Abstract
Current knowledge of wild Lactuca L. species, their taxonomy, biogeography, gene-pools, germplasm collection quality and quantity, and accession availability is reviewed in this paper. Genetic diversity of Lactuca spp. is characterized at the level of phenotypic and phenological variation, variation in karyology and DNA content, biochemical traits, and protein and molecular polymorphism. The reported variation in reaction to pathogens and pests of wild Lactuca spp. is summarized, including the viral pathogens (Lettuce mosaic virus-LMV, Mirafiori lettuce virus/Lettuce big vein virus-LBV, Beet western yellows virus-BWYV, Tomato spotted wilt virus-TSWV, Cucumber mosaic virus-CMV, Lettuce necrotic stunt virus-LNSV), bacterial pathogens (corky root-Rhizomonas suberifaciens, bacterial leaf spot-Xanthomonas campestris pv. vitians), fungal pathogens (downy mildew-Bremia lactucae, powdery mildew-Golovinomyces cichoracearum, anthracnose-Microdochium panattoniana, stemphylium leaf spot-Stemphylium spp., sclerotinia drop-Sclerotinia spp., verticillium wilt-Verticillium dahliae, fusarium wilt-Fusarium spp., pythium wilt-Pythium tracheiphylum, P. uncinulatum), nematodes (potato cyst nematode-Globodera rostochiensis, root-knot nematode-Meloidogyne spp., incognita, hapla, javanica, enterolobii), insects and mites (the green lettuce aphid-Nasonovia ribisnigri, the green peach aphid-Myzus persicae, the potato aphid-Macrosiphum euphorbiae, leafminer-Liriomyza spp., L. langei). The approaches used to exploit wild Lactuca spp. in lettuce breeding (interspecific hybridization, cell and tissue culture, transformation) are dicussed, and known examples of lettuce cultivars with traits derived from wild Lactuca spp. are described.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The potential of wild Lactuca species to be used in lettuce breeding is being demonstrated by means of classical biology and modern approaches and the study of their diversity has been a subject of theoretical research and practical application during the last 25 years (Ryder 1999; Lebeda et al. 2007c; Mou 2008). The currently available knowledge of wild Lactuca species as donors (sources) of traits important in lettuce breeding was thoroughly analyzed in our previous paper focused on wild Lactuca germplasm (Lebeda et al. 2009a, b). The main aim of this paper is to critically summarize the available information about interactions between wild Lactuca species and the most important lettuce pathogens and pests from the viewpoint of their resistance and their potential exploitation and utilization in lettuce breeding.
Taxonomy of Lactuca spp.
The genus Lactuca L. (family Compositae/Asteraceae) is composed of one cultivated species-lettuce (Lactuca sativa L.), and about 100 wild Lactuca spp. of which nearly 95 % are autochtonous in Asia and Africa (Lebeda et al. 2007c). Species are arranged into seven sections and two geographic groups (Table 1). This broader generic concept summarized by Lebeda et al. (2007c) should be critically re-considered with regard to the molecular data on phylogenetic relationships among Lactuca species (Koopman et al. 1998, 2001).
Eco-geographic characteristics
The genus Lactuca L. comprises annual, biennial or perennial herbs and rarely shrubs with various ecological requirements. The species L. serriola, L. saligna and L. virosa are weedy and occur on waste places and ruderal habitats, along roads, highways and ditches, L. perennis, L. viminea, L. graeca, L. tenerrima are calciphilous plants and colonise limestone and dolomite areas, mostly rocky slopes. Endemic lianalike species are found in rain forests of East Africa (Lebeda et al. 2001, 2004b, 2007c).
The greatest diversity of Lactuca species is confined to the Mediterranean basin and Southwest Asia (Doležalová et al. 2001; Beharav et al. 2008; Kitner et al. 2008; Lebeda et al. 2001, 2009a, b). The occurrence of valuable germplasm is expected in the Central and South Africa, Southwest and Central Asia, and North America regions (Lebeda et al. 2007c, 2011, 2012a).
Lactuca germplasms and their availability
The concept of conservation and management of wild crop relatives, and conservation priorities were proposed by Maxted et al. (2008) and Ford-Lloyd et al. (2008). The linking of in-situ and ex-situ conservation with the use of wild crop relatives is the leading principle of their conservation and management (Maxted and Kell 2008). Access to wild genetic resources and the possibility to explore and exploit them depend upon the successful and reasonable protection of wild species in–situ, i.e. in their natural habitats (Iriondo and De Hond 2008), upon the complex study of wild species in natural habitats and upon the possibility to exchange information and biological material (Azzu and Collette 2008). These research activities are regulated by national policies and the international conventions and protocols, e.g. the recent “Nagoya Protocol on Access to Genetic Resources and the Fair and Equitable Sharing of Benefits Arising from their Utilization to the Convention on Biological Diversity” adopted by the Conference of the Parties to the Convention on Biological Diversity at its tenth meeting on 29 October 2010 in Nagoya, Japan (http://www.cbd.int/abs/about/ from 16 January 2013).
A recent inventory of The International Lactuca Database (ILDB) with passport data for 11,643 Lactuca accessions, and of the Dutch national Lactuca germplasm collection (van Treuren and van Hintum 2009; van Treuren et al. 2011) confirmed the conclusions regarding gaps in collection structures reported previously by Lebeda and Boukema (2001) and Lebeda et al. (2004a, 2009a, b). Wild Lactuca germplasms are not adequately conserved by official gene banks and the species spectrum and world geographic distribution of the genus are not adequately represented in their germplasm collections (Lebeda et al. 2004a, 2007b, c).
Basic errors in the taxonomic status of wild Lactuca accessions as declared by gene banks were found during recent studies (Doležalová et al. 2004, Lebeda et al. 2007b) and duplicates between and within germplasm collections were identified (van Hintum and Boukema 1999, Doležalová et al. 2007; Sretenović-Rajičić et al. 2008). In addition, we are missing basic information on wild Lactuca germplasm resistance to the most important diseases and pests of lettuce (Lebeda et al. 2009a, b).
Gene pools of Lactuca spp.
The categorization of many Lactuca spp. into gene pools based on their crossing ability and fertility of F1 hybrids is still questionable and needs to be clarified. The primary gene pool of cultivated lettuce L. sativa comprises its cultivars and landraces, wild L. serriola, L. aculeata, L. altaica, L. azerbaijanica, L. georgica, L. scarioloides, and L. dregeana (Lebeda et al. 2007c). The categorization of L. saligna and L. virosa to secondary and tertiary gene pools is not resolved yet. Koopman et al. (1998) suggested that section Lactuca subsection Lactuca comprises the primary and secondary gene pool, while the sections Phaenixopus, Mulgedium and Lactucopsis include the tertiary gene pool (Table 1). Modern lettuce breeding has been mainly based on the utilization of wild Lactuca germplasm from the primary gene pool (L. serriola), however more recently it has shifted to the exploitation of secondary and tertiary Lactuca germplasm (Maisonneuve et al. 1995; Jeuken and Lindhout 2004). The main reason for this strategy is to broaden the genetic variation of cultivated lettuce by interspecific hybridization (Chupeau et al. 1994; Jeuken et al. 2001), including the introduction of a new and broader spectrum of resistances to diseases and pests (Jeuken and Lindhout 2002; Lebeda et al. 2002, 2007c, 2009a, b)
Genetic diversity of Lactuca spp.
Phenotypic variation
Descriptor lists as a tool for the correct taxonomic determination of wild Lactuca genetic resource accessions and for a definition of both interspecific and intraspecific variation have been produced at the national (Boukema et al. 1990; McGuire et al. 1993) and international levels (Doležalová et al. 2002, 2003a).
A large degree of variation in plant phenotypes has been described in greenhouse experiments among samples of two world-wide distributed species, Lactuca serriola (Doležalová et al. 2005; Lebeda et al. 2007a, 2011; Novotná et al. 2011), and Lactuca saligna (Křístková et al. 2007a; Beharav et al. 2008). This results from the evolutionary adaptation of plants under different climatical and ecological condition in their original habitats in different countries from Europe, Near East and North America. In contrast, a low level of phenotypic variation within Lactuca aculeata reflects the relatively limited distribution area of this species (Beharav et al. 2010a).
A high level of intraspecific variation is reported for many other species, e.g. for L. virosa (Feráková 1977) and has also been recently observed by the authors of this paper. However, this variation was not described in relation to the ecogeographic conditions and distribution of the accessions. The intraspecific classification of this and many other Lactuca species has not as yet been critically described. Recent broad application of wild Lactuca species in lettuce breeding and their influence on L. sativa phenotypic variation need a new treatment arrising from the previous one (Lebeda et al. 2007c) based on application of various approaches (i.e. phenotyping, digital image analysis, numerical taxonomy, molecular polymorphism etc.).
Variation in phenology features
A high level of variation in phenological characteristics within the genus Lactuca was recorded among accessions grown in greenhouse experiments. Substantial differences in the time of flowering were recorded between samples of L. serriola originating from various countries (Doležalová et al. 2005; Lebeda et al. 2007b, c, 2011, unpublished results). A similar phenomenon was recorded for L. saligna samples (Křístková et al. 2011). Differences in developmental rates of plants, which are influenced by the original eco-geographic conditions of samples (Lebeda et al. 2001), persist when plants are cultivated in uniform environmental conditions and have a genetic basis (Křístková et al. 2007b).
Karyology and DNA contents variation
Perennial wild Lactuca species of Europe and the Himalayas have haploid chromosome number n = 8; the haploid chromosome number n = 9 characterizes the majority of European and Mediterranean species, and species from the Middle East, Africa and India; species autochtonous from Canada to Florida possess the haploid chromosome number of n = 17 (Feráková 1977, Lebeda and Astley 1999). However, the chromosome numbers of numerous Lactuca species are not known (Lebeda and Astley 1999) and the actual chromosome numbers of many North American species may differ from the reported data (Doležalová et al. 2003b).
Chromosomal studies (Matoba et al. 2007) and approaches combining analysis of karyotype and variation in relative DNA content serve as tools for distinguishing some Lactuca species (Koopman 1999, 2000; Doležalová et al. 2003b), characterization of their evolutionary relationships (Koopman and De Jong 1996), and intraspecific variation (Koopman 2002). The relative DNA content was analysed for large sets of L. serriola and L. saligna samples originating from different eco-geographical conditions in Europe, Near East and North America (Lebeda et al. 2004c, 2007c, 2011). However, there was little variation and it seems that Lactuca species are highly conservative in DNA content at the intraspecific level.
Biochemical trait variation
Nearly 10 % of plant species, including Lactuca spp. produce latex which contains complex mixtures of terpenoids, phenolics, proteins, glycosides and alkaloids (Agrawal and Konno 2009). The most important subgroups of sesquiterpenoids within the tribe Cichorieae (Asteraceae) are costus lactone type quaianolides and lactucin derivates, and they make up large numbers of the total of 360 different sesquiterpene lactones and precursorss reported in the tribe (Zidorn 2008). Based on sesquiterpene profiles the 31 genera of the tribe Cichorieae form seven main clusters, and within the second group with eleven genera the genus Lactuca is very close to the genera Notoseris and Cichorium. The integration of chemosystematic data to the botanical systematics is limited by the lack of standard specimens for verifying the identity of plant material (Zidorn 2008). When correctly identified plant material is available, the results of chemical analyses bring new light to evolutionary concepts and taxonomical relationships between different Lactuca spp. (Sessa et al. 2000; Kisiel and Michalska 2009; Michalska and Kisiel 2009, 2010; Lebeda et al. 2009a, b; Michalska et al. 2009; Beharav et al. 2010b).
The HPLC profile of sesquiterpene lactones from latex of several L. sativa cultivars differed to those of L. serriola and L. virosa genotypes resistant to important races of lettuce downy mildw (Bremia lactucae). Although this resistance was not corrrelated to the “SL” profile, heritability of this profile was demonstrated by analysis of progeny from a cross between L. sativa and L. virosa (Sessa et al. 2000).
Pharmacological exploitation of some chemical compounds in wild Lactuca species (e.g. sesquiterpene lactones, phenolics and glucosides, flavonoids) (Rees and Harborne 1984; Kisiel and Barszcz 1998; Kisiel and Zielinska 2000; Chen et al. 2007; Kim et al. 2007) or plant-produced antigens (Pniewski 2013) is another goal of studies aimed at their detection and characterization. This research is stimulated by the promising results of analgesic and sedative activities of lactucin from Cichorium intybus, a species chemotaxonomically closely related to lettuce (Wesolowska et al. 2006).
Most studies have focused on latex as a substance that reduces herbivory or the preference or performance of herbivores (Agrawal and Konno 2009). Latex from the resistant variety of lettuce ‘Valmain’ inhibited feeeding of Diabrotica balteata when painted on leaves of lima bean, conversely the latex from the susceptible variety ‘Tall Guzmaine’ did not inhibit feeding (Huang et al. 2003). To our knowledge, there is no report of antiherbivoral activities of latex from wild Lactuca species.
Protein and molecular polymorphism
Genotyping with molecular markers for genetic diversity detection, assesment of population structure, selection of desirable genotypes for lettuce breeding, mapping of genes, identification and variation of resistance genes has become common place in modern exploitation of wild lettuce progenitors. The studies related to use of protein and molecular markers in Lactuca spp. germlasm collections have been reviewed by Dziechciarková et al. (2004). In the present contribution we want to summarise recent progress on the comprehensive molecular based research on the genus Lactuca spp. The survey of studies including wild Lactuca species as well as studies using offspring from crosses between cultivated lettuce and wild progenitors and mapping populations derived from these crosses are presented in Table 2.
In general, in the first decade of 21st century there has been a dramatic shift in the number of papers based on isozyme studies (Lebeda et al. 2009a, b, 2012a, b) to more advanced studies utilising microsatellite and AFLP markers (e.g. Kitner et al. 2008; van de Wiel et al. 2010; Lebeda et al. 2009a, b), NBS profiling (e.g. van Treuren and van Hintum 2009) and high-resolution DNA melting analysis (Simko et al. 2009, 2010). However, during the last 2 years there has been an increase in the number of papers using new high throughput marker technologies based on single nucleotide polymorphism (SNP) or single position polymorphism (SPP) arrays on Affymetrix and Illumina genechips (e.g. Kwon et al. 2012; Stoffel et al. 2012; Uwimana et al. 2012b, c).
There are several studies by Koopman et al. (1998, 2001) and Koopman (2002) using molecular markers primarily describing relationships among Lactuca species and related genera, which are based on ITS1 sequencing and AFLP’s. Several other studies have commented on the phylogenetic relationships between Lactuca species as well, but either this has not been a primary aim of the study, or has been restricted to a more narrow frame of Lactuca genetic pools (Matoba et al. 2007; Yang et al. 2007; Stoffel et al. 2012) (Table 2).
Several types of molecular markers have been applied in studies of genetic variation in natural populations and for germplasm maintenance and characterisation (Table 2). These studies were frequently carried out with microsatellites (Lu et al. 2007; Riar et al. 2011; Uwimana et al. 2012a) or AFLP markes (Kitner et al. 2008; Kuang et al. 2008; Lebeda et al. 2009a, b) or combination of both methods, sometimes extended with additional markers (van Treuren and van Hintum 2009; van de Wiel et al. 2010; Hooftman et al. 2011). Several studies describing genetic diversity were primarily foccused on marker development, espetially on EST- or genomic-microsatellite design (Simko 2009; Rauscher and Simko 2013). These SSR-based markers with publicly available primer sequences provide an important tool for researchers for future studies of wild lettuce populations. Some of these SSR markers are linked to herbicide resistance genes 2,4-D and ALS resistance (Riar et al. 2011). In fact, the number of population-based studies is limited and was mainly performed with AFLP’s or non-publically available SSR’s, on lettuce germplasm pseudopopulations originating from a larger geographical scale and/or with a sampling period of several years (van de Wiel et al. 2010; Lebeda et al. 2009a, b, 2012b; Uwimana et al. 2012a) than e.g. comparisons of true populations of local character originating from several regions/countries and sampled within a short period of time. To conclude, we expect that current advances in lettuce genomics (Kwon et al. 2012; Stoffel et al. 2012) will stimulate researchers to use these comercially available genechips based on SNP or SPP features for fast and cost-efficient population-genetic studies in the near future.
Several genetic linkage maps have been published for lettuce. Truco et al. (2007) presented a consensus map of 2,744 markers integrating seven intra- and inter-specific mapping populations and included information from five previously published genetic maps (Kesseli et al. 1994; Witsenboer et al. 1997; Waycott et al. 1999; Johnson et al. 2000; Jeuken et al. 2001). These maps are continuously updated according to marker developments (Schwember and Bradford 2010; Argyris et al. 2011; Aruga et al. 2012; Rauscher and Simko 2013). Such studies are closely associated to QTL studies, design of molecular markers for marker-assisted selection (Simko et al. 2009, 2010, 2011), detection of interspecific hybrids, distribution of crop alleles in natural populations (Uwimana et al. 2012b, c), and finally studies analysing and describing the background of resistance gene clusters, their identification and variability screening (Kuang et al. 2008; McHale et al. 2009).
To conclude, the genome of lettuce has been sequenced using ‘next-generation’ DNA sequencing, the sequenced genome has been assembled and annotation is underway (Michelmore 2012).
Variation of wild Lactuca spp. in reaction to pathogens and pests
Viral pathogens
The viral pathogens are after fungi the most economically important pathogens of plants. The impact of virus infection is seen in a reduction of yield, decrease quality of size, shape, taste, structure, composition (sugar content) of plants, decrease of viability (sensitivity to dry, cold), predisposition to infection of other pathogens, short shelf life and decrease of fertility. All viruses are obligate parasites that depend on the cellular machinery of their hosts to reproduce (Gergerich and Dolja 2006). Most plant viruses are transmitted by passive transmission from plant to plant and active transmission from infected to healthy plants by a living organism termed a vector. Plant-feeding arthropods, nematodes and plant-parasitic fungi are the major types of vector organisms for plant viruses (Walkey 1991). All types of plant viruses are important disease causing agents and are responsible for losses in crop yields and quality in all parts of the world. Among the most serious lettuce viruses are: Lettuce mosaic virus, Mirafiori (big vein) lettuce virus, Beet western yellows virus, Tomato spotted wilt virus, Cucumber mosaic virus and Lettuce necrotic stunt virus. Lettuce is at risk of infection by these viruses and production of new resistant cultivars is a priority. In breeding programmes crossess between cultivars and wild species such as Lactuca serriola, L. saligna, L. virosa and L. perennis offer a way of introgressing new resistances.
Lettuce mosaic virus (LMV)
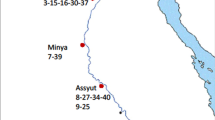
Lettuce mosaic virus has been a serious worldwide disease problem in lettuce and a few other leafy vegetable species, for a long time (Ryder 2002). It was as first described in Florida (Jagger 1921) and now is distributed worldwide, probably because the virus is seed transmitted and lettuce seeds have been exchanged internationally over many years (Dinant and Lot 1992). The worldwide distribution of LMV includes Europe, North and South America (Mexico, USA, Argentina, Brazil, Uruguay), the West Indies (Bermuda), Africa, the Middle East (Egypt, Israel, Iraq, Iran, Jordan and Turkey), Asia (China and Japan) and Oceania (Australia, Tasmania, New Zealand) (German-Retana et al. 2008).
LMV belongs to the genus Potyvirus of the family Potyviridae, which is seed-borne in lettuce and disseminated by aphid vectors-Myzus persicae, Macrosiphum euphorbiae, Aphis gossipii. The characteristic symptoms on susceptible lettuce cultivars are dwarfism, mosaic, distortion and yellowing of the leaves with sometimes a much reduced heart (failure to form heads). The differences in virus strains, cultivars and the physiological stage of the host at the moment of the attack cause different symptom severity; from a very slight discoloration of the veins to severe necrosis leading to death of the plant (German-Retana et al. 2008). The genomic organization of LMV is typical of potyviruses, with a single positive-sense genomic RNA of 10,080 nucleotides encapsidated as flexuous rods (Krause-Sakate et al. 2002). The viral genomic RNA has a viral encoded protein covalently linked at the 5′end, a poly-A tail at the 3′end, and contains a single open reading frame (ORF) which encodes a large polyprotein with 3,255 amino acids (Revers et al. 1997a). This polyprotein undergoes self-cleavage as it is translated, generating 8–10 viral proteins (Shukla et al. 1994; Revers et al. 1997b).
Three phylogenetic groups of LMV isolates were discriminated, correlating with geographical origin of the isolates rather than with their pathogenicity. The largest group includes isolates from western Europe and California. A second group includes three isolates from Greece whereas the third group consists, so far, of a single isolate from the Yemen Arab Republic (Revers et al. 1997b). LMV isolates have been classified into four pathotypes, according to their virulence on lettuce varieties carrying the three resistance or tolerance genes mo (mo11, mo12) and Mo2, which were identified in L. sativa cultivars (Pink et al. 1992a; Bos et al. 1994) and LMV genes Mo3 and Mo4, which are described in L. virosa sources but which are difficult to introgress and not well-characterized. These dominant genes have not currently been used in the field.
In Brazil, and in most European countries, LMV has been controlled through the use of resistant cultivars (Krause-Sakate et al. 2001). Two sources of resistance were identified in the late 1960’s – the first recessive gene mo1 1 (formely named g) in Argentina, in a Latin- type cultivar Gallega de Invierno (Bannerot et al. 1969) and European lettuce breeders used the Gallega source of resistance to incorporate the g gene in numerous varieties of lettuce, including butterhead, Batavia, cos and crisphead types (Pink et al. 1992b). Later in the USA the recessive gene mo1 2 (previously mo) was identified in three Egyptian wild Lactuca sativa lines (Ryder 1970), this recessive gene mo has been used by North American breeders, who introduced it into crisphead and cos types of lettuce. Two of these genes mo1 1 and mo1 2, are recessive and are believed to be either closely linked or allelic (Pink et al. 1992b) and which encode alleles of the cap-binding protein, elF4E. ElF4E-this identification was based on three converging lines of evidence: (1) allelic sequence co-variation between the elF4E gene and mol 1 and mol 2 resistance status of plants; (2) co-segregation of mutations in the elF4E gene and the mol 1 and mol 2 resistance status and finally (3) functional complementation using a viral transient expression vector to vector to restore LMV susceptibility in mol 1 -or mol 2 carrying lettuce plants using the elF4E allele from susceptible plants (Nicaise et al. 2003). The resistant alleles of the eIF4E gene in lettuce, mo1 1 and mo1 2, are currently the only genetic determinants used to protect lettuce crops from LMV; the third resistance (dominant) gene Mo2, found in the cv. Ithaca (Pink et al. 1992a, b) a is not effective in practice for LMV control, because it is overcome by most LMV isolates.
