Abstract
Chromium (Cr) pollution is an emerging environmental problem. The present study was carried out to isolate Cr-resistant bacteria and characterize their Cr detoxification and resistance ability. Bacteria screened by exposure to chromate (Cr6+) were isolated from Mandovi estuary Goa, India. Two isolates expressed high resistance to Cr6+ (MIC ≥ 300 µg mL−1), Cr3+ (MIC ≥ 900 µg mL−1), other toxic heavy metals and displayed a pattern of resistance to cephalosporins and ß-lactams. Biochemical and 16 S rRNA gene sequence analysis indicated that both isolates tested belonged to the Staphylococcus genus and were closely related to S. saprophyticus and S. arlettae. Designated as strains NIOER176 and NIOER324, batch experiments demonstrated that both removed 100% of 20 and 50 µg mL−1 Cr6+ within 4 and 10 days, respectively. The rate of reduction in both peaked at 0.260 µg mL−1 h−1. ATP-binding cassette (ABC) transporter gene involved in transport of a variety of substrates including efflux of toxicants was present in strain NIOER176. Through SDS-PAGE analysis, whole-cell proteins extracted from both strains indicated chromium-induced specific induction and up-regulation of 24 and 40 kDa proteins. Since bacterial ability to ameliorate Cr6+ is of practical significance, these findings demonstrate strong potential of some estuarine bacteria to detoxify Cr6+ even when its concentrations far exceed the concentrations reported from many hazardous effluents and chromium contaminated natural habitats. Such potential of salt tolerant bacteria would help in Cr6+ bioremediation efforts.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Rise in industrialization and other anthropogenic practices over the past few decades have led to increased use and reliance on chromium (Singh et al. 2015). Cr occurs naturally in the earth’s crust, but its indiscriminate use and unregulated discharge from human practices have resulted in elevated environmental concentrations (Pereira 2019). Of the two commercially important states, Cr6+ (chromate) is more mobile and water-soluble than Cr3+. Its ability to rapidly traverse biological membranes through sulfate transport systems also makes it considerably more toxic than Cr3+ (Thatoi et al. 2014). Being immutable, Cr6+ has a tendency to persist, bioaccumulate and contaminate food chains. On entering animal and human tissue, several toxicity symptoms have been observed ranging from abdominal pain to brain dysfunction (McLaughlin et al. 1999). Therefore, Cr6+ is designated as a priority pollutant by the US Environmental Protection Agency (USEPA).
As other components of the biota, bacteria are also sensitive to Cr6+ pollution (Sathicq and Gómez 2018). Cr6+ exerts toxicity by oxidatively damaging DNA, proteins and lipids inhibiting metabolic function (Nies 2003). In order to survive, bacteria have developed Cr-ion homeostasis and Cr6+-resistance. Reduced accumulation either by reduced uptake or by active efflux, precipitation and alteration in cell morphology are some general mechanisms (Shamim et al. 2012, Ahemad 2014). These also include Cr6+ removal either by cell-surface binding (biosorption), intracellular sequestration/translocation and direct or indirect reduction (Joutey et al. 2015). Bacterial species with a potential to tolerate, reduce or detoxify Cr6+ are thus ideal candidates for environmental pollution cleanup. Being natural, unmodified entities, they offer an eco-friendly, cost-effective metal bioremediation/detoxification technology (Bolan et al. 2014, Akcil et al. 2015). Bacterial bioremediation thus forms a promising alternative to conventional physicochemical methods of metal remediation.
To exploit this feature, many researchers using bacterial strains have demonstrated remediation of industrial wastewater. Several isolates indigenous to wastewater and contaminated soil have been assessed for their ability to reduce Cr6+concentrations (Batool et al. 2012, Pulimi et al. 2012, Salamanca et al. 2013, García et al. 2016). However, few reports address the utilization of marine/estuarine strains (Mohapatra et al. 2017; Pereira et al. 2017) for the same. These microbial strains are variously useful as they are continuously exposed to adverse conditions. They are thus naturally suited for bioremediation and may possess new information on resistance mechanisms.
Studies on metal and antibiotic resistant bacteria have shown that resistance systems share structural and functional characteristics (Bontidean et al. 2000, Nies 2003). This implicates the co-selection of Cr6+ resistant bacteria and their potential to (i) resist multiple metals (Kamika and Momba 2013) and (ii) maintain a pool of antibiotic-resistance genes (Baker-Austin et al. 2006). Subsequently, specific patterns of antibiotic resistance may suggest which resistance response is activated. Similarly, expression of bacterial proteins in response to stress stimuli (Congeevaram et al. 2007) has great potential to explore different peptides and proteins involved in heavy metal resistance. This may provide insight into strategies for Cr6+ removal from contaminated environments.
