Abstract
Lettuce (Lactuca spp.) is an annual and self-pollinating crop that belongs to the Asteraceae (Compositae) family. It is one of the most globally essential commercial vegetable crops, used in salads and sandwiches. The lettuce leaves are used to make a cigarette without nicotine. Seeds and stems contain edible oil and dried latex. Gene banks have conserved a large pool of lettuce's genetic resources, including wild Lactuca species with the same chromosome numbers 2n = 2x = 18. Lactuca species vary greatly in terms of geographical distribution and morpho-agronomic characteristics. By crossing commercial varieties with locally adapted varieties, novel alleles can be introduced, increasing genetic diversity and making preselection for desirable traits easier. Lettuce breeders and geneticists' main objectives are to improve lettuce for various desirable traits, including tolerance to abiotic and biotic stress and high yield. These targets accomplished with modern genomics tools together with traditional breeding methods. This chapter discusses lettuce conservation and biodiversity, stages of lettuce breeding, agriculture practices, and conventional breeding techniques and their restrictions. It also includes modern plant breeding tools and marker-assisted breeding, editing of genome, and genetic engineering.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lactuca sativa L. (lettuce) is a fresh leafy crop commonly consumed in salad mixtures and sandwiches. It is grown in many climatic and soil conditions in the Mediterranean region and extend to Europe (Lebeda et al. 2007; Rubatzky and Yamaguchi 1997). Middle East (eastern Mediterranean basin) was the center of origin of lettuce as mentioned by Ryder (1986), while recently, Pitrat (2012) proved that the main center of origin of lettuce is in Europe and Southwest Asia. There are one hundred species in the genus Lactuca. Four of them are wild species (L. sativa L., L. saligna L., L. serriola L. and L. virosa L.) can be crossed by classical methods (Bremer et al. 1994; Lebeda and Astley 1999; Lebeda et al. 2007). They are diploids with the same chromosome numbers that have been self-fertilized. (2n = 2x = 18) (Doležalová et al. 2002; Grulich 2004; Rubatzky and Yamaguchi 1997). Lettuce is a self-pollinated annual crop. From the taxonomic side, it belongs Lactuca genus, tribe Cichoreae, family Asteraceae (Compositae) and order Asterales. However, there is a cross-pollination of up to 5% among lettuce varieties (George 1999). It has a deep horizontal lateral root system, the densest near the soil surface for water and nutrients absorbance. The arrangement of leaves is in a thick rosette form. The stem is often shortening. According to lettuce, there is a great diversity in plant morphology, such as color, margin, surface, texture, and shape of the leaves.
The leaf surface can be smooth or wrinkled. The leaves ranges in color from yellow to green to dark purple. Anthocyanins can be mixed to cover all or part of the leaves. Elongation of the stem is the end signal of the vegetative phase, and beginning to the reproductive phase. Usually, a single stem is formed to endure flowering. A dense scarlet complexion made up of many capitula, each one consisting of many flowers. The small flowers usually vary in number ranging from 12 to 20, but their number can be less than 7 and up to 35. The seeds have many colors, including white, gray, yellow, brown and black. The freshly collected seeds enter a dormancy period, and many varieties vary in thermal dormancy levels. Aside from being an important vegetable crop, lettuce has economic value for numerous commercial applications, including cigarette production free of nicotine from the leaves, edible oil extracted from seeds, as well as a sedative derived from dried latex found in stems and other tissues (Dupont et al. 2000). It can be cultivated using seeds under rain-fed and irrigated conditions. Conventional breeding has a leading role in integrating novel alleles by crossing genotypes from various plant genetic resources to enhance many traits; e.g., modern varieties hybrid with locally adapted ones, to obtain early maturity, high yield, enhance quality and resistance to abiotic as well as biotic stresses. New biotechnology approaches such as using advanced DNA sequences and molecular methods are essential to develop new lettuce cultivars with desirable characters. This chapter aims to introduce a general overview of lettuce's importance and its economic and medicinal values. Also, it provides different methods to obtain new lettuce cultivars (e.g., conventional breeding, mutagenesis, genetic engineering tools either by gene transformation or genome editing), germplasm diversity and conservation, genetic resource conservation, developments on biotechnology, crop cultivation practices, molecular biology and their application for crop improvement concerning conventional breeding methods of lettuce.
Origin and distribution
Lettuce's main center of origin (L. sativa L.) is in Europe and Southwest Asia (Pitrat 2012). The origin of this crop is the Western south of Asia, while the primary essence of its culture is the Mediterranean region. It has been cultivated since 2000 BC (Hancock 2004). According to Whitaker (1969), lettuce ancestors are native to the Eastern Mediterranean region, most likely Egypt. According to Lindqvist (1960b), a significant centre of L. sativa origin is located in the European Siberian region. Rather than Egypt, Rulkens (1987) assumed that cultivated lettuce originated in Kurdistan—Mesopotamia. Boukema et al. (1990) claimed that lettuce was domesticated in Southwest Asia, in the region between Iran and Egypt, whereas De Vries (1997a, b) claimed that cultivated lettuce originated in Southwest Asia, near the Tigris and Euphrates rivers. In ancient Egyptian agriculture, lettuce was cultivated for several purposes, including oil production from seeds, leaf consumption, or religious purposes (Pitrat 2012). The lettuce crop has a long history, as Lindqvist (1960a) mentioned in Egyptian civilization (e.g., mural paintings of lettuce in Egyptian temples and tombs), as shown in Fig. 1. On the walls of ancient Egyptian tombs, a long-leafed lettuce type was depicted (Harlan 1986). Lindqvist (1960b) confirmed some primitive L. sativa forms in Egypt and considered them to be semi-wild and not cultivated. The genus Lactuca contains about 100 species, 17 species in Europe, 10 species in North America, 33 species in tropical East Africa and 40 species in Asia (Lindqvist 1960a; Rulkens 1987). Figure 2 indicated the center of origin of lettuce and its distribution around the world. Table 1 showed some species of Lactuca according to Kew Botanical Garden (Bano and Qaiser 2011; Lebeda et al. 2019).

Source: Harlan (1986)
Inscriptions on the wall of the tomb of the Church of Choi in Abydos (circa 1800 BC. Photograph of E. Oost Archaeological Museum, Leiden).

Source: Harlan (1986)
Origin and distribution of lettuce around the world. Text by Wolfgang Schuchert; Adapted to HTML by R. Saedler (http://www1.biologie.uni-hamburg.de/b-online/schaugarten/LactucasativaL/Lettuce.html).
Health benefits and economic importance
Lettuce is considered a common food crop that belongs to the earliest domesticated vegetable crops (about 8,000 to 4,000 years before now) (Hancock 2004). It is commonly used in salads that are eaten up abundantly because they are considered from the "healthy" foods (DuPont et al. 2000). The healthy characters are rich in fiber, antioxidants, antiproliferative compounds (vitamins C, E, polyphenols and carotenoids) and phytochemicals like (anthocyanins and chlorophylls) (Chu and Liu 2002; Li et al. 2010; Llorach et al. 2008; Mulabagal et al. 2010; Nicolle et al. 2004; Noumedem et al. 2017; Serafini et al. 2002). Llorach et al. (2008) mentioned that red genotypes have anthocyanins in abundance. Lettuce is rich in minerals like calcium, iron, magnesium, potassium, sodium and vitamins A and C. However, it nutritional value varies depending on the genotype (Romani et al. 2002). Also, It is rich in vitamins A and C. However, vitamin C is lost if it is overcooked. It is usually grown for its soft leaves and heads, which are cut and used as a salad with salt and vinegar. In general, lettuce is a popular leafy crop. Also, the stalk is eaten, and the seeds can be used for the oil production. Lettuce is eaten all year because it is cultivated both outside and, in the greenhouse (Vries 1997a, b). This is summarized in Fig. 3 and Table 2.

Source: Noumedem et al. (2017)
The nutritional value of lettuce.
L. sativa is an essential leafy vegetable crop and has medicinal properties. According to traditional culture, it is used in the treatment of a variety of health disorders, including insomnia, neuropathy, dry cough, rheumatic pain (Araruna and Carlos 2010; Katz and Weaver 2003) and anxiety (Harsha and Kumar 2012, Harsha and Kumar 2013).
The presence of bioactive compounds in lettuce has been shown to have medicinal properties (e.g., anti-inflammatory, cholesterol-lowering, and anti-diabetic) in vitro and in vivo studies. The phenolic compounds in red lettuce genotypes are higher than in green lettuce genotypes. The vitamin C content of baby green romaine was particularly high (Kim et al. 2016). Lettuce foliage is rich in folic acid content. Also, lettuce is considered to be an analgesic, diuretic and expectorant. Lettuce contains a large proportion of water (95%) and low calories and the nutritional value were indicated in Table 2.
