Abstract
Ecophysiological differences between ammonia-oxidizing bacteria (AOB) and ammonia-oxidizing archaea (AOA) enable them to adapt to different niches in complex freshwater wetland ecosystems. The community characters of AOA and AOB in the different niches in a freshwater wetland receiving municipal wastewater, as well as the physicochemical parameters of sediment/soil samples, were investigated in this study. AOA community structures varied and separated from each other among four different niches. Wetland vegetation including aquatic macrophytes and terrestrial plants affected the AOA community composition but less for AOB, whereas sediment depths might contribute to the AOB community shift. The diversity of AOA communities was higher than that of AOB across all four niches. Archaeal and bacterial amoA genes (encoding for the alpha-subunit of ammonia monooxygenases) were most diverse in the dry-land niche, indicating O2 availability might favor ammonia oxidation. The majority of AOA amoA sequences belonged to the Soil/sediment Cluster B in the freshwater wetland ecosystems, while the dominant AOB amoA sequences were affiliated with Nitrosospira-like cluster. In the Nitrosospira-like cluster, AOB amoA gene sequences affiliated with the uncultured ammonia-oxidizing beta-proteobacteria constituted the largest portion (99 %). Moreover, independent methods for phylogenetic tree analysis supported high parsimony bootstrap values. As a consequence, it is proposed that Nitrosospira-like amoA gene sequences recovered in this study represent a potentially novel cluster, grouping with the sequences from Gulf of Mexico deposited in the public databases.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Ammonia can be oxidized to nitrite under aerobic condition by aerobic ammonia-oxidizing bacteria (AOB) (Winogradsky 1890). Until recently, the discovery and isolations of ammonia-oxidizing archaea (AOA) capable to carry out chemoautotrophic ammonia oxidation, challenge the traditional view that ammonia oxidation is restricted to AOB alone (Könneke et al. 2005; Venter et al. 2004). The aerobic ammonia-oxidizing prokaryotes (including AOA and AOB) play an important role in nitrogen removal as they carry a unique amo gene encoding for the enzyme ammonia monooxygenase, including three small subunits, amoA, amoB and amoC (Norton et al. 2002).
The amoA genes in both AOA and AOB have been used as molecular biomarkers to investigate the corresponding community structures and distribution in diverse ecosystems, including soils (Pett-Ridge et al. 2013; Taylor et al. 2012; Tourna et al. 2011; Wang et al. 2013a, 2013b, 2014b; Zhang et al. 2012; Zhou et al. 2014), oceans (Beman et al. 2008; Hollibaugh et al. 2002; Pitcher et al. 2011; Santoro and Casciotti 2011; Walker et al. 2010; Wuchter et al. 2006), coastal wetlands (Cao et al. 2012; Wang et al. 2013c; Wang and Gu 2013, 2014; Wang et al. 2014a), estuaries (Beman and Francis 2006; Dang et al. 2008), lakes (Jiang et al. 2009), freshwaters (Herrmann et al. 2008, 2009), salts marsh (Dorador et al. 2008; Moin et al. 2009). However, the role of AOA in nitrogen cycle of the freshwater habitat such as freshwater marsh remains unclear (Laanbroek and Speksnijder 2008). Most of the studies showed that AOA amoA gene is predominant over that of AOB in terms of both the richness and abundance in various ecosystems (Adair and Schwartz 2008; Dang et al. 2008; Herrmann et al. 2009; Prosser and Nicol 2008), but some reports showed contradictory results in environments such as activated wastewater treatment bioreactor (Wells et al. 2009), lake sediment (Jiang et al. 2009) and estuary sediment (Mosier and Francis 2008). AOB isolates are divided into two monophyletic groups: beta- and gamma-proteobacteria (Kowalchuk and Stephen 2001). The beta-AOB communities are simply further divided into two genera: Nitrosomonas and Nitrosospira (Purkhold et al. 2000; 2003).
Wetlands are a complex ecosystem consisting of marshes, swamps and bogs and these different environmental types offer a favorable condition for the development of common wetland plants such as Phragmites australis and P. communis and create different niches due to vegetation types, oxygen bioavailability, water coverage, and nutrient variation (Ruiz-Rueda et al. 2009; You et al. 2009). It is suggested that both nitrification and denitrification could be achieved in wetlands but at low rates (Clément et al. 2002; Verhoeven and Meuleman 1999). However, to our knowledge, there is no information about the niche specificity of AOA communities in freshwater marsh although the potential niche differentiation may drive AOB communities (Laanbroek and Speksnijder 2008). It is hypothesized that the wetland vegetation of common reed and the environment physicochemical parameters contribute to the species or ecotypes differentiation of AOA and AOB communities in a freshwater wetland.