More of studies were done on cultivars of L. sativa (the most frequently tested cultivars were: Trocadéro, Mantilia, Floribibb, Ithaca, Salinas 88, and Vanguard). Among the most frequently tested isolates of lettuce mosaic virus on these cultivars were isolates LMV- 0, LVM-1, LMV-9, LMV-E, LMV-13 and AF-199. Additional sources of resistance to lettuce mosaic virus are known in accessions of Lactuca virosa and Lactuca serriola (Table 3).
Mirafiori lettuce virus (Lettuce big vein virus, LBV)
Lettuce big vein disease (LBVD) was first described in California (Jagger and Chandler 1934), and it occurs widely in regions of the world with temperate or Mediterranean-type climates (Coutts et al. 2004). LBVD is associated with a complex of two viruses, Lettuce big-vein associated virus (LBVV and LBVaV, genus Varicosavirus) and Mirafiori lettuce big vein virus (MLBVV, Ophiovirus) (Rogero et al. 2000). Its natural host range is limited to lettuce (Lactuca sativa), endivie (Cichorium endivia) and sow thistle (Sonchus oleraceus). The vector for both viruses is the root-infecting fungus Olpidium brassicae (Coutts et al. 2004).
Both LBVaV and MLBVV have segmented ssRNA genomes, and their virions contain RNA molecules of both polarities. The LBVaV geonome contains two RNA segments-RNA1 is 6797 nucleotide length with a single large open reading frame (ORF), and RNA2 has a slightly smaller size than RNA1 (6081 nt) having coding capacities for five ORFs (Navarro et al. 2005). LBVV particles of virus are fragile, rather rigid rods 320 to 360 nm in length and 18 nm in diameter, with central canal and an obvious helix of pitch = 5 nm, MiLV particles, like those of recognized ophioviruses, are highly kinked filaments =3 nm in diameter that form masses of two distinct sizes but of undetermined contour length; they probably form closed circles, because free DNAs are very seldom seen (van Regenmortel et al. 2000)
The mechanism of resistance in cultivated lettuce is not known, and more research is needed to determine the relative role of virus resistance and symptom expression in big vein resistance. Among wild relatives of lettuce, only accesions of L. virosa have demonstrated a complete lack of symptom expression in inoculation trials (Bos and Huijberts 1990) (Table 3). L. virosa accession IVT280 was identified as 100 % asymptomatic in the greenhouse inoculation trials. Analysis by RT-PCR demonstrated no viral amplification, indicating apparent immunity in this accession (Hayes et al. 2006) Currently, no genotype of L. sativa has been identified as immune to big vein (Ryder and Robinson 1995), cultivars Pacific, Thompson, Margarita and Pavane are considered resistant (Ryder and Robinson 1995; Hayes et al. 2006). Crossing between L. virosa and L. sativa cultivars was dificult to perform (Hayes et al. 2004), nevertheless introgression of big vein tolerance from L. virosa to cultivars of lettuce has been successful (Hayes et al. 2004; Hayes and Ryder 2007).
Beet western yellows virus (BWYV), Turnip yellows virus (TuYV)
Beet western yellows virus (BWYV) was originally identified in the USA during the late 1950s as an important virus causing stunting and chlorosis in a wide range of plant species resulting in yield losses in crops such as sugar beet, spinach, lettuce and turnip (Duffus 1961). Beet western yellows virus has been associated with lettuce production since at least the 1950s, when it and the complex of virus diseases affecting spring crops of lettuce were referred to as June Yellows (Davis et al. 1997). BWYV belongs to the genus Polerovirus in the family Luteoviridae and recently a BWYV isolate, which does not infect sugar beet was renamed Turnip yellows virus (TuYV) (Stevens et al. 2005).
BWYV is transmitted by aphid vectors-especially Myzus persicae (Sulz.), Macrosiphum euphorbiae (Thos.) and induced symptoms are chlorotic (yellow) symptoms, which are first observable at the tips or margins of leaves and soon spread to cover whole leaves. Interveinal chlorosis first occurs in older leaves and progress acropetally, followed by necrosis (Maisonneuve et al. 1991; Hampton et al. 1998).
The genome of BWYV is composed of a single-stranded plus-sense RNA, approximately 5,6 kb in length. The genome contains six large open reading frames (ORFs, ORF0-ORF5), a short 5′- untranslated region (UTR), a 3′-UTR without tRNA-like or poly (A) structure, and an intergenic non-coding region (NVR) between ORF2 and ORF3 about 200nt (Stevens et al. 2005).
Lettuce necrotic yellows virus (LNYV)
LNYV was first found in 1954 in Australia (Stubbs and Grogan 1963). LNYV is the type species of the genus Cytorhabdovirus, members of which are characterised by accumulation of enveloped virions, which is transmitted by Hyperomyzus lactucae L.
Lettuce plants naturally infected with LNYV acquire a dull green appearance, the young leaves developing bronzing and necrosis, especially along the veins, and older leaves become chlorotic or mottled and plants often die (Fry et al. 1972). Among the tested wild species are L. serriola and L. saligna (Table 3) and some accessions show a resistant response.
The LNYV genome consists of a monopartite, negative-sense, single-stranded RNA of 12–15 kb, which encodes five functionally conserved proteins (Dietzgen et al. 2006). The physical map of the LNYV genome is 3′ leader – N – P- 4b – M- G – L – 5′trailer, where N is the nucleocasid gene, P is phosphoprotein gene, 4b encodes a putative movement protein, M is the matrix protein gene, G is the glycoprotein gene and L is the polymerase gene (Wetzel et al. 1994).
Lettuce chlorotic virus (LCV)
Lettuce chlorotic virus (LCV) is a member of the rapidly emerging genus Crinivirus, family Closteroviridae (Duffus et al. 1996). It is transmitted by silverleaf whitefly Bemisia tabaci and B. argentifolii with about the same efficiency. This is a major difference between LCV and Lettuce infectious yellows virus (LIYV) since LIYV is transmitted very inefficiently by B. argentifolii (Davis et al. 1997; Wintermantel 2004).
LCV has a large bipartite RNA genome encoding several open reading frames (at least 13 ORFs). RNA1 encodes functions involved in virus replication, while RNA2 encodes up to 7 ORFs involved in virion assembly, vector transmission and other functions, many of which remain to be determined (German-Retana et al. 1999). Virions are encapsidated into long flexuous rods averaging between 650 to 900 nm in length.
Lettuce chlorotic virus resistance has only been assessed in L. sativa accessions.
Tomato spotted wilt virus (TSWV)
Tomato spotted wilt virus (TSWV) belongs to the genus Tospovirus, family Bunyaviridae, it is a one of the most widely spread plant viruses and the causal agent of economically important yield losses in many crops. TSWV was first found in 1915 in Australia (Brittlebank 1919). Since then, its known host range has increased to over 900 dicotyledonous and monocotyledonous plant species worldwide (Peters and Goldbach 1998). Many horticultural crops and weeds are hosts. TSWV is transmitted by several thrips species, of which the western flower thrips (Frankliniella occidentalis) is the most efficient vector (Hobbs et al. 1993). Infection reservoirs from which TSWV spreads to susceptible crops include nearby plantings of TSWV-susceptible crops, volunteer crop plants and weeds (Cho et al. 1989; Groves et al. 2002). TSWV is readily transmitted mechanically from sap of naturally infected plants. For manual inoculation Nicotiana tabacum, N. glutinosa and N. bethamiana, which develope large necrotic local lesions followed by systemic mosaic and necrosis are used to provide inoculum (Parella et al. 2003).
TSWV virions are 80–120 nm diameter, spherical, enveloped, and studded with surface projections composed of two glycoproteins G1 and G2. Virion composition is 5 % nuclei acid (RNA), 70 % protein, 5 % carbohydrate, and 20 % lipid. The genome consists of three negative or ambisense ssRNA species designated as S (2.9 kb), M (4.8 kb) and L (8.9 kb) (Parella et al. 2003).
Introgression of genes for resistance into to lettuce cultivars is a possible strategy for control of Tomato spotted wilt virus. However, screening of some wild Lactuca species (L. serriola, L. virosa and L. floridana) have shown only susceptible reaction (Hobbs et al. 1993; Parella et al. 2003). Resistance was recorded only in one accession of L. serriola by Groves et al. (Groves et al. 2002; Table 3), but without any detailed specification.
Cucumber mosaic virus (CMV)
Cucumber mosaic virus (CMV) is the type species of the genus Cucumovirus, family Bromoviridae. CMV occurs worlwide and is a very important disease agent in temperate, tropic and subtropic regions of the world. It is a virus with a very wide host range including plants from approximately 365 genera and at least 85 families (Roossinck et al. 1999). CMV is an important pathogen of many vegetable crops and is the target of breeding programs for resistance. The Cucumoviruses are transmitted by aphids (especially Myzus persicae, Aphis gossypii), which ensures a multiplicity of inoculation sufficient to reliably establish infection. Symptoms of CMV infection in lettuce consist of leaf mottling, severe roughness of the leaf and occasional necrosis within the leaf tissue. Plants are usually stunted if infected at an early stage of development (Zitter and Murphy 2009).
CMV consists of three spherical particles, each approximately 28 nm in diameter (Zitter and Murphy 2009). The genome is divided into three plus-sense, single-stranded, RNA molecules, designated RNA 1, RNA 2 and RNA 3. Each RNA molecule is enclosed within a protective protein coat with each being a distinct single spherical-shaped particle. CMV contains five open reading frames (ORFs). These can be used for phylogeny estimation of the species of the Cucumovirus genus (indicating evolutionary histories for each RNA strongly supporting the occurrence of re-assortment in the evolutionary history of the genus (Roossinck 2002).
CMV resistance has been derived from L. saligna (Table 3), introgression of resistance in to lettuce was done by backcrossing with L. sativa. By the F7 generation of cultivar Montello × (Vanguard 75 × L. saligna PI 261653) (Tamaki et al. 1995), and Lactuca saligna × L. sativa (Saladcrisp) (Provvidenti et al. 1980) the introgression was successful, and these lettuce lines are resistant to CMV.
Lettuce necrotic stunt virus (LNSV)
LNSV causes lettuce dieback, a disease resulting in stunting, necrosis, and lack of marketability in lettuce, it is likely that it has been present under the name brown blight since the 1920s (Wintermantel and Anchieta 2012). LNSV can infect lettuce through the soil in the absence of fungal vectors. Fields with high disease incidence are usually poorly drained and variation in soil salinity influences LNSV infection of lettuce (Wintermantel et al. 2003). Lettuce necrotic stunt virus is caused by several members of the soilborne virus family Tombusviridae, including the type member, Tomato bushy stunt virus (TBSV), and Lettuce necrotic stunt virus (LNSV) (Obermeier et al. 2001). Tombusviridae is a relatively large and diverse family of soil-borne viruses that have single-stranded, positive-sense, RNA (ribonucleic acid) genomes and that share morphological, structural, molecular and genetic features. Resistance against LNSV is conferred by Tvr1-a single, dominant gene that provides durable resistance (Grube et al. 2005b; Simko et al. 2009). Table 3 shows the resistance to LNSV of accessions of the wild species L. serriola (UC96US23; PI 491178; PI 271940), L. virosa (PI 273597; IVT 280) and L. saligna (PI 271940; PI 490999).
Bacterial pathogens
Corky root (Sphingomonas suberifaciens, formerly Rhizomonas suberifaciens)
Corky root of lettuce has been observed in several major lettuce producing areas of the world, including North America, Western Europe, Australia and New Zealand. The symptoms of disease are dark discolouration and longitudinal cracks on the taproot, penetrating to the cortical region and the disease causes slow progressive deterioration of the root system of infected plants and lettuce seedlings and plants wilt under water stress. In severly infested fields of California and Florida, yield losses from reduced head size can reach 30 % to 70 % (Mou et al. 2007); the reduced development of heads is correlated with reduced root growth. The pathogen most commonly isolated from diseased roots is the bacterium Sphingomonas suberifaciens (Yabuuchi et al. 1999), formerly Rhizomonas suberifaciens (van Bruggen 1997).
The use of resistant cultivars is the most efficient strategy to avoid economic losses (Mou 2011a). The first resistant lettuce cultivars Marquette, Montello and Green Lake developed by Sequiera (1970, 1978) were released from crosses with a resistant line PI 171669. This line was identified by Dickson (1963) as a local lettuce landrace from Turkey.
The resistance to corky root is conferred by a recessive allele (cor) at a single locus (Brown and Michelmore 1988), which is present in many modern crisphead lettuce cultivars, e.g. Bronco, Cannery Row, Glacier, Premiere, Misty Day, Sharp Shooter, Sniper (Mou et al. 2007), however there are few leaf lettuce varieties with this resistance (Mou 2011a). Recently two breeding lines 06–831 and 06–833 of greeen leaf lettuce were released from the cross between green leaf cultivar Waldmann’s and the crisphead cultivar Glacier (Mou 2011a).
Brown and Michelmore (1988) identified resistant lines within the wild species L. serriola, L. saligna, L. dentata, L. virosa and Lactuca spp. (Table 4). Mou and Bull (2004) identified three L. serriola and one L. virosa accessions consistently resistant to corky root in growth chamber, greenhouse and field experiments (Table 4), and they demonstrated significant genotype by environment interactions for corky root severity. Moreover, none of these four resistant lines possessed the two molecular markers closely linked to the cor allele suggesting that they may be sources of a new resistance factor (Mou and Bull 2004).
Bacterial leaf spot (Xanthomonas campestris pv. vitians)
Bacterial leaf spot of lettuce caused by Xanthomonas campestris pv. vitians has been reported from different countries, since the beginning of the 20th century (Toussaint et al. 2012). On romaine cos type lettuce, symptoms typically appear at the leaf margin as water-soaked lesions which become black after a few days and may be surrounded by a chlorotic halo. Later on they enlarge and coalesce, and large necrotic areas on leaves may develop (Toussaint et al. 2012). Lesions may expand towards veins, resulting in V-shaped lesions. Small individual black spots on the leaf surface may also be observed (Sahin and Miller 1997). Seed collected from infected plants were found to be colonized by bacteria externally, but no bacteria were recorded from within the seed (Barak et al. 2002).
When the infection remains restricted to the older leaves, no economic losses occur, however, in severe epidemics, the inner leaves are infected and the lettuce is then unmarketable (Toussaint et al. 2012).
Populations of X. campestris pv. vitians can survive on lettuce plant debris and infect subsequent lettuce crops (Barak et al. 2001). The pathogen has also been recovered from leaves of several symptomless weed species collected around infested commercial fields, but not from weeds collected around previously infested fields during fallow periods. Thus, weeds may not be an important long-term source of X. campestris pv. vitians, possibly due to the lack of stable epiphytic populations on weedy plants (Barak et al. 2001). Xanthomonas campestris pv. vitians can infect not only cultivated lettuce but also the wild Lactuca species, L. serriola and L. biennis, and these species may serve as a reservoir for this pathogen (Toussaint et al. 2012).
The possibilities of chemical control of bacterial diseases on lettuce are limited (Toussaint et al. 2012) and so host resistance is the most likely means of controlling the disease. Several commercial cultivars of lettuce have been screened for resistance to this bacterial pathogen (Sahin and Miller 1997, Carisse et al. 2000) and activities aimed at the development of lettuce breeding lines resistant to bacterial leaf spot have been briefly reported (Anonymous 2005). However, the genetics of resistance to X. campestris pv. vitians and the response of wild Lactuca species to this bacterial pathogen have not yet been published.
Fungal pathogens
Downy mildew (Bremia lactucae)
Lettuce downy mildew (Bremia lactucae) has a very high economic impact on lettuce production (Crute 1992), and the study of its biology and epidemiology, sources of resistance, mechanisms and genetic control of resistance in Lactuca species, including germplasm evaluations have been a high priority of researchers and breeders in many countries (Lebeda et al. 2002, 2009a, b). The interaction between cultivars of L. sativa and B. lactucae is clearly race-specific (Crute and Johnson 1976; Lebeda 1984; Farrara and Michelmore 1987).
Of the 100 wild Lactuca species described (Lebeda et al. 2004b) only 14 are definitely known as natural hosts of B. lactucae (Lebeda et al. 2002). L. serriola, is the most common wild Lactuca spp. occurring around the world (Lebeda et al. 2004b), and could be an important weedy host. However, except for the Czech Republic, there is no detailed information on the natural occurrence of lettuce downy mildew and its epidemiological impact on this species (Lebeda et al. 2002, 2008a). There is only limited knowledge of virulence variation of B. lactucae in wild pathosystems (Lebeda et al. 2008a). Only isolates originating from natural populations of L. serriola have been investigated for specific virulence variation (Lebeda et al. 2002, 2008a; Lebeda and Petrželová 2004). Generally, B. lactucae isolates from wild pathosystem are characterized in terms of v-factors mostly matching Dm genes or R-factors located or derived from L. serriola (Lebeda and Petrželová 2004).
Currently, searching for new sources of resistance and genes suitable for practical lettuce breeding (Lebeda and Zinkernagel 2003a; Beharav et al. 2006; Petrželová and Lebeda 2011; Petrželová et al. 2011; van Treuren et al. 2013) is considered to be very important. Accessions of L. serriola (reported as PI 91532 but subsequently shown to be PI 104584 and PI 167150) originating from Russia and Turkey were used in the 1930s in the USA as sources of resistance against B. lactucae (Lebeda et al. 2002). These sources created the breeding pool for a new generation of lettuce cultivars (Imperial 410, Calmar, Valmaine) for outdoor cropping which were introduced in the 1940s and 1950s (Whitaker et al. 1958). All of these cultivars have race-specific resistance (Table 5).
In Europe, the utilization of wild Lactuca germplasm was based on two different strategies (the Netherlands and Great Britain). In the 1950s, genes originating from old German and French cultivars of L. sativa were used mostly (Crute 1992). At the end of the 1960s in the Netherlands an interspecific hybrid between L. sativa (cv. Hilde) and an accession of L. serriola, described as H × B, Hilde × L. serriola was released. Resistance derived from this material was assigned to the race-specific gene Dm11 (Lebeda et al. 2002) (Table 5).
In the 1970s and 1980s other sources of resistance to B. lactucae, derived from L. serriola with resistance genes (factors) described as Dm16 and R18 (Table 5), were used in the Netherlands. All of these genes have been used frequently in breeding programs in Europe during the last 20 years. However, resistance based on these genes is no longer effective against many B. lactucae isolates (Lebeda and Zinkernagel 2003b). From the end of the 1980s there was increasing interest (esp. in the Netherlands and U.K.) for the utilization of resistance located in the hybrid line L. serriola (Swedish) × L. sativa (Brunhilde) and line CS-RL (Lebeda and Blok 1991) was derived from this material. This line was highly resistant for a long time. However recently a new race overcoming the resistance was described (Lebeda and Zinkernagel 2003a).
L. saligna and L. virosa may possess novel and very interesting resistance to B. lactucae (Lebeda et al. 2002). As a result of studies in the 1990s a new lettuce cultivar Titan (Sluis & Groot) with the race-specific gene Dm6 plus resistance derived from L. saligna (pers. comm., K. Reinink, Rijk Zwaan, the Netherlands) was released in the Netherlands (Lebeda et al. 2002). However, this resistance is no longer effective (Lebeda and Zinkernagel 2003a, b). Recently, a very intensive program of lettuce breeding based on introduction of newly located sources and genes of resistance from L. serriola, L. saligna and L. virosa was developed in the USA (Michelmore et al. 2005). Also new sources of resistance were located in wild Lactuca spp. originating mostly from the Middle East (Beharav et al. 2006; Petrželová et al. 2011).
The effectiveness of the expression of some Dm genes located in L. serriola can be dependent on environmental factors. Judelson and Michelmore (1992) showed that resistance (assessed as the absence of sporulation) based on Dm6, Dm7, Dm11, Dm15, and Dm16 became less effective or ineffective at temperatures below 10 °C. The ecological and epidemiological consequences of this effect are not known (Lebeda et al. 2002).
The occurrence of race-specificity in other wild Lactuca species and related genera has not been analysed in detail. However, recent analyses (Lebeda et al. 2002) have shown that the occurrence of race-specific resistance in wild Lactuca species is a common phenomenon. In the section Lactuca, all of the species studied express race-specificity after inoculation with isolates of B. lactucae from L. sativa and L. serriola. The presence of race-specific resistance in L. saligna was described as questionable because most of the screened accessions exhibited complete or incomplete resistance at both the seedling and adult stage (Lebeda et al. 2002; Beharav et al. 2006), and recent results (Petrželová et al. 2011) showed that L. saligna may possess non-host resistance. A race-specific response was also confirmed in some species from other sections of the genus Lactuca (L.viminea, L. tatarica, L. quercina, L. indica, L. biennis) (Lebeda et al. 2002) and there is clear evidence of the occurrence of a race-specific response in some species of related genera (e.g. Cicerbita, Mycelis) (Lebeda et al. 2002).
Other types of resistance (race-nonspecific, field, non-host; for detailed description see Lebeda et al. (2002)) of Lactuca species against B. lactucae are not as well understood. There is only limited information available about race-nonspecific resistance in wild Lactuca spp. germplasm. The presence of race-nonspecific resistance has only been reported in L. serriola (Lebeda et al. 2002). Currently only two L. serriola accessions can be considered as potential sources of this type of resistance. It was recognised that accessions PI 281876 and PI 281877 at the seedling stage were infected by some B. lactucae isolates. However, the intensity of sporulation was mostly very low and in some interactions was followed by expression of a necrotic response (Lebeda 1986). Current thoughts are that this resistance is based on some major gene(s) and modifiers (Lebeda et al. 2002). L. serriola (PI 281876) has been used frequently in practical breeding programs (Lebeda and Pink 1998).
The most comprehensive experiments focused on field resistance of wild Lactuca spp. were carried out by Lebeda (1990). In total, thirty-one accessions of four Lactuca species (L. serriola, L. saligna, L. aculeata, L. indica/syn. L. squarrosa/) and one L. serriola × L. sativa hybrid (line CS-RL) were studied in 3 years of field experiments. The disease incidence was significantly different across species and accessions. L. saligna, L. aculeata accessions and the L. serriola × L. sativa hybrid were free of infection during the observation period. This reaction implies the presence of effective unknown R-factors (Lebeda et al. 2002) in these genotypes. In the L. serriola accessions, significant differences in the level of field resistance were observed (Lebeda 1990). Some accessions were highly susceptible (e.g. PI 204753, PI 253468, PI 273596, PI 273617, PI 274359), in contrast, accessions PI 281876 and PI 253467 were free of disease symptoms (again implying the presence of effective unknown R-factors). However, the possible race-nonspecific resistance in PI 281876 is also likely to be expressed as field resistance (Lebeda 1990).