We have previously investigated possible relationships between Cr6+ resistance and the remediation of Cr6+ contaminated soil (Pereira et al. 2017). In the present investigation, we describe the identification, characterization and comparison of two estuarine bacterial isolates NIOER176 and NIOER324. Our findings report the removal of Cr6+ under varying temperature and pH, by antibiotic and multi-heavy metal resistant strains including the detection of ABC transporter gene and protein profiles in response to Cr6+ stress.
Materials and methods
Isolation and identification of chromate resistant bacterial strains
Chromate resistant bacteria were isolated from surface water samples collected (serially diluted) along the Mandovi estuary Goa, India and plated on nutrient agar (NA, HiMedia M001, India) supplemented with 100 µg mL−1 potassium dichromate (K2Cr2O7, Merck). Bacterial colonies that appeared on the NA plate were further streaked on NA supplemented with higher concentrations of K2Cr2O7 (200–600 µg mL−1). Two discrete bacterial colonies that grew at highest Cr6+ level were selected for biological characterization. Selected isolates were characterized morphologically and biochemically according to Bergey’s Manual of Systematic Bacteriology (Holt et al. 1994) and were designated as NIOER176 and NIOER324.The isolates were subjected to partial 16S rRNA gene sequencing using primers 27F and 1492R (Table S1). PCR amplification of the reaction mixture (Table S2) was performed using a Thermocycler (Applied Biosystems, USA) and PCR protocol (Table S1). PCR products purified using GenElute PCR Clean-Up Kit (Sigma-Aldrich) were sequenced in an ABI 3130XL genetic analyzer (Applied Biosystems, California, USA) using primers, 27F and 1492R. Sequences obtained were assembled into contigs using DNA Baser sequence assembly software ver.2 and analysed against the NCBI database using BLAST program packages (https://blast.ncbi.nlm.nih.gov/Blast.cgi) for matching to known 16S rRNA gene sequences. Sequences were submitted to the GenBank database under accession numbers MG205901 and MG206032, for NIOER176 and NIOER324 respectively.
Growth characteristics under chromate stress
Growth characteristics of both isolates were evaluated in nutrient broth (NB, HiMedia M002, India) with constant shaking at 120 rpm at 28 °C and pH 7 ± 0.2 (in triplicate). The minimum inhibitory concentration (MIC, defined as the lowest concentration of Cr6+ at which no visible turbidity/ growth was observed) was determined by broth dilution method in NB supplemented with different concentrations of Cr6+, with 0.1% (v/v) inoculum volume for 24–48 h (Pereira et al. 2017). The growth kinetic studies of both isolates were performed in NB with and without (control) 100 µg mL−1Cr6+ by measuring optical density (O.D) at 600 nm using a UV-Vis spectrophotometer (Shimadzu, model UV-1800) at hourly intervals up to 36 h. Two physiological parameters, temperature and pH were considered for optimizing growth in NB with and without 100 µg mL−1 Cr6+. Isolates were incubated at 20, 24, 28, 32, 37 or 42 °C in a temperature controlled orbital shaker. Similarly, initial pH of separate sets was adjusted to 4.0, 5.0, 6.0, 7.0, 8.0, 9.0, or 10.0 with conc. HNO3 or 1 N NaOH. All sets were incubated for 24 h and final O.D observed. Resistance to other heavy metals (chromium trichloride, arsenic trioxide, mercury chloride, cadmium chloride, copper sulphate, zinc sulphate and nickel chloride) was determined similarly by broth dilution method in NB amended with appropriate stock solutions of each metal salt.
Antibiotic susceptibility test
Susceptibility of bacterial strains to common antibiotics was tested by Kirby–Bauer disc diffusion method (Bauer et al. 1966). Antibiotic discs were placed on NA plates spread with 0.1 mL of undiluted overnight grown bacterial culture (~1 × 108 cells mL−1) to form lawn cultures, and incubated for 24 h at 28 °C. Zones of inhibition were measured and sensitivity to an antibiotic was evaluated as per zone size interpretive criteria provided by the manufacturer (HiMedia, India).