Longfeng Freshwater Wetland Nature Reserve (E125°07′–125°15′; N46°28′–46°32′) is one of the main local provincial level marshes in Daqing, Heilongjiang Province, China (Xu and Zhang 2009). This wetland includes 314 species of protected wild plants, 207 species of wild land animals, 45 species of wild provincial capital level protected fish, 24 species of wild state level protected birds, 98 species of wild provincial capital level birds and 19 Heilongjiang province protected animal species in an area of 6,211 hm2 (Hou 2009). However, human disturbance to the wetlands such as Daqing oil field development, urban construction, agriculture activities and other factory discharge may threaten the wetland. Because of that, this wetland could serve as a prime study site for examination of nitrogen removal by ammonia-oxidizing microorganisms. The objectives of this study were to investigate the potential driving factors on the community characteristics of AOA and AOB in the freshwater wetland close to the Daqing city.
Materials and methods
Sampling site description and collecting method
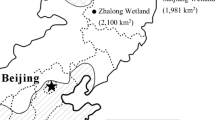
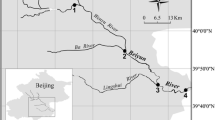
Surface and bottom soil/sediment samples were collected from different sites in the Daqing Longfeng Freshwater Wetland Natural Reserve, Heilongjiang, China in summer of 2008 (Fig. 1). The sampling sites were classified as four types of niches: (1) Samples 23 and 24 sediments were collected from a wetland covered with reeds and the surface sediment was characterized of rich nitrate. This site was labeled as R; (2) Sample 25 and 26 sediments were collected from a wetland near the niche R, covered with standing water but without any plantations. This site served as the reed control and was labeled as R-Con; (3) Sample 2S and 2B sediments were collected from a marsh characterized of relatively high concentration of ammonia, located near a main highway and further apart from niches R and R-Con. This site was labeled RW; (4) Sample 3S and 3B soils were collected from a dry land nearby the niche RW with mixed vegetation of mainly common poplar trees and labeled as D. The soils/sediments between 0.5 cm and 10 cm were classified as surface while that below 10 cm were classified as bottom. All samples were transported to the laboratory on the dry-ice in a cooling box.
Analysis of chemical characteristics
Concentrations of ammonium-N, nitrate–N, and nitrite-N were measured by extracting sediments in 2.0 M KCl (Zuber et al. 2005) and the extracts were analyzed by an auto-analyzer (QuikChem, Milwaukee, WI) using standard flow injection analysis (FIA) techniques according to the standard extraction procedures. Replicate chemical tests were performed for each sample.
DNA extraction and PCR amplification
DNA was extracted from 100 to 200 mg of soil/sediment using the SoilMaster™ DNA Extraction Kit according to the manufacturer’s protocol (Epicentre Biotechnologies, Madison, WI). DNA extracts were stored at −20 °C before using as a template for PCR reaction.
The archaeal amoA genes were amplified using the PCR primers Arch-amoAF (5′–STAATGGTCTGGCTTAGACG–3′) and Arch-amoAR (5′–GCGGCCATCCATCTGTATGT–3′). Based on the standard procedures described in the manufacturer’s instructions and results of previous studies (Francis et al. 2005), the optimized PCR reaction mixture in a 25 μl volume contained 5 μl of 5× colorless GoTaq®Flexi Buffer, 12.5 mM MgCl2, 5 mM of each deoxyribonucleotide triphosphate, 0.5 μM of each primer, 1.25U GoTaq® DNA Polymerase and 3 μl of purified DNA as template (10–100 ng). PCR conditions were set as follows: 94 °C for 3 min; 40 cycles of 94 °C for 30 s, 53 °C for 1 min, and 72 °C for 1 min; and finally 72 °C for 15 min.
The bacterial amoA genes were amplified using the PCR primers amoA-1F (5′–GGGGGTTTCTACTGGTGGT–3′) and amoA-2R (5′–CCCCTCKGSAAAGCCTTCTTC–3′). The optimized PCR reaction mixture in a 25 μl volume contained 5 μl of 5× colorless GoTaq®Flexi Buffer, 12.5 mM MgCl2, 5 mM of each deoxyribonucleotide triphosphate, 0.5 μM of each primer, 1.25U GoTaq® DNA Polymerase and 3 μl of purified DNA as template (10–100 ng). PCR conditions were set as follows: 94 °C for 3 min; 37 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 45 s; and finally 72 °C for 10 min.