Nonhost resistance should be very effective, durable and not influenced by changes of environmental conditions (Lebeda et al. 2002). It was hypothetized that some L. saligna accessions may possibly possess nonhost resistance (Lebeda 1986). Recent experimental results with new highly virulent isolates of B. lactucae originating from L. sativa have not confirmed the presence of race-specific resistance in L. saligna (Lebeda and Zinkernagel 2003a). However, recent findings indicate that, at least some L. saligna accessions possess race-specific resistance factors (Jeuken and Lindhout 2002), in addition to possible non-host resistance to B. lactucae (Beharav et al. 2006; Petrželová et al. 2011).
There is only limited information available on the histological, cytological, biochemical and molecular background of resistance to lettuce downy mildew in L. sativa and wild Lactuca species. Some basic ideas and conclusions related to this subject were summarized by Lebeda et al. (2002, 2006, 2008b), Jeuken and Lindhout (2002, 2004). Data obtained in histological studies of resistance in wild Lactuca spp. suggest there are a wide range of resistance mechanisms in Lactuca spp. against B. lactucae (Lebeda et al. 2008b).
Powdery mildew (Golovinomyces cichoracearum)
Lettuce powdery mildew is considered as a disease of increasing importance (Lebeda and Mieslerová 2011). The ascomycete Golovinomyces cichoracearum V.P. Gelyuta (previously Erysiphe cichoracearum DC. s.str. (Lebeda and Mieslerová 2011)) is the predominating powdery mildew species, however, another powdery mildew species, Podosphaera fusca was collected and described on Lactuca sativa in Korea (Shin et al. 2006). Great progress in the research of the taxonomy, distribution and biology of lettuce powdery mildew (Golovinomyces cichoracearum sensu stricto) has been achieved during the last 15 years (Lebeda and Mieslerová 2011).
Natural hosts of powdery mildew include L. muralis, L. perennis, L. quercina, L. serriola, L. saligna, L. sibirica, L. viminea, L. virosa (Lebeda 1985a, b; Lebeda and Mieslerová 2011), and L. aculeata (Lebeda unpubl.). One of the most common species in Europe is L. serriola (prickly lettuce) which also could be considered as a common host of G. cichoracearum. Substantial variation in expression of the degree of infection between different sites and/or populations was recognized. It was concluded that L. serriola could act as a reservoir of inoculum for lettuce infection (Lebeda et al. 2012c, 2013).
Lebeda (1985c) demonstrated in a set of 25 lettuce (L. sativa) cultivars substantial differences in disease severity. Only two cultivars (Amanda Plus, Bremex) were free of natural infection.
The screening of more than one hundred accessions of wild representatives of the genus Lactuca (L. aculeata, L. dentata, L. perennis, L. saligna, L. serriola, L. tatarica, L. tenerrima, L. viminea, L. virosa) under conditions of natural infection by G. cichoracearum revealed high variability in resistance (Lebeda 1985b, 1994). The accessions of L. serriola were attacked most severely and L. saligna showed highly variable levels of resistance, while the lowest levels of infection were found in accessions of L. virosa, L. viminea, L. tenerrima and L. tatarica. In some species (e.g. L. saligna, L. serriola) the interaction with the pathogen is probably based on race-specific resistance (Lebeda and Mieslerová 2011; Lebeda et al. 2012c, 2013) (Table 6).
Some L. saligna accessions are potentially useful sources of resistance, especially where they carry resistance to Bremia lactucae as well (Lebeda 1985b). L. virosa could be also considered as a suitable donor of resistance; however its resistance seems to depend on a certain stage of the ontogenetic development (Lebeda 1985a).
Anthracnose (Microdochium panattoniana)
Anthracnose (shothole disease, ringspot) caused by Microdochium panattoniana (Berl.) Sutton, Galea & Price is manifested as small circular brown spots primarily on the lower leaf blades. The centers of these spots dry and fall out. Lesions on the midrib become necrotic, sunken, and elongated. The initial infection may be soilborne or seedborne, the anthracnose conidia are spread in lettuce crops by splashes of rain or irrigation water (Galea et al. 1986). Lettuce ringspot causes serious damage of lettuce crops in the southern states of Australia, in California and throughout Europe especially under cool wet conditions when application of fungicides is difficult (Galea and Price 1988). Sources of resistance have been identified in wild Lactuca species L. angustana, L. livida, L. perennis, L. serriola, L. saligna, and L. virosa (Ochoa et al. 1987; Galea and Price 1988) (Table 7).
Stemphylium leaf spot (Stemphylium spp.)
Symptoms of Stemphylium botryosum f. lactucum Wallr. on lettuce leaves are small, round, and brown spots, which may appear sunken because the tissue becomes necrotic (Netzer et al. 1985). This fungal disease has been reported in many parts of the world (Raid 1997), but it has relatively small economic impact. The only known source of resistance to the disease is an unspecified line of L. saligna collected in Israel (Table 7). Resistance is controlled by two genes, with one allele dominant Sm 1 for resistance and the other recessive sm 1 (Netzer et al. 1985).
Sclerotinia drop-lettuce drop (Sclerotinia spp.)
Two fungal species, Sclerotinia minor and Sclerotinia sclerotiorum cause sclerotinia drop of lettuce, one of the most widespread and destructive disease worldwide in lettuce production (Purdy 1979; Subbarao 1998). Both S. minor and S. sclerotiorum survive mainly as sclerotia in soil. S. minor primarily infects lettuce by direct eruptive germination of soilborne sclerotia. This mode of infection is less frequent in S. sclerotiorum. The primary inoculum source of S. sclerotiorum is airborne ascospores from carpogenic germination of sclerotia (Abawi and Grogan 1979). Sclerotinia is difficult to control with cultural methods (Lebeda et al. 2007c), and it is difficult to elaborate protocols of resistance screening (Grube and Ryder 2004).
Extensive evaluation of lettuce germplasm has been carried out either for resistance to S. sclerotiorum (Chupp and Sherf 1960; Elia and Piglionica 1964; Whipps et al. 2002) or S. minor (Abawi et al. 1980; Subbarao 1998; Grube and Ryder 2004) but no complete resistance has been identified, and it is unknown whether resistance to the two species is correlated (Lebeda et al. 2007c). Wild Lactuca species were included in these tests but the numbers of accession was relatively low (Abawi et al. 1980; Whipps et al. 2002).
An accession of primitive oilseed lettuce L. sativa (PI 251246) may have partial resistance to Sclerotinia sp. infection (Whipps et al. 2002; Hayes et al. 2010), but this is very likely associated with its primitive growth habit (Grube 2004). L. dentata (PI 234204, later named as Sonchus oleraceus (Doležalová et al. 2004; Lebeda et al. 2007c)), Lactuca sp. (PI 274376) and L. serriola (PI 271938) were shown to be highly resistant to S. minor (Abawi et al. 1980), and the latter accession was also resistant to S. sclerotiorum (Whipps et al. 2002) (Table 7). In the S. sclerotiorum-infested field experiments, three L. virosa accessions (SAL 012, IVT 280 and IVT 1398) demonstrated high levels of resistance (Table 7), although further analysis is needed to determine the role of the slow bolting/biennial nature of L. virosa in resistance (Hayes et al. 2010).
Verticillium wilt (Verticillium dahliae)
Verticillium wilt, is a relatively newly recognised lettuce disease caused by the soilborne fungus Verticillium dahliae Kleb. It was reported for the first time in a lettuce crop in the Pajaro Valley (California) in 1995 (Subbarao et al. 1997; Bhat and Subbarao 1999), in 1999 it was first observed on lettuce in the Salinas Valley (Atallah et al. 2011), in 2006 in northern Italy (Garibaldi et al. 2007). In 2009, this disease appeared in commercial fields in Japan (Usami et al. 2012). Losses of up to 100 % may occur in head lettuce: smaller losses occur in other lettuce types (Lebeda et al. 2007c).
V. dahliae was first isolated from L. serriola during a field survey carried out in Crete in 1992–2000 (Ligoxigakis et al. 2002). Disease symptoms and recovery of V. dahliae are known in wild L. serriola and other L. serriola–like species, L. saligna, and L. virosa (Hayes et al. 2009).
Two pathogenic races of V. dahliae were described, and currently race 2 predominates as a result of worldwide cultivation of lettuce cultivars resistant to race 1 (Hayes et al. 2006, 2007a, b, 2011b; Atallah et al. 2011). Despite widespread screening, complete resistance to race 2 has yet to be identified (Grube et al. 2005a; Attalah et al. 2011). Partial resistance to race 2, in the form of reduced disease incidence or delayed expression of symptoms, has been found in four PI accessions L. sativa (Hayes et al. 2011a).
The L. virosa accession IVT 280 (Table 7) and some other accessions have shown high levels of resistance in field tests (Grube et al. 2005a).
Fusarium wilt (Fusarium spp.)
Fusarium wilt was first reported on lettuce in Japan in 1955 (Matuo and Matahashi 1967), but it was not until many years later that its widespread occurrence and potential for economic damage was fully recognized (Fujinaga et al. 2003; Hubbard and Gerik 1993; Garibaldi et al. 2004). The disease is caused by Fusarium oxysporum f. sp. lactucae n.f. (same as f. sp. lactucum (Hubbard and Gerik 1993; Fujinaga et al. 2003). Three races of the fungus have been identified (Fujinaga et al. 2003). It is a disease of the root vascular system. As one of several wilting diseases exhibiting yellowing and wilting of leaves and stunting and plant death, the principal diagnostic symptom is a reddish brown discoloration of the cortex and upper crown (Matheron and Koike 2003). Higher temperatures tend to increase the severity of fusarium wilt in lettuce (Scott et al. 2010).
Resistance sources have been identified for races 1 and/or 2, but not for race 3 (Garibaldi et al. 2004; Tsuchiya et al. 2004). All sources are cultivars of the various lettuce types.
Pythium wilt (Pythium spp.)
Soilborne pathogens Pythium tracheiphylum and P. uncinulatum were reported as causing vascular wilt and stem rot of lettuce in Italy in 1965 (Matta 1965) and subsequently in other parts of Europe (Blok and van der Plaats-Niterink 1978), North America (Tortolero and Sequeira 1978), Australia (Kumar et al. 2007) and Japan (Matsuura et al. 2010). Yield reductions up to 30 % have been recorded (Davis et al. 1995). In spite of the economic importance of these pathogens, the screening of wild Lactuca species for resistance is not reported.
Nematodes
Root-knot nematode (Meloidogyne spp., incognita, hapla, javanica, enterolobii)
Nematodes occuring on lettuce can be classified into 23 genera: Aphelenchoides (da Silveira 1990), Meloidogyne (e.g. Viaene and Abawi 1996, Blancard 2011), Pratylenchus (Moretti et al. 1981; Mani et al. 1997), Rotylenchulus, Tetylenchus, Tylenchorhynchus (e.g. Radewald 1969a; Philis 1995; Koenning et al. 1999; Kohl 2011; Pedroche et al. 2012), Helicotylenchus (Anwar and McKenry 2012), Criconemoides, Heterodera, Hoplolaimus, Paratylenchus (Machado and Inomoto 2001; Bao and Neher 2011), Paratrichodorus (Boydston et al. 2004), Hemicycliophora (Chitambar 1993; Blancard 2011), Rotylenchoides, Tylenchus, (Addoh 1971), Longidorus (Radewald 1969b, c; MacGowan 1982; Huang and Ploeg 2001), Mesocriconema (DAFF 2012), Nacobbus, Paralongidorus, Xiphinema (Sikora and Fernández 2005), Aglenchus (Ökten 1988), Belonolaimus (Chitambar 2007), Radopholus (Ferris 2013), Merlinius (Bridge 1976).
However the most important nematodes with documented impact on lettuce growth and yield include the needle nematode (Longidorus africanus), root-knot nematode (Meloidogyne spp.), root lesion nematode (Pratylenchus penetrans), and the spiral nematode (Rotylenchus robustus) (Davis et al. 1997). There are several other nematodes associated with lettuce in the field-stubby root nematode Paratrichodorus minor, the reniform nematode Nacobbus aberrans, and stunt nematodes Tylenchorhynchus clarus and Merlineus spp. (Davis et al. 1997).
The number of papers reporting occurrence of nematode infection/diseases on lettuce wild relatives is limited. The majority of reports are based on field observation of nematode infection on L. serriola (Table 8). There are just two reports related to identification of resistant accessions to root-knot nematode (M. hapla) in a larger germplasm collections. Abawi and Robinson (1991) evaluated 85 genotypes (including accessions of L. serriola, L. virosa and L. saligna) in two greenhouse tests, and in a recent study Kaur and Mitkowski (2010) analysed 494 lettuce accessions, including 36 L. serriola, 7 L. virosa and 8 L. saligna genotypes. Genotypes with moderate to highly resistant reaction to M. hapla inoculation were observed in these studies (Table 8). The occurrence of root-knot nematodes on weed hosts was investigated by Gowda et al. (1995), who reported light root gall intensity and small size of galls indicating resistance to Meloidogyne incognita. Davis and Venette (2004) considered Meloidogyne falax as potential risk (potential hosts) for several threatened and/or endangered wild Lactuca species (L. floridana, L. hirsuta, L. tatarica var. pulchella).
The information on the genetic basis of nematode resistance in wild Lactuca species is missing. In cultivated lettuce the resistance genes for Meloidogyne appears to be under control of a single gene locus, with predominantly additive gene action (for M. incognita races 1, 2, 3 and 4, and M. kabanica) (Gomes et al. 2000; Maluf et al. 2002; Cavalhi Filho et al. 2008). On the other hand de Carvalho et al. (2011) proved that two different genes are involved in control of resistance to M. incognita race 1 in lettuce cultivars Grand Rapids and Salinas-88. Further, they reported that lines with higher levels of nematode resistance than either Grand Rapids or Salinas-88 could be selected in the F4 generation of the cross between these resistant parental lines indicating that the two parental cultivars possess different genetic factors for resistance.
There are also reports on transgenic lettuce linies bearing tomato root-knot resistance gene Mi-1 (Zhang et al. 2010) and the linkage of tomato resistance genes to root-knot nematode (Meloidogyne spp) to leaf mold (Cladosporium fulvum) (Dickinson et al. 1993; Jones et al. 1993).
Insects and mites
There are a number of aphid species, occuring both on cultivated lettuce and its wild relatives, these belong to the following genera-Acyrthosiphon, Aulacorthum, Aphis, Dysaphis, Hyperomyzus, Macrosiphum, Myzus, Nasonovia, Neomyzus, Pemphigus, Protrama, Sitobion, Trama, Uroleucon (Blackman and Eastop 2000), Eucarazzia (Stoetzel 1985) and Rhopalosiphum (Sangün and Satar 2012). However, we present here a review of papers using wild lettuce species as parents for resistance breeding to three aphid species-the green lettuce aphid (Nasonovia ribisnigri), the potato aphid (Macrosiphum euphorbiae) and lettuce root aphid (Pemphigus bursarius) and the leafminer (Liriomyza langei). There are other important aphid species (e.g. Myzus persicae, the green peach aphid, possibly the most important leaf-feeding pest on lettuce because of its ability to transmit several important viruses - see above), however no information related to resistance screening studies (or breeding) using wild lettuce species has been published so far. For a detailed survey of occurence of various aphid species on other Lactuca spp. (see Blackman and Eastop 2000).
Nasonovia ribisnigri Mosley [the „green lettuce aphid“(GLA) or „currant-lettuce aphid“(CLA)] is commercially the most important lettuce pest (Martin et al. 1996) with a worldwide distribution (Blackman and Eastop 2000). It colonizes the interior of the lettuce head, making its control difficult both with contact insecticides (Liu 2004) and biological control (Mackenzie and Vernon 1988). The use of resistant cultivars is therfore the best option to protect lettuce from this pest. Resistance to biotype 0 (Nr:0) was first reported in Lactuca virosa accession IVT 280 (Eenink et al. 1982a,b) and characterized as complete (i.e. virtually no aphids survived), and genetically dominant to the partial resistance found in L. virosa accession IVT 273. Complete and partial resistances to Nr:0 were conditioned by two alleles, Nr (complete resistance) and nr (partial resistance). McCreight and Liu (2012) proposed the following system of allelic designation: Nr:0 C for complete resistance and Nr:0 P for partial resistance, with their relationships: Nr:0 C (in IVT 280, ‘Barcelona’) > Nr:0 P (in PI 491093) > nr (in susceptible genotypes) (McCreight and Liu 2012).
Subsequently, resistance in IVT 280 was successfully transferred to lettuce by a bridging cross to L. serriola (e.g. Eenink et al. 1982a, b; van der Arend et al. 1999). There are a large number of modern lettuce cultivars with GLA resistance e.g. cvs. Barcelona, Campionas, Dynamite, Elenas, Fortunas, Irina, Krinas, Veronas, 83-67RZ (van der Arend et al. 1999; Liu and McCreight 2006) and in 2010 there were 88 comercially available lettuce varieties in Australia (McDougall and Troldahl 2010), produced by several breeding companies including: RijkZwaan, Nunhems, South Pacific Seeds, Lefroy Valley, Seminis Vegetable Seeds, Terranova Seeds. However, this widespread deployment of a single dominant gene for resistance has exerted a high selection pressure for the emergence of a resistance breaking phenotype and several N. ribisnigri populations feeding on resistant cultivars were detected in 2007. These were subsequently characterized as a resistance-breaking biotype of N. ribisnigri designated Nr:1 (Sauer 2008). This biotype has spread throughout the European continent (Cid et al. 2012).
Wild progenitors of cultivated lettuce appear to be a valuable source of resistance to GLA, as evident from the results of recent large germplasm screenings. McCreight (2008) identified two new potential sources for resistance to Nr:0 in L. serriola acc. PI 491093 (partial resistance) and L. virosa PI 274378 (complete resistance). This was a result of a large greenhouse screening of 1203 lettuce accessions (included 7 L. perennis, 18 L. saligna, 125 L. serriola, and 6 L. virosa accessions). Sixty-four L. serriola and L. virosa accessions (see Table 9) resistant to Nr:1 were reported in CGN germplasm collection (the Center for Genetic Resources, the Netherlands) (Anonymous 2008). For some of these (CGN13361, CGN16266, CGN16272), all five replicate plants were resistant while in other accessions (CGN04757, CGN04930, CGN04973) resistance segregated in the tested plants. Several accessions were resistant to both Nr:0 and Nr:1 (Anonymous 2008). Dominant resistance to Nr:0 and Nr:1 was also reported to have been found in the L. serriola accession 10G.913571 by Thabuis et al. (2011).
More recently, Cid et al. (2012) performed two tests: a greenhouse screening of 264 lettuce accessions including 40 accessions closely related to L. sativa (3 L. perennis, 6 L. virosa, 13 L. serriola and 18 L. sativa × L. serriola) and laboratory screening of 40 L. virosa accessions against both N. ribisnigri biotypes (Nr:0, Nr:1) and against a clone of M. euphorbiae. Three L. virosa accessions showed (Table 9) resistance against N. ribisnigri, two (CGN16272 and CGN13361) partial resistance to the Nr:1 biotype of N. ribisnigri and to M. euphorbiae. While near complete resistance to M. euphorbiae was found in CGN13355 but this was susceptible to N. ribisnigri (Cid et al. 2012).
The „lettuce root aphid“(LRA), Pemphigus bursarius (L.) is one of several aphid species that feed on cultivated lettuce (Blackman and Eastop 2000) and can cause severe damage to crops (Dunn, 1959). It is considered as an occasional pest of lettuce crops but can also cause severe losses in lettuce seed production. Additionally, it is able to colonise a variety of non-crop species, largely within the Compositae (Dunn 1959; Alleyne and Morrison 1977). This aphid is regarded as holocyclic, alternating annually between sexual reproduction on a primary woody host, poplar (Populus nigra, L.) and parthenogenesis on its secondary hosts such as lettuce (see Dunn 1959; Miller et al. 2003, 2008). Lettuce root aphid is one of the first examples of successful insect resistance breeding in vegetable crop (Reinink 1999). As the most effective control of root aphids, highly resistant cultivars eliminating the aphid colonisation on lettuce roots. This resistance is controlled by one or two genes as described by Ellis et al. (1994, 2002). At least one of these genes is not allelic to the existing Lra gene, which can be linked to downy mildew resistance gene DM6 (e.g. linked in cv. Avonscrisp but not in cv. Lakeland). There is only a single paper focussed on LRA resistance screening in germplasm collections by Ellis et al. (2002). This describes the testing of 55 Lactuca spp. accessions for resistance to P. bursarius and the identification of extremely high levels of resistance in accessions of the wild species L. saligna, L. perennis, L. virosa, and in the variety Grand Rapids. Cole et al. (1991) described the use of allozyme analysis to detect bands related to LRA resistance in a screening of four Lactuca species (L. serriola, L. virosa, L. saligna and primitive L. sativa, eight accessions per species). Out of the forty samples tested, only ten accessions were resistant to colonisation by the pest (L. serriola—001562, HRIGRU1606, HRIGRU1573, HRIGRU7145; L. saligna—006186, 001627, HRIGRU1630 and L. sativa 006779, 001886, 006612).
Leafminer (Liriomyza langei Frick) is a major insect pest of many important agricultural crops including lettuce (Mou and Liu 2003, 2004; Lebeda et al. 2007c). Succesfull breeding for resistance to leafminer in lettuce was reported by Mou and Ryder (2010). However, studies exploring genetic variation of leafminer resistance in lettuce germplasm, including wild progenitors, are limited (Mou and Liu 2003, 2004). Also the mechanism of leafminer resistance in lettuce is unknown (Mou and Liu 2004). Mou and Liu (2003) performed screening of 46 Lactuca accessions, including 2 accessions of L. serriola, 1 acc. of L. saligna, and 1 acc. of L. virosa. High levels of resistance were discovered in these wild genotypes. In a subsequent study by Mou and Liu (2004) fifty-four lettuce genotypes and 232 F2 plants of crosses were evaluated for leafminer resistance, again a significantly lower occurence of leafminers were found on the accessions of the wild species (L. serriola, L. saligna, and L. virosa) compared to the cultivars.
Approaches to exploitation of wild Lactuca spp. in lettuce resistance breeding
Interspecific hybridization
Autogamy is the predominating breeding system within the genus Lactuca, especially in the marginal parts of its distribution area (Feráková 1977). Stebbins (1957) estimated a higher occurrence of allogamy in the centre of distribution. Lindqvuist (1960) proved experimentally that all species belonging to the “serriola” group were self-fertile. Spontaneous cross-pollination occurs through activity of various insect from the Hymenoptera and Diptera groups (Feráková 1977). In the commercial seed production of L. sativa, up to 5 % of cross-pollination has been observed (George 1999). Hybridization can occur not only within one species, but also between species. Lactuca altaica hybridizes spontaneously with L. saligna and L. serriola; L. aculeata hybridizes with L. sativa and with L. serriola; L. serriola hybridizes with L. dregeana (Zohary Zohary 1991). The close relationship of serriola-like species L. serriola, L. dregeana, L. altaica, and L. aculeata to L. sativa is supported not only by the same chromosome number but also by molecular (AFLP) markers (Koopman et al. 1998, 2001), and by DNA content (Koopman 2002).
