W (1993) Genetic resources of lettuce and Lactuca species in California. An assessment of the USDA and UC collections and recommendations for long-term security. Report no. 12. University of California, Genetic Resources Conservation Program, Davis
Michalska K, Stojakowska A, Malarz J, Doležalová I, Lebeda A, Kisiel W (2008) Systematic implications of sesquiterpene lactones in Lactuca species. Biochem Syst Ecol (in press)
Michelmore RW (2002) Genetic variation in lettuce. In: Kurtz E (ed) Annual Report 2001–2002. California Lettuce Research Board, Salinas, pp 77–86
Michelmore RW, Ochoa OE (2005) Breeding crisphead lettuce. California Lettuce Research Board, Annual Report, April 1, 2004 through March 31, 2005. California Research Board, Salinas, pp 68–78
Mieslerová B, Petrželová I, Lebeda A, Česneková E (2007) Occurrence of lettuce downy mildew and powdery mildew in natural populations of prickly lettuce. In: Lebeda A, Spencer-Phillips PTN (eds) Advances in downy mildew research, vol 3. Proceedings of the 2nd international downy mildews symposium, Palacký University in Olomouc and JOLA, v.o.s., Kostelec na Hané, Czech Republic, pp 59–64
Mikel MA (2007) Genealogy of contemporary North American lettuce. HortScience 42:489–493
Mizutani T, Tanaka T (2003) Genetic analyses of isozyme in lettuce, Lactuca sativa, and its relatives. J Jap Soc Hortic Sci 72:122–127
Mou B (2008) Lettuce. In: Prohens J, Nuez F (eds) Handbook of plant breeding. Vegetables I. Asteraceae, Brassicaceae, Chenopodiaceae, and Cucurbitaceae. Springer Science, New York, pp 75–116
Mou BQ, Bull C (2004) Screening lettuce germplasm for new sources of resistance to corky root. J Am Soc Hortic Sci 129:712–716
Netzer D, Globerson D, Sacks J (1976) Lactuca saligna L. a new sources of resistance to downy mildew (Bremia lactucae Reg). Hortic Sci 11:612–613
Netzer D, Globerson D, Weintal C, Elyassi R (1985) Sources and inheritance of resistance to Stemphylium leaf spot of lettuce. Euphytica 34:393–396. doi:10.1007/BF00022934
Norwood JM, Crute IR, Lebeda A (1981) The location and characteristics of novel sources of resistance to Bremia lactucae Regel (downy mildew) in wild Lactuca L. species. Euphytica 30:659–668. doi:10.1007/BF00038794
Pandey A, Tomer AK, Bhandari DC, Pareek SK (2008) Towards collection of wild relatives of crop plants in India. Genet Resour Crop Evol 55:187–202. doi:10.1007/s10722-007-9227-4
Petrželová I, Lebeda A (2003) Distribution of compatibility types and occurrence of sexual reproduction in natural populations of Bremia lactucae on wild Lactuca serriola plants. Acta Phytopathol Entomol Hung 38:43–52. doi:10.1556/APhyt.38.2003.1-2.6
Petrželová I, Lebeda A (2004a) Comparison of virulence of Bremia lactucae isolates originating from Lactuca sativa and Lactuca serriola. Acta fytotech zootech 7:248–250
Petrželová I, Lebeda A (2004b) Occurrence of Bremia lactucae in natural populations of Lactuca serriola. J Phytopathol 152:391–398. doi:10.1111/j.1439-0434.2004.00859.x
Petrželová I, Lebeda A (2004c) Temporal and spatial variation in virulence of natural populations of Bremia lactucae occurring on Lactuca serriola. In: Spencer-Phillips PTN, Jeger M (eds) Advances in downy mildew research, vol 2. Kluwer, Dordrecht, pp 141–163
Petrželová I, Lebeda A, Nevo E, Beharav A (2007) Variation of response against Bremia lactucae in natural populations of Lactuca saligna. In: Lebeda A, Spencer-Phillips PTN (eds) Advances in downy mildew research, vol 3. Proceedings of the 2nd international downy mildews symposium, Palacký University in Olomouc and JOLA, v.o.s., Kostelec na Hané, Czech Republic, pp 169–173
Provvidenti R, Robinson RW, Shail W (1980) A source of resistance to a strain of cucumber mosaic virus in Lactuca saligna L. HortScience 15:528–529
Rajicic TS, Dehmer KJ (2008) Analysis of wild Lactuca gene bank accessions and implications for wild species conservation. In: Maxted N, Ford-Lloyds BV, Kell SP, Iriondo JM, Dulloo ME, Turok J (eds) Crop wild relative conservation and use. CABI International, Wallingford, pp 429–436
Rees S, Harborne J (1984) Flavonoids and other phenolics of Cichorium and related members of the Lactuceae (Compositae). Bot J Linn Soc 89:313–319. doi:10.1111/j.1095-8339.1984.tb02563.x
Reuveni R, Shimoni M, Crute IR (1991) An association between high peroxidase activity in lettuce (Lactuca sativa) and field resistance to downy mildew (Bremia lactucae). J Phytopathol 132:312–318. doi:10.1111/j.1439-0434.1991.tb00126.x