Cloning, sequencing and phylogenetic analysis
Purified amoA DNA fragments were ligated into a pMD18-T vector (TaKaRa, Japan) and transformed into competent Escherichia coli DH5α cells. The library clones were screened directly by PCR for the presence of inserts using two M13 universal primers: M13F (5′-GTTTCCCAGTCACGAC-3′) and M13R (5′-TCACA CAGGAAACAGCTATGAC-3′). The positive clones from each library were randomly selected and the PCR products were purified using a PCR Purification Kit (Qiagen, USA). The PCR products were sequenced (Tech Dragon Ltd, Hong Kong). DNA sequences were examined and edited using BioEdit (Tom Hall, North Carolina State University, NC) and MEGA, version 5.0 (Tamura et al. 2011). NCBI BLAST (http://www.ncbi.nih.gov) was used to find the closest related amoA gene sequences in the GenBank. The multiple alignments of amoA gene sequences were considered and neighbor-joining phylogenetic trees with bootstrap values based on 1,000 replications (Felsenstein 1985) were produced using MEGA program, version 5.0.
Data statistical analysis
The DOTUR (Distance-Based OTU and Richness) program was employed to compare diversity for amoA sequences from each sample (Schloss and Handelsman 2005). Operational taxonomic units (OTU) for community analysis were defined by a 5 and 1 % sequence variation in detecting AOA and AOB respectively, as determined using the furthest neighbor algorithm in DOTUR. DOTUR was also used to generate diversity analyses such as coverage, Chao1, Simpson index (D) and Shannon index (H) numbers for each site. The H index and the D index were calculated as the diversity indices by the program (Beman and Francis 2006; Schloss and Handelsman 2005). Another recently developed and more advanced community analysis program MOTHUR was employed to conduct the UniFrac Environment Clustering Analysis (Schloss et al. 2009). The estimated coverage of the constructed archaeal amoA gene and bacterial amoA gene libraries were calculated using the formula of C = [1−(n1/N)] × 100, where n1 represents the number of OTUs detected in one clone library and N strands for the total number of clones in that particular library. This coverage estimates the probability that all the unique sequences present in a given sample were represented at least once in the library (Dang et al. 2008; Mullins et al. 1995). Both AOA and AOB community differences between the niches were determined using non-weighted principal co-ordinates analysis (PCoA) and Jackknife Environment Clustering Analysis (Lozupone et al. 2007). Both the PCoA and Jackknife Environment Clustering Analysis were generated using online UniFrac tool program (http://bmf2.colorado.edu/unifrac/index.psp) according to the sequences molecular evolutionary distances and microbial community similarity based on the environmental occurrences. Correlation analysis between the diversity of archaeal and bacterial amoA genes and environmental variables were conducted using Microsoft Excel program.
Nucleotide sequence accession numbers
All the partial archaeal and bacterial amoA gene sequences of ammonia-oxidizing bacteria and archaea determined in this study were deposited in GenBank with accession numbers HM537746 to HM537915 and HM537916 to HM538185, respectively.
Results and discussion
Physicochemical characteristics of sediments/soils
Sediment/soil cores were obtained from four selected locations and the concentrations of N-associated parameters were measured (Table 1). Samples 2S and 2B, classified as ammonia rich niches (RW), had highest concentration of ammonia. In contrast, samples 25 and 26, defined as non-vegetated ones (R-Con), had the lowest except the sample 24. In general, surface samples had higher concentrations of ammonium and nitrate than bottom ones except in the RW niche. Nitrite concentrations in the soils were low in non-vegetated niche, but slightly higher in vegetated niches. Nitrate concentrations in all samples were generally low and even below detection limit (0.1 µg g−1 DW) in the RW niches and the bottom soil of the dry-land samples.