PCR amplification and phylogenetic analysis of ABC transporter gene
Molecular characterization of ABC transporter was performed by PCR amplification of the reaction mixture (Table S2) with genomic DNA from NIOER176 and NIOER324 using primer set: ABC1 and ABC2, and PCR protocol (Table S1) set with a lower annealing temperature (54 °C). Electrophoresis and ethidium bromide staining confirmed positive amplicons from each DNA sample. Purified PCR products were sequenced using primers, ABC1 and ABC2. Sequences homology search was performed using NCBI BLASTn, BLASTx and aligned using CLUSTALW algorithms (Thompson et al. 1997). Phylogenetic analysis was conducted with MEGA 6 software package and phylogenetic trees constructed using neighbour joining (NJ) algorithm (Tamura et al. 2004; 2011).
Optimization of temperature and initial pH for chromate removal assay
Batch tests were performed (in triplicate) to evaluate bacterial chromate removal ability under static condition. Strains were grown in series in 50 mL NB amended with either 20 or 50 µg mL−1 Cr6+ for 4 and 10 days, respectively. Effect of temperature and pH (Section 2.2) on Cr6+ removal was also tested. All sets were incubated along with a control, and residual Cr6+ in culture was determined by 1, 5-diphenylcarbazide (DPC) method (APHA 1998). Aliquots (1 mL) taken at regular intervals were centrifuged (10000 rpm) for 5 min and 100 µL of cell-free supernatant was added to 10 mL of acidified glass-distilled water with 1 mL of DPC. The absorbencies measured at 540 nm were compared with Cr6+ standards and residual Cr6+ concentration was estimated (Pereira et al. 2017).
SDS PAGE analysis of bacterial whole-cell proteins
Protein profiling of bacterial isolates NIOER 176 and NIOER324 exposed to Cr6+ stress was determined by whole-cell protein extraction and sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) analysis (Laemmli 1970). Bacterial cells were harvested at mid-log phase for protein extraction (Pereira et al. 2018) and electrophoresed on a 12% SDS-PAGE gel using Bio-Rad Mini-Protean electrophoresis unit at 90 V (Bio-Rad Laboratories, USA). Cells grown without Cr6+ served as control. Following electrophoresis, protein gels were treated with fixative for 2 h and stained following silver nitrate procedure (Chevallet et al. 2006).
Results
Identification of chromate resistant bacterial strains
Bacterial isolates NIOER176 and NIOER324 were isolated from water samples collected from Divar (15°30′ 31.4606″ N, 73°54′ 43.8876″ E) and Dias beach, Dona Paula (15°27′ 12. 5496″ N, 73°48′ 4.49283″ E) respectively, along the Mandovi estuary Goa, India. Both isolates were gram positive cocci with round cream-colored colonies producing catalase. They were negative for citrate utilization, indole, methyl red, Voges Proskauer tests and did not show presence of nitrate reductase, amylase and caseinase (Table 1). NIOER324 produced gelatinase and fermented only glucose. NIOER176 did not elaborate gelatinase and fermented lactose and sucrose. Based on these biochemical characteristics according to Holt et al. (1994) and 16S rRNA gene sequencing, the isolates NIOER176 and NIOER324 were identified as Staphylococcus saprophyticus and Staphylococcus arlettae respectively.
Growth characteristics under chromate stress
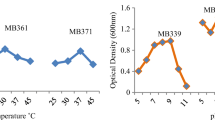
Isolates S. saprophyticus NIOER176 and S. arlettae NIOER324 were resistant to Cr6+, both having a MIC of 300 µg mL−1. Tolerance to other heavy metals revealed Hg as most toxic and As the least, in the order of tolerance As>Cr>Ni>Cu>Zn>Cd>Hg. Also, Cr3+was much less toxic than Cr6+ (Table 2). From growth kinetics (O.D6oo) (Fig. 1), rapid growth (log phase) was observed between 6–8 h with growth maxima within 24 h. Optimal growth was observed between a range of 28–37 °C and pH 7–9 (Fig. 2).
In the presence of Cr6+, both strains had prolonged lag phases, longer generation times, shorter durations of log phase and reduced growth maxima. Optimal growth was observed at pH 9.
Optimal temperature and pH for chromate removal
Batch tests with initial concentrations of 20 and 50 µg mL−1 revealed that strains completely reduced Cr6+ to non-detectable levels within 4 and 10 days respectively, at a rate of 0.260 µg mL−1 h−1 (Fig. 3). These strains were also found to remove Cr6+ over a wide range of temperatures (20–42 °C) and pH (6–9) (Table 3). At acidic pH 5 and 6 and alkaline pH 10, Cr6+ removal was drastically inhibited. No removal of Cr6+ was observed at pH 4 even after 144 h of incubation.