Lettuce is cultivated for commercial purposes in many countries worldwide and is grown as a garden vegetable. In 2018, China, the United States, India, Spain, Italy, Japan, Iran, Turkey, Mexico and Germany were the top ten producers of lettuce worldwide (FAO 2018). It is important commercially in Asia, America, and Europe (FAO 2018) (Fig. 4). Most of them are xerophytes, Liana-like endemic species in the central African mountains (Stebbins 1937). Lettuce is grown in cold areas and can be grown in hot weather with special care. It is preferred to be cultivated in sandy loam soils rich in organic matter and lime. Also, lettuce is sensitive to excess soil salinity, especially in the germination stage.

Source: FAO (2018)
Important region of lettuce.
Lettuce has valuable economic importance as a vegetable crop. Lettuce and chicory's world production is 24.32 million tons over the total area of approximately 1.11 million hectares.
Current cultivation practices and challenges
Current cultivation practices
Standard cultural management practices were employed on land preparation, planting distances, seedling medium, pricking time, which was 10 days after sowing, irrigation and harvesting (Cervantes et al. 2017). Cultivation requires a well-drained fertile soil of pH (6–7.5). Lettuce is moderately salt tolerant, needs a mean temperature of (10–20 °C). On a large-scale, cultivation takes place without soil with the help of hydroponic techniques or protective cultivation. Fertilizers of Nitrogen, Phosphorus and Potassium are used in the ratio of 5:10:10, respectively. These beds are 100 cm wide and 30 cm in height. After that, fertilizers are mixed, 100 g of sulphate is added on each bed.
Seeds are sown in lines at a spacing 5 cm and 1.5–2 cm depth. Then, covered with sand and ready to plant in 21–28 days. When the plants have 2–3 leaves, they should be foliated to a final spacing of 25–45 cm. Young plantlets can be grown in larger pots or cell trays at about 2 weeks old (Williams 2012). The seeds are kept dry at room temperature and viable for 5–7 years (Cumo 2013).
Agricultural challenges
One of the biggest challenges is delivering sufficient, diverse, and safe food to satisfy a global need. Climatic changes such as rainfall, temperature fluctuation and CO2 concentration cause abiotic stress and less productivity. Salt stress increases ionic concentration causing less water uptake and therefore results in drought. The water deficit reduces water, potentially leading to osmotic and oxidative stress within plants, especially lettuce (Dun-Chun et al. 2016).
Genetic improvement objectives
Recently, many strategies have evolved. To face these challenges, new tolerant cultivars are needed to handle stress. The development of an adaptive mechanism requires morphological, physiological and metabolic processes dependent on multiple genes. To overcome salt and drought, water-saving irrigation and soil management are used. Adaptations to environmental stresses are essential to cope with different environmental conditions (Bohnert et al. 1995, Hartman et al. 2014). The abiotic approach involves the use of stress-tolerant genotypes (Rai et al. 2011).
Germplasm diversity and conservation
Germplasm diversity
The role of genetic resources in developing probable solutions to major crop limitation has been suggested on and off. These genetic resources cannot be fully exploited due to their inherent large-scale problems and lack of adequate assessment and classification. Conservation, evaluation, and characterization of economically important traits are essential requirements for a genetic improvement program for any crop. Both quantitative and qualitative traits can be picked out to be hybridized to achieve heterosis or select desirable segregates in advanced offspring. The association of various parameters provides the basis of selection for yield and its improvement. Genetic variability is an essential need for any vegetable crop breeding program. Analysis of variance showed significant variations between all traits’ genotypes studied (e.g., days of maturity, gross head weight, equatorial diameter, seed germination and seed weight. These differences indicated variability and opportunity for improvement in the production and quality traits of lettuce (Kumar et al. 2016). Molecular markers are effective for the identification of genetic diversity in L. sativa. Table 3 summarizes using different molecular markers techniques used for various genetic purposes for L. sativa species. These purposes include genetic variation, taxonomy and genetic diversity. Both Fig. 5 and Table 4 show the morphology of some important common lettuce species and some of their relative plants, which is considered common lettuce in some cultures (Mao et al. 2014).
Morphology of some common Lactuca species and their relatives. Botanical names are listed in Table 5.
Genetic resources conservation approaches
It is essential to gather collections of leafy vegetable germplasm, which would measure the current situation for crops and other plants. Lactuca sativa is propagated vegetatively and not by seed. This makes lettuce has different approaches for propagation rather than seed plants. Křístková et al. (2008) mentioned that researches in germplasm (including conservation, evaluation and utilization) of lettuce resources are presided by leafy vegetable crop germplasm committees (CGC) with the sponsor of the US Department of Agriculture – Agricultural Research Service’s National Plant Germplasm System. Nybom et al. (2014) reported that Lactuca species have more variation among populations than within them. The regional variation has a major effect on this variable. There are four different conservation approaches for germplasm. They are in situ, ex-situ, in vitro and DNA bank.
In situ conservation
The most appropriate way for in situ conservation of lettuce plants is on the farm. This can assist in identifying the specific demands of an on-farm conservation plan. This plan has 6 main advantages that make it unparalleled among the choices available to environmentalists. Those requirements relate to ecosystem health, genetic diversity and human luxury. This benefit is based on in situ conservation of current germplasm and the conditions that develop new germplasm. This estimation of dynamic conservation expands to all facets of the farming system, like the wild species that interact with their cultivated relatives (Maxted et al. 1997).
Ex situ conservation
The European Cooperative Program for Plant Genetic Resources (ECPGR) Vegetable Network is the top priority in participating responsibilities for ex-situ conservation in the European vegetable crops and genetic resources in the integrated system of the European Gene bank (AEGIS) Plant Genetic Resources for Food and Agriculture. The challenge now is identifying genetically vital and unique accessions among the same genetic materials. The Most Appropriate Accessions (MAA) for each crop will be admitted into the decentralized European collection. As for lettuce, the comprehensive approach involves: (a) morphological comparison of accessions within and among groups, (b) obtaining basic missing passport data on accessions and (c) the use of biochemical and molecular techniques for the genetic distinction of genetic resources (Sochor et al. 2019).
In vitro conservation
The in vitro conservation provides an essential tool for the production of L. sativa plants able to produce new crops with high yields under abiotic stress. For example, Pileggi et al. (2001) examined proline which is osmolyte produced during stress conditions. They transform mutated P5CS (D1-pyrroline-5-carboxylate synthetase gene) carried on pBIF414 plasmid within Agrobacterium tumefaciens (LBA4404). They obtained the lettuce plant with high shooting productivity even though it was grown in the rooting medium (Fig. 6).

Source: Ismail et al. (2016)
Explants with strong shoots in rooting medium. callus culture of lettuce, b regenerated shots of lettuce.
DNA banks
Only twenty-two of the approximately one hundred Lactuca species are preserved in the world gene bank's collection, according to Dolealová et al. (2002). As a result, the taxonomy of the entire genus Lactuca is still being developed. Furthermore, according to Fukuda et al. (2017), many database tools (e.g., the Basic Local Alignment Search Tool (BLAST) in the National Center for Biotechnology Information (NCBI) database (ncbi.nlm.nih.gov/BLAST/) were used to determine the sequences of some genes (e.g., flowering-related genes). Many software programmes, such as Clustal Omega (ebi.ac.uk/Tools/msa/clustalo/) and GENETYX, are used to align it with homologous sequences (GENETYX CORPORATION, Tokyo, Japan). The accessions LC164345 (LsLFYL), LC164344 (LsAP1L), LC164348 (LsFVEL), and LC164347 (LsFVEL) have been submitted to the DNA Data Bank of Japan (LsFLDL).
Cytogenetics
The haploid type will provide a good foundation for producing a pure line of doubled haploids (DHLs) plants, allowing for the regeneration of new varieties. Piosik et al. (2015) devised a method for efficiently producing haploid plants of economically important species. In vivo distant pollination with fresh pollen grains of Helianthus annuus L. or Helianthus tuberosus L. was used to stimulate the development of haploid lettuce embryos. Callus tissue formation was observed in the laboratory, and 23 haploid L. sativa plants were regenerated. Several methods have been used to confirm the haploid status of regenerated plantlets, including I flow cytometry to estimate genome size, ii) counting chromosome numbers in root tips, iii) stomata cell size, and iv) pollen grain formation disturbances caused by abnormal microsporogenesis. The ideal method for obtaining haploid plants in L. sativa is detailed in this report. The difference between diploid and haploid L. sativa was shown in Fig. 7.