AOA community composition and phylogenetic affiliation
Seven archaeal amoA gene libraries among four locations were constructed except the bottom sample in the eutrophic reed site with water covering (RW), which failed to amplify amoA gene successfully. Possible reasons might be that AOA were either absent or under the PCR amplification detection limit. Among these seven clone libraries, 270 clones were obtained and 26 OTUs were identified. A total of 6–14 OTUs were observed in each niche type using a 5 % cutoff at the nucleotide level (Table 2). The high OTUs value indicated that the four niches harbored diverse AOA. The estimated coverage of all libraries was quite high (Table 2), indicating the clone libraries represented majority of archaeal amoA gene sequences present in the samples, and this information is in agreement with rarefaction analyses (Fig. S1a). The dry-land site (D) had the highest Shannon index values while non-vegetated reed site (R-Con) had the lowest. These results may raise a possibility that the terrestrial soil supports a higher AOA diversity than the freshwater covered niches, where terrestrial soil has a higher oxygen concentration and hence facilitate the ammonia oxidation (Christensen et al. 1990). The presence of plants may release oxygen in the rhizosphere through the roots (Herrmann et al. 2008; Laanbroek and Speksnijder 2008) and provide aerobic zones sustaining the growth of AOA communities. Hence common weed plant species in vegetated niches except the R-Con might favor the development of various AOA species.
The phylogenetic tree of AOA (Fig. 2) showed that all the archaeal amoA gene sequences were grouped into two main clusters: Water column/sediment clade (Cluster A1) and Soil/sediment clade (Cluster B) defined earlier (Francis et al. 2005). In the phylogenetic tree, Cluster A2 consisted of several amoA gene sequences from estuarine samples and soils. Only a small proportion (about 2 %) of sequences fell into Cluster A1 while the rest majority fell into Cluster B. It is reasonable that the majority of archaeal amoA gene sequences recovered from the freshwater wetland soils are associated with the Soil/sediment AOA sequences. In Cluster A1, all the sequences were retrieved from the niche R-Con except one clone from the niche common reed (R). The closest representative relative in Cluster A1 was Nitrosopumilus Maritimus, which was firstly found in substratum of a tropical marine tank and inhabited at oligotrophic environment (Könneke et al. 2005).
Phylogenetic tree of AOA constructed with amoA gene fragments (594 bp). The numbers in the brackets are the clone number of the related phylotype which S represents surface sediment and B represents bottom sediment. The numbers at the nodes are percentages that indicate the levels of bootstrap support based on 1,000 resample data sets (only values greater than 50 % are shown). Branch lengths correspond to sequence differences as indicated by the scale bar. The clustering reference was defined by Francis et al. (2005), Herrmann et al. (2008), and (2009), and modified with several sub-clusters in Cluster B in this study
AOA community distribution in response to niches and physicochemical parameters
The Jackknife Environment Clusters Analysis (Fig. S2a) showed that the vegetated niches RW, D and R were grouping together, and the non-vegetated niche R-Con was distinct from vegetated ones. This supported the previous studies in that plantation could stimulate the rhizosphere-association nitrification, in turn cause the community shift of AOA (Herrmann et al. 2008; Laanbroek and Speksnijder 2008; Ottosen et al. 1999; Riis and Sand-Jensen 1998). PCoA analysis of AOA communities showed that individual niches did not group together (Fig. 3a), indicating that unique patterns of AOA community structures occurred in each niche independently. The results are consistent with the idea proposed by Herrmann et al. (2009). The phylogeny of AOA in freshwater marsh environment might be associated with different niche characteristics such as presence of water and plantation. Environmental factors might also contribute to the community diversity of AOA in the wetland. The results of Pearson moment correlation test further supported this suggestion (Table 3). Shannon index of AOA had significant positive correlation to the concentration of ammonia in soils (Pearson moment correlation: r = 0.74, P < 0.05, n = 6), suggesting that ammonia input could be a pivotal factor enhancing AOA diversity in freshwater wetland. Previous studies showed that a higher abundance and diversity of archaeal amoA genes were detected under higher total ammonium concentrations (Park et al. 2006; Treusch et al. 2005). In contrast, some studies found that low ammonium concentrations might restrict the growth of AOA in some low N-containing environments (Erguder et al. 2009; Könneke et al. 2005; Reigstad et al. 2008).
Ordination diagrams of the sedimentary a archaeal amoA and b bacterial amoA assemblages calculated with unweighted UniFrac PCoA analyses using deduced amoA protein sequences. See Table 1 for detailed explanation for niche codes
AOB community composition and phylogenetic affiliation
The phylogenetic analysis was based on bacterial amoA gene with representative sequences retrieved from the GenBank database (Fig. 4). Eight bacterial amoA libraries were constructed among four niches, a total of 170 clones were obtained and 13 OTUs were identified. The estimated coverage of all libraries was quite high (Table 3), which was in agreement with the result of rarefaction analyses (Fig. S1b). Similar to the Shannon index of AOA, the dry-land site had the highest Shannon index values of AOB than other niches and the non-vegetated reed site had the lowest.