Antibiotic susceptibility test
Susceptibility of isolates S. saprophyticus NIOER176 and S. arlettae NIOER324 to a variety of broad and narrow spectrum antibiotics were tested (penicllins, aminoglycosides, tetracyclines, macrolides, cephalosporins, fluoroquinolones and sulfonamides). Both exhibited resistance to cephalosporins (cefotaxime, cefoxitin, ceftazidime, ceftriaxone, cefuroxime) and ß-lactams (ampicillin/sulbactam, aztreonam) but were sensitive to aminoglycosides (gentamycin, netillin, tobramycin) and fluoroquinolones (ciprofloxacin, levofloxacin, ofloxacin) (Table 4).
PCR amplification and phylogenetic analysis of ABC transportergene
PCR product was not detected in S. arlettae NIOER324. The PCR product from S. saprophyticus NIOER176 (Fig. 4) was sequenced and BLASTn search showed the sequence had the highest sequence similarity to its homologous genes from Staphylococci species. BLASTx searches identified full predicted protein functions (belonging to Staphylococci in databases) encoded by the partial DNA sequence. The partial sequence of ABC transporter gene (GenBank accession no.MH792160) isolated in this study shared highest homology (94%) with ABC transporter ATP-binding protein in Staphylococcus saprophyticus (WP 069838101). Figure 5 shows the phylogenetic relationship between Staphylococcus ABC transporter ABCT_NIOER176 and members belonging to ABC transporters from Staphylococcus species.
Phylogenetic analysis of ABT_ NIOER176 encoded ABC transporter (YP 300821.1) from S. saprophyticus. ABC transporter ATP-binding proteins S. saprophyticus (WP 002482678, WP 108866158, WP 103388803), S. aureus (CXL69696, WP 043039715, WP 000909234) and MULTISPECIES: ABC transporter ATP-binding protein Staphylococcus (WP 047528421) were used for comparison. The tree was constructed by the neighbor-joining method using Mega ver. 5
SDS PAGE analysis of bacterial whole-cell proteins
The electrophoretic profile displayed significant change in protein expression pattern when the bacterial isolates were grown in Cr6+stress conditions. A number of proteins were repressed and down-regulated in strain S. saprophyticus NIOER176 (15, 34, 49, 55, 65 and 99 kDa) and S. arlettae NIOER324 (20, 34. 44, 49, 55, 60, 65, 99 and 120 kDa) when exposed to varying concentrations of Cr6+. Both isolates revealed chromium-induced specific induction and up-regulation of 24 and 40 kDa proteins (Fig. 6). Significant diversity was also observed when comparing strain-specific patterns although both isolates displayed common and reproducible strain-specific distortions.
SDS-PAGE protein expression profile from chromate-resistant bacterial isolates a NIOER 176 and b NIOER 324 at mid-log growth phase: Lane (1) protein molecular weight marker (medium range); (2) protein isolated from bacterial strain in the absence of chromate exposure; (3) protein isolated from bacterial strain that grew in 20 µg mL−1 chromate as supplement; (4) protein isolated from bacterial strain that grew in 50 µg mL−1 chromate as supplement; (5) protein isolated from bacterial strain that grew in 100 µg mL−1 chromate as supplement
Discussion
Bacteria are exposed to heavy metals such as Cr6+, widespread in environmental settings. Once intracellular, Cr6+ binds to -SH groups and interacts with physiological ions inhibiting enzyme activity and physiological function of respective cations. This causes considerable oxidative stress. As a result, Cr6+ is toxic even at micromolar levels. Several mechanisms of bacterial Cr6+ resistance have been described (Ahemad 2014). Unlike organic toxic compounds which can be broken down or inactivated by enzyme action, Cr6+ is not degradable. In order to exert toxicity, it has to first interact with bacterial metal import systems, porins or metal ion chaperones (Hobman and Crossman 2015). Therefore primary general mechanisms of resistance involve metal/multidrug efflux and/or ion-specific efflux systems.