Source: Mousavi et al. (2013)
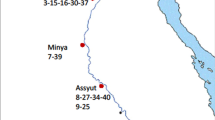
Representative mitotic metaphase cells in some Iranian lettuce. The numbers indicate the origin: 1 Abtavil, 2 Borazjan, 3 Ahvaz, 4 Karaj, 5 Qom, 6 Neishaboor, 7 Gorgan, 8 Babol, 9 Varamin, 10 Shiraz, 11 Kazeroon, 12 Hamadan, 13 Parsabad, 14 Jahrom and 15 Fasa.
Regarding Poisik et al. (2015), the relative DNA content (2 C) in haploid plants was 2.87 ± 0.01 pg. However, at control plants, the DNA content was 5.74 pg, which confirmed that the regenerated plantlets are haploid (Fig. 8a, b). The haploid number of chromosomes (n = 9; Fig. 8a) was observed in the metaphase of the regenerated plant individuals' root cells, and a diploid number in the control individuals was (2n = 18; Fig. 8b).

Source: Piosik et al. (2015)
Plant habit chromosome numbers and histograms of relative DNA content of a haploid and b diploid L. sativa plants.
Also, Mousavi et al. (2013) showed that the basic chromosome number in the studied accessions was n = 9. Probably due to the fact that it has a long history. There were also morphological differences and heteromorphisms between chromosomes and their types in all accessions (e.g., metacentric, submetacentric and subtelocentric). The results of an analysis of variance (ANOVA) revealed that the chromosomal characteristics differ significantly. The genome estimated to be the largest was that of the Qom accession (62.45 m). The Babol accession had the smallest size (19.94 m). Figure 7 shows the symmetrical and asymmetrical karyotypes between the accessions.
Traditional breeding
Improvement strategies
Although current breeding methods that use DNA technology advances have attracted lettuce breeders, classical breeding methods are still essential to improve new lettuce cultivars with useful features. The main goal of conventional lettuce breeding is to create a lettuce plant that can easily grow and thrive in a variety of environments. With emerging global issues, changes in farmable lands, biotic and abiotic stress, improved quality, and increased yield should all be taken into account (Hartman et al. 2014; McCabe et al. 2001). Plant breeding methods allow for the selection of genotypes with desirable genes that control important traits, which are then processed in lettuce (Hartman et al. 2014). (Fig. 9)

Source: Simko et al. (2018)
Flow chart illustrating linkage mapping of QTLs performed for two populations genotyped of lettuce with molecular markers and phenotyped environments.
Traditional breeding methodologies and limitations
In 1900 Mendel’s law provided the basis of plant breeding science. Plant breeding is conducted by combining the identified and selected traits in only one individual plant. These traits may increase the head size and weight, higher seed yield, improved color, tolerance to heat and drought, resistance to insects and diseases and better agronomic quality. To develop any novel plant, it should follow several steps to define and analyze the current germplasm problem, then establish the goals and objectives of breeding. Traditional plant breeding is the manipulation of how chromosomes combine. Generally, there are two main procedures to control plant chromosome combinations. Firstly, selecting the germplasm that meets the breeding goals selection in plant breeding is considered the most ancient process. Generally, it involves three different steps starting with primary selection of a large number of the genetically variable original population. Next, for observational purposes progeny rows are grown from the selected individual plants. After elimination, the selected plants are grown over many years to allow performance observations under different environmental conditions for making further eliminations. Finally, compare the selected and inbred lines to existing commercial varieties in their yielding performance and other agronomic importance aspects (Gupta et al. 2008; Ragheb 2015; Souza et al. 2008; Tashi et al. 2010). Secondly, hybridization of desired traits found in different plants together to obtain plants that obtain the desired traits. This aims to bring the desired traits together into one plant line via cross-pollination. Primarily generate homozygous inbred lines, which is done by self-pollinating plants in the same plants' male flowers pollen pollinate female flowers. As soon as a pure line is generated, another inbred line is combined with it resulting in progeny is chosen for combination of the desired traits. These strategies aim to maintain desirable traits, high yield, good quality, tolerance to abiotic and abiotic stresses (Michelmore 1995; Ryder 1986, 1991). There are some limitations for a traditional breeding technique such as difficulty maintaining the purity of offspring, time consuming, not producing many offspring, and the appearance of unfavorable traits that can be removed by time-consuming backcrossing.
Role of biotechnology
The main focus of breeding L. sativa is to improve its quality and resistance to early wilting, pests and diseases. To obtain such characteristics, biotechnology and genetic engineering methods are used, including gene transfers. Using those modern techniques saved different genotypes and produced a healthy environment and overcame the limitations of in vivo techniques by shortening breeding programs. Various methods are now used, including haploidization, shoot organogenesis, somatic hybridization and others. Haploidization can produce pure lines. At the same time, shoot organogenesis improves important characteristics of lettuce. We can access exotic and sexually incompatible germplasm through somatic hybridization via protoplast fusion. Another source of genetic variation is somaclonal variation. Sariçam et al. (2017a) found that regeneration via somatic embryogenesis is also beneficial for lettuce production.
Molecular breeding
Molecular marker-assisted breeding
Quantitative trait loci (QTL) related to biotic (Christopoulou et al. 2015; Simko et al. 2013) and abiotic stresses have been identified in lettuce breeding (Hartman et al. 2014; Jenni et al. 2013), allowing for the improvement of lettuce tolerance to various stresses. Molecular breeding has become a practise since the development of DNA markers in the 1980s (Rafalski and Tingey 1993). Many molecular markers have been developed to construct genetic maps for crop improvement, including amplified fragments length polymorphism (AFLP), random amplified polymorphic DNA (RAPD), restriction fragment length polymorphism (RFLP), simple sequence repeat (SSR), sequence characterised amplified region (SCAR), and single nucleotide polymorphism (SNP). For marker-assisted selection (MAS) in plant breeding, QTLs related to important agronomical characters and their highly related DNA markers have been used (Collard and Mackill 2008). In lettuce seedlings, quantitative trait loci linked to root growth (Roberts et al. 2020). Reyes-Chin-Wo et al. recently published the annotated genome sequence assembly of lettuce species at the Lettuce Genome Resource (2017) (https://lgr.genomecenter. ucdavis.edu).
Biochemical markers were also used to investigate the genetic differences between lettuce and wild lettuce genotypes (Cole et al. 1991; Collard and Mackill 2008; Dziechciarková et al. 2004).
In breeding studies, molecular markers are used in a variety of ways. Landry et al. (1987) used forty-one RFLP molecular markers to create a linkage map of lettuce. Using two molecular markers, Kesseli (1994) discovered a genetic linkage map for L. sativa (RAPD and RFLP). Jeuken et al. (2001) used two L. sativa x L. saligna F2 populations to create an integrated interspecific AFLP map for lettuce species. Truco et al. (2007) combined seven lettuce linkage maps into a high-density one with 2,744 DNA markers. Truco et al. (2013) used F7:8 recombinant inbred lines (RILs) derived from L. sativa x L. serriola to create an extremely high-density genetic map (13,943 markers) of lettuce. Simko et al. (2018) created a flow chart to explain how to use QTLs as a molecular marker and different environmental factors to perform linkage mapping of two genotyped lettuce populations.
Functional genomics
Few works have been carried out in functional genomics in lettuce plants. Functional genomics has been used to improve the expression of genes responsible for stress tolerance. Reyes-Chin-Wo et al. (2017) published the genome of L. sativa, and reference genomes of other lettuce species (L. serriola, L. saligna, and L. virosa) are expected to be available soon. Lactuca germplasm is undergoing large-scale DNA resequencing projects. The enriched DNA sequencing data is expected to lead to functional genomics approaches, which will eventually lead to more efficient lettuce breeding with higher production and nutritional quality. However, these methods can only be successful if phenotype data is collected in a comprehensive and efficient manner (Still 2007).
Bioinformatics
The bioinformatics for L. sativa is available on the web and can be easily accessed. There are many databases available for this plant that include Kyoto Encyclopedia for Genes and Genomes (KEGG) where transcriptomic and proteomic data can be explored. For example, a statistical summary shows that the number of protein genes is 35603 and the number of RNA genes is 2587 (genome.jp/dbget-bin/www_bget?gn:T05352). A genetically validated reference assembly with several genomic features is also provided. Kinases, Cycloidea-like transcription factors, disease-resistant proteins, and 21 novel microRNAs are just a few examples (Chin-Wo et al. 2017). In addition, the database Uniprot gives information on Gene Ontology of different proteins showing their function and catalytic activity (uniprot.org/uniprot/?query = lactuca + sativa). In NCBI, the assembly reports history for L. sativa, started in April 2011 to the latest project in April 2020 can be found in the assembly section (ncbi.nlm.nih.gov/assembly/GCA_002870075.2).