Phylogenetic tree of AOB constructed with amoA gene fragments (450 bp). The numbers in the brackets are the clone number of the related phylotype which S represents surface sediment and B represents bottom sediment. The numbers at the nodes are percentages that indicate the levels of bootstrap support based on 1,000 resample data sets (only values greater than 50 % are shown). Branch lengths correspond to sequence differences as indicated by the scale bar. The clustering reference was defined by Avrahami et al. (2002), Purkhold et al. (2000), and Purkhold et al. (2003), and modified in this study
Compared to AOA, a relatively lower diversity of AOB was found in different niches, for both surface and bottom sediment/soil. All the sequences were classified and grouped into two β-proteobacteria: Nitrosospira-like and Nitrosomonas-like sequences. Most of the sequences (about 62 %) belonged to the Nitrosospira-like cluster while the Nitrosomonas-like cluster contained the rest clone sequences (about 38 %). Among the Nitrosospira-like cluster, 99 % sequences were similar to Nitrosospira-like sequences retrieved from sea water column in Gulf of Mexico. As defined by the previous studies, a sequence identity of less than 80 % (partial amoA gene sequence) and 88 % (whole amoA gene sequence) respectively has been suggested as a novel AOB clusters/lineages (Kim et al. 2008; Purkhold et al. 2000). The 105 amoA sequences found in this study had 75 % DNA identity to the referencing sequences Nitrosospira multiformis (DQ228454) and Nitrosomonas marina Nm22 (AF272405). As a consequence, these sequences together with the amoA sequences from Gulf of Mexico are proposed to be a novel independent cluster. This proposed cluster was found from all tree methods independently and supported by high parsimony bootstrap analysis (100 %). According to the phylogenetic tree, this novel cluster consisted of amoA sequences from all niches, indicating that it might be widely distributed in the freshwater marsh. One sequence from eutrophic reed (RW) niche was related to the Nitrosospira briensis Nsp10 and C-57 (AY123821 and Z97858 respectively) and grouped into cluster 3b. This finding was consistent with the dominance of Nitrosospira cluster 3 found in the relatively high ammonium concentrations in soils (Avrahami et al. 2002; Bruns et al. 1999; Kowalchuk et al. 2000).
Around 70 % sequences in the Nitrosomonas-like clade were affiliated to the Nitrosomonas communis cluster and had 98 % identity similar to an uncultured ammonia-oxidizing bacterium, isolate S38 (AM851051) retrieved from the environmental clone in rice field soil affected by nitrogen fertilization (Wang et al. 2009). Among the published AOB isolated species, these amoA sequences in the N. communis cluster were the most similar to the Nitrosomonas nitrosa. Studies showed that N. nitrosa prefer to inhabit in eutrophic freshwater environment, while N. communis prefer to inhabit in soils (Koops and Pommerening-Röser 2001). Two sequences from the non-vegetated niche R-Con were related to a sequence found in the coastal aquaculture environment (EU156173). In the eutrophic reed niche RW, 14 clone sequences were related to the N. marina Nm22 (AF272405). All these 16 clones were clustered into the Nitrosomonas oligotropha/Nitrosomonas marina lineage. The isolates among this lineage were retrieved from either wastewater or freshwater habitats, and they had low ammonium affinity constants (Koops and Pommerening-Röser 2001; Pommerening-Röser et al. 1996; Purkhold et al. 2003; Stehr et al. 1995). Four sequences in dry-land (D) niche were related to the Nitrosomonas halophila (AF272398) and uncultured AOB clone 17-11 (EU116356), with 96 % identities. These four sequences fell into the Nitrosomonas europaea/Nitrosococcus mobilis lineage whose members preferred to inhabit in sewage disposal plants, eutrophic freshwater and brackish water with relatively high ammonium substrate (Koops and Pommerening-Röser 2001).
The AOB detected in the freshwater wetland soils were mainly Nitrosospira-like amoA sequences, which was consistent with the results of other wetland studies that Nitrosospira species dominated in the wetland system (Ibekwe et al. 2003; Laanbroek and Speksnijder 2008; Moin et al. 2009). The phylogenetic tree showed a high AOB community composition overlapping among different niches from the freshwater marsh (Fig. 4). In the present study, AOB communities in the freshwater wetland sediments consisted of five stable lineages: Nitrosospira cluster 3b, Nitrosomonas marina/Nitrosomonas oligotropha cluster, N. communis cluster, Nitrosomonas europaea/Nitrosococcus mobilius cluster, and a proposed novel Nitrosospira cluster. The discovery of the novel potential AOB cluster suggested that a more wide distribution of AOB communities is present in natural environments.