PCR amplification of target DNA revealed the presence of chromosomally encoded ABC transporter gene in NIOER176. The ABC transporters are a group of integral membrane proteins (part of the P-loop ATPases super-family) that translocate a variety of extracellular molecules across the membrane (Chang 2003). As a consequence of sulfate limitation during Cr6+ stress, these transporter-proteins act as importers of sulfate. They also play a major role in the iron transport system (Mazmanian et al. 2003). Known to provide Cr6+ resistance (Claude and Adolhe 2013), these transporters are abundantly expressed during Cr6+ stress and further confer resistance to structurally diverse antibiotics (Blanco et al. 2016) as expressed by multi-antibiotic resistance in S. saprophyticus NIOER176. Notably, phylogentic analysis revealed, NIOER176 encoded ABCT_NIOER176 placed in the same clade as ABC transporter ATP-binding protein from Staphylococcus saprophyticus. This is confirming its origin from staphylococcal genome. The PCR negative result in S. arlettae 324 did not allow detection of this sequence. Claude and Adolhe (2013) similarly reported detection in S. saprophyticus and S. aureus, but not in S. arlettae strains. The PCR negative result in S. arlettae NIOER324 is likely due to its absence and reliance on mediation by other mechanisms. Further, primers used in this study may not have picked up its analog owing to species gene diversity and difference-based degeneracy of ABC transporter genes (Achour et al. 2007).
Resistance was not confined to chromium alone. Both isolates were tolerant of different heavy metals and were resistant to many antibiotics. Similarly, Safari Sinegani and Younessi (2017) also reported a close association of multiple metals and antibiotic resistance. This shared structural and functional characteristic of bacterial antibiotic and metal resistance systems indicate overlapping mechanisms of action. As Baker-Austin et al. (2006) suggested, a specific pattern of high resistance to ß-lactams and cephalosporins (observed in this study) indicates (i) reduction in Cr6+ permeability, (ii) ability for drug/metal alteration or efflux, and/or (iii) modification of cellular targets as shared mechanisms for metal and antibiotic resistance. Several components underlie this co-selection such as (i) the close proximity of genes encoding resistance on the same genetic element (co-resistance) (Summers et al. 1993), (ii) the same genetic determinants responsible for resistance to both (cross-resistance), or (iii) indirect but shared responses such as biofilm formation or efflux pumps that target both toxicants (Baker-Austin et al. 2006). Thus, exposure to a single toxic agent may select for strains resistant to multiple toxicants. Such microorganisms are important in carrying out biological function in metal contaminated environs and may have application in their remediation.
Reduction of Cr6+ to less toxic Cr3+ can be performed by bacterial mediated processes of Cr resistance. The removal of Cr6+ in culture supernatants of bacteria in this study is consistent with earlier reports of Wang and Xiao (1995) and Song et al. (2009). Isolates studied, removed Cr6+ from solution rapidly over a wide range of temperatures, concentrations (20–50 µg mL−1) and a pH range of 6–9. Camargo et al. (2003) and Congeevaram et al. (2007) similarly demonstrated optimal removal at near-natural range of pH. They suggested that beyond the pH range of 6–9, inability to remove or reduce chromium was attributed to osmotic changes, hydrolyzing effect and/or change in solution chemistry of Cr6+ metal ions such as Cr2O72−, HCrO−4 and Cr2O42−. This, and optimal removal observed at pH 9 indicates that Cr6+ removal is subject to organism-specific biological/physiological factors. The range of optimal temperature values (24–37 °C) reflects the mesophilic nature of the strains. At temperature extremes (20 and 42 °C) it is likely that ionization of related moieties, cell wall stability and its configuration are affected thus reducing Cr6+ removal (Congeevaram et al. 2007). Although such details were not carried out for this study, the results do connotes a positive relation between Cr6+ removal and cellular growth independent of varied ranges of temperature or pH used in the experiments.
Protein profiling of Cr6+exposed NIOER 176 and NIOER324 indicated inverse decrease in protein content with increase in Cr6+. This highlighted cellular proteins as a main target of metal toxicity as reported by Novo et al. (2000), Congeevaram et al. (2007) and Shamim et al. (2012). As a result, both high and low- molecular weight proteins were repressed. Estuarine strains in this study also exhibited several other responses to sustain in Cr6+ stress exposure. Expression patterns (disappearance of bands) indicated some proteins were asymmetrically down-regulated. Reports of proteins actively repressed by bacteria (Chandrangsu et al. 2017, Quintana et al. 2017) especially those involved in metal uptake and transport have been described as an adaptation to limit intracellular entry of toxic metal ions. It could be speculated that some of these proteins down-regulated play a similar role. Chromate induced specific up-regulation of unidentified 40 kDa protein represents a regulatory response that may confer protection, possibly as a metal efflux protein (Haritha et al. 2009, Khan et al. 2016). Similarly, the unidentified 24 kDa protein may provide membrane-associated function (Kassab and Roane 2006). Further studies are required to identify the putative protein and to elucidate its role, if any, in Cr6+ resistance.