Hybridization data on the species belonging to the different sections or groups of the genus Lactuca are limited to L. viminea and L. tatarica. Groenwold (1983) reported partly fertile hybrids between L. viminea (section Phoenixopus) and L. virosa (section Lactuca).
Natural hybridization
D’Andrea et al. (2008) proved in a field experiment conducted in Switzerland that natural hybridization between lettuce and L. serriola occurred up to the maximal distance tested (40 m), and hybridization rates varied between 0 to 26 %, decreasing with distance. More than 80 % of the wild plants produced at least one hybrid at within 1 m and 4 to 5 % at 40 m. In sympatric crop-wild populations, cross-pollination between cultivated lettuce and its wild relative has to be seen as the rule rather than the exception for short separation distances. However, in the northern parts of Europe, where expansion of prickly lettuce (L. serriola) took place, only a few putative hybrids with L. sativa were found. So, very probably mechanisms other than crop/wild gene flow, such as those connected to the human activities around building and transport are more likely explanations for this phenomenon (Hooftman et al. 2009; Uwimana et al. 2012a).
The phenotype of putative natural interspecific hybrids was recorded for primarily self- pollinated Lactuca species acquired during collecting missions in natural habitats: Lactuca serriola (× L. sativa) acquired from natural populations L. serriola f. serriola in northern Moravia in the year 1995 (Křístková et al. 2012). The hybrid character of plants L. aculeata (× L. serriola) from a natural population L. aculeata collected in Israel in 2005 was confirmed by allozyme markers (Lebeda et al. 2012b). Hybrid characteristics of L. saligna (× L. serriola) were observed on plants raised from L. saligna achenes collected in Jordan in 2007. Putative hybrid plants showed a low level of self-fertility (Křístková et al. 2012). The molecular profile of plants of L. serriola collected in Israel in 2009 suggests natural hybridization of L. serriola (× L. saligna) (Křístková et al. 2012).
The phenotype of interspecific hybrids was recorded also on several Lactuca spp. accessions obtained from world germplasm collections. Species L. sativa, L. serriola, L. saligna, L. dregeana and L. virosa probably participated in the hybridization (Doležalová et al. 2007).
Managed hybridization
Hybridization experiments have been used to: reveal evolutionary relations among Lactuca species (de Vries 1990) and aspects of domestication process of lettuce; serve as a base for plant genetic resources management (van de Wiel et al. 2010) and the practical application in breeding programmes; and recently for the assessment of ecological risk of transgenes (Giannino et al. 2008; Hooftman et al. 2011).
Three approaches are employed in order to prevent self-pollination and to perform sexual hybridization: manual removal of anthers, spraying with water of the self-pollen from the stigma prior the cross-pollination, and the exploitation of male-sterile lines as pollen recipients (Davey and Anthony 2011). Examples of interspecific hybridization with potential economic impact are given in this paper below in following chapters.
Hybridization experiments of lettuce with L. serriola and QTL analysis identified genomic regions with major QTL effects important for breeding programmes and plant transformation (Hartman et al. 2012, 2013a). Hartman et al. (2013b) show that there is a high likehood in lettuce for novel crop-wild hybrids to arise with a higher fitness than the wild parent through combination of heterosis, linkage and transgressive segregation.
The genomic analysis of plants derived from the hybridization of L. sativa × L. serriola and their backrosses proved that the domesticated parent contributed QTLs with either a positive or a negative effect on plant vigour, and there are genomic locations where transgenes could be preferably located to mitigate (reduce) their persistance in natural populations in occassional crop-wild hybridization (Uwimana et al. 2012b, c).
Interspecific hybrids between the species with a low sexual compatibility have been obtained by using a bridging species (e.g. L. serriola) i.e. crossing one parent to the bridging species and then crossing the resultant F1 with the other parent (Thompson and Ryder 1961; Eenink et al. 1982b; Lebeda et al. 2007c).
Cell and tissue culture
Tissue culture based procedures for the regeneration of fertile plants from tissue explants, cells and isolated protoplasts technologies are the basis for their exploitation in lettuce improvement. Techniques and approaches used for in vitro culture of lettuce have been reviewed thoroughly (e.g. Michelmore and Eash 1986; Alconero 1988; Pink and Keane 1993; Davey et al. 2007a, b; Lebeda et al. 2007c), however there is no recent reports of tissue culture technologies exploring wild Lactuca species in lettuce improvement.
In vitro rescue of immature embryos was used successfully for sexual hybridization between L. sativa and L. virosa (Maisonneuve et al. 1995; Maisonneuve 2003).
Protoplast fusion permitted the regeneration of somatic hybrids between L. sativa and either L. tatarica or L. perennis (Chupeau et al. 1994; Maisonneuve et al. 1995). Somatic hybrids between cultivated lettuce and L. virosa were produced by protoplast electrofusion (Matsumoto 1991). Hybrids had normal flower morphology, but all were sterile. L. indica (section Tuberosae) can be somatically hybridized with L. sativa to produce a viable callus (Mizutani et al. 1989).
So far, fertile interspecific hybrids have only been produced between species L. sativa (section Lactuca) and L. tatarica (section Mulgedium) by using somatic hybridization (Chupeau et al. 1994; Maisonneuve et al. 1995).
Transformation
During the last 10 years, there are numerous reports on the gene introduction into lettuce by transformation (Davey et al. 2002, 2007a, b; Klocke et al. 2010). Agrobacterium inoculation of leaf explants is a universal approach for inducing genes into lettuce, and although this approach has been focused, to date, on L. sativa, wild Lactuca species could be exploited in a similar way (Davey and Anthony 2011).
Transformation of the plastid genome has several advantages over conventional nuclear transformation, mainly by the high level of transgene expression in chloroplasts, this is opening new possibilities for metabolic engineering, resistance management, and production of biopharmaceuticals (Davey and Anthony 2011). Plastid transformation was applied on the lettuce cv. ‘Cisco’ (Kanamoto et al. 2006) and cv. ‘Grand Rapid’ (Pileggi et al. 2001).
There are two ways that transgenic approaches could allow better access to the tertiary gene pool (L. virosa, L. saligna and others). One way would be through directly transforming L. sativa with characterized genes from wild species. The second method is less direct, and would be used in conjunction with somatic hybridization techniques. Transformation would be used to create a universal hybridizer by combining dominant antibiotic resistance and recessive albinism markers in the same genotype (Chupeau et al. 1994). Another potentially valuable use of this technology is to increase or decrease the expression of endogenous genes, or genes already present within cultivated lettuce.
Construction of transgenic plants follows various aims, e.g. production of vaccines, ascorbic acid, tocopherols, increase of iron uptake, decrease of nitrate accumulation, as reviewed by Davey and Anthony (2011) and Davey et al. (2002), Lebeda et al. (2007c), to induce the herbicide resistance (McCabe et al. 1999; Mohapatra et al. 1999; Torres et al. 1999; Dubois et al. 2005) or to delay leaf senescence (McCabe et al. 2001).
Transgenic male-sterile plants are valuable as recipient of donor pollen during hybridization (Curtis et al. 1996; Takada et al. 2007).
Viral genes to confer LMV resistance have been introduced into lettuce (Dinant et al. 1993; Dinant 1997; Gilberton, 1996). Kawazu et al. (2009, 2010) generated lettuce plants resistant to LBVaV, MLBVV by introducing coat protein in anti-sense orientation.
The cloning of genes (Dm3) for resistance to the lettuce downy mildew (Bremia lactucae) performed by Okubara et al. (1997) have led to clarification of the molecular basis of resistance to this pathogen,
Despite the potential value of some of these traits, no transgenic lettuce has yet been commercialized. In some cases the transgenes did not have the expected desirable effects on the plant phenotype possibly due to geen silencing mechanisms such as methylation (Dinant et al. 1993; McCabe et al. 1999). However, the main reasons for the lack of commercial application of GM technology in lettuce is due to the currently high costs associated with the regulatory procedures for release of a transgenic cultivar and a worldwide lack of public acceptance of the technology in a fresh vegetable crop. Biotechnology-derived vegetables, including lettuce, will only succeed if clear advantages and safety are demonstrated to both growers and more importantly to consumers (Dias and Ortiz 2012a, b).
Mutations
Induced mutation has led to the creation of lettuce lines with interesting traits, like dwarfing, early flowering, male sterility or herbicide tolerance (Mou 2011b). Lettuce mutants are useful in genomic studies of resistance to Bremia lactucae, and for gene cloning; combined with genomic advances and new technologies like TILLING (targeted induced local lesions in genome), mutagenesis is becoming more useful tool for lettuce breeders (Mou 2011b).
Known examples of lettuce cultivars issued from exploitation of wild Lactuca spp.
Lettuce breeders have increased genetic diversity and achieved disease resistance through crossing cultivated lettuce with non-cultivated or wild lettuce types (Mikel 2007). Breeding aims, strategies and issues have been reviewed by Ryder (2001). Reviews of wild Lactuca species used in the lettuce breeding and description of breeding approaches and methods were given by Lebeda et al. (2007c), Mou (2008), and Davey and Anthony (2011).
Lebeda and Blok (1991) reported downy mildew resistance in the hybrid of L. sativa × L. serriola. More details about this and the historical consequences about the influence of L. serriola on lettuce resistance breeding to B. lactucae were summarized by Lebeda et al. (2002).
Lactuca saligna is known to produce, hybrids with L. sativa and L. serriola when used as the female parent (Pink and Keane 1993). L. saligna was crossed to a cultivated iceberg type by R.W. Robinson (Provvidenti et al. 1980) who developed the cultivar ‘Salad Crisp’ from that cross. Jeuken and Lindhout (2004) developed backcross inbred lines in which chromosome segments from L. saligna were introgressed into cultivated lettuce and used for genetic analysis of resistance to Bremia lactucae.
Crosses between L. sativa as one parent and L. virosa as the other parent are made with a great difficulty, resulting in low seed set, unviable seeds, stunted plants and/or sterile hybrids (Lindqvist 1960). Viable hybrid plants were obtained only when L. serriola was used as a bridging species (Thompson and Ryder 1961; Eenink et al. 1982b). In 1958, the cultivar Vanguard was developed from a cross between a L. sativa × L. serriola which was then crossed to L. virosa. However, with some manipulations, crosses have been made and have led to development of cultivars (Ryder 1999).
During at least the last two decades there has been an increasing private sector breeding effort in lettuce cultivar development with a concomitant decrease in public sector breeding in many countries. In the USA the legal protection of cultivars is accomplished by their registration supplemented by their pedigree which are made publically available (Mikel 2007), in contrast to lettuce cultivars bred in Europe where such pedigrees are not released in to the public domain.
The pedigree analysis of 328 proprietary and publicly developed lettuce cultivars registered in the USA from 1970 through to 2004 showed that 1 % of these cultivars were developed from interspecific crosses (Mikel 2007). Three wild Lactuca species, L. serriola, L. saligna and L. virosa were involved in this process (Mikel 2007). A further analysis of pedigree history of 146 lettuce cultivars registered in the U.S. by Plant Variety Protection and/or utility patent of the era from 2000 through 2010 have led to the determination of a coefficient of parentage among these cultivars (Mikel 2013). Among three crisphead lettuce ancestors, cultivars ‘Vanguard’ and ‘Salinas’ descend from the interspecific cross of L. sativa with L. virosa and L. serriola. Among these three cultivars, ‘Vanguard’ is the major ancestor contributing 23.8 % of the genes to crisphead lettuce. Cultivars ‘Vanguard’ and ‘Salinas’ followed by the cultivar ‘Calmar’ (which has no L. virosa in its ancestry) were determined as elite programme cultivars, frequently used in lettuce breeding (Mikel 2007). Of the 37 progenitor cultivars, breeders at public institutions and private companies in the USA developed 31 new cultivars in the period of 1970–2004, and ten of these progenitors of today’s lettuce germplasm were developed before 1960 (Mikel 2007). The cultivar ‘Salinas’ was frequently crossed with romaine lettuce types and the romain parental cultivar ‘Paris Island Cos’ was crossed with leaf types contributing to romaine and leaf lettuce genetic diversity (Mikel 2013).
Pedigree analysis of 146 lettuce cultivars registered in the USA in the period 2000–2010 demonstrated that leaf lettuce cultivars were more genetically diverse than romaine and crisphead cultivars (Mikel 2013). The lower diversity among romaine and crisphead cultivars is due to recurrent recycling of related cultivars in breeding programmes (Mikel 2013). This however, means that the percentage of cultivars possessing genes from wild species given above are likely to be underestimates as many modern day cultivars will have gained such genes through this recycling of parental cultivars most of which will have Vanguard and/or Salinas in their pedigrees both of which possess L. virosa genes.
Conclusions and future prospects
Lettuce (Lactuca sativa) is one of the oldest domesticated plants and vegetable crops (Hancock 2012). However, our knowledge about the origin, process of domestication, diversification and spread of lettuce around the globe is still quite fragmentary (Lebeda et al. 2007c). The genus Lactuca L. comprises approximately 100 wild species, however, detailed information about the biogeography and ecobiology of most of these species is not available (Lebeda et al. 2004b, 2007c, 2009a, 2012a). The taxonomy of wild Lactuca species and related genera is currently unclear (Lebeda et al. 2007c; Funk et al. 2009), and needs more detailed elaboration at the level of the genus, involving all known described species (Lebeda et al. 2007c, 2009a). Also the collection, conservation and evaluation of wild Lactuca germplasm is incomplete. Most (ca 90 %) of the currently available wild Lactuca genetic resources in world genebank collections is represented by accession of only three species (L. serriola, L. saligna, L. virosa) originating mostly from Europe and from the primary center of origin (Lebeda et al. 2004a, 2007c, 2009a, b). Although some collections have been made for example in the territory of North America (Lebeda et al. 2011, 2012a) in general, field studies and collecting activities have been reduced and neglected in the last few decades (Lebeda et al. 2009a). In some areas of the world (Africa, Asia) local landraces are still grown but the breeding and widespread marketing of modern cultivars often on a regional basis by multinational companies is leading to a loss of genetic diversity and local adaptation in the crop (Lebeda et al. 2007c). For many traits (e.g. Bremia resistance) extensive screening for new resistance genes/factors in gene bank accessions of the cultivated crop has led to to a diminshing return in terms of new resistances. This is leading to an increas in the use of wild Lactuca species germplasm as sources of new beneficial alleles for a range of valuable characters for future breeding progress (Lebeda and Boukema 2005).
Lettuce is one of the main horticultural crops where a strategy of wild related germplasm exploitation and utilization in breeding programmes is most commonly used with very high practical impact. During the last 70 years, the genus Lactuca has been intensively studied with respect to exploitation of wild relatives in commercial lettuce breeding (Pink and Keane 1993; Lebeda et al. 2007c; Mou 2008). In the last three decades, significant progress has been made in germplasm enhancement and the introduction of novel traits into cultivated lettuce, mostly from the primary lettuce gene-pool, however more recently the secondary and tertiary gene-pools have been accessed, particularly for disease and insect resistances (Lebeda et al. 2007c, 2009a). Unfortunately, the process of resistance breeding is complicated because the variation in host plant resistance is mirrored by the diversity (emergence of new different strains, pathotypes and races) within pathogen/pest populations (Lebeda et al. 2007c). Nevertheless, as is evident from this review, wild Lactuca germplasm are being increasingly studied and used in lettuce breeding. Valuable sources of resistance have been located in many accessions of different Lactuca species and successfully introduced to recent commercial lettuce cultivars. However, despite the progress that has been made, there are still many questions and problems which must be solved by close cooperation between plant scientists, ecologists, plant pathologists, geneticists and lettuce pre-breeders and breeders.
Current areas of weakness are the lack of detailed floristic, bio-geographic and ecologic delimitation of the distribution of known Lactuca spp., and few recent collecting and exploration missions, especially to the areas of high species richness and diversity (e.g. South Africa and Asia) (Lebeda et al. 2009a). Such activities need to be linked with detailed observations and recording of the occurrence of diseases and pests on weedy growing Lactuca species (e.g. Lebeda et al. 2008a, 2011, 2012a,2013) to provide a better understanding of host-pathogen/pest interactions in natural habitats and aid the exploitation of wild Lactuca germplams in lettuce resistance breeding.
Another relatively under researched area is the agro-ecological interface (Burdon and Thrall 2008), i.e. interactions between wild (weedy growing Lactuca spp.) and crop (lettuce, Lactuca sativa) pathosystems. For Lactuca spp. and lettuce this has only really been considered for interactions with Bremia lactucae (Lebeda and Petrželová 2004; Lebeda et al. 2002, 2008a) and Golovinomyces cichoracearum (Lebeda and Mieslerová 2011; Lebeda et al. 2012c, 2013). This type of knowledge may yield a better understanding about the maintenance of a dynamic balance (homeostasis) between hosts and pathogens in wild pathosystems which could inform deployment staregies for resistance genes in the cultivated crop as well as identify the potential risks of pathogen transfer from wild to crop pathosystems from the viewpoint of potential breakdown of resistance introduced to lettuce from wild Lactuca progenitors as demonsrated by the example of Lactuca serriola/L. sativa-Bremia lactucae interactions studied by Lebeda et al. (2008a). In addition the potential transfer of genes from lettuce to weedy growing Lactuca species (Hartman et al. 2012; Uwimana et al. 2012a, b) should be investigated from the viewpoint of stability of both the wild and cultivated systems.
From this review, and previous reports (Lebeda et al. 2007c, 2009a), it is clearly evident that wild Lactuca germplasm are highly valuable sources of genetic variation and resistance to different biotic stresses. However, it is also evident that current knowledge about these interactions is fragmentary covering only a limited part of the potential variation in the host-pathogen/pest interaction. This can be addressed by screening large collections of well defined wild Lactuca germplasm for resistance to the most important lettuce pathogens and pests, followed by detailed genetical studies of the inheritance of resistance. Where knowledge of pathogen effectors is available these can be substituted for the pathogen isolates, with the aim of finding recognition factors which recognise effectors which are highly conserved within the pathogen population. However, again knowledge of the variation in effectors in wild Lactuca spp/pathogen interactions is necessary to inform this strategy.
One of the main goals of current lettuce breeding is multiple disease and pest resistance. A detailed characterisation of wild Lactuca germplasm resistances can contribute to this goal. At least some Lactuca accessions have been shown to possess multiple resistances which can be introduced to cultivated lettuce and combined with agronomically important traits. The use of wild Lactuca spp. as sources of resistance has been hampered in the past by loss of quality characteristics, this has been particularly problematic when ‘poor’ quality loci have been linked to resistance loci e.g. stunting linked to Nasonovia resistance. However, many lettuce breeding companies have now addressed this by instigating ‘pre-breeding’ programes which aim to produce resistant lines of sufficient quality to be included in a commercial lettuce breeding programme. The use of markers to select for recombinants where undesirable linkages have been broken has been a great step forward in this respect.
New technologies and knowledge offer new ways to screen germplams collections for novel alleles. Currently 52 phenotypic loci that confer resistance in lettuce to 8 diseases (Bremia lactucae, Sphingomonas suberifaciens, Microdochium panattonianum, Fusarium oxysporum and Verticillium dahliae, lettuce root aphid, lettuce mosaic virus, and lettuce big vein) have been reported as being mapped in lettuce. For some of these diseases candidate genes have been identified and causality demonstrated for a subset of them using RNAi (Truco et al. 2013). This opens up a new more efficient strategy for assessing the value of Lactuca germplasm through resequencing to identify allelic variation to provide a library of alleles which can then be tested against the variation present in the pathogen either in the form of diverse isolates or where available, effectors. Those alleles which determine resistance against a broad spectrum of the variation in individual pathogens can then be combined using MAS to produce advanced breeding lines with multiple resistance (Michelmore 2013, pers. commun.).
In addition to the combination and pyramiding of race-specific resistance genes/factors (Lebeda et al. 2002; Pink 2002), exploitation of non-host resistance in lettuce breeding remains a challenge (Lebeda et al. 2002). In relation to lettuce non-host resistance was hypothetised for first time in the response of Lactuca saligna to infection by Bremia lactucae (Lebeda 1986). L. saligna is sexually compatible with L. sativa (Lebeda et al. 2007c). The concept of non-host resistance in L. saligna was later studied in detail from a mechanistic and population viewpoint (see e.g. Lebeda et al. 2002, 2008b; Petrželová et al. 2011), as well as a genetical viewpoint (Jeuken et al. 2008, 2009; Zhang et al. 2009). This resistance is hypothesised to be durable (Lebeda et al. 2002). However, the use of this type of resistance in lettuce breeding is still at the early stages and requires further research to develop the tools and knowledge for its exploitation (Jeuken 2012).
Future developments in methodology of interspecific hybridization, as well as transfer of resistance genes are playing important role in accessing the genetic variation present in wild Lactuca germplasm collections (Lebeda et al. 2007c; Davey and Anthony 2011). Both conventional breeding and genetic manipulation approaches will be very likely applied for the improvement of the crop. Conventional breeding augmented by marker assisted selection will continue to play a key role in the introgression of genetic material from wild Lactuca species into cultivated lettuce, particularly where quantitaitive resistance under the control of several loci is involved or where the aim is to pyramid several resistance genes. Somatic hybridization will enable novel nuclear-cytoplasmic combinations, and the introgression of beneficial alleles from wild Lactuca species which are sexually incompatible with lettuce (Davey and Anthony 2011). Improvements in transformation technologies are opening up the possibilities of deploying resistance genes in multilines to provide a more durable resistance crop phenotype as described by Pink and Puddephat (1999) and Pink (2002).
It is also evident that our knowledge of the mechanisms of resistance in various Lactuca-pathogen/pest interactions is limited. Only in a few plant-pathogen interactions (e.g. Lactuca-Bremia lactucae) is more detailed information about the type and mechanism of resistance, including features of inheritance available (Lebeda et al. 2002, 2008b). For many lettuce/disease interactions the basic information about the type of resistance (non-host versus host, race-specific vs. race-nonspecific, field or durable resistance etc.) is not available. Again this knowledge will improve our efficiency in exploiting the Lactuca genepool for lettuce crop improvement and allow the combination of different resistance mechanisms in a single cultivar in order to provide a potentially more durable resistance phenotype.