Ryder EJ (2002) A mild systemic reaction to lettuce mosaic virus in lettuce (Lactuca sativa), inheritance and interaction with an allele for resistance. J Am Soc Hortic Sci 127:814–818
Sedlářová M, Lebeda A (2001) Histochemical detection and role of phenolic compounds in defence response of Lactuca spp. to lettuce downy mildew (Bremia lactucae). J Phytopathol 149:1–5. doi:10.1046/j.1439-0434.2001.00698.x
Sedlářová M, Lebeda A, Pink DAC (2001) The early stages of interaction between effective and non-effective race-specific genes in Lactuca sativa, wild Lactuca spp. and Bremia lactucae (race NL16). J Plant Dis Prot 108:477–489
Sedlářová M, Luhová L, Petřivalský M, Lebeda A (2007a) Localisation and metabolism of reactive oxygen species during Bremia lactucae pathogenesis in Lactuca sativa and wild Lactuca spp. Plant Physiol Biochem 45:607–616. doi:10.1016/j.plaphy.2007.05.010
Sedlářová M, Výtisková M, Doležal K, Lebeda A (2007b) Pre-incubation with cytokinins delays chlorophyll degradation in Lactuca spp. tissues and reduces Bremia lactucae sporulation. In: Lebeda A, Spencer-Phillips PTN (eds) Advances in downy mildew research, vol 3. Proceedings of the 2nd international downy mildews symposium, Palacký University in Olomouc and JOLA, v.o.s., Kostelec na Hané, Czech Republic, pp 185–194
Sicard D, Woo SS, Arroyo-Garcia R, Ochoa O, Nguyen D, Korol A, Nevo E, Michelmore RW (1999) Molecular diversity at the major cluster of disease resistance genes in cultivated and wild Lactuca spp. Theor Appl Genet 99:405–418. doi:10.1007/s001220051251
Spooner D, van Treuren R, de Vicente MC (2005) Molecular markers for genebank management. IPGRI Technical Bulletin 10, International Plant Genetic Resources Institute, Rome
van de Wiel CCM, Arens P, Vosman B (1998) Microsatellite fingerprinting in lettuce (Lactuca sativa L.) and wild relatives. Plant Cell Rep 17:837–842. doi:10.1007/s002990050494
van de Wiel CCM, Arens P, Vosman B (1999) Microsatellite retrieval in lettuce (Lactuca sativa L.). Genome 42:139–149. doi:10.1139/gen-42-1-139
van de Wiel CCM, Flavell A, Syed N, Antonise R, van der Voort JR, van der Linden G (2004) Analysis of gene flow in the lettuce crop-weed complex. In: den Nijs HCM, Bartsch D, Sweet J (eds) Introgression from genetically modified plants into wild relatives. CABI Publishing, Wallingford, pp 163–171
van Hintum TJL, Boukema IW (1999) Genetic resources of leafy vegetables. In: Lebeda A, Křístková E (eds) Eucarpia leafy vegetables ′99. Proceedings of the Eucarpia meeting on leafy vegetables genetics and breeding, Palacký University in Olomouc, Olomouc, pp 59–72
Vermeulen A, Desprez B, Lancelin D, Bannerot H (1994) Relationship among Cichorium species and related genera as determined by analysis of mitochondrial RFLPs. Theor Appl Genet 88:159–166. doi:10.1007/BF00225892
Walkey DGA, Pink DCA (1990) Studies on resistance to beet western yellows virus in lettuce (Lactuca sativa) and the occurrence of field sources of the virus. Plant Pathol 39:141–155. doi:10.1111/j.1365-3059.1990.tb02485.x
Wang M, Cho JJ, Provvidenti R, Hu JS (1992) Identification of resistance to tomato spotted wilt virus in lettuce. Plant Dis 76:642
Welch JE, Zink FW, Grogan RG (1965) Calmar. Calif Agric 19:3–4
Witsenboer H, Vogel J, Michelmore RW (1997) Identification, genetic localization, and allelic diversity of selectively amplified microsatellite polymorphic loci in lettuce and wild relatives (Lactuca spp.). Genome 40:923–936. doi:10.1139/g97-119
Zhang N (2008) Genetic dissection of nonhost resistence of wild lettuce, Lactuca saligna, to downy mildew. PhD Thesis, Wageningen University, Wageningen
Acknowledgments
Critical reading and valuable remarks by Dr. M. P. Widrlechner (USDA-ARS, Iowa State University, North Central Regional Plant Introduction Station, Ames, Iowa, USA) are gratefully acknowledged. The research was supported by grant MSM 6198959215 (Ministry of Education, Youth and Sports of the Czech Republic).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lebeda, A., Doležalová, I., Křístková, E. et al. Wild Lactuca germplasm for lettuce breeding: current status, gaps and challenges. Euphytica 170, 15–34 (2009). https://doi.org/10.1007/s10681-009-9914-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-009-9914-7