Resistance to multiple toxicants has been observed in many bacterial systems and characterized at the physiological and molecular level. Chihomvu et al. (2015) has reported of genetic determinants that confer resistance to heavy metals. It is likely that multi-metal, multi-drug resistant isolates NIOER176 and NIOER324 possess the genetic machinery for targeting other toxic metal ions. Biological approaches utilizing such microbes offer the potential for highly selective removal of toxicants along with added operational flexibility. Such isolates originating from habitats with low or no contamination of toxic metals are useful tools for molecular characterization of multiple metal resistance mechanisms for delineating metal microbe-mediated ecologic processes and in devising bioremediation of toxicants especially from environments contaminated by many metals. Both isolates being resistant to many tested antibiotics, prior confirmation of their non-virulence would be essential to ensure their inclusion into bioremediation is safe and sound and health-risk free.
References
Achour AR, Bauda P, Billard P (2007) Diversity of arsenite transporter genes from arsenic-resistant soil bacteria. Res Microbiol 158(2):128–37. https://doi.org/10.1016/j.resmic.2006.11.006
Ahemad M (2014) Bacterial mechanisms for Cr(VI) resistance and reduction: an overview and recent advances. Folia Microbiol (Praha) 59(4):321–32. https://doi.org/10.1007/s12223-014-0304-8
Akcil A, Erust C, Ozdemiroglu S, Fonti V, Beolchini F (2015) A review of approaches and techniques used in aquatic contaminated sediments: metal removal and stabilization by chemical and biotechnological processes. J Clean Prod 86:24–36. https://doi.org/10.1016/j.jclepro.2014.08.009
APHA, AWWA, WEF (1998) Standard methods for the examination of water and wastewater. 20th American Public Health Association, American Water Works Association, Water Environment Federation. Washington, DC
Baker-Austin C, Wright MS, Stepanauskas R, McArthur JV (2006) Co-selection of antibiotic and metal resistance. Trends Microbiol 14(14):176–82
Batool R, Yräla K, Hasnain S (2012) Hexavalent chromium reduction by bacteria from tannery effluent. J Microbio lBiotechnol 22(4):547. https://doi.org/10.4014/jmb.1108.08029
Bauer A, Kirby W, Sherirs J, Turch M (1966) Antibiotic susceptibility testing by standard single disk method. Am J Clin Pathol 45(4):493–6. PMID: 5325707
Blanco P, Hernando-Amado S, Reales-Calderon JA, Corona F, Lira F, Alcalde-Rico M, Bernardini A, Sanchez MB, Martinez JL (2016) Bacterial multidrug efflux pumps: much more than antibiotic resistance determinants. Microorganisms 4(1):pii: E14. https://doi.org/10.3390/microorganisms4010014
Bolan N, Kunhikrishnan A, Thangarajan R, Kumpiene J, Park J, Makino T, Kirkham MB, Scheckel K (2014) Remediation of heavy metal(loid)s contaminated soils—to mobilize or to immobilize? J Hazard Mater 266:141–66. https://doi.org/10.1016/j.jhazmat.2013.12.018
Bontidean I, Lloyd JR, Hobman JL, Wilson JR, Csöregi E, Mattiasson B, Brown NL (2000) Bacterial metal-resistance proteins and their use in biosensors for the detection of bioavailable heavy metals. J Inorg Biochem 79(1-4):225–9. https://doi.org/10.1016/S0162-0134(99)00234-2
Camargo FAO, Bento FM, Okeke BC, Frankenberger WT (2003) Chromate reduction by chromium resistant bacteria isolated from soils contaminated with dichromate. J Environ Qual 32(4):1228–1233. https://doi.org/10.2134/jeq2003.1228
Chandrangsu P, Rensing C, Helmann JD (2017) Metal homeostasis and resistance in bacteria. Nat Rev Microbiol 15(6):338–50. https://doi.org/10.1038/nrmicro.2017.15
Chang G (2003) Multidrug resistance ABC transporters. FEBS Lett 555(1):102–5. https://doi.org/10.1016/S0014-5793(03)01085-8
Chevallet M, Luche S, Rabilloud T (2006) Silver staining of proteins in polyacrylamide gels. Nat Protoc 1(4):1852–58. https://doi.org/10.1038/nprot.2006.288