References
Abawi, G. S., & Grogan, R. G. (1979). Epidemiology of diseases caused by Sclerotinia species. Phytopathology, 69, 899–904.
Abawi, G. S., & Robinson, R. W. (1991). Reaction of selected lettuce germplasm to artificial inoculation by Meloidogyne hapla in the greenhouse. Journal of Nematology, 23, 519.
Abawi, G. S., Robinson, R. W., Cobb, A. C., & Shail, J. W. (1980). Reaction of lettuce germplasm to artificial inoculation with Sclerotinia minor under greenhouse conditions. Plant Disease, 64, 668–671.
Addoh, P. G. (1971). The distribution and economic importance of plant parasitic nematodes in Ghana. Ghana Journal of Agricultural Science, 4, 21–32.
Agrawal, A. A., & Konno, K. (2009). Latex: a model for understanding mechanisms, ecology, and evolution of plant defense against herbivory. The Annual Review of Ecology, Evolution, and Systematics, 40, 311–331.
Alconero, R. (1988). Lettuce (Lactuca sativa L.). In Y. P. S. Bajaj (Ed.), Biotechnology in agriculture and forestry (pp. 351–369). Berlin: Springer.
Alleyne, E. H., & Morrison, F. O. (1977). The lettuce root aphid, Pemphigus bursarius (L.) (Homoptera: Aphidoidea) in Quebec, Canada. Annals of Entomological Society Quebec, 22, 171–180.
Anonymous (2005). Development of lettuce breeding lines resistant to bacterial leaf spot. HortScience, 40, 1098.
Anonymous (2008). Resistance to the lettuce leaf aphid Nasonovia ribisnigri. Disclosure Number IPCOM000176078D dated 4 Nov. 2008. IP.com Prior Art Database Disclosure. <http://ip.com/IPCOM/000176078>.
Anwar, S. A., & McKenry, M. V. (2012). Incidence and reproduction of Meloidogyne incognita on vegetable crop genotypes. Pakistan Journal of Zoology, 44, 327–333.
Argyris, J., Truco, M. J., Ochoa, O., Knapp, S. J., Still, D. V., Lenssen, G. M., et al. (2005). Quantitative trait loci associated with seed and seedling traits in Lactuca. Theoretical and Applied Genetics, 111, 1365–1376.
Argyris, J., Truco, M. J., Ochoa, O., McHale, L., Dahal, P., van Deynze, A., et al. (2011). A gene encoding an abscisic acid biosynthetic enzyme (LsNCED4) collocates with the high temperature germination locus Htg6.1 in lettuce (Lactuca sp.). Theoretical and Applied Genetics, 122, 95–108.
Aruga, D., Tsuchiya, N., Matsumura, H., Matsumoto, E., & Hayashida, N. (2012). Analysis of RAPD and AFLP markers linked to resistance to Fusarium oxysporum f. sp lactucae race 2 in lettuce (Lactuca sativa L.). Euphytica, 187, 1–9.
Attalah, Z. K., Hayes, R. J., & Subbarao, K. V. (2011). Fifteen years of Verticillium wilt of lettuce in America’s Salad Bowl: a tale of immigration, subjugation, and abatment. Plant Disease, 95, 784–792.
Azzu, N., & Collette, L. (2008). Addressing the conservation and sustainable utilization of crop wild relatives: the international policy context. In N. Maxted, B. V. Ford–Lloyd, S. P. Kell, J. M. Iriondo, E. Dulloo, & J. Turok (Eds.), Crop wild relative conservation and use (pp. 31–40). Wallingford: CABI.
Bannerot, H., Boulidard, L., Marron, J., & Duteil, M. (1969). Etude de la tolerance au virus de la mosaïque de laitue chez la variété Gallega de invierno. Annales Phytopathologie, 1, 219–226.
Bao, Y., & Neher, D. A. (2011). Survey of lesion and northern root-knot nematodes associated with vegetables in Vermont. Nematropica, 41, 100–108.
Barak, J. D., Koike, S. T., & Gilbertson, R. L. (2001). Role of crop debrits and weeds in the epidemiology of bacterial leaf spot of lettuce in California. Plant Disease, 85, 169–178.
Barak, J. D., Koike, S. T., & Gilbertson, R. L. (2002). Movement of Xanthomonas campestris pv. vitians in the stems of lettuce and seed contamination. Plant Pathology, 51, 506–512.
Barbosa, P. (Ed.). (1998). Conservation biological control. New York: Academic.
Beharav, A., Ben-David, R., Doležalová, I., & Lebeda, A. (2008). Eco-geographical distribution of Lactuca saligna natural populations in Israel. Israel Journal of Plant Sciences, 56, 195–206.
Beharav, A., Ben-Roi, R., Doležalová, I., & Lebeda, A. (2010a). Eco-geographical distribution of Lactuca aculeata natural population in northeastern Israel. Genetic Resources and Crop Evolution, 57, 679–686.
Beharav, A., Ben-David, R., Malarz, J., Stojakowska, A., Michalska, K., Doležalová, I., et al. (2010b). Variation of sesquiterpene lactones in Lactuca aculeata natural population from Israel, Jordan and Turkey. Biochemical Systematics and Ecology, 38, 602–611.
Beharav, A., Lewinsohn, D., Lebeda, A., & Nevo, E. (2006). New wild Lactuca genetic resources with resistance against Bremia lactucae. Genetic Resources and Crop Evolution, 53, 467–474.
Bhat, R. G., & Subbarao, K. V. (1999). Host range specificity in Verticillium dahliae. Phytopathology, 89, 1218–1225.
Blackman, R. L., & Eastop, V. F. (2000). Aphids on the world’s crops. Chichester: John Wiley & Sons.
Blancard, D. (2011). Meloidogyne spp. (Galles racinaires des salades). INRA <(http://ephytia.inra.fr/salade/salade_utilisateur/index_appli.php?portail=legumes&produit=salade&main=1&ssrub1=8&ssrub2=10&ssrub3=16&ssrub4=81&id_fiche=28&theme=96> [last accessed 2013-02-07]
Blok, I., & van der Plaats-Niterink, A. J. (1978). Pythium uncinulatum sp. nov. and P. tracheiphilum pathogenic to lettuce. Netherlands Journal of Plant Pathology, 84, 135–147.
Bos, L., & Huijberts, N. (1990). Screening for resistance to big-vein disease of lettuce (Lactuca sativa). Crop Protection, 9, 446–452.
Bos, L., Huijberts, N., & Cuperus, C. (1994). Further observations on variation of lettuce mosaic virus in relation to lettuce (Lactuca sativa) and a discussion of resistance terminology. European Journal of Plant Pathology, 100, 293–314.
Boukema, I. W., Hazekamp, T., & van Hintum, T. J. L. (1990). The CGN collection reviews: The CGN lettuce collection. Wageningen: Centre for Genetic Resources, Netherlands.
Boydson, R. A., Mojtahedi, H., Crosslin, J. M., Thomas, P. E., Anderson, T., & Riga, E. (2004). Evidence for the influence of weeds on corky ringspot persistence in alfalfa and Scotch spearmint rotations. Američan Journal of Potato Research, 81, 215–225.
Bridge, J. (1976). Plant parasitic nematodes from the lowlands and highlands of Equador. Nematropica, 6, 18–23.
Brittlebank, C. C. (1919). Tomato disease. Journal of Department of Agriculture of Victoria Australia, 17, 231–235.
Brown, P. R., & Michelmore, R. W. (1988). The genetics of corky root resistance in lettuce. Phytopathology, 78, 1145–1150.
Burdon, J. J., & Thrall, P. H. (2008). Pathogen evolution across the agro-ecological interface: implications for disease management. Evolutionary Applications, 1, 57–65.
Carvalho Filho, J. L. S., Gomes, L. A. A., Westerich, J. N., Maluf, W. R., Campos, V. P., & Ferreira, S. (2008). Inheritance of resistance of ‘Salinas 88’ lettuce to the root-knot nematode Meloidogyne incognita (Kofoid & White) Chitwood. Revista Brasileira de Agrociência, 14, 279–289.
Carisse, O., Ouimet, A., Toussaint, V., & Phillon, V. (2000). Evaluation of the effect of seed treatments, bactericides, and cultivars on bacterial leaf spot of lettuce caused by Xanthomonas campestris pv. vitians. Plant Disease, 84, 295–299.
Chen, Y. H., Vhen, H. Y., Hsu, C. L., & Yen, G. C. (2007). Induction of apoptosis by the Lactuca indica L. in human leucemia cell line and its active components. Journal of Agricultural and Food Chemistry, 55, 1743–1749.
Chitambar, J. J. (1993). Host range of Hemicycliophora poranga and its pathogenicity on tomato. Fundamental and Applied Nematology, 16, 557–561.
Chitambar, J. J. (2007). Status of ten quarantine “A” nematode pests in California. California Plant Pest & Disease Report, 2, 62–75.
Cho, J. J., Mau, R. F. L., German, T. L., Hartmann, R. W., Yudin, L. S., Gonsalves, D., et al. (1989). A multidisciplinary approach to management of Tomato spotted wilt virus in Hawaii. Plant Disease, 73, 375–383.
Chupp, C., & Sherf, A. F. (1960). Vegetable diseases and their control. New York: Ronald Press.
Chupeau, M. C., Maisonneuve, B., Bellec, Y., & Chupeau, Y. (1994). A Lactuca universal hybridizer, and its use in creation of fertile interspecific somatic hybrids. Molecular Genetics and Gemonics, 245, 139–145.
Cid, M., Ávila, A., Garía, A., Abad, J., & Fereres, A. (2012). New sources of resistance to letuce aphids in Lactuca spp. Arthropod-Plant Interactions, 6, 655–669.
Cole, R. A., Sutherland, R. A., & Riggall, W. E. (1991). The use of polyacrylamide gradient gel electrophoresis to identify variation in isozymes as markers for Lactuca species and resistance to the lettuce root aphid Pemphigus bursarius. Euphytica, 56, 237–242.
Coutts, B. A., Thomas-Carroll, M. L., & Jones, R. A. C. (2004). Analysing spatial patterns of spread of Lettuce necrotic yellows virus and lettuce big-vein disease in lettuce field plantings. Annals of Applied Biology, 145, 339–343.
Crute, I. R. (1992). From breeding to cloning (and back again) a case-study with lettuce downy mildew. Annual Review of Phytopathology, 30, 485–506.
Crute, I. R., & Johnson, A. G. (1976). The genetic relationship between races of Bremia lactucae and cultivars of Lactuca sativa. Annals of Applied Biology, 83, 125–137.
Curtis, I. S., Caiping, H., Scott, R., Power, J. B., & Davey, M. R. (1996). Genomic male sterility in lettuce, a baseline for the production of F1 hybrids. Plant Science, 113, 113–119.
DAFF—Department of Agriculture, Fisheries and Forestry Biosecurity. (2012). Draft import risk analysis report for fresh ginger from Fiji. Canberra: Department of Agriculture, Fisheries and Forestry. CCBY 3.0.
D’Andrea, L., Felber, F., & Guadagnuolo, R. (2008). Hybridization rates between lettuce (Lactuca sativa) and its wild relative (L. serriola) under field conditions. Environmental Biosafety Research, 7, 61–71.
da Silveira, S. G. P. (1990). Two hosts of Aphelenchoides besseyi in Brazil. Nematologia Brasileira, 14, 146–150.
Davey, M. R., & Anthony, P. (2011). Lactuca. In C. Kole (Ed.), Wild crop relatives: genomic and breeding resources (pp. 115–128). Berlin: Heidelberg: Springer-Verlag.
Davey, M. R., Anthony, P., Power, J. B., & Lowe, K. C. (2007a). Leafy vegetables. In C. Kole & T. C. Hall (Eds.), Compendium of transgenic crop plants, Vol. 6, Transgenic vegetable crops (pp. 217–248). Chichester: Willey-Blackwell.
Davey, M. R., Anthony, P., Van Hooff, P., Power, J. B., & Lowe, K. C. (2007b). Lettuce. In E. C. Pua & M. R. Davey (Eds.), Biotechnology in agriculture and forestry, Vol. 59, Transgenic crops IV (pp. 221–249). Berlin: Springer.
Davey, M. R., McCabe, M. S., Mohapatra, U., & Power, J. B. (2002). Genetic manipulation of lettuce. In G. G. Khachatourians, A. McHughen, R. Scorza, W. K. Nip, & Y. H. Hui (Eds.), Transgenic plants and crops (pp. 613–635). New York: Marcel Dekker, Inc.
Davis, R. M., Subbarao, K. V., Raid, R. N., & Kurtz, E. A. (1997). Compendium of lettuce diseases. St. Paul: APS Press, The American Phytopathologica Society.
Davis, E. E., & Venette, R. C. (2004). Mini risk assessment false Columbia root-knot nematode: Meloidogyne fallax Karssen [Nematoda: Heteroderidae]. Department of Entomology, University of Minnesota <http://www.aphis.usda.gov/plant_health/plant_pest_info/pest_detection/downloads/pra/mfallaxpra.pdf> [last accessed 2013-01-22]
Davis, R. M., Winterbottom, C. Q., & Aguiar, J. L. (1995). First report of Pythium uncinulatum on romaine lettuce in California. Plant Disease, 79, 642.
de Carvalho, J. L. S., Gomes, L. A. A., Maluf, W. R., Oliveira, R. R., Costa, D. S., Fereira, S., et al. (2011). Resistance to Meloidogyne incognita race 1 in the lettuce cultivars grand rapids and Salinas-88. Euphytica, 182, 199–208.
de Vries, I. M. (1990). Crossing experiments of lettuce cultivars and species (Lactuca sect. Lactuca, Compositae). Plant Systematics and Evolution, 171, 233–248.
Dietzgen, R. G., Callaghan, B., Wetzel, T., & Dale, J. L. (2006). Completion of the genome sequence of Lettuce necrotic yellows virus, typespecies of the genus Cytorhabdovirus. Virus Research, 118, 16–22.
Dias, J. S., & Ortiz, R. (2012a). Transgenic vegetable breeding for nutritional quality and health benefits. Food and Nutrition Sciences, 3, 1209–1219.
Dias, J. S., & Ortiz, R. (2012b). Transgenic vegetable crops: progress, potentials and prospects. Plant Breeding Rewievs, 35, 151–246.
Dickinson, M. J., Jones, D. A., & Jones, J. D. G. (1993). Close linkage between the Cf-2/Cf-5 and Mi resistance loci in tomato. Molecular Plant-Microbe Interactions, 6, 341–347.
Dickson, M. H. (1963). Resistance to corky root rot in head lettuce. Americain Society for Horticultural Science, 82, 388–390.
Dinant, S. (1997). Coat protein mediated protection in Lactuca sativa against lettuce mosaic potyvirus strains. Molecular Breeding, 3, 75–86.
Dinant, S., Blaise, F., Kusiak, C., Astier-Manifacier, S., & Albouy, J. (1993). Heterologous resistance to potato virus Y in transgenic tobacco plants expressing the coat protein gene of lettuce mosaic potyvirus. Phytopathology, 83, 818–824.
Dinant, S., & Lot, H. (1992). Lettuce mosaic virus: a review. Plant Pathology, 41, 528–542.
Doležalová, I., Křístková, E., Lebeda, A., & Vinter, V. (2002). Description of morphological characters of wild Lactuca L. spp. genetic resources (English-Czech version). Horticultural Science (Prague), 29, 56–83.
Doležalová, I., Křístková, E., Lebeda, A., Vinter, V., Astley, D., & Boukema, I. W. (2003a). Basic morphological descriptors for genetic resources of wild Lactuca spp. Plant Genetic Resources Newsletter, 134, 1–9.
Doležalová, I., Lebeda, A., Dziechciarková, M., Křístková, E., Astley, D., & van de Wiel, C. C. M. (2003b). Relationships among morphological characters, isozymes polymorphism and DNA variability-the impact on Lactuca germplasm taxonomy. Czech Journal of Genetics and Plant Breeding, 39, 59–67.
Doležalová, I., Lebeda, A., & Křístková, E. (2001). Prickly lettuce (Lactuca serriola L.) germplasm collecting and distribution study in Slovenia and Sweden. Plant Genetic Resources Newsletter, 128, 41–44.
Doležalová, I., Lebeda, A., Křístková, E., & Novotná, A. (2005). Morphological variation of Lactuca serriola populations from some European countries. In XVII International Botanical Congress, Vienna, Austria, 17–23 July 2005, Abstracts. (p. 458).
Doležalová, I., Lebeda, A., Křístková, E., & Novotná, A. (2007). Relevance of morphologic assessment of wild Lactuca spp. germplasm for their taxonomic determination. Bulletin of Botanical Gardens, Museums & Collections, Polish Botanical Society, 16A, 22.
Doležalová, I., Lebeda, A., Tiefenbachová, I., & Křístková, E. (2004). Taxonomic reconsideration of some Lactuca spp. germplasm maintained in world genebank collections. Acta Horticulturae, 634, 193–201.
Dubois, V., Botton, E., Meyer, C., Rieu, A., Bedu, A., Maisonneuve, B., et al. (2005). Systematic silencing of a tobacco nitrate reductase transgene in lettuce (Lactuca sativa L.). Journal of Experimental Botany, 56, 2379–2388.
Duffus, J. E. (1961). Economic significance of beet western yellows (radish yellows) on sugar beet. Phytopathology, 51, 605–607.
Duffus, J. E., Liu, H. Y., Wisler, G. C., & Li, R. (1996). Lettuce chlorosis virus-a new whitefly trasmitted closterovirus. European Journal of Plant Pathology, 102, 591–596.
Dunn, J. A. (1959). The biology of the lettuce root aphid. Annals of Applied Biology, 47, 475–491.
Dziechciarková, M., Lebeda, A., Doležalová, I., & Astley, D. (2004). Characterization of Lactuca spp. germplasm by protein and molecular markers-a review. Plant Soil Environment, 50, 47–58.
Edwards, M. C., Gonsalves, D., & Provvidenti, R. (1983). Genetic analysis of cucumber mosaic virus in relation to host resistence: location of determinants for pathogenicity to certain legumes and Lactuca saligna. Phytopathology, 73, 269–273.
Eenink, A. H., & Dieleman, F. L. (1983). Inheritance of resistance to the leaf aphid Nasonovia ribis-nigri in the wild lettuce species Lactuca virosa. Euphytica, 32, 691–695.
Eenink, A. H., Dieleman, F. L., & Groenwold, R. (1982a). Resistance of lettuce (Lactuca) to the leaf aphid Nasonovia ribis-nigri. 2. Inheritance of the resistance. Euphytica, 31, 301–304.
Eenink, A. H., Groenwold, R., & Dieleman, F. L. (1982b). Resistance of lettuce (Lactuca) to the leaf aphid Nasonovia ribis-nigri. 1: transfer of resistance from L. virosa to L. sativa by interspecific crosses and selection of resistant breeding lines. Euphytica, 31, 291–300.
Elia, M., & Piglionica, V. (1964). Preliminary observations on the resistance of some lettuce cultivars to collar rot caused by Sclerotinia spp. Phytopathologia Mediterranea, 3, 37–39.
Ellis, P. R., McClement, S. J., Saw, P. L., Phelps, K., Vice, W. E., Kift, N. B., et al. (2002). Identification of sources of resistance in lettuce to the lettuce root aphid, Pemphigus bursarius-Resistance to lettuce root aphid. Euphytica, 125, 305–315.
Ellis, P. R., Pink, D. A. C., & Ramsey, A. D. (1994). Inheritance of resistance to lettuce root aphid in the lettuce cultivars ‘Avoncrisp’ and ‘Lakeland’. Annals of Applied Biology, 124, 141–151.
Farrara, B. F., & Michelmore, R. W. (1987). Identification of new sources of resistance to downy mildew in Lactuca spp. HortScience, 22, 647–649.
Feráková, V. (1977). The genus Lactuca L. in Europe. Bratislava: Univerzita Komenského.
Ferris, H. (2013). “The Nematode-plant expert information system”. A Virtual Encyclopedia on Soil and Plant Nematodes “NEMAPLEX”. Department of Nematology, University of California. <http://plpnemweb.ucdavis.edu/nemaplex>. [last accessed 2013-01-22]
Ford-Lloyd, B., Kell, S. P., & Maxted, N. (2008). Establishing conservation priorities for crop wild relatives. In N. Maxted, B. V. Ford–Lloyd, S. P. Kell, J. M. Iriondo, E. Dulloo, & J. Turok (Eds.), Crop wild relative conservation and use (pp. 110–119). Wallingford: CABI.
Fujinaga, M., Ogiso, H., Tsuchiya, N., Saito, H., Yamanaka, S., Nozue, M., et al. (2003). Race 3, a new race of Fusarium oxysporum f. sp. lactucae determined by a differential system with commercial cultivars. Journal of General Plant Pathology, 69, 23–28.
Fry, P. R., Close. R C., Procter, C. H., & Sunde, R. (1972). Lettuce necrotic yellows virus in New Zealand. Journal of Agricultural Research, 16, 143–146.
Funk, V. A., Susanna, A., Stuessy, T. F., & Bayer, R. J. (Eds.). (2009). Systematics, evolution and biogeography of compositae. Vienna: International Association for Plant Taxonomy.
Galea, V. J., & Price, T. V. (1988). Resistance of lettuce and related species to anthracnose (Microdochium panattonianum) in Australia. Plant Pathology, 37, 363–372.
Galea, V. J., Price, T. V., & Sutton, B. C. (1986). Taxonomy and biology of the lettuce anthracnose fungus. Transactions of the British Mycological Society, 86, 619–628.
Garibaldi, A., Gilardi, G., & Gullino, M. L. (2004). Varietal resistance of lettuce to Fusarium oxysporum f. sp. lactucae. Crop Protection, 23, 845–851.
Garibaldi, A., Gilardi, G., & Gullino, M. L. (2007). First report of Verticillium wilt caused by Verticillium dahliae on lettuce in Italy. Plant Disease, 91, 990.
Gaskin, T. A. (1958). Weed hosts of Meloidogyne incognita in Indiana. Plant Disease Reporter, 42, 802–803.
George, R. A. T. (1999). Vegetable seed production (2nd ed.). Wallingford: CABI Publishing.
Gergerich, R. C., & Dolja, V. V. (2006). Introduction to plant viruses, the invisible Foe. The Plant Health Instructor. doi:10.1094/PHI-I-2006-0414-01.
German-Retana, S., Walter, J., & Le Gall, O. (2008). Lettuce mosaic virus: from pathogen diversity to host interactors. Molecular Plant Pathology, 9, 127–136.
German-Retana, S., Candresse, T., & Martelli, G. (1999). Closteroviruses (Closteroviridae). In Encyclopedia of virology (2nd ed., pp. 266–273). San Diego: Academic.