Genetic engineering and gene editing
Methodologies and enhanced traits
Genetic engineering is used to manipulate lettuce genes directly. By developing high tolerance lines under a variety of conditions, it is expected to support traditional breeding methods by increasing lettuce production efficiency and avoiding losses due to biotic and abiotic stresses. Using advances in molecular techniques and genome sequencing, genetic engineering provides tools to improve lettuce yield and quantity. It leads to highly effective and robust transformation systems for nucleases that target specific sequences. In lettuce, transformation systems such as Agrobacterium tumefaciens have been successful. The successes proceeded in genome editing using technologies (e.g., technique of Cas9 is promising for performance and yield-boosting). As a result, lettuce genetic engineering must outperform traditional transformation methods. Several studies on lettuce have been published, including one that used Agrobacterium (LBA4404) to transform the Pta gene from Pinellia ternate to L. sativa using leaf explants in tissue culture and confirmed the transformation with RT-PCR; the resulting transgenic lines were resistant to insecticide-released toxic compounds (Ahmed et al. 2007; Chen et al. 2018; Curtis et al. 1994; Woo et al. 2015).
Transgenic cultivars
Gene transformation processes is applied to extend the germplasm genetic base and is available for classical breeding. It is used to minimize the time required to insert a single gene into the essential economic crop. L. sativa is one of the most widely cultivated leafy vegetable crops on the planet. It responds to a wide range of growth regulators in tissue culture, with shoot regeneration achieved from a wide range of cultivars representing different genotypes. Lettuce genetic manipulation necessitates a genotype-independent transformation method that is both reliable and efficient (Chen et al. 2018; Curtis et al. 1994; Dan et al. 2014). Plant biotic stresses (e.g., diseases and insect pests) are barriers for the cultivation and production of many lettuce plants. Plant diseases seriously affect lettuce crop yield and quality. For example, the disease of Lactuca sativa caused by Sclerotinia fungi has occurred in lettuce and cause serious harm in basal part of stems and leaves. Thus, many experts committed themselves to study transgenic lettuce. Diverse types of genes that were transferred to lettuce, being expressed and stably inherited in progenies (Dan et al. 2014). Many processes of gene transformation are to resist much biotic or abiotic stress. Also, genetic transformation in lettuce plants was for the enhancement of vegetative characters. All that are summarized in Table 5.
Modern lettuce breeding objectives start developing genotypes that can resist insects and disease and improve quality and high yield. For example, lettuce has been injected with the ipt gene, which significantly slows down the development and senescence of mature leaves (McCabe et al. 2001). Mohebodini et al. (2011) used MS medium supplemented with some growth hormones (0.1 mg/L 6-benzylaminopurine (BA) and 0.1 mg/L indole-3-butyric acid IBA) to induce callus in cotyledons from 48-h germinated seeds. Following that, the callus was placed in MS medium containing 0.1 mg/L BA for indirect shoot regeneration. Two original species genotypes from central (Yazd) and southwest (Ahvaz) were used in this experiment, both of which had good potential for biotic and abiotic stress. The symptoms of extending and swelling appeared three days after the culture was started. The average callus percentage is affected by genotype explant age and different concentrations of growth regulators. Callus formation began within 7 days of the start of the culture and was observed in all media. The effects of genotype, explant age, and various growth regulator concentrations on direct shoot regeneration. Within fourteen days of culture, direct shoots had regenerated. On cotyledon explants, shoot buds emerged directly from the end of the region near the petiole. Within 21 days of starting the culture, multiple shoots appeared (Fig. 10).

Source: Mohebodini et al. (2011)
Direct shoot regeneration on cotyledon explants of lettuce near to petiole of cotyledons. a Left arrow is middle of cotyledon; right arrow is the region near to petiole of cotyledon, b Left arrow is the region near to petiole of cotyledon; right arrow is middle of cotyledon.
Breeding programmes' main goal is to produce a high yield of cultivars that are resistant to both abiotic and biotic stresses. So, using various techniques such as chemical mutagens, T-DNA insertions, zinc finger nucleases (ZFNs), and TAL effector nucleases, breeders attempted to edit the plant genome by inducing mutations or editing specific gene/s in the genome (TALENs). Several obstacles appeared in using these methods, including time-consuming and high costs because protein engineering is required. The new genome-editing technique called clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein 9 (Cas 9), has been produced to overcome the difficulties of the previous methods. The CRISPR/Cas9 nuclease system becomes popular in the precise genome editing of living organisms for accuracy and ease of manipulation. The CRISPR genome editing technique of lettuce requires four main steps: (a) purification of the Cas effector proteins and guide RNA, (b) protoplasts preparation, (c) transfection of pre-assembled RNPs into protoplasts and (d) regeneration of whole plants from engineered tissues (Park et al. 2019). Kozai et al. (2018) summarized the different methods used to obtain new lettuce cultivars. These methods were illustrated in Fig. 11: crossbreeding, mutation, gene transformation and genome editing.
Mutation breeding
Koornneef (2002) defines mutation as the process by which genes are permanently altered, either naturally or artificially. It's found on the chromosome or in the genes. Because mutations can cause a trait to gain or lose, plant breeders use mutations to create new cultivars with specific characteristics. Mutations can be induced by either chemical or physical agents. The physical mutagens are the most commonly used, such as X-rays and ultraviolet (UV) illumination. Till et al. (2007) list nitric acid, sodium azide, ethyl methane sulfonate (EMS), methyl methane sulfonate (MMS), and diethyl sulphate as examples of chemical mutagens (DES). They've also been used in the induction of mutations in various plants. CRISPR/Cas9 is a transformative technique for making targeted genetic mutations, according to Bertier et al. (2018). Some lines have been reported to have high mutation efficiencies in various plants.
Conventional mutagenesis
Genetic differences are significant for plant breeding programs. When compared to cross-pollinated crops, lettuce's inherited habitat orders a low genetic variance in the crop. Inherited and caused mutations are two types of mutations that can be used to create new traits in lettuce plants. Natural mutation occurs in crops and wild plants, though the lead to beneficial characters can only be selected for human needs on a small scale. Mutagenic agents (chemical and physical) are used to increase mutation rates or produce mutations not found in nature (Mou 2011). EMS, fast neutron, and gamma rays were used to treat seeds of lettuce varieties 'Diana' and 'Saffier' in an attempt to find mutants resistant to herbicides such as rimsulfuron, imazamox, glufosinate, and glyphosate, according to Michelmore et al. (2002).
The frequency of the mutation that comes from chemical mutagenesis in the physical compartment is 103 times higher than naturally occurring variations. Lettuce breeding studies focus on leaf-shaped, tight heads, presence or absence of anthocyanins, resistance to diseases and dirtballs, good quality under stress conditions, short growth period, yield, adaptation to different areas and environmental conditions (De Vries 1997; Mou 2011). This work was carried out to produce a database of mutation breeding studies on lettuce CV. Cervantes. For this purpose, Gray (Gy) doses of Co60 (gamma rays) of 0, 50, 100, 200, 300, 400, 500 and 600 were applied on lettuce seeds as a physical mutagen. Thirty seeds were used per dose. After a month of treatment, germination and shoot development were recorded. The Effective Mutagen Does was calculated by the method of linear regression analyzes. According to the results, a dose of 372.66 Gy was found as EMD50 (Sariçam et al. 2017b).
In vitro mutagenesis and selection
According to Mou (2011), both L. sativa and L. serriola species can be hybridised because their chromosomes are morphologically similar. Both of these species are subspecies of the same species. Mutations can cause changes in L. serriola, which has resulted in the appearance of desirable traits in humans. These characters are number of spines on stems, leaves and seeds. They were then selected for reproduction and further modified to meet human needs. These early forms may be suitable for the production of oil from seeds for domestic purposes. The majority of this grows and develops quickly, with unreflective materials to prevent seed shattering, large seeds, and a high oil content in seeds (35%). Franco et al. (2015) experimented the mutagenic effect of radiation on seeds of lettuce. The aim was to estimate the potential growth parameters of lettuce. Gurdon et al. (2019) worked with ethyl methyl sulphonate (EMS) to produce red L. sativa mutants with high flavonoids and phenolic compounds. These mutants are used medicinally as anti-diabetic.