Chihomvu P, Stegmann P, Pillay M (2015) Characterization and structure prediction of partial length protein sequences of pcoA, pcoR and chrB genes from heavy metal resistant bacteria from the Klip River, South Africa. Int J MolSci 16(4):7352–74. https://doi.org/10.3390/ijms16047352
Claude KZG, Adolhe Z (2013) Proteomic analysis of chromate response in Staphylococcus saprophyticus isolated from a fly ash dumping site. Afr J Biotechnol 12(18):2322–30. https://doi.org/10.5897/AJB12.127
Congeevaram S, Dhanarani S, Park J, Dexilin M, Thamaraiselvi K (2007) Biosorption of chromium and nickel by heavy metal resistant fungal and bacterial isolates. J Harzard Mater 19(1-2):270–7. https://doi.org/10.1016/j.jhazmat.2006.12.017
García R, Campos J, Cruz JA, Calderón ME, Raynal ME, Buitrón G (2016) Biosorption of Cd, Cr, Mn and Pb from aqueous solutions by Bacillus sp strains isolated from industrial waste activate sludge. TIP 19:5–14. https://doi.org/10.1016/j.recqb.2016.02.001
Haritha A, Sagar KP, Tiwari A, Kiranmayi P, Rodrigue A, Mohan PM, Singh SS (2009) MrdH, a novel metal resistance determinant of Pseudomonas putida KT2440, is flanked by metal-inducible mobile genetic elements. J Bacteriol 191(19):5976–87. https://doi.org/10.1128/JB.00465-09
Hobman JL, Crossman LC (2015) Bacterial antimicrobial metal ion resistance. J Med Microbiol 64(Pt 5):471–97. https://doi.org/10.1099/jmm.0.023036-0
Holt JG, Krieg NR, Sneath PHA, Staley JT, Williams ST (1994) Bergey’s Manual of Determinative Bacteriology. Baltimore, MD
Joutey NT, Sayel H, Bahafid W, El Ghachtouli N (2015) Mechanisms of hexavalent chromium resistance and removal by microorganisms. Rev Environ Contam Toxicol 233:45–69. https://doi.org/10.1007/978-3-319-10479-9_2
Kamika I, Momba MN (2013) Assessing the resistance and bioremediation ability of selected bacterial and protozoan species to heavy metals in metal-rich industrial wastewater. BMC Microbiol 13:28. https://doi.org/10.1186/1471-2180-13-28
Kassab DM, Roane TM (2006) Differential responses of a mine tailings Psuedomonas isolate to cadmium and lead exposure. Biodegradation 17(4):379–87. https://doi.org/10.1007/s10532-005-9010-1
Khan Z, Rehman A, Hussain SZ, Nisar MA, Zulfiqar S, Shakoori AR (2016) Cadmium resistance and uptake by bacterium, Salmonella enterica 43C, isolated from industrial effluent. AMB Express 6(1):54. https://doi.org/10.1186/s13568-016-0225-9
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of the bacteriophage T4. Nature 227(5259):680–685
Mazmanian SK, Skaar EP, Gaspar AH, Humayun M, Gornicki P, Jelenska J, Joachmiak A, Missiakas DM, Schneewind O (2003) Passage of heme-iron across the envelope of Staphylococcus aureus. Science 299(5608):906–09. https://doi.org/10.1126/science.1081147
McLaughlin MJ, Parker DR, Clark JM (1999) Metal and micronutrients-food safety issues. Field Crops Res 60(1-2):143–63. https://doi.org/10.1016/S0378-4290(98)00137-3
Mohapatra RK, Parhi PK, Thatoi H, Panda CR (2017) Bioreduction of hexavalent chromium by Exiguobacterium indicum strain MW1 isolated from marine water of Paradip Port, Odisha, India. Chem Ecol 33(2):114–130
Nies DH (2003) Efflux-mediated heavy metal resistance in prokaryotes. FEMS Microb Rev 27(2-3):313–39. https://doi.org/10.1016/S0168-6445(03)00048-2
Novo MTM, Da Silva AC, Ronaldo M, Paula C, Antonia C, Oswaldo JG, Ottoboni LMM (2000) Thiobacillus ferrooxidans response to copper and other heavy metals: growth, protein synthesis and protein phosphorylation. Antonie Van Leeuwenhoek 77(2):187–95. https://doi.org/10.1023/A:1002462701671
Pereira EJ (2019) Molecular and physiological analyses of hexavalent chromium biotransformation by marine bacteria. Ph D Thesis, Goa University