Gharabadiyan, F., Jamali, S., Yazdi, A. A., Hadizadeh, M. H., & Eskandari, A. (2012). Weed hosts of root-knot nematodes in tomato fields. Journal of Plant Protection Research, 52, 230–234.
Giannino, D., Nicolodi, C., Testone, G., Di Giacomo, E., Iannelli, M. A., Frugis, G., et al. (2008). Pollen-mediated transgene flow in lettuce (Lactuca sativa L.). Plant Breeding, 127, 308–314.
Gilberton, R. L. (1996). Management and detection of LMV: production of LMV resistant lettuce and LMV coat protein antibodies (pp. 78–81). Iceberg Lettuce Advisory Board Annual Report.
Gomes, L. A. A., Maluf, W. R., & Campos, V. P. (2000). Inherintance of the resistance reaction of the lettuce cultivar ‘Grand Rapids’ to the southern root-knot nematode Meloidogyne incognita (Kofoid & White) Chitwood. Euphytica, 114, 34–46.
Gowda, D. N., Kurdikeri, C. B., & Gowda, C. K. (1995). Weeds as hosts of root-knot nematodes. Indian Journal of Nematology, 25, 215–216.
Groenwold, R. (1983). Onderzoek van de relatie tussen Lactuca en Bremia lactucae. Verslag van een voorlichtingsbijeenkomst voor slaveredelaars (vervolg). Zaadbelangen, 37, 132.
Groves, R. L., Walgenbach, J. F., Moyer, J. W., & Kennedy, G. G. (2002). The role of weed hosts and tobacco thrips, Frankliniella fusca, in the epidemiology of tomato spotted wilt virus. Plant Disease, 86, 573–582.
Grube, R. C. (2004). Genetic analysis of resistance to lettuce drop caused by Sclerotinia minor. Acta Horticulturae, 637, 49–53.
Grube, R. C., Hayes, R., Mou, B., & McCreight, J. D. (2005a). Lettuce breeding, USDA-ARS. California Lettuce Research Board Annual Report, 2004–2005.
Grube, R., & Ryder, E. (2004). Identification of lettuce (Lactuca sativa L.) germplasm with genetic resistance to drop caused by Sclerotinia minor. Journal of the American Society for Horticultiral Science, 129, 70–76.
Grube, R. C., Wintermantel, W. M., Hand, P., Aburomia, R., Pink, D. A. C., & Ryder, E. J. (2005b). Genetic analysis and mapping of resistance to lettuce dieback: a soilborne disease caused by tombusviruses. Theoretical and Applied Genetics, 110, 259–268.
Haley, V., & McCreight, J. D. (1990). Resistance of wild lettuce (Lactuca saligna L.) to lettuce infectious yellows virus. HortScience, 25, 1163. Abstract.
Hampton, R. O., Keller, K. E., & Baggett, J. R. (1998). Beet western yellows luteovirus in Western Oregon. Plant Disease, 82, 140–148.
Hancock, J. F. (2012). Plant evolution and the origin of crop species (3rd ed.). Wallingford: CABI.
Hartman, Y., Hooftman, D. A. P., Schranz, M. E., & van Tienderen, P. H. (2013a). QTL analysis reveals the genetic architecture of domestication traits in Crisphead lettuce. Genetic Resources and Crop Evolution, 60, 1487–1500.
Hartman, Y., Hooftman, D. A., Uwimana, B., van de Wiel, C. C. M., Smulders, M. J., Visser, R. G., et al. (2012). Genomic regions in crop-wild hybrids of lettuce are affected differently in different environments: implications for crop breeding. Evolutionary Applications, 5, 629–640.
Hartman, Y., Uwimana, B., Hooftman, D. A. P., Schranz, M. E., van de Wiel, C. C. M., Smulders, M. J. M., et al. (2013b). Genomic and environmental selection patterns in two distinct lettuce crop-wild hybrid crosses. Evolutionary Applications. doi:10.1111/eva.12043.
Hayes, R. J., Maruthachalam, K., Vallad, G. E., Klosterman, S. J., & Subbarao, K. V. (2011a). Selection for resistance to Verticillium wilt caused by race 2 isolates of Verticillium dahliae in accessions of lettuce (Lactuca sativa L.). HortScience, 46, 201–206.
Hayes, R. J., & Ryder, E. J. (2007). Introgression of novel alleles for partial resistance to big vein disease from Lactuca virosa into Cultivated Lettuce. HortScience, 42, 3539.
Hayes, R. J., Ryder, E. J., & Robinson, B. (2004). Introgression of big vein tolerance from Lactuca virosa L. into cultivated lettuce (Lactuca sativa L.). HortScience, 39, 881.
Hayes, R. J., Ryder, E. J., & Wintermantel, W. M. (2008). Genetic variation for big-vein symptom expression and resistance to Mirafiori lettuce big vein virus in Lactuca virosa L., and wild relative of cultivated lettuce. Euphytica, 164, 493–500.
Hayes, R. J., McHale, L. K., Vallad, G. E., Truco, M. J., Michelmore, R. W., Klosterman, S. J., et al. (2011b). The inheritance of resistance to Verticillium wilt caused by race 1 isolates of Verticillium dahliae in the lettuce cultivar La Brillante. Theoretical and Applied Genetics, 123, 509–517.
Hayes, R. J., Vallad, G. E., McHale, L. K., Truco, M. J., Ochoa, O. E., Michelmore, R. W., et al. (2009). Breeding for resistance-new approaches and challenges. Phytopathology, 99, S168.
Hayes, R. J., Vallad, G. E., Qin, Q. M., Grube, R. C., & Subbarao, K. V. (2007a). Variation for resistance to Verticillium wilt in lettuce (Lactuca sativa L). Plant Disease, 91, 439–445.
Hayes, R. J., Vallad, G. E., & Subbarao, K. V. (2007b). The inheritance of resistance to race 1 isolates of Verticillium dahliae in lettuce. HortScience, 37, 1015–1022.
Hayes, R. J., Wintermantel, W. M., Nicely, P. A., & Ryder, E. J. (2006). Host resistance to mirafiori lettuce big-vein virus and lettuce big-vein associated virus and virus sequence diversity and frequency in California. Plant Disease, 90, 233–239.
Hayes, R. J., Wu, B. M., Pryor, B. M., Chitrampalam, P., & Subbarao, K. V. (2010). Assessment of resistance in lettuce (Lactuca sativa L.) to mycelial and ascospore infection by Sclerotinia minor Jagger and S. sclerotiorum (Lib.) de Bary. HortScience, 45, 333–341.
Hill, M., Witsenboer, H., Zabeau, M., Vos, P., Kesseli, R., & Michelmore, R. (1996). PCR-based fingerprinting using AFLPs as a tool for studying genetic relationships in Lactuca spp. Theoretical and Applied Genetics, 93, 1202–1210.
Hobbs, H. A., Black, L. L., Story, R. N., Valverde, R. A., Bond, W. P., Gatti, J. M., et al. (1993). Transmission of tomato spotted wilt virus from pepper and three weed hosts by Frankliniella fusca. Plant Disease, 77, 797–799.
Hooftman, D. A. P., Flavell, A. J., Jansen, H., den Nijs, H. C. M., Syed, N. H., Sørensen, A. P., et al. (2011). Locus-dependent selection in crop-wild hybrids of lettuce under field conditions and its implication for GM crop development. Evolutionary Applications, 4, 648–659.
Hooftman, D. A. P., Hartman, Y., Oostermeijer, J. G. B., & den Nijs, J. C. M. (2009). Existence of vigorous lineages of crop-wild hybrids in lettuce under field conditions. Environmental Biosafety Research, 8, 203–217.
Hu, J. G., Ochoa, O. E., Truco, M. J., & Vick, B. A. (2005). Application of the TRAP technique to lettuce (Lactuca sativa L.) genotyping. Euphytica, 144, 225–235.
Huang, J., McAuslane, H. J., & Nuessly, G. S. (2003). Resistance in lettuce to Diabrotica balteata (Coleoptera: Chrysomelidae): the roles of latex and inducible defense. Environmental Entomology, 32, 9–16.
Huang, X., & Ploeg, A. T. (2001). Effect of plant age and Longidorus africanus on the growth of lettuce and carrot. Journal of Nematology, 33, 2–3.
Hubbard, J. C., & Gerik, J. S. (1993). A new wilt disease of lettuce incited by Fusarium oxysporum f. sp. lactucum forma specialis nov. Plant Disease, 77, 750–754.
Iriondo, J. M., & De Hond, L. (2008). Crop wild relative in-situ management and monitoring: the time has come. In N. Maxted, B. V. Ford–Lloyd, S. P. Kell, J. M. Iriondo, E. Dulloo, & J. Turok (Eds.), Crop wild relative conservation and use (pp. 319–330). Wallingford: CABI.
Jagger, I. C. (1921). A transmissible mosaic disease of lettuce. Journal of Agricultural Research, 20, 737–741.
Jagger, I. C., & Chandler, N. (1934). Big vein, a disease of lettuce. Phytopathology, 24, 1253–1256.
Jeuken, M. (2012). Industry highlights. Breeding for durable resistance against an oomycete in lettuce. In G. Acquaah (Ed.), Principles of plant genetics and breeding. Second edition (chapter 14) (pp. 273–276). Chichester: Wiley-Blackwell.
Jeuken, M., & Lindhout, P. (2002). Lactuca saligna, a non-host for lettuce downy mildew (Bremia lactucae), harbors a new race-specific Dm gene and three QTLs for resistance. Theoretical and Applied Genetics, 105, 384–391.
Jeuken, M., & Lindhout, P. (2004). The development of lettuce backcross inbred lines (BILs) for exploitation of the Lactuca saligna (wild lettuce) germplasm. Theoretical and Applied Genetics, 109, 394–401.
Jeuken, M. J. W., Pelgrom, K., Stam, P., & Lindhout, P. (2008). Efficient QTL detection for nonhost resistance in wild lettuce: backcross inbred lines versus F2 population. Theoretical and Applied Genetics, 116, 845–857.
Jeuken, M., van Wijk, R., Peleman, J., & Lindhout, P. (2001). An integrated interspecific AFLP map of lettuce (Lactuca) based on two L. sativa × L. saligna F-2 populations. Theoretical and Applied Genetics, 103, 638–647.
Jeuken, M. J. W., Zhang, N. W., McHale, L. K., Pelgrom, K., den Boer, E., Lindhout, P., et al. (2009). Rin4 causes hybrid necrosis and race-specific resistance in an interspecific lettuce hybrid. Plant Cell, 21, 3368–3378.
Johnson, W. C., Jackson, L. E., Ochoa, O., van Wijk, R., Peleman, J., St. Clarir, D. A., et al. (2000). Lettuce, a shallowrooted crop, and Lactuca serriola, its wild progenitor, differ at QTL determining root architecture and deep soil water exploitation. Theoretical and Applied Genetics, 101, 1066–1073.
Jones, D. A., Dickinson, M. J., Balint-Kurti, P. J., Dixon, M. S., & Jones, J. D. G. (1993). Two complex resistance loci revealed in tomato by classical and RFLP mapping of the Cf-2, Cf-4, Cf-5 and Cf-9 genes for resistance to Cladosporium fulvum. Molecular Plant-Microbe Interactions, 6, 348–357.
Judelson, H. S., & Michelmore, R. W. (1992). Temperature and genotype interactions in the expression of host resistance in lettuce downy mildew. Physiological and Molecular Plant Pathology, 40, 233–245.
Kanamoto, H., Yamashita, A., Asao, H., Okumura, S., Takase, H., Hattori, M., et al. (2006). Efficient and stable transformation of Lactuca sativa L. cv. ‘Cisco’ (lettuce) plastids. Transgenic Research, 15, 205–217.
Kaur, P., & Mitkowski, N. A. (2010). Evaluation of Lactuca germplasm for resistance to the northern root-knot nematode (Meloidogyne hapla Chitwood). International Journal of Vegetable Science, 17, 26–36.
Kawazu, Y., Fujiyama, R., & Noguchi, Y. (2009). Transgenic resistance to Mirafiori lettuce virus in lettuce carrying inverted repeats of the viral coat protein gene. Transgenic Research, 18, 113–120.
Kawazu, Y., Fujiyama, R., Noguchi, Y., Kjubota, M., Ito, H., & Fukuoka, H. (2010). Detailed characterization of Mirafiori lettuce virus-resistant transgenic lettuce. Transgenic Research, 19, 211–220.
Kesseli, R., Ochoa, O., & Michelmore, R. (1991). Variation at RFLP loci in Lactuca spp. and origin of cultivated lettuce (L. sativa). Genome, 34, 430–436.
Kesseli, R. V., Paran, I., & Michelmore, R. W. (1994). Analysis of a detailed genetic linkage map of Lactuca sativa (Lettuce) constructed from RFLP and RAPD markers. Genetics, 136, 1435–1446.
Kim, K. H., Kim, Y. H., & Lee, K. R. (2007). Isolation of quinic acid derivatives and flavonoids from the aerial parts of Lactuca indica L. and their hepatoprotective activity in vitro. Bioorganic and Medicinal Chemistry Letters, 17, 6739–6743.
Kisiel, W., & Barszcz, B. (1998). A germacrolide glucoside from Lactuca tatarica. Phytochemistry, 48, 205–206.
Kisiel, W., & Michalska, K. (2009). Lignans ansd sesquiterpenoids from Lactuca sibirica. Fitoterapia, 79, 241–244.
Kisiel, W., & Zielinska, K. (2000). Sesquiterpenoids and phenolics from Lactuca perennis. Fitoterapia, 71, 86–87.
Kitner, M., Lebeda, A., Doležalová, I., Maras, M., Křístková, E., Nevo, E., et al. (2008). AFLP analysis of Lactuca saligna germplasm collections from four European and three Middle Eastern countries. Israel Journal of Plant Sciences, 56, 185–193.
Klocke, E., Nothnagel, T., & Schumann, G. (2010). Vegetables. In F. Kempken & C. Jung (Eds.), Genetic modification of plants, agriculture, horticulture and forestry (pp. 449–552). Berlin: Springer.
Koenning, S. R., Overstreet, C., Noling, J. W., Donald, P. A., Becker, J. O., & Fortnum, B. A. (1999). Survey of crop losses in response to Phytoparasitic Nematodes in the United States for 1994. Supplement to the Journal of Nematology, 31, 587–618.
Kohl, L. M. (2011). Astronauts of the Nematode World: An Aerial View of Foliar Nematode Biology, Epidemiology, and Host Range. APSnet Features. doi:10.1094/APSnetFeature-2011-0111.
Koopman, W. J. M. (1999). Plant systematics as useful tool for plant breeders, examples from lettuce. In A. Lebeda & E. Křístková (Eds.), Eucarpia leafy vegetables ′99, proceedings of the eucarpia meeting on leafy vegetables genetics and breeding (pp. 95–105). Olomouc: Palacký University in Olomouc.
Koopman, W. J. M. (2000). Identifying lettuce species (Lactuca subs. Lactuca, Asteraceae). A practical application of flow cytometry. Euphytica, 116, 151–159.
Koopman, W. J. M. (2002). Zooming in on the lettuce genome: Species relationships in Lactuca s.l. inferred from chromosomal and molecular characters. Ph.D. diss., Wageningen University, The Netherlands.
Koopman, W. J. M., & de Jong, H. J. (1996). A numerical analysis of karyotypes and DNA amounts in lettuce cultivars and species (Lactuca subs. Lactuca, Compositae). Acta Botanica Neerlandica, 45, 211–222.
Koopman, W. J. M., Guetta, E., Van de Wiel, C. C. M., Vosman, B., & Van den Berg, R. G. (1998). Phylogenetic relationships among Lactuca (Asteraceae) species and related genera based on ITS-1 DNA sequences. American Journal of Botany, 85, 1517–1530.
Koopman, W. J. M., Zevenbergen, M. J., & Van den Berg, R. G. (2001). Species relationships in Lactuca s.l. (Lactuceae, Asteraceae) inferred from AFLP fingerprints. American Journal of Botany, 88, 1881–1887.
Krause-Sakate, R., Le Gall, O., Fakhfakh, H., Peypelut, M., Marrakchi, M., Varveri, C., et al. (2002). Molecular and biological characterization of Lettuce mosaic virus (LMV) isolates reveals a distanct and widespread type of resistance-breaking isolate: LMV-Most. Phytopathology, 92, 563–572.
Krause-Sakate, R., Mello, N. R., Pavan, A. M., Zambolim, M. E., Carvalho, G. M., Le Gall, O., et al. (2001). Molecular characterization of two brazilian isolates of Lettuce mosaic virus with distinct biological properties. Phytopathologia Brasileira, 26, 153–157.
Křístková, E., Lebeda, A., & Doležalová I. (2007a). Phenotypic variability of Lactuca saligna germplasm collected in Italy and France. In EUCARPIA Leafy Vegetables 2007, Conference Abstracts (p. 15). Warwick: University of Warwick.
Křístková, E., Lebeda, A., Doležalová, I., Vinter, V., & Křístková, A. (2007b). Variation in developmental stages of Lactuca serriola L. (prickly lettuce) germplasm from different European countries. In EUCARPIA Leafy Vegetables 2007, Conference Abstracts (p. 16). Warwick: University of Warwick.
Křístková, E., Lebeda, A., Kitner, M., Vafková, B., Matoušková, Z., Doležalová, I., et al. (2012). Phenotypes of the natural interspecific hybrids in the genus Lactuca. Úroda, 60, 28–31.
Křístková, E., Tvardková, M., & Lebeda, A. (2011). Characterization of developmental stages in Lactuca saligna germplasm from Europe and USA. In Eucarpia Leafy Vegetables 2011. Abstract (p.78). Villneuve d’ Ascq: Université Lille Nord de France.
Kuang, H., Ochoa, O. E., Nevo, E., & Michelmore, R. W. (2006). The disease resistance gene Dm3 is infrequent in natural populations of Lactuca serriola due to deletions and frequent gene conversions at the RGC2 locus. Plant Journal, 47, 38–48.
Kuang, H., van Eck, H. J., Sicard, D., Michelmore, R., & Nevo, E. (2008). Evolution and genetic population structure of prickly lettuce (Lactuca serriola) and its RGC2 resistance gene cluster. Genetics, 178, 1547–1558.
Kumar, S., Ray, J., Davison, E. M., Cunnington, J. H., & de Alwis, S. (2007). First record of Pythium tracheiphilum associated with lettuce wilt and leaf blight in Australia. Australasian Plant Disease Notes, 2, 7–9.
Kwon, S. J., Truco, M. J., & Hu, J. G. (2012). LSGermOPA, a custom OPA of 384 EST-derived SNPs for high-throughput lettuce (Lactuca sativa L.) germplasm fingerprinting. Molecular Breeding, 29, 887–901.
Lebeda, A. (1984). Race-specific factors of resistance to Bremia lactucae in the world assortment of lettuce. Scientia Horticulturae, 22, 23–32.
Lebeda, A. (1985a). Differences in resistance of wild Lactuca species to natural infection of lettuce powdery mildew (Erysiphe cichoracearum). Euphytica, 34, 521–523.
Lebeda, A. (1985b). Susceptibility of some lettuce cultivars to natural infection by powdery mildew. Tests of Agrochemicals and Cultivars No. 6 (Annals of Applied Biology 106, Suppl.), 158–159.
Lebeda, A. (1985c). Auftreten der Natürlichen Infektion durch den Echten Mehltau (Erysiphe cichoracearum) bei der Gattung Lactuca in der Tschechoslowakei. Acta Phytopathologica Academiae Scientarum Hungaricae, 20, 149–162.
Lebeda, A. (1986). Specificity of interactions between wild Lactuca spp. and Bremia lactucae isolates from Lactuca serriola. Journal of Phytopathology, 117, 54–64.
Lebeda, A. (1990). The location of sources of field resistance to Bremia lactucae in wild Lactuca species. Plant Breeding, 105, 75–77.
Lebeda, A. (1994). Evaluation of wild Lactuca species for resistance of natural infection of powdery mildew (Erysiphe cichoracearum). Genetic Resources and Crop Evolution, 41, 55–57.
Lebeda, A., & Astley, D. (1999). World genetic resources of Lactuca spp., their taxonomy and biodiversity, In A. Lebeda, & E. Křístková (Eds.), Eucarpia Leafy Vegetables ′99, Proceedings of the Eucarpia Meeting on Leafy Vegetables Genetics and Breeding. (pp. 81–94). Olomouc: Palacký University in Olomouc.
Lebeda, A., & Blok, I. (1991). Race-specific resistance genes to Bremia lactucae in new Czechoslovak lettuce cultivars and location of resistance in a Lactuca serriola × Lactuca sativa hybrid. Archiv für Phytopathologie und Pflanzenschutz, 27, 65–72.
Lebeda, A., & Boukema, I. W. (2001). Leafy vegetables genetic resources. In L. Maggioni, & O. Spellman (Eds.), Report of a Network Coordinating Group on Vegatables; Ad hoc meeting, 26–27 May 2000, Vila Real, Portugal. (pp. 48–57). Rome: IPGRI.
Lebeda, A., & Boukema, I. W. (2005). Ad Hoc meeting on leafy vegetables. In G. Thomas, D. Astley, I. Boukema, M. C. Daunay, A. Del Greco, M. J. Diez, W. van Dooijeweert, J. Keller, T. Kotlinska, A. Lebeda, E. Lipman, L. Maggioni, & E. Rosa (Eds.), Report of a Vegetables Network (pp. 82–94). Joint meeting with and ad hoc group on leafy vegetables, 22–24 May 2003. Skierniewice: International Plant Genetic Resources Institute.
Lebeda, A., Doležalová, I., & Astley, D. (2004a). Representation of wild Lactuca spp. (Asteraceae, Lactuceae) in world genebank collections. Genetic Resources and Crop Evolution, 51, 167–174.
Lebeda, A., Doležalová, I., Feráková, V., & Astley, D. (2004b). Geographical distribution of wild Lactuca spp. (Asteraceae, Lactuceae). Botanical Reviews, 70, 328–356.
Lebeda, A., Doležalová, I., Janeček, J. & Gasmanová, N. (2004c). Differences in relative DNA content od Lactuca serriola germplasm collected in Europe. In Summaries and Program, 17th International lettuce and leafy vegetable conference, 28–31 August 2004, Sandman Hotel, Montreal-Longueuil, Agriculture and Applied Food, Canada, Montreal, pp. 29–30.
Lebeda, A., Doležalová, I., Kitner, M., Novotná, A., Šmachová, P., & Widrlechner, M. P. (2011). North American continent-a new source of wild Lactuca spp. germplasm variability for future lettuce breeding. Acta Horticulturae, 918, 475–482.