Molecular analysis
Two main molecular mutation analysis methods have been used in the molecular analysis of mutation (Candela et al. 2015), (a) sequencing of bulk DNA and identification of mutation by crosses to polymorphic mutants, and (b) sequencing bulk DNA and identification of mutation using backcross isogenic mutants.
Huo et al. (2016) mentioned an example of molecular analysis of mutagenic lettuce lines: two independent heat-tolerant lettuce seeds mutant lines, TG01 and TG10, were generated through ethyl methane sulfonate (EMS) mutagenesis. These two mutations were allelic and recessive, according to physiological and genetic analyses. Apply bulk segregant analysis to whole-genome sequencing to find the causal gene/s. For each mutant line, bulked DNA samples from segregating heat-tolerant (mutant) seeds were sequenced and homozygous single nucleotide polymorphisms (SNPs) were looked for. Traditional genetic mapping confirmed the presence of causal mutations near the ZEP/ABA1 gene, but whole-genome sequencing is more effective at identifying specific genes responsible for the phenotype.
Enhanced traits and improved cultivars
Mutations have an essential role in domesticating many crops. Many physiological and genetic studies benefit from characters resulting from both induced and natural mutations (e.g., early flowering, dwarfing, chlorophyll deficiency, and male sterility). Mutations have also been used to create new lettuce varieties, including herbicide-tolerant cultivars. In genomic studies of lettuce, such as the identification and cloning of disease-resistant genes, mutation analysis was crucial. sThey can also have a significant influence on the evolution and improvement of a self-pollinated crop such as lettuce. It is a method to make genetic variation, which can then be used in physiological or genetic studies, as well as cultivar development. It is more efficient to change qualitative traits like disease resistance and quality under the control of essential genes than it is to change quantitative traits like yield and adaptation (Mou 2011). Different mutagens were used for the enhancement of lettuce characters such as EMS, gamma rays, and fast neutrons (Michelmore et al. 2002).
EMS mutagen was used to characterize biosynthetic mutants of flavonoids in lettuce with medicinal uses. The modified flavonoid is distinguished by a high concentration of naringenin chalcone and kaempferol, allowing lettuce to be used in foods with health benefits. To back up the health benefits of the high flavonoid lettuce varieties described here, animal studies and human clinical trials will be needed. Innovative mutations and selection plans to increase the levels of beneficial phytochemicals in common crops could be a crucial strategy (Gurdon et al. 2019).
Hybridization
Conventional hybridization
Producing F1 hybrids has been unsuccessful because the pollen of L. sativa’s is heavy, sticky and not easily transferred. As a result, three principles guided hybridization: (a) pedigree method, (b) backcross, and (c) single-plant or mass selection. Different parents are staggered in sunny and warm days to ensure that all plants flower. In the morning, the flowers bloom (Acquaah 2012). On cloudy and cool days, it takes longer. The capitulum of lettuce is made up of 10–20 florets (Mou 2008). The crossing was carried out according to Hooftman et al. (2005) and Ryder (1997). The second generation (F2) seeds can be produced using the selfing of a single F1 plant. F2 seeds are sown and 200 seedlings are randomly selected, transplanted and genotyped. The plants are selfed and the third generation (F3) seeds are harvested for each second-generation plant (Uwimana 2011).
Somatic cell hybridization
Electric methods and polyethylene glycol (PEG) can both be used to obtain lettuce protoplast fusion easily (Siddiqui 2014; Taniguchi 1990). Protoplasts can be obtained from lettuce developed in greenhouses and growth chambers, and from shoots, roots, and cotyledons of seedlings developed in vitro (Berry 1982). Somatic hybridization was conducted so that the investigation on the culture could be carried out in four media types by polyethylene glycol (PEG) and electrofusion. While other treatments were observed to damage the protoplast, it was shown that both micro and macro colonies' chemical and electro-fusion resulted in significant outcomes for fused protoplasts when treated with different treatment for each replication. Additionally, electrofusion resulted in a protoplast fusion of 40.51% frequency over PEG (Siddiqui 2014).
Hybrid cultivars
Lettuce comprises seven main groups of genotypes cultivated in many locations in a wide range of land heights and old varieties held in the world's gene banks (Lebeda et al. 2007). They are usually described as morphotypes. New cultivars are being developed using both traditional and modern breeding methods to meet the specific needs of producers and consumers. For example, Uwimana et al. (2012) used a crop-wild hybridization for lettuce to improve the colour of the leaves. The cosmopolitan (L. serriola, L. aculeata, L. scarioloides, L. azerbaijanica, L. georgica, L. taica) and the cosmopolitan (L. azerbaijanica, L. georgica, L. taica) occurring in Asia, as well as L. dregeana from South Africa, make up the primary gene pool of L. sativa (Zohary 1991).
Climate change implication and mitigation
Change in climate is controlling crop cultivation and productivity. Lee et al. (2009) explained that negative shocks of global warming which lead to decrease in crop quality and quantity. This is due to agricultural crops having a short growth period after exposure to high temperatures, as well as low sugar content, poor coloration, and decreased storage stability. Lee et al. (2009) also mentioned that lettuce may have problems at high temperatures, as it causes flowers to break up. When the temperature arises, it is possible to preserve energy for heating greenhouses in winter.
Conclusions and prospects
Lettuce (L. sativa) is a common economic vegetable crop plant. It has many centers of origins. It was mentioned in the old Egyptian civilization. There are many nutrients in lettuce include vitamin A, potassium, calcium and folate. Due to the economic importance of lettuce, a lot of studies have focused on some cultivars’ domestication. Quantitative trait loci (QTLs) related to biotic and abiotic stresses have been identified in lettuce breeding, allowing for the improvement of lettuce's tolerance to various stresses.
There are still numerous reasons for more investigations aimed at lettuce development. These reasons focused only on Food sufficiency and vitamin supplementation. Work is needed to improve the quantitative and qualitative traits of lettuce. These traits could include enhancing and increasing vitamin content and regulating mineral uptake through an osmotic balance between minerals outside and inside the cells. There may also be further cross-section studies to link between many vegetables for consumption. It may also enhance consuming this nutritious vegetable as lettuce contains many fibers, sugars, proteins, vitamins (C, E, K) and minerals (Ca, Mg, Fe, P).
Author Contributions Statement
Conceptualization: [Sara A. Mekkawy, Khaled F. M. Salem and Eman Tawfik]; Methodology: [Mohamed N. Hassan, Sara A. Mekkawy, Mayada Mahdy, Khaled F. M. Salem and Eman Tawfik]; Writing—original draft preparation: [Mohamed N. Hassan, Sara A. Mekkawy, Mayada Mahdy, Khaled F. M. Salem and Eman Tawfik]; Writing—review and editing: [Mohamed N. Hassan, Sara A. Mekkawy and Eman Tawfik]; Supervision: [Eman Tawfik].
Availability of data and material
All data materials are available in manuscript.