Pereira EJ, Fonseca S, Meena RM, Ramaiah N (2017) Improved sprouting and growth of mung plants in chromate contaminated soils treated with marine strains of Staphylococcal species. Indian J Microbiol 57(4):400–408. https://doi.org/10.1007/s12088-017-0668-y
Pereira EJ, Ramaiah N, Damare S, Furtado B (2018) Differential protein analysis of hexavalent chromium stress response in marine Staphylococcus xylosus. Curr Proteomics 15:42–54
Pulimi M, Jamwal S, Samuel J, Chandrasekaran N, Mukherjee A (2012) Enhancing the hexavalent chromium bioremediation potential of Acinetobacter junii VITSUKMW2 using statistical design experiments. J Microbiol Biotechnol 22(12):1767–75. https://doi.org/10.4014/jmb.1203.03063
Quintana J, Novoa-Aponte L, Arguello JM (2017) Copper homeostasis networks in the bacterium Pseudomonas aeruginosa. J Biol Chem 292(38):15691–704. https://doi.org/10.1074/jbc.M117.804492
Safari Sinegani AA, Younessi N (2017) Antibiotic resistance of bacteria isolated from heavy metal-polluted soils with different land uses. J Glob Antimicrob Resist 10:247–255. https://doi.org/10.1016/j.jgar.2017.05.012
Salamanca D, Strunk IN, Engesser KH (2013) Chromate reduction in anaerobic systems by bacterial strain Pseudomonas aeruginosaCRM100. Chemie Ingenieur Technik 85(10):1575–80. https://doi.org/10.1002/cite.201200144
Sathicq MB, Gómez N (2018) Effects of hexavalent chromium on phytoplankton and bacterioplankton of the Río de la Plata estuary: an ex-situ assay. Environ Monit Assess 190(4):229. https://doi.org/10.1007/s10661-018-6619-1
Shamim K, Naik M, Pandey A, Dubey SK (2012) Isolation and identification of Aeromonas caviae strain KS-1 as TBTC- and lead-resistant estuarine bacteria. Environ Monit Assess 185(6):5243–9. https://doi.org/10.1007/s10661-012-2940-2
Singh S, Parihar P, Singh R, Singh VP, Prasad SM (2015) Heavy metal tolerance in plants: role of transcriptomics, proteomics, metabolomics, and ionomics. Front Plant Sci 6:1143. https://doi.org/10.3389/fpls.2015.01143
Song H, Liu Y, Xu W, Zeng G, Aibibu N, Xu L, Chen B (2009) Simultaneous Cr(VI) reduction and phenol degradation in pure cultures of Pseudomonas aeruginosa CCTCC AB91095. Bioresour Technol 100(21):5079–84. https://doi.org/10.1016/j.biortech.2009.05.060
Summers AO, Wireman J, Vimy MJ, Lorscheider FL, Marshall B, Levy SB, Bennett S, Billard L (1993) Mercury released from dental silver fillings provokes an increase in mercury-resistant and antibiotic resistant bacteria in oral and intestinal floras of primates. Antimicrob Agents Chemother 37(4):825–834
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci USA 101:11030–35
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–39
Thatoi H, Das S, Mishra J, Rath BP, Das N (2014) Bacterial chromate reductase, a potential enzyme for bioremediation of hexavalent chromium: a review. J Environ Manage 15(146):383–9. https://doi.org/10.1016/j.jenvman.2014.07.014
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–82
US Environmental Protection Agency (1998) Toxicological review on hexavalent chromium. CAS No. 18540-29-9. Washington, DC
Wang YT, Xiao C (1995) Factors affecting hexavalent chromium reduction in pure cultures of bacteria. Water Res 29(11):2467–74. https://doi.org/10.1016/0043-1354(95)00093-Z
Acknowledgements
We thank the Director, CSIR-NIO for support and facilities to carry out this study. The authors acknowledge the funding support from CSIR project PSC0206. The first author acknowledges research fellowship by the University Grant Commission (UGC) (Sanction letter No: 18-12/2011(ii) EU-V). This is NIO’s contribution No.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
About this article
Cite this article
Pereira, E.J., Ramaiah, N. Chromate detoxification potential of Staphylococcus sp. isolates from an estuary. Ecotoxicology 28, 457–466 (2019). https://doi.org/10.1007/s10646-019-02038-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10646-019-02038-w