Lebeda, A., Doležalová, I., Křístková, E., Dehmer, K. J., Astley, D., van de Wiel, C. C. M., et al. (2007a). Acquisition and ecological characterization of Lactuca serriola L. germplasm collected in the Czech Republic, Germany, the Netherlands and United Kingdom. Genetic Resources and Crop Evolution, 54, 555–562.
Lebeda, A., Doležalová, I., Křístková, E., Kitner, M., Petrželová, I., Mieslerová, B., et al. (2009a). Wild Lactuca germplasm for lettuce breeding: recent status, gaps and challenges. Euphytica, 170, 15–34.
Lebeda, A., Doležalová, I., Křístková, E., & Mieslerová, B. (2001b). Biodiversity and ecogeography of wild Lactuca spp. in some European countries. Genetic Resources and Crop Evolution, 48, 153–164.
Lebeda, A., Doležalová, I., Křístková, E., Mieslerová, B., Kitner, M., Navrátilová, B., Duchoslav, M., Havránek, P., & Vondráková, D. (2007b). Germplasm collections of crop wild relatives – research, study and use on the Department of Botany, Palacký University in Olomouc (Czech Republic). In P. Hauptvogel, D. Benediková, R. Hauptvogel (Eds.), Plant Gentic Resources and their Exploitation in the Plant Breeding for Food and Agriculture, Book of abstracts, 18 th Eucarpia Genetic Resources Section Meeting. (pp. 94–95). Piešťany: NP print s.r.o. Piešťany.
Lebeda, A., Doležalová, I., & Novotná, A. (2012a). Wild and weedy Lactuca species, their distribution, ecogeography and ecobiology in USA and Canada. Genetic Resources and Crop Evolution, 59, 1805–1822.
Lebeda, A., Kitner, M., Křístková, E., Doležalová, I., & Beharav, A. (2012b). Genetic polymorphism in Lactuca aculeata populations and occurrence of natural putative hybrids between L. aculeata and L. serriola. Biochemical Systematics and Ecology, 42, 113–123.
Lebeda, A., Kitner, M., Dziechciarková, M., Doležalová, I., Křístková, E., & Lindhout, P. (2009b). An insight into the genetic polymorphism among European populations of Lactuca serriola assessed by AFLP. Biochemical Systematics and Ecology, 37, 597–608.
Lebeda, A., & Mieslerová, B. (2011). Taxonomy, distribution and biology of lettuce powdery mildew (Golovinomyces cichoracearum sensu stricto). Plant Pathology, 60, 400–415.
Lebeda A., Mieslerová, B., Petrželová, I., & Korbelová, P. (2013). Host specificity and virulence variation in populations of lettuce powdery mildew pathogen (Golovinomyces cichoracearum s. str.) from prickly lettuce (Lactuca serriola). Mycological Progress, 12, 533–545.
Lebeda, A., Mieslerová, B., Petrželová, I., Korbelová, P., & Česneková, E. (2012c). Patterns of virulence variation in the interaction between Lactuca spp. and lettuce powdery mildew (Golovinomyces cichoracearum). Fungal Ecology, 5, 670–682.
Lebeda, A., & Petrželová, I. (2004). Variation and distribution of virulence phenotypes of Bremia lactucae in natural populations of Lactuca serriola. Plant Pathology, 53, 316–324.
Lebeda, A., Petrželová, I., & Maryška, Z. (2008a). Structure and variation in the wild-plant pathosystem: Lactuca serriola—Bremia lactucae. European Journal of Plant Pathology, 122, 127–146.
Lebeda, A., & Pink, D. A. C. (1998). Histological aspects of the response of wild Lactuca spp. and their hybrids, with L. sativa to lettuce downy mildew (Bremia lactucae). Plant Pathology, 47, 723–736.
Lebeda, A., Pink, D. A. C., & Astley, D. (2002). Aspects of the interactions between wild Lactuca spp. and related genera and lettuce downy mildew (Bremia lactucae). In P. T. N. Spencer-Phillips, U. Gisi, & A. Lebeda (Eds.), Advances in downy mildew research (pp. 85–117). Dordrecht: Kluwer.
Lebeda, A., Ryder, E. J., Grube, R., Doležalová, I., & Křístková, E. (2007c). Lettuce (Asteraceae; Lactuca spp.). In R. J. Singh (Ed.), Genetic resources, chromosome engineering, and crop improvement, Vol. 3, Vegetable crops (pp. 377–472). Boca Raton: CRC Press, Taylor and Francis Group.
Lebeda, A., Sedlářová, M., Lynn, J., & Pink, D. A. C. (2006). Phenotypic and histological expression of different genetic backgrounds in interactions between lettuce, wild Lactuca spp., L. sativa × L. serriola hybrids and Bremia lactucae. European Journal of Plant Pathology, 115, 431–441.
Lebeda, A., Sedlářová, M., Petřivalský, M., & Prokopová, J. (2008b). Diversity of defence mechanisms in plant-oomycete interactions: a case study of Lactuca spp. and Bremia lactucae. European Journal of Plant Pathology, 122, 71–89.
Lebeda, A., & Zinkernagel, V. (2003a). Characterization of new highly virulent German isolates of Bremia lactucae and efficiency of resistance in wild Lactuca spp. germplasm. Journal of Phytopathology, 151, 274–282.
Lebeda, A., & Zinkernagel, V. (2003b). Evolution and distribution of virulence in the German population of Bremia lactucae. Plant Pathology, 52, 41–51.
Ligoxigakis, E. K., Vakalounakis, D. J., & Thanassoulopoulos, C. C. (2002). Weed hosts of Verticillium dahliae in Crete: susceptibility, symptomatoology and significance. Phytoparasitica, 30, 511–518.
Lindqvist, K. (1960). Cytogenetic studies in the Serriola group of Lactuca. Hereditas, 46, 75–151.
Liu, Z. B. (2004). Distribution and population development of Nasonovia ribisnigri (Homoptera: Aphididae) in iceberg lettuce. Journal of Economic Entomology, 97, 883–890.
Liu, Y. B., & McCreight, J. D. (2006). Responses of Nasonovia ribisnigri (Homoptera: Aphididae) to susceptible and resistant lettuce. Journal of Economic Entomology, 99, 972–978.
Lu, Q. Y., Baker, J., & Preston, C. (2007). The spread of resistence to acetolactate synthase inhibiting herbicides in a wind borne, self-pollinated weed species, Lactuca serriola L. Theoretical and Applied Genetics, 115, 443–450.
MacGowan, J. B. (1982). Needle nematodes: Longidorus spp.. Nematology Circular No. 89. Florida Department of Agriculture and Consumer Services, Contribution No. 250.
Machado, A. C. Z., & Inomoto, M. M. (2001). Host status of eighteen vegetable crops for Pratylenchus brachyurus. Nematropica, 31, 259–265.
Mackenzie, J. R., & Vernon, R. S. (1988). Sampling for distribution of the lettuce aphid, Nasonovia ribisnigri (Homoptera: Aphididae), in fields and within heads. Journal of the Entomological Society of British Columbia, 85, 10–14.
Maisonneuve, B. (2003). Lactuca virosa, a source of disease resistance genes in lettuce breeding: results and difficulties for gene introgression. In Th. J. L. van Hintum, A. Lebeda, D. Pink, & J.W. Schur (Eds.), Eucarpia Leafy Vegetables Conference (pp. 61–67). Noordwijkerhout, Netherlands, 19–23 May 2003.
Maisonneuve, B., Bellec, Y., Souche, S., & Lot, H. (1999). New resistance against downy mildew and lettuce mosaic potyvirus in wild Lactuca spp. In A. Lebeda & E. Křístková (Eds.), Eucarpia leafy vegetables ′99 (pp. 191–197). Olomouc: Palacky University, Olomouc.
Maisonneuve, B., Chupeau, M. C., Bellec, Y., & Chupeau, Y. (1995). Sexual and somatic hybridization in the genus Lactuca. Euphytica, 85, 281–285.
Maisonneuve, B., Chovelon, V., & Lot, H. (1991). Inheritance of resistance to beet western yellows virus in Lactuca virosa L. HortScience, 26, 1543–1545.
Maluf, W. R., Azevedo, S. M., Gomes, L. A. A., & de Oliveira, A. C. B. (2002). Inheritance of resistance to the root-knot nematode Meloidogyne javanica in lettuce. Genetics and Molecular Research, 1, 64–71.
Mani, A., Al Hinai, M. S., & Handoo, Z. A. (1997). Occurrence, population density, and distribution of root-lesion nematodes, Pratylenchus spp., in the Sultanate of Oman. Nematropica, 27, 209–219.
Martin, C., Schoen, L., Rufingier, C., & Pasteur, N. (1996). A contribution to the integrated pest management of the aphid Nasonovia ribisnigri in salad crops. Bulletin OILB-SROP, 19, 98–101.
Matheron, M. E., & Koike, S. T. (2003). First report of fusarium wilt of lettuce caused by Fusarium oxysporum f. sp. lactucae in Arizona. Plant Disease, 87, 1265.
Matoba, H., Mizutani, T., Nagano, K., Hoshu, Y., & Uchiyama, H. (2007). Chromosomal study of lettuce and its allied species (Lactuca spp., Asteraceae) by means of karyotype analysis and fluorescence in situ hybridization. Hereditas, 144, 235–243.
Matsumoto, E. (1991). Interspecific somatic hybridization between lettuce (Lactuca sativa) and wild species L. virosa. Plant Cell Reports, 9, 531–534.
Matsuura, K., Kanto, T., Uzuhashi, S., & Kakishima, M. (2010). Pythium wilt of lettuce caused by Pythium uncinulatum in Japan. Journal of General Plant Pathology, 76, 320–323.
Matta, A. (1965). Una malattia della lattuga prodotta da una nuova specie di Pythium. Phytopathologia Mediterranea, 4, 48–53.
Matuo, T., & Matahashi, S. (1967). On Fusarium oxysporum f. sp. lactucae causing root rot of lettuce. Transactions of Mycological Society of Japan, 32, 13–15.
Maxted, N., & Kell, S. P. (2008). Linking in-situ and ex-situ conservation with use of crop wild relatives. In N. Maxted, B. V. Ford–Lloyd, S. P. Kell, J. M. Iriondo, E. Dulloo, & J. Turok (Eds.), Crop wild relative conservation and use (pp. 450–470). Wallingford: CABI.
Maxted, N., Kell, S. P., & Ford-Lloyd, B. V. (2008). Crop wild relative conservation and use: establishing the context. In N. Maxted, B. V. Ford–Lloyd, S. P. Kell, J. M. Iriondo, E. Dulloo, & J. Turok (Eds.), Crop wild relative conservation and use (pp. 3–30). Wallingford: CABI.
Mazier, M., German-Retana, S., Flamain, F., Dubois, V., Botton, E., Sarnette, V., et al. (2003). A simple and efficient method for testing Lettuce mosaic virus resistence in in vitro cultivated lettuce. Journal of Virological Methods, 116, 123–131.
McCabe, M. S., Garratt, L. C., Schepers, F., Jordi, W., Stoopen, G. M., Davelaar, F., et al. (2001). Effects of P-SAG12-IPT gene expression on development and senescence in transgenic lettuce. Plant Physiology, 127, 505–516.
McCabe, M. S., Schepers, F., van der Arend, A., Mohapatra, U., de Laat, A. M. M., Power, J. B., et al. (1999). Increased stable inheritance of herbicide resistance in transgenic lettuce carrying a petE promoter-bar gene compared with a CaMV 35S-bar gene. Theoretical and Applied Genetics, 99, 587–592.
McCreight, J. D. (2008). Potential sources of genetic resistance in Lactuca spp. the lettuce aphid Nasanovia ribisnigri (Mosely) (Homoptera: Aphididae). Hortscience, 43, 1355–1358.
McCreight, J. D., & Liu, Y. B. (2012). Resistance to lettuce aphid (Nasonovia ribisnigri) biotype 0 in wild lettuce accessions PI 491093 and PI 274378. Hortscience, 47, 179–184.
McDougall, S., & Troldahl, R. (2010). Current lettuce aphid resistant varieties available in Australia. NSW Department of Primary Industries. <http://www.dpi.nsw.gov.au/agriculture/horticulture/vegetables/diseases/currant-lettuce-aphid/resistant>[last accessed 2013-04-27]
McGuire, P. E., Ryder, E. J., Michelmore, R. W., Clark, R. L., Antle, R., Emery, G., Hannan, R. M., Kesseli, R. V., Kurtz, E. A., Ochoa, O., Rubatzky, V. E., & Waycott, W. (1993). Genetic Resources of Lettuce and Lactuca species in California. An Assessment of the USDA and UC Collections and Recommendations for Long-term Security. Report No. 12. Davis: University of California, Genetic Resources Conservation Program.
McHale, K. L., Truco, J. M., Kozik, A., Wroblewski, T., Ochoa, E. O., Lahre, A. K., et al. (2009). The genomic architecture of disease resistance in lettuce. Theoretical and Applied Genetics, 118, 565–580.
Michalska, K., & Kisiel, W. (2009). Root constituents of Lactuca sibirica and a comparison of metabolite profiles of L. sibirica and L. tatarica. Acta Societatis Botanicorum Poloniae, 78, 25–27.
Michalska, K., & Kisiel, W. (2010). Sesquiterpene lactones from roots of Lactuca aculeata. Biochemical Systematics and Ecology, 38, 830–832.
Michalska, K., Stojakowska, A., Malarz, J., Doležalová, I., Lebeda, A., & Kisiel, W. (2009). Systematic implication of sesquiterpene lactones in Lactuca species. Biochemical Systematics and Ecology, 37, 174–179.
Michelmore, R. W. (2012). California leafy greens research program. <http://calgreens.org/control/uploads/Genetic_Variation_in_Lettuce.pdf> [last accessed 2013-02-01]
Michelmore, R. W., & Eash, J. A. (1986). Lettuce. In Handbook of plant cell culture, vol. 4 (pp. 512–551). New York: Macmillan.
Michelmore, R. W., Ochoa, O. E., Truco, M. J., Grube, R., & Gates, R. (2005). Breeding crisphead lettuce. USDA-ARS. California Lettuce Research Board Annual Report (pp. 68–78), 2004–2005.
Mikel, M. A. (2007). Genealogy of contemporary North American lettuce. HortScience, 42, 489–493.
Mikel, M. A. (2013). Genetic composition of contemporary proprietary U.S. lettuce (Lactuca sativa L.) cultivars. Genetic Resources and Crop Evolution, 60, 89–96.
Miller, N. J., Birley, A. J., Overall, A. D. J., & Tatchell, G. M. (2003). Population genetic structure of the lettuce root aphid, Pemphigus bursarius (L.), in relation to geographic distance, gene flow and host plant usage. Heredity, 91(3), 217–223.
Miller, N. J., Kift, N. B., & Tatchell, G. M. (2008). Host-associated populations in the lettuce root aphid, Pemphigus bursarius (L.). Heredity, 94, 556–564.
Mizutani, T., Liu, X. J., Tashiro, Y., Miyazaki, S., & Shimazaki, K. (1989). Plant regeneration and cell fusion of protoplasts from lettuce cultivars and related wild species in Japan. Bulletin of Faculty of Agriculture, Saga University, 67, 109–118.
Mohapatra, U., McCabe, M. S., Power, J. B., Schepers, F., Van der Arend, A., & Davey, M. R. (1999). Expression of the bar gene confers herbicide resistance in transgenic lettuce. Transgenic Research, 8, 33–44.
Mojtahedi, H., Boydson, R. A., Thomas, P. E., Crosslin, J. M., Santo, G. S., Roga, E., et al. (2003). Weed hosts of Paratrichodorus allius and tobacco rattle virus in the Pacific Northwest. American Journal of potato Research, 80, 379–385.
Moretti, F., Cotroneo, A., & Mancini, G. (1981). The reproduction and pathology of Pratylenchus penetrans on some varieties of lettuce. Revue de Nématologie, 4, 271–276.
Mou, B. (2008). Lettuce. In J. Prohens & F. Nuez (Eds.), Handbook of plant breeding. Vegetables I. Asteraceae, brassicaceae, chenopodiaceae, and cucurbitaceae (pp. 75–116). New York: Springer Science.
Mou, B. (2011a). Green leaf lettuce breeding lines with resistance to corky root, 06–831 and 06–833. HortScience, 46, 1324–1325.
Mou, B. (2011b). Mutations in lettuce improvement. International Journal of Plant Genomics, Volume 2011, Article ID 723518, doi: 10.1155/2011/723518.
Mou, B., & Bull, C. (2004). Screening lettuce germplasm for new sources of resistance to corky root. Journal of the American Society for Horticultural Science, 129, 712–718.
Mou, B., Hayes, R. J., & Ryder, E. J. (2007). Crisphead lettuce breeding lines with resistance to corky root and lettuce mosaic virus. HortScience, 42, 701–703.
Mou, B., & Liu, Y. (2003). Leafminer resistance in lettuce. Hortscience, 38, 570–572.
Mou, B., & Liu, Y. (2004). Host plant resistance to leafminers in lettuce. Journal of the American Society for Horticultural Science, 129, 383–388.
Mou, B., & Ryder, E. J. (2010). MU06-857, a green leaf lettuce breeding line with resistance to leafminer and lettuce mosaic virus. Hortscience, 45, 666–667.
Navarro, J. A., Torok, V. A., Vetten, H. J., & Pallás, V. (2005). Genetic variability in the coat protein genes of lettuce big-vein associated virus and Mirafiori lettuce big-vein virus. Archives for Virology, 150, 681–694.
Netzer, D., Globerson, D., Weintal, C., & Elyassi, R. (1985). Sources and inheritance of resistance to Stemphylium leaf spot of lettuce. Euphytica, 34, 393–396.
Nicaise, V., German-Retana, S., Sanjuán, R., Dubrana, M. P., Mazier, M., Maisonneuve, B., et al. (2003). The eukaryotic translation initiation factor 4E controls lettuce susceptibility to the potyvirus Lettuce mosaic virus. Plant Physiology, 132, 1272–1282.
Novotná, A., Doležalová, I., Lebeda, A., Kršková, M., & Berka, T. (2011). Morphological variability of achenes of some European populations of Lactuca serriola L. Flora, 206, 473–483.
Obermeier, C., Sears, J. L., Liu, H. Y., Schlueter, K. O., Ryder, E. J., Duffus, J. E., et al. (2001). Characterization of distinct tombusviruses that cause disease of lettuce and tomato in the western United States. Phytopathology, 91, 797–806.
Ochoa, O., Delp, B., & Michelmore, R. W. (1987). Resistance in Lactuca spp. to Microdochium panattoniana (lettuce anthracnose). Euphytica, 36, 609–614.
Okubara, P. A., Arroyo-Garcia, R., Shen, K. A., Mazier, M., Meyers, B. C., Ochoa, O. E., et al. (1997). A transgenic mutant of Lactuca sativa (lettuce) with a T-DNA tightly linked to loss of downy mildew resistance. Molecular Plant Microbe Interactions, 10, 970–977.
Ökten, M. E. (1988). Some species of Tylenchidae (Tylenchida: Nematoda) from the Istanbul province. Türkiye Entomoloji Dergisi, 12, 209–214.
Parella, G., Gognalons, P., Gebre-Selassie, K., Vovlas, C., & Marchoux, G. (2003). An update of the host range of Tomato spotted wilt virus. Journal of Plant Pathology, 85, 227–264.
Pedroche, N. B., Villaneuva, L. M., & de Waele, D. (2012). Plant parasitic nematodes associated with semi-temperate vegetables in Benguet Province, Philippines. Archives of Phytopathology and Plant Protection, iFirst article 1–17.
Peters, D., & Goldbach, R. (1998). An updated list of plant species susceptible to tospovirus. Wageningen Agricultural University, Section Virology, The Netherlands.
Petrželová, I., & Lebeda, A. (2011). Distribution of race-specific resistance against Bremia lactucae in natural populations of Lactuca serriola. European Journal of Plant Pathology, 129, 233–253.
Petrželová, I., Lebeda, A., & Beharav, A. (2011). Resistance to Bremia lactucae in natural populations of Lactuca saligna from some Middle Eastern countries and France. Annals of Applied Biology, 159, 442–455.
Philis, J. (1995). An up-dated list of plant parasitic nematodes from Cyprus and their economic importance. Nematologia Mediterranea, 23, 307–314.
Pileggi, M., Mielniczki Pereira, A. A., dos Santos Silva, J., Veiga Pileggi, S. A., & Verma, D. P. S. (2001). An improved method for transformation of lettuce by Agrobacterium tumefaciens with a gene that confers freezing resistance. Brazilian Archives of Biology and Technology, 44, 191–196.
Pink, D. A. C. (2002). Strategies using genes for non-durable resistance. Euphytica, 124, 227–236.
Pink, D. A. C., & Keane, E. M. (1993). Lettuce: Lactuca sativa L. In G. Kalloo & B. O. Bergh (Eds.), Genetic improvement of vegetable crops (pp. 543–571). Oxford: Pergamon Press.
Pink, D. A. C., Kostova, D., & Walkey, D. G. A. (1992a). Differentiation of pathotypes of lettuce mosaic virus. Plant Pathology, 41, 5–12.
Pink, D. A. C., Lot, H., & Johnson, R. (1992b). Novel pathotypes of lettuce mosaic virus-breakdown of a durable resistance. Euphytica, 63, 169–174.
Pink, D. A. C., & Puddephat, I. J. (1999). Deployment of disease resistance genes by plant transformation – a “mix and match” approach. Trends in Plant Science, 4, 71–75.
Pniewski, T. (2013). The twenty-year story of a plant-based vaccine against hepatitis B: stagnation or promising prospects? International Journal of Molecular Sciences, 14, 1978–1998.
Provvidenti, R., Robinson, R. W., & Shail, J. W. (1980). A source of resistance to a strain of cucumber mosaic virus in Lactuca saligna L. HortScience, 15, 528–529.
Purdy, L. H. (1979). Sclerotinia sclerotiorum: history, disease and symptomatology, host range, geographical distribution and impact. Phytopathology, 69, 875–880.
Radewald, J. D., Mowbray, P. G., Paulus, A. O., Shibuya, F., & Rible, J. M. (1969a). Preplant soil fumigation for California head lettuce. Plant Disease Reporter, 53, 385–389.
Radewald, J. D., Osgood, J. W., Mayberry, K. S., Paulus, A. O., & Shibuya, F. (1969b). Longidorus africanus a pathogen of head lettuce in the Imperial Valley of southern California. Plant Disease Reporter, 53, 381–384.
Radewald, J. D., Osgood, J. W., Mayberry, K. S., Paulus, A. O., Otto, H. W., & Shibuya, F. (1969c). Longidorus africanus merny; nematode found pathogen of imperial lettuce. California Agriculture, 23, 10–13.