References
Acquaah G (2012) Principles of plant genetics and breeding, 2nd edn. John Wiley and Sons, USA, pp 131–145
Ahmed MB, Akhter MS, Hossain M et al (2007) An efficient Agrobacterium-mediated genetic transformation method of lettuce (Lactuca sativa L.) with an Aphidicidal gene, Pta (Pinellia ternata Agglutinin). Middle East J Sci Res 2(2):155–160
Araruna K, Carlos B (2010) Anti-inflammatory activities of triterpene lactones from Lactuca sativa. Phytopharmacology 1(1):1–6
Bano R, Qaiser M (2011) A taxonomic revision of the genus Lactuca L. Cichorieae-Asteraceae from Pakistan and Kashmir. Pak. J. Bot 43(5):2259–2268
Berry SF, Lu DY, Pental D, Cocking EC (1982) Regeneration of plants from protoplasts of Lactuca sativa L. Z Pflanzenphysiol 108:31–38
Bertier LD, Ron M, Huo H et al (2018) High-resolution analysis of the efficiency, heritability and editing outcomes of CRISPR/Cas9-induced modifications of NCED4 in lettuce (Lactuca sativa). G3 8:1513–1521
Bohnert HJ, Nelson DE, Jensenay RG (1995) Adaptations to environmental stresses. Plant Cell 7:1099–1111
Boukema IW, Hazekamp Th, van Hintum Th JL (1990) The CGN collection reviews: the CGN lettuce collection. Wageningen, Centre for Genetic Resources, pp 2–5
Bremer K, Anderberg AA, Karis PO et al (1994) Asteraceae: cladistics and classification. Portland, Oregon, Timber Press
CABI Crop Protection Compendium (2008) Lactuca sativa (lettuce) datasheet. Available at: http://www.cabi.org/cpc/datasheet/29609
Candela H, Casanova Saez R, Micol JL (2015) Getting started in mapping by sequencing. J Integr Plant Biol 57:606–612
Cervantes CN, Laynesa FP, Pacis JB (2017) Validation and documentation of organic production systems for lettuce (Lactuca sativa) Camarines sur, Philippines. Int J Agric Tech 13:1277–1284
Chen Z, Han Y, Ning K et al (2018) Inflorescence development and the role of LsFT in regulating bolting in lettuce (Lactuca sativa L). Front Plant Sci 8:2248. https://doi.org/10.3389/fpls.2017.02248
Christopoulou M, McHale LK, Kozik A et al (2015) Dissection of two complex clusters of resistance genes in lettuce (Lactuca sativa). Mol Plant Microbe Interact 28:751–765
Chu YF, Sun J, Liu RH (2002) Antioxidant and antiproliferative activities of common vegetables. J Agri Food Chem 50:6910–6916
Cole RA, Sutherland RA, Riggall WE (1991) The use of polyacrylamide gradient gel electrophoresis to identify variation in isozymes as markers for Lactuca species and resistance to the lettuce root aphid Pemphigus bursarius. Euphytica 56:237–242
Collard BC, Mackill DJ (2008) Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Philosophical transactions of the Royal Society of London Series B. Biol Sci 363:557–572
Cumo C (2013) Encyclopedia of cultivated plants: from Acacia to Zinnia, Kindle. ABC-CLIO Publisher, Santa Barbara, California, pp 577–578
Curtis IS, Power JB, Blackhall NW et al (1994) Genotype independent transformation of lettuce using Agrobacterium tumefaciens. J Exp Bot 45(279):1441–1449
Dan S, Qiang H, Zhaonan D, Zhengquan H (2014) Genetic transformation of lettuce (Lactuca sativa): a review. Afr J Biotech 13(16):1686–1693
De Vries IM (1990) Crossing experiments of lettuce cultivars and species (Lactuca sect. Lactuca, Compositae). Plant Syst Evol 171:233–248
De Vries IM (1997a) Origin and domestication of Lactuca sativa L. Genet Resour Crop Evol 44:165–174
de Vries IM (1996) Characterization and identification of Lactuca sativa cultivars and wild relatives with SDS-electrophoresis (Lactuca sect. Lactuca, Compositae). Genet Resour Crop Evol 43:193–202
Dias BBA, Cunha WG, Morais LS et al (2006) Expression of an oxalate decarboxylase gene from Flammulina sp. in transgenic lettuce (Lactuca sativa) plants and resistance to Sclerotinia sclerotiorum. Plant Pathol 55:187–193
Doležalová I, Křístková E, Lebeda A, Vinter V (2002) Description of morphological characters of wild Lactuca L. spp. genetic resources (English-Czech version). Hort Sci (PRAGUE) 29(2):56–83
Dun-Chun H, Jia suiLian hui ZX (2016) Problems, challenges and future of plant disease management: from an ecological point of view. J Integr Agric 15(4):705–715
Dupont S, Mondi Z, Willamson G, Price K (2000) Effect of variety, processing and storage on the flavonoid glycoside and composition of lettuce and cichory. J Agric Food Chem 48:3957–3964
Dziechciarková M, Lebeda A, Doležalová I, Astley D (2004) Characterization of Lactuca spp. germplasm by protein and molecular markers–a review. Plant Soil Environ 50:47–58
El-Esawi MA (2015) Molecular genetic markers for assessing the genetic variation and relationships in Lactuca Germplasm. ARRB 8(5):1–13
Fallah-Ziarani M, Haddad R, Garoosi G, Jalali M (2013) Agrobacterium-mediated transformation of cotyledonary leaf of lettuce (Lactuca sativa L.) by the GCHI gene. J Genet Plant Breed 2(2):47–55
FAO (2018) FAOSTAT crops. http://www.fao.org/faostat/en/#data/QC/visualize
Franco CH, Santos HM, Silva LP, Arthur V, Silva RGM (2015) Potential of lettuce grown from irradiated seeds. Sci Hortic 182:27–30
Fukuda M, Yanai Y, Nakano Y et al (2017) Isolation and gene expression analysis of flowering-related genes in lettuce (Lactuca sativa L). Hortic J 86(3):340–348
George RAT (1999) Compositae. In: George RAT (ed) Vegetable seed production. CAB International, Wallingford, pp 122–1353
Grulich V (2004) Lactuca L. In: Slavík B, Štěpánková J (eds) Květena České Republiky 7. Academia, Praha, pp 487–497
Gupta A, Tashi D, Chattoo M, Yasmin S (2008) Estimation of genetic variability and heritability in lettuce (Lactuca sativa L.). Indian J Plant Genet Resour 21(2):138–140
Gurdon C, Poulev A, Armas I et al (2019) Genetic and phytochemical characterization of lettuce flavonoid biosynthesis mutants. Sci Rep 9:3305. https://doi.org/10.1038/s41598-019-39287-y
Hancock JF (2004) Plant evolution and the origin of crop species, 2nd edn. CABI Publishing, Walling ford
Harlan JR (1986) Lettuce and the sycamore: sex and romance in ancient Egypt. Econ Bot 40:4–15
Harsha SN, Kumar AKR (2012) Effects of Lactuca sativa extract on exploratory behavior pattern, locomotor activity and anxiety in mice. Asian Pac J Trop Dis 2:S475-479
Harsha SN, Kumar AKR (2013) Anxiolytic property of hydo-alcohol extract of Lactuca sativa and its effect on behavioral and biochemical activity. J Biomed Res 27(1):37–42
Hartman Y, Hooftman DAP, Uwimana B et al (2014) Abiotic stress QTL in lettuce crop-wild hybrids: comparing greenhouse and field experiments. Ecology Evolution 4:2395–2409
Hill M, Witsenboer H, Zabeau M et al (1996) PCR-based fingerprinting using AFLPs as tool for studying genetic relationship in Lactuca spp. Theor Appl Genet 93:1202–1210
Van Hintum T (2009) Molecular characterization of a lettuce germplasm collection. Eucarpia Leafy Vegetables 99–104
Hooftman DA, Oostermeijer JGB, Jacobs MM, Den Nijs HC (2005) Demographic vital rates determine the performance advantage of crop–wild hybrids in lettuce. J Appl Ecol 42(6):1086–1095
Huo H, Henry IM, Coppoolse ER et al (2016) Rapid identification of lettuce seed germination mutants by bulked segregant - analysis and whole genome sequencing. Plant J 88(3):345–360. https://doi.org/10.1111/tpj.13267
Ichikawa Y, Tamoi M, Sakuyama H et al (2010) Generation of transplastomic lettuce with enhanced growth and high yield. GM Crops 1(5):322–326
Ismail H, Dilshad E, Mirza B (2016) Transformation of Lactuca sativa L. with rol C gene results in increased antioxidant potential and enhanced analgesic, anti-inflammatory and antidepressant activities in vivo. 3 Biotech 6(2) doi:https://doi.org/10.1007/s13205-016-0533-4
Jackson M, Ekkehard H (2008) Transgenic lettuce seedlings carrying hepatitis B virus antigen HBsAg. Braz J Infect Dis 12(6):469–471
Jansen J, Verbakel H, Peleman J, van Hintum TJL (2006) A note on the measurement of genetic diversity within gene bank accessions of lettuce (Lactuca sativa L.) using AFLP markers. Theor Appl Genet 112:554–561
Jenni S, Truco M, Michelmore R (2013) Quantitative trait loci associated with tipburn, heat stress-induced physiological disorders, and maturity traits in crisphead lettuce. Theor Appl Genet 126:3065–3079
Jeuken M, van Wijk R, Peleman J, Lindhout P (2001) An integrated interspecific AFLP map of lettuce (Lactuca) based on two L. sativa × L. saligna F2 populations. Theor Appl Genet 103:638–647
Kanamoto H, Yamashita A, Asao H et al (2006) Efficient and stable transformation of Lactuca sativa L. cv. Cisco (lettuce) plastids. Trans Res 15:205–217
Katz SH, Weaver WW (2003) Encyclopedia of food and culture. Schribner, New York
Kesseli R, Ochoa O, Michelmore R (1991) Variation at RFLP loci in Lactuca spp. and origin of cultivated lettuce (L. sativa). Genome 34:430–436
Kesseli RV, Paran I, Michelmore RW (1994) Analysis of a detailed genetic linkage map of Lactuca sativa (Lettuce) constructed from RFLP and RAPD markers. Genetics 136:1435–1446
Kim MJ, Moon YY, Tou JC et al (2016) Nutritional value, bioactive compounds and health benefits of lettuce (Lactuca sativa L.). J Food Compos Anal 49:19–34
Koopman WJM, Zevenbergen MJ, Ronald G, Van den Berg RG (2001) Species relationship in Lactuca s. l. (Lactuceae, Asteraceae) inferred from AFLP fingerprints. Am J Bot 88:1881–1887
Koornneef M (2002) Classical mutagenesis in higher plants. In: Gilmartin PM (ed) Molecular plant biology. Oxford University Press, Oxford, UK, pp 1–11
Kozi T (2018) Smart plant factory. the next generation indoor vertical farms. Springer Nature, Singapore
Křístková E, Doležalová I, Lebeda A et al (2008) Description of morphological characters of lettuce (Lactuca sativa L.) genetic resources. Hort Sci (Prague) 35(3):113–129
Kumar R, Kaushal S, Kumar S et al (2016) Morphological characterization of newly introduced lettuce (Lactuca sativa L.) germplasm through principal component and regression analyses. Elect J Plant Breed 7(3):742–749
Landry BS, Kesseli R, Farrara B, Michelmore RW (1987) A genetic map of lettuce (Lactuca sativa L.) with restriction fragment length polymorphism, isozymes, disease resistance and morphological markers. Genetics 116:331–337
Lebeda A, Křístková E, Doležalová I, Kitner M, Widrlechner MP (2019) Wild Lactuca Species in North America. Iowa State University, Horticulture Publication
Lebeda A, Astley D (1999) World genetic resources of Lactuca spp. their taxonomy and biodiversity. In: Lebeda A, Křístková E (eds) Eucarpia leafy vegetables 99. Proceedings of the Eucarpia Meeting of Leafy Vegetables Genetics and Breeding, Palacký University, Olomouc, Czech Republic, pp 81–94
Lebeda A, Ryder EJ, Grube R et al (2007) Lettuce (Asteraceae; Lactuca spp.). In: Singh RJ (ed) Genetic resources, chromosome engineering and crop improvement, vol 3. Vegetable crops. CRC Press, Tailor and Francis Group, Boca Raton, pp 377–472
Lettuce Genome Resource (2017) https://lgr.genomecenter.ucdavis.edu
Li Z, Zhao X, Sandhu AK, Gu L (2010) Effects of exogenous abscisic acid on yield, antioxidant, capacities and phytochemical contents of greenhouse grown lettuces. J Agric Food Chem 58:6503–6509
Lee JH, Felipe P, Yang YH, Kim MY, Kwon OY, Sok D, Kim HC, Kim MR (2009) Effects of dietary supplementation with red-pigmented leafy lettuce (Lactuca sativa) on lipid profiles and antioxidant status in C57BL/6J mice fed a high-fat high-cholesterol diet. Br J Nutr 101(8):1246–1254. https://doi.org/10.1017/S0007114508073650
Lindqvist K (1960a) Cytogenetic studies in the srriola group of Lactuca. Hereditas 46:75–151
Lindqvist K (1960b) On the origin of cultivated lettuce. Hereditas 46:319–350
Liu L, Liu Z, Chen H et al (2011) SRAP markers and morphological traits could be used in test of distinctiveness, uniformity, and stability (DUS) of Lettuce (Lactuca sativa) Varieties. J Agric Sci 4(3):227–236
Llorach R, Martínez-Sánchez A, Tomas-Barberán IA et al (2008) Characterisation of polyphenols and antioxidant properties of five lettuce varieties and escarole. Food Chem 108:1028–1038
Mao Y, Wu F, Yu X et al (2014) microRNA 319 a-targeted brassica Rapa ssp. pekinensis TCP genes modulate head shape in Chinese cabbage by differential cell division arrest in leaf regions. Plant Physiol 164(2):710–720 doi: https://doi.org/10.1104/pp.113.228007
Matvieieva NA, Vasylenko MYu, Shakhovsky AM, Kuchuk NV (2009) Agrobacterium-mediated transformation of lettuce (Lactuca sativa L.) with genes coding bacterial antigens from mycobacterium tuberculosis. Cytol Genet 43(2):94–98
Maxted N, Hawkes JG, Ford-Lloyd BV, Williams JT (1997) A practical model for in situ genetic conservation complementary conservation strategies. Chapman and Hall, London pp 339–367
McCabe MS, Garratt LC, Schepers F et al (2001) Effects of PSAG12-IPT gene expression on development and senescence in transgenic lettuce. Plant Physiol 127:505–516
Michelmore RW (1995) Isolation of disease resistance genes from crop plants. Curr Opin Biotech 6:145–152
Michelmore RW, Ochoa OE, Truco MJ (2002) Breeding crisphead lettuce. California Lettuce Research Board Annual Report The Board. Pennsylvania State University pp 51–54
Mohebodini M, Javaran MJ, Mahboudi F, Alizadeh H (2011) Effects of genotype, explant age and growth regulators on callus induction and direct shoot regeneration of lettuce (Lactuca sativa L.). Aust J Crop Sci 5(1):92–95
Mohebodini M, Jalali-Javaran M, Alizadeh H et al (2014) Agrobacterium-mediated transformation of lettuce (Lactuca sativa L.) to express IgG-binding protein A and human pro-insulin as a fusion protein. J Hortic Sci Biotech 89(6):719–725
Moreno-Vázquez S, Ochoa OE, Faber N et al (2003) SNP-based codominant markers for a recessive gene conferring resistance to corky root rot (Rhizomonas suberifaciens) in lettuce (Lactuca sativa). Genome 46:1059–1069
Mou B (2011) Review article. mutations in lettuce improvement. Int J Plant Genom 2011:1–7. https://doi.org/10.1155/2011/723518
Mou B (2008) Lettuce. In: Prohens J, Neuz F (eds) Handbook of plant breeding. Vegetables I: Asteraceae, Brassicaceae, Chenopodicaceae, and Cucurbitaceae. New York: Springer, pp 75–116
Mousavi SH, Hassandokht MR, Choukan R et al (2013) Genetic diversity of Iranian lettuce (Lactuca sativa L.) accessions revealed by cytological traits. Caryologia 66(1):41–48. https://doi.org/10.1080/00087114.2013.780440
Mulabagal V, Ngouajjo M, Nair A et al (2010) In vitro evaluation of red and green lettuce (Lactuca sativa) for functional food properties. Food Chem 118:300–306
Nicolle C, Cardinault N, Gueux E et al (2004) Health effect of vegetable-based diet: Lettuce consumption improves cholesterol metabolism and antioxidant status in the rat. Clin Nutr 23:605–614
Noumedem JAK, Dieussi DE, Hritcu L et al (2017) Lactuca sativa Kuete V (ed) Medicinal spices and vegetables from Africa: Therapeutic potential against metabolic, inflammatory, infectious and systematic diseases, US: Academic Pres, pp 437–449
Nybom H, Weising K, Rotter B (2014) DNA fingerprinting in botany: past, present, future. Investig Genet 5:1
Park J, Choi S, Park S et al (2019) DNA-free genome editing via ribonucleoprotein (RNP) delivery of CRISPR/Cas in lettuce. In: Kuete V (ed) Plant genome editing with CRISPR systems: methods and protocols. Springer, New York, pp 337–354
Pileggi M, Pereira AAM, Silva JS et al (2001) An improved method for transformation of Lettuce by Agrobacterium tumefaciens with a gene that confers freezing resistance. Braz Arch Biol Technol 44(2):191–196
Piosik L, Zenkteler E, Zenkteler M (2015) Development of haploid embryos and plants of Lactuca sativa induced by distant pollination with Helianthus annuus and H. tuberosus. Euphytica 208:439–451
Pitrat M (2012) Vegetables crops in the mediterranean basin with an overview of virus resistance. Adv Virus Res 84:1–29