Raid, R. N. (1997). Stemphylium leaf spot. In R. M. Davis, K. V. Subbarao, R. N. Raid, & E. A. Kurtz (Eds.), Compendium of lettuce diseases (pp. 25–26). St. Paul: APS Press.
Rauscher, G., & Simko, I. (2013). Development of genomic SSR markers for fingerprinting lettuce (Lactuca sativa L.) cultivars and mapping genes. BMC Plant Biology, 13, 11.
Ray, D. T., McCreight, J. D., McGrady, J. J., & Brown, J. K. (1989). Resistance in cultivated and wild lettuce to lettuce infectious yellows virus. Vegetable Report, 78, 73–77.
Rees, S., & Harborne, J. (1984). Flavonoids and other phenolics of Cichorium and related members of the Lactuceae (Compositae). Botanical Journal of the Linnean Society, 89, 313–319.
Reinink, K. (1999). Lettuce resistance breeding. In A. Lebeda & E. Křístková (Eds.), Eucarpia leafy vegetables ′99. Proceedings of the eucarpia meeting on leafy vegetables genetics and breeding (pp. 139–147). Olomouc: Palacký University in Olomouc.
Reinink, K., & Dieleman, F. L. (1989). Comparison of sources of resistance to leaf aphids in lettuce (Lactuca sativa L.). Euphytica, 40, 21–29.
Revers, F., Lot, H., Souche, S., Le Gall, O., Candresse, T., & Dunez, J. (1997a). Biological and molecular variability of lettuce mosaic virus isolates. Phytopathology, 87, 397–403.
Revers, F., Yang, S. J., Walter, J., Souche, S., Lot, H., Le Gall, O., et al. (1997b). Comparison of the complete nucleotide sequences of two isolates of lettuce mosaic virus differing in their biological properties. Virus Research, 47, 167–177.
Riar, D. S., Rustgi, S., Burke, I. C., Gill, K. S., & Yenish, J. P. (2011). EST-SSR development from 5 Lactuca species and their use in studying genetic diversity among L. serriola biotypes. Journal of Heredity, 102, 17–28.
Rich, J., Brito, J., Ferrell, J., & Kaur, R. (2010). Weed hosts of root-knot nematodes common to Florida. University of Florida, IFAS, ENY-060 <http://edis.ifas.ufl.edu/pdffiles/IN/IN84600.pdf>. [last accessed 2013-02-01]
Roggero, P., Ciuffo, M., Vaira, A. M., Accotto, G. P., Masenka, V., & Milne, R. G. (2000). An Ophiovirus isolated from lettuce with big-vein symptoms. Archives of Virology, 145, 2629–2642.
Roossinck, M. J. (2002). Evolutionary history of Cucumber mosaic virus deduced by phylogenetic analyses. Journal of Virology, 76, 3382–3387.
Roossinck, M. J., Bujarski, J., Ding, S. W., Hajimorad, R., Hanada, K., Scott, S., & Tousignant, M. (1999). Family Bromoviridae. In M. H. V. van Regenmortel, C. M. Fauquet, & D. H. L. Bishop (Eds.), Virus Taxonomy (pp. 923–935). Seventh Report of the International Committee on Taxonomy of Viruses. San Diego, California: Academic Press.
Ryder, E. J. (1970). Inheritance of resistance to common lettuce mosaic. Journal of the American Society of Horticultural Science, 95, 378–379.
Ryder, E. J. (1999). Lettuce, endive and cichory. Wallingford: CABI Publishing.
Ryder, E. J. (2001). Current and future issues in lettuce breeding. In J. Janick (Ed.), Plant breeding reviews (Vol. 20, pp. 105–134). San Francisco: Willey.
Ryder, E. (2002). A mild systemic reaction to lettuce mosaic virus in lettuce: inheritance and interaction with an allele for resistance. Journal of the American Society of Horticultural Science, 127, 814–818.
Ryder, E. J., Grube, R. C., Subbarao, K. V., & Koike S. T. (2003). Breeding for resistance to diseases in lettuce: successes and challenges. In Th.J.L. van Hintum, A. Lebeda, D. A. Pink, J. W. Schut (Eds.), Eucarpia Leafy Vegetables 2003, Proceedings of the Eucarpia Meeting on Leafy Vegetables Genetics and Breeding (pp. 25–30). Wageningen, The Netherlands: Centre for Genetic Resources (CGN).
Ryder, E. J., & Robinson, B. J. (1995). Big-vein resistance in lettuce – identifying, selecting, and testing resistant cultivars and breeding. Journal of the American Society for Horticultural Science, 120, 741–746.
Sahin, F., & Miller, S. A. (1997). Identification of the bacterial leaf spot pathogen of lettuce, Xanthomonas campestris pv. vitians, in Ohio, and assessment of cultivar resistance and seed treatment. Plant Disease, 81, 1443–1446.
Sangün, O., & Satar, S. (2012). Aphids (Hemiptera: Aphididae) on lettuce in the Eastern Mediterranean Region of Turkey: incidence, population fluctuations, and flight activities. Turkiye Entomoloji Dergisi - Turkish Journal of Entomology, 36, 443–454.
Sauer, C. (2008). Salatanbau-noch kein genereller Durchbruch der Nasonovia-Resistenz. Auszug aus Gemusebau-Info 10/2008. - Agroscope Changins-Wädenswil ACW. <http://www.agroscope.admin.ch/publikationen/einzelpublikation/index.html?aid=824&lang=fr&pid=9386> [last accessed 2013-04-27]
Schwember, A. R., & Bradford, K. J. (2010). Quantitative trait loci associated with longevity of lettuce seeds under conventional and controlled deterioration storage conditions. Journal of Experimental Botany, 61, 4423–4436.
Scott, J. C., Kirkpatrick, S. C., & Gordon, T. R. (2010). Variation in susceptibility of lettuce cultivars to fusarium wilt caused by Fusarium oxysporum f.sp. lactucae. Plant Pathology, 59, 139–146.
Sequiera, L. (1970). Resistance to corky root rot in lettuce. Plant Disease Reports, 54, 754–758.
Sequiera, L. (1978). Two root rot resistant varieties of head lettuce. Ohio Agricultural Experimental Research Station Bulletin, 359, 197–214.
Sessa, R., Bennett, M. H., Lewin, M. J., Mansfield, J. W., & Beale, M. H. (2000). Metabolite profiling of sesquiterpene lactones from Lactuca species. Journal of Biological Chemistry, 275, 26877–26884.
Shin, H. D., Jee, H. J., & Shin, C. K. (2006). First report of powdery mildew caused by Sphaerotheca fusca on Lactuca sativa in Korea. Plant Pathology, 55, 814.
Shukla, D. D., Ward, C. W., & Brunt, A. A. (1994). The potyviridae. Wallingford: CAB International.
Sicard, D., Woo, S. S., Arroyo-Garcia, R., Ocho, O., Nguyen, D., Korol, A., et al. (1999). Molecular diversity at the major cluster of disease resistance genes in cultivated and wild Lactuca spp. Theoretical and Applied Genetics, 99, 405–418.
Sikora, R. A., & Fernández, E. (2005). Nematode parasites of vegetables. In M. Luc, R. A. Sikora, & J. Bridge (Eds.), Plant parasitic nematodes in subtropical and tropical agriculture (2nd ed., pp. 319–392). Wallingford: CABI Publishing.
Simko, I. (2009). Development of EST-SSR markers for the study of population structure in lettuce (Lactuca sativa L.). Journal of Heredity, 100, 256–262.
Simko, I., Hayes, R. J., Truco, M. J., & Michelmore, R. W. (2011). Mapping a dominant negative mutation for triforine sensitivity in lettuce and its use as a selectable marker for detecting hybrids. Euphytica, 182, 157–166.
Simko, I., & Hu, J. (2008). Population structure in cultivated lettuce and its impact on association mapping. Journal of the American Society for Horticultural Science, 133, 61–68.
Simko, I., Pechenick, D. A., McHale, L. K., Truco, M. J., Ochoa, O. E., Michelmore, R. W., et al. (2009). Association mapping and marker-assisted selection of the lettuce dieback resistance gene Tvr1. BMC Plant Biology, 9, 135.
Simko, I., Pechenick, D. A., McHale, L. K., Truco, M. J., Ochoa, O. E., Michelmore, R. W., et al. (2010). Development of molecular markers for marker-assisted selection of dieback resistance in lettuce (L. sativa. Acta Horticulturae (ISHS), 859, 401–408.
Sretenović-Rajičić, T., van Hintum, T. J. L., Lebeda, A., & Dehmer, K. (2008). Analysis of wild Lactuca accessions: conservation and identification of redundancy. Plant Genetic Resources: Characterization and Utilization, 6, 153–163.
Stebbins, G. L. (1957). Self-fertilization and population variability in the higher plants. American Naturalist, 91, 418–428.
Stevens, M., Freeman, B., Liu, H., Herrbach, E., & Lemaire, O. (2005). Beet poleroviruses: close friends or distant relatives? Molecular Plant Pathology, 6, 1–9.
Stoetzel, M. B. (1985). Eucarazzia elegans (Ferrari) an aphid new to the Western hemisphere, with archival data (Homoptera: Aphididae). Proceedings of the Entomological Society of Washington, 87, 44–48.
Stoffel, K., van Leeuwen, H., Kozik, A., Caldwell, D., Ashrafi, H., Cui, X., et al. (2012). Development and application of a 6.5 million feature Affymetrix Genechip® for massively parallel discovery of single position polymorphisms in lettuce (Lactuca spp.). BMC Genomics, 13, 185.
Stubbs, L. L., & Grogan, R. G. (1963). Necrotic yellows: a newly recognised virus disease of lettuce. Australian Journal of Agricultural Research, 14, 439–459.
Subbarao, K. V. (1998). Progress towards integrated management of lettuce drop. Plant Disease, 80, 28–33.
Subbarao, K. V., Hubbard, J. C., Greathead, A., & Spencer, G. A. (1997). Verticillium wilt. In R. M. Davis, K. V. Subbarao, R. N. Raid, & E. A. Kurtz (Eds.), Compendium of lettuce diseases (pp. 26–27). St. Paul: American Phytopathological Society.
Syed, H. N., Sørensen, P. A., Antonise, R., van de Wiel, C., van der Linden, G. C., van ‘t Westende, W., et al. (2006). A detailed linkage map of lettuce based on SSAP, AFLP and NBS markers. Theoretical and Applied Genetics, 112, 517–527.
Takada, K., Watanabe, S., Sano, T., Ma, B., Kamada, H., & Ezura, H. (2007). Heterologous expression of the mutated melon ethylene receptor gene Cm-ERS1/H70A produces stable sterility in transgenic lettuce (Lactuca sativa). Journal of Plant Physiology, 164, 514–520.
Tamaki, H., Robinson, R. W., Anderson, J. L., & Stoewsand, G. S. (1995). Sesquiterpene lactones in virus-resistant lettuce. Journal of Agricultural and Food Chemistry, 43, 6–8.
Thabuis, A. P. P., Teekens, K. C., & van Herwijnen, Z. O. (2011). Lettuce that is resistant to the lettuce aphid Nasonovia ribisnigri biotype 1. World Intellectual Property Organization. PCT/EP2010/067588.
Thompson, R. C., & Ryder, E. J. (1961). Descriptions and pedigrees of nine varieties of lettuce. U.S. Department of Agriculture Technical Bulletin, 1244.
Torres, A. C., Nagata, R. T., Ferl, R. J., Bewick, T. A., & Cantliffe, D. J. (1999). In vitro assay selection of glyphosate resistance in lettuce. Journal of the American Society for Horticultural Science, 124, 86–89.
Tortolero, O., & Sequeira, L. (1978). A vascular wilt and leaf blight disease of lettuce in Wisconsin caused by a new strain of Pythium tracheiphilum. Plant Disease Reporter, 62, 616–620.
Toussaint, V., Benoit, D. L., & Carisse, O. (2012). Potential of weed species to serve as a reservoir for Xanthomonas campestris pv. vitians, the causal agent of bacterial leaf spot of lettuce. Crop Protection, 41, 64–70.
Truco, M. J., Antonise, R., Lavelle, D., Ochoa, O., Kozik, A., Witsenboer, H., et al. (2007). A high-density integrated genetic linkage map of lettuce (Lactuca spp.). Theoretical and Applied Genetics, 115, 735–746.
Truco, M. J., Ashrafi, H., Kozik, A., van Leeuwen, H., Bowers, J., Chin Wo, S.R., Stoffel, K., Xu, H., Hill, T., Van Deynze, A., & Michelmore, R. (2013). An ultra high-density, transcript-based, genetic map of lettuce. G3: Genes, Genomes and Genetics (submitted)
Tsuchiya, N., Fujinaga, M., Ogiso, H., Usui, T., & Tsukada, M. (2004). Resistance tests and genetic resources for breeding fusarium root rot resistant lettuce. Journal of Japanese Society of Horticultural Science, 73, 105–113.
Usami, T., Itoh, M., Morii, S., Miyamoto, T., Kaneda, M., Ogawara, T., et al. (2012). Involvment of two different types of Verticillium dahliae in lettuce wilt in Ibaraki Prefecture, Japan. Journal of General Plant Pathology, 78, 348–352.
Uwimana, B., d’Andrea, L. D., Felber, F., Hooftman, D. A., den Nijs, H. C. M., Smulders, M. J. M., et al. (2012a). A Bayesian analysis of gene flow from crops to their wild relatives: cultivated (Lactuca sativa L.) and prickly lettuce (L. serriola L.) and the recent expansion of L. serriola in Europe. Molecular Ecology, 21, 2640–2654.
Uwimana, B., Smulders, M. J., Hooftman, D. A., Hartman, Y., van Tienderen, P. H., Jansen, J., et al. (2012b). Crop to wild introgression in lettuce: following the fate of crop genome segments in backcross populations. BMC Plant Biology, 12, 43.
Uwimana, B., Smulders, M. J., Hooftman, D. A., Hartman, Y., van Tienderen, P. H., Jansen, J., et al. (2012c). Hybridization between crops and wild relatives: the contribution of cultivated lettuce to the vigour of crop-wild hybrids under drought, salinity and nutrient deficiency conditions. Theoretical and Applied Genetics, 125, 1097–1111.
van Bruggen, A. H. C. (1997). Corky root. In R. M. Davis, K. V. Subbarao, R. N. Raid, & E. A. Kurtz (Eds.), Compendium of lettuce diseases (pp. 28–29). St. Paul: APS Press.
van der Arend, A. J. M., Ester, A., & Schijndel, J. T. V. (1999). Developing an aphid-resistant butterhead lettuce “Dynamite”. In A. Lebeda & E. Křístková (Eds.), EUCARPIA leafy vegetables′99 (pp. 149–157). Olomouc: Palacky University in Olomouc.
van de Wiel, C., Arens, P., & Vosman, B. (1998). Microsatellite fingerprinting in lettuce (Lactuca sativa L.) and wild relatives. Plant Cell Reports, 17, 837–842.
van de Wiel, C., Arens, P., & Vosman, B. (1999). Microsatellite retrieval in lettuce (Lactuca sativa L.). Genome, 42, 139–149.
van de Wiel, C. C. M., Sretenović-Rajičić, T., van Treuren, R., Dehmer, K. J., van der Linden, C. G., & van Hintum, T. J. L. (2010). Distribution of genetic diversity in wild European populations of prickly lettuce (Lactuca serriola): implications for plant genetic resources management. Plant Genetic Resources: Characterization and Utilization, 8, 171–181.
van Hintum, T. J. L. (2003). Molecular characterisation of a lettuce germplasm collection. In T. J. L. van Hintum, A. Lebeda, D. A. Pink, & J. W. Schut (Eds.), Eucarpia leafy vegetables 2003, proceedings of the eucarpia meeting on leafy vegetables genetics and breeding (pp. 19–21). Wageningen: Centre for Genetic Resources (CGN).
van Hintum, Th. J. L. & Boukema, I. W. (1999). Genetic resources of leafy vegetables. In A. Lebeda, & E. Křístková (Eds.), Eucarpia Leafy Vegetables ′99, Proceedings of the Eucarpia Meeting on Leafy Vegetables Genetics and Breeding. (pp. 59–72). Olomouc: Palacký University in Olomouc.
van Leeuwen, H., Stoffel, K., Kozik, A., Cui, X., Ashrafi, H., McHale, L., Lavelle, D., Wong, G., Chen, F., Truco, M. J., Van Deynze, A., & Michelmore, R. W. (2009). High-density mapping of the lettuce genome with SFP markers in over 15,000 unigenes. Plant and Animal Genome Conference XVII, San Diego, USA.
van Regenmortel, M. H. V., Fauquet, C. M., Bishop, D. H. L., Carstens, E. B., Estes, M. K., Lemon, S. M., et al. (2000). Virus taxonomy: classification and nomenclature of viruses (7th report ICTV). San Diego: Academic.
van Treuren, R., van der Arend, A. J. M., & Schut, J. W. (2013). Distribution of downy mildew (Bremia lactucae Regel) resistances in a genebank collection of lettuce and its wild relatives. Plant Genetic Resources: Characterization and Utilization, 11, 15–25.
van Treuren, R., & van Hintum, T. J. L. (2009). Comparison of anonymous and targeted molecular markers for the estimation of genetic diversity in ex situ conserved Lactuca. Theoretical and Applied Genetics, 119, 1265–1279.
van Treuren, R., Coquin, P., & Lohwasser, U. (2011). Genetic resources collections of leafy vegetables (lettuce, spinach, chicory, artichoke, asparagus, lamb’s lettuce, rhubarb and rocket salad): composition and gaps. Genetic Resources and Crop Evolution, 59, 981–997.
Vanstone, V. A., & Russ, M. H. (2001). Ability of weeds to host the root lesion nematodes Pratylenchus neglectus and P. thornei II*. Broad-leaf weeds. Australasian Plant Pathology, 30, 251–258.
Vermeulen, A., Desprez, B., Lancelin, D., & Bannerot, H. (1994). Relationships among Cichorium species and related genera as determined by analysis of mitochondrial RFLPs. Theoretical and Applied Genetics, 88, 159–166.
Viane, N. M., & Abawi, G. S. (1996). Damage threshold of Meloidogyne hapla to lettuce in organic soil. Journal of Nematology, 28, 537–545.
Walkey, D. (1991). Applied plant virology (2nd ed.). London: Chapman and Hall.
Waycott, W., Fort, S. B., Ryder, E. J., & Michelmore, R. W. (1999). Mapping morphological genes relative to molecular markers in lettuce (Lactuca sativa L.). Heredity, 82, 245–251.
Wesolowska, A., Nikiforuk, A., Michalska, K., Kisiel, W., & Chojnacka-Wójcik, E. (2006). Analgesic and sedative activities of lactucin and some lactucin-like guaianolides in mice. Journal of Ethnofarmacology, 107, 254–258.
Wetzel, T., Dietzgen, R. G., & Dale, J. L. (1994). Genomic organization of lettuce necrotic yellows rhabdovirus. Virology, 200, 401–412.
Whipps, J. M., Budge, S. P., McClement, S., & Pink, D. A. C. (2002). A glasshouse cropping method for screening lettuce lines for resistance to Sclerotinia sclerotiorum. European Journal of Plant Pathology, 108, 373–378.
Whitaker, T. W., Bohn, G. W., Welch, J. F., & Grogan, R. G. (1958). History and development of head lettuce resistant to downy mildew. Proceedings of the American Society for Horticultural Science, 72, 410–416.
Wintermantel, W. M. (2004). Emergence of greenhouse whitefly (Trialeurodes vaporariorum) transmitted Criniviruses as threats to vegetable and fruit producion in North America. APSnet Features. Online. doi:10.1094/APSnetFeature-2004-0604.
Wintermantel, W. M., & Anchieta, A. G. (2012). The genome sequence of lettuce necrotic stunt virus indicates a close relationship to Moroccan pepper virus. Archiv of Virology, 157, 1407–1409.
Wintermantel, W. M., Anchieta, A. G., Obermeier, C., & Wisler, G. C. (2003). Tombusvirus infection of lettuce is influenced by soil enviroment. Phytopathology, 93, 101.
Witsenboer, H., Michelmore, R. W., & Vogel, J. (1997). Identification, genetic localization and allelic diversity of selectively amplified microsatellite polymorphic loci in lettuce and wild relatives (Lactuca spp.). Genome, 40, 923–936.
Yabuuchi, E., Kosako, Y., Naka, Y., Suzuki, S., & Yano, I. (1999). Proposal of Sphingomonas suberifaciens (van Bruggen, Jochimsen and Brown 1990) Comb. Nov., Sphingomonas natatoria (Sly 1985) Comb Nov., Sphingomonas ursincola (Yurkov et al. 1997) Comb. Nov., and emendation of the genus Sphingomonas. Microbiology and Immunology, 43, 339–349.
Yang, T. J., Jang, S. W., & Kim, W. B. (2007). Genetic relationships of Lactuca spp. revealed by RAPD, Inter-SSR, AFLP, and PCR-RFLP analyses. Journal of Crop Science and Biotechnology, 10, 29–34.
Zhang, N. W., Lindhout, P., Niks, R. E., & Jeuken, M. J. W. (2009). Genetic dissection of Lactuca saligna nonhost resistance to downy mildew at various lettuce developmental stages. Plant Pathology, 58, 923–932.
Zhang, L. Y., Zhang, Y. Y., Chen, R. G., Zhang, J. H., Wang, T. T., Li, H. X., et al. (2010). Ectopic expression of the tomato Mi-1 gene confers resistance to root knot nematodes in lettuce (Lactuca sativa). Plant Molecular Biology Reporter, 28, 204–211.
Zidorn, C. (2008). Sesquiterpene lactones and their precursors as chemosystematic markers in the tribe Cichorieae of the Asteraceae. Phytochemistry, 69, 2270–2296.
Zitter, T. A., & Murphy, J. F. (2009). Cucumber mosaic virus. The Plant Health Instructor. doi:10.1094/PHI-I-2009-0518-01.
Zohary, D. (1991). The wild genetic resources of cultivated lettuce (Lactuca sativa L.). Euphytica, 53, 31–35.
Acknowledgments
The research was supported by grant MSM 6198959215 (Ministry of Education, Youth and Sports of the Czech Republic) and by the internal grant of Palacký University in Olomouc (IGA_PrF_2013_001) and by IF0157 Leafy Vegetable Genetic Improvement Network (VeGIN): Pre-breeding research to support sustainable farming of leafy vegetables and salads (UK Department of Environment Food and Rural Affairs)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lebeda, A., Křístková, E., Kitner, M. et al. Wild Lactuca species, their genetic diversity, resistance to diseases and pests, and exploitation in lettuce breeding. Eur J Plant Pathol 138, 597–640 (2014). https://doi.org/10.1007/s10658-013-0254-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-013-0254-z