Porcel R, Aroca R, Azco´n R, Ruiz-Lozano JM, (2006) PIP aquaporin gene expression in arbuscular mycorrhizal Glycine max and Lactuca sativa plants in relation to drought stress tolerance. Plant Mol Biol 60:389–404
Rafalski J, Tingey S (1993) Genetic diagnostics in plant breed: RAPDs, microsatellites and machines. Trends Genet 9:275–280
Ragheb E (2015) Mass selection and individual plant selection as two breeding methods for improving lettuce (Lactuca sativa L.). Alex J Agric Sci 3:213–220
Rai MK, Kalia RK, Singh R et al (2011) Developing stress tolerant plants through in vitro selection-an overview of the recent progress. Environ Exp Bot 71(1):89–98
Rauscher G, Simko I (2013) Development of genomic SSR markers for fingerprinting lettuce (Lactuca sativa L.) cultivars and mapping genes. BMC Plant Biol 13:1–11
Reyes-Chin-Wo S, Wang Z, Yang X et al (2017) Genome assembly with in vitro proximity ligation data and whole-genome triplication in lettuce. Nature Commun 8:14953. https://doi.org/10.1038/ncomms14953
Riar DS, Rustgi S, Burke IC et al (2011) EST-SSR Development from 5 Lactuca Species and Their Use in Studying Genetic Diversity Among L. serriola Biotypes. J Hered 102 (1):17–28
Roberts J, Broadley MR, Pink D et al (2020) Quantitative trait loci (QTLs) linked with root growth in lettuce (Lactuca sativa) seedlings. Mol Breed 40(1):8
Romani A, Pinelli P, Galardi C et al (2002) Polyphenols in greenhouse and open air-grown lettuce. Food Chem 79:337–342
Rubatzky VE, Yamaguchi M (1997) World Vegetables. Chapman and Hall, New York
Ruhlman T, Ahangari R, Devine A et al (2007) Expression of cholera toxin B-proinsulin fusion protein in lettuce and tobacco chloroplasts-oral administration protects against development of insulitis in non-obese diabetic mice. Plant Biotech J 5(4):495–510
Rui S, Qi G, Shuangxi F et al (2020) Analysis of genetic diversity in purple lettuce (Lactuca sativa l.) by SSR markers. Pak J Bot 52(1):181–196
Rulkens AJH (1987) DECGN sla collectie: inventarisatie, paspoort gegevens en enkele richtlijnen voor de toekomst. CGN report, CGN-T, CGN, Wageningen:51
Ryder EJ (1986) Lettuce breeding. In: Basset MJ (ed) Breeding vegetable crops. The AVI Publishing Company Inc, Westport, pp 433–474
Ryder EJ (1991) Salinas 88 lettuce. HortSci 26:439–440
Ryder EJ (1997) Introduction. In: Davis RM, Subbarao KV, Raid RN, Kurtz EA (eds) Compendium of lettuce diseases. Aps Press, St Paul, Minnesota, pp 1–8
Sariçam SK, Kantoğlu YŞ, Ellialtioğl SU (2017) Tissue culture applications in lettuce (Lactuca sativa L). Afro J Pharm Pharmacol 1(2):88–95
Sariçam S, Kantoğlu KY, Ellialtioğlu SS (2017b) Determination of effective mutagen dose for lettuce (Lactuca sativa var. longifolia cv. Cervantes) Seeds. Eurasian J Agric Res 1(2):96–101
Serafini M, Bugianes R, Salucci M et al (2002) Effect of acute ingestion of fresh and stored lettuce (Lactuca sativa) on plasma total antioxidant levels in human subjects. Br J Nutr 88:615–623
Siddiqui MR (2014) Somatic hybridization via protoplasts fusion in Lactuca sativa (Lettuce) and it’s fused product response to culture media. J Agric Res 52(1):1–9
Simko I, Atallah AJ, Ochoa OE et al (2013) Identification of QTLs conferring resistance to downy mildew in legacy cultivars of lettuce. Sci Rep 3:2875
Simko I, Hayes RJ, Truco M, Antonise MRW, R, Massoudi M, (2018) Molecular markers reliably predict postharvest deterioration of fresh-cut lettuce in modified atmosphere packaging. Horticulture Res 5(21):1–13
Simko I, Pechenick DA, McHale L et al (2010) Development of molecular markers for marker-assisted selection of dieback disease resistance in lettuce (Lactuca sativa). Acta Hort 401–408
Simko I (2008) Development of EST-SSR markers for the study of population structure in lettuce (Lactuca sativa L.). J Hered 100(2):256–262
Sochor M, Jemelková M, Doležalová I (2019) Phenotyping and SSR markers as a tool for identification of duplicates in lettuce germplasm. CJGPB 55(3):110–119
Souza MM, Resende LV, Menezes D et al (2008) Genetic variability for agronomic characteristics in lettuce progenies with heat tolerance. Hortic Bras 26:354–358
Stebbins GL (1937) The scandent species of Prenanthes and Lactuca in Africa. Bulletin Du Jardin Botanique De L’état a Bruxelles 14:333–352
Still DW (2007) Lettuce. In: Kole C (ed) Genome mapping and molecular breeding in plants, vol 5. vegetables. Springer, Berlin, pp 127–140
Taniguchi T, Sato T, Maeda K, Maeda E (1990) Microscopic observations of fusion process of rice and lettuce protoplasts. Curr Plant Sci Biotech Agric 8:281–298
Tardin FD, Júnior ATA, Pereira MG et al (2003) Genetic diversity and determination of the optimum number of RAPD markers in lettuce (Lactuca sativa L.). Acta Scientiarum: Agronomy Maringá 25(1):1–5
Tashi D, Gupta AJ, Ahmed N (2010) Variability, heritability and genetic advance in lettuce. Indian J Hortic 67:193–196
Till BJ, Cooper J, Tai TH et al (2007) Discovery of chemically induced mutations in rice by TILLING. BMC Plant Biol 7:19
Truco MJ, Antonise R, Lavelle D et al (2007) A high-density, integrated genetic linkage map of lettuce (Lactuca spp.). Theor Appl Genet 115:735–746
Truco MJ, Ashrafi H, Kozik A et al (2013) An ultra-high-density, transcript-based, genetic map of lettuce. G3 (Bethesda) 3:617–631
Uwimana B, Smulders MJM, Hooftman DAP et al (2012) Hybridization between crops and wild relatives: the contribution of cultivated lettuce to the vigour of crop–wild hybrids under drought, salinity and nutrient deficiency conditions. Theor Appl Genet 125:1097–1111
Uwimana B (2011) A genetic analysis of the introgression process from cultivated lettuce (Lactuca sativa L.) to wild prickly lettuce (L. serriola L). PhD thesis, Wageningen University, The Netherlands
Van de Wiel C, Arens P, Vosman B (1998) Microsatellite fingerprinting in lettuce (Lactuca sativa L.) and wild relatives. Plant Cell Rep 17:837–842
Van de Wiel C, Arens P, Vosman B (1999) Microsatellite retrieval in lettuce (Lactuca sativa L.). Genome 42:139–149
Van Treuren R, Van Hintum JL (2009) Comparison of anonymous and targeted molecular markers for the estimation of genetic diversity in ex-situ conserved Lactuca. Theor Appl Genet 119:1265–1279
Vermeulen A, Desprez B, Lancelin D, Bannerot H (1994) Relationship among Cichorium species and related genera as determined by analysis of mitochondrial RFLPs. Theor Appl Genet 88:159–166
Walley PG, Hough G, Moore JD et al (2017) Towards new sources of resistance to the currant-lettuce aphid (Nasonovia ribisnigri). Mol Breed 37(4):1–18
Whitaker TW (1969) Salads for everyone - a look at the lettuce plant. Econ Bot 23:261–264
Williams M (2012) Organic lettuce and leafy greens. University of Kentucky Cooperative Extension service.
Witsenboer H, Vogel J, Michelmore RW (1997) Identification, genetic localization, and allelic diversity of selectively amplified microsatellite polymorphic loci in lettuce and wild relatives (Lactuca spp.). Genome 40:923–936
Woo JW, Kim J, Kwon SI, Corvalan C et al (2015) DNA-free genome editing in plants with preassembled CRISPRCas9 ribonucleoproteins. Nat Biotechnol 33:1162–1164. https://doi.org/10.1038/nbt.3389
Yamamoto T, Nishikawa A, Oeda K (1994) DNA polymorphisms in Oryza sativa L. and Lactuca sativa L. amplified by arbitrary primed PCR. Euphytica 78:143–148
Zohary D (1991) The wild genetic resources of cultivated lettuce (Lactuca sativa L.). Euphytica 53:31–35
Acknowledgements
The authors would like to thank bot Botany and Microbiology Department, and Biotechnology Department, Faculty of Science, Helwan University.
Funding
No funding received for this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have nothing to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hassan, M.N., Mekkawy, S.A., Mahdy, M. et al. Recent molecular and breeding strategies in lettuce (Lactuca spp.). Genet Resour Crop Evol 68, 3055–3079 (2021). https://doi.org/10.1007/s10722-021-01246-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-021-01246-w