AOB community distribution in response to depths and niches
A number of studies found that Nitrosospira dominated over Nitrosomonas in terrestrial environments (Avrahami and Conrad 2003; Avrahami et al. 2002; Kowalchuk et al. 2000; Mintie et al. 2003; Wang et al. 2009), which were consistent with this study showing a majority of Nitrosospira-like amoA sequences occurred in this wetland. Nitrosomonas were often found in high nitrogen or contaminated environments and even predominated in wastewater treatment plants (Dionisi et al. 2002; Geets et al. 2006; Wagner et al. 1995; Wang et al. 2009). A portion of Nitrosomonas-like amoA gene sequences were also detected in this study might give evidence that domestic discharge contamination from the nearby city influenced the ammonia-oxidizing oxidizers community ecology.
UniFrac environmental clustering of the bacterial amoA gene clone libraries showed clear grouping of separate AOB communities in surface and bottom sediments/soils except the samples 2S and 3B (Fig. S3). The surface samples 23, 25 and 3S were grouped together while the bottom samples 24, 26 and 2B were grouped together. In this study, more Nitrosospira-like sequences inhabited the bottom soils whereas more Nitrosomonas-like sequences were detected in upper soils (data not shown). This finding suggested a depth separation of the AOB community in the freshwater wetland. The factor of O2 availability might partly govern the AOB community distribution in soil system (Briones et al. 2002; Nicolaisen et al. 2004). Previous reports showed that oxygen concentration across soil depth gradient showed an inverse proportion (Freney et al. 1992; Lüdemann et al. 2000; Wang et al. 2009). Nitrosospira had a relatively lower O2 affinity compared to Nitrosomonas (Schramm et al. 1998) so that the former strains probably preferred partially oxic surface soils and the latter had better adaption of relatively less oxic deeper soil layer. The finding of depth separation of AOB community was also supported by Wang et al. (2009).
For AOB distribution in response to niches, niches R and R-Con were clustered together; niches D and RW formed the second and third groups respectively (Fig. S2b). Relatively high sequence similarity between niches R and R-Con suggested that these two niches had low within-group variation of AOB. That is to say, the presence of plants did not affect the AOB community composition since the niche R was grown with reeds while the niche R-Con had no plants. This finding was in agreement with previous studies that plantation had little influence on the composition of AOB community (Herrmann et al. 2009; Laanbroek and Speksnijder 2008). In contrast, the results from Jackknife Environment Clusters Analysis (Fig. S2b) supported that ammonium might be an important factor on the AOB community structure since niche RW was separated from other three niches which grouped together. Because the niche RW had a significant higher ammonium concentration than the other three niches, it is proposed that the different community composition of AOB in RW niche from other three niches was due to the effect of ammonium.
References
Adair KL, Schwartz E (2008) Evidence that ammonia-oxidizing archaea are more abundant than ammonia-oxidizing bacteria in semiarid soils of northern Arizona, USA. Microb Ecol 56:420–426. doi:10.1007/s00248-007-9360-9
Avrahami S, Conrad R (2003) Patterns of community change among ammonia oxidizers in meadow soils upon long-term incubation at different temperatures. Appl Environ Microbiol 69:6152–6164. doi:10.1128/aem.69.10.6152-6164.2003
Avrahami S, Conrad R, Braker G (2002) Effect of soil ammonium concentration on N2O release and on the community structure of ammonia oxidizers and denitrifiers. Appl Environ Microbiol 68:5685–5692. doi:10.1128/aem.68.11.5685-5692.2002
Beman JM, Francis CA (2006) Diversity of ammonia-oxidizing archaea and bacteria in the sediments of a hypernutrified subtropical estuary: Bahía del Tóbari, Mexico. Appl Environ Microbiol 72:7767–7777. doi:10.1128/aem.00946-06
Beman JM, Popp BN, Francis CA (2008) Molecular and biogeochemical evidence for ammonia oxidation by marine Crenarchaeota in the Gulf of California. ISME J 2:429–441
Briones AM, Okabe S, Umemiya Y, Ramsing N-B, Reichardt W, Okuyama H (2002) Influence of different cultivars on populations of ammonia-oxidizing bacteria in the root environment of rice. Appl Environ Microbiol 68:3067–3075. doi:10.1128/aem.68.6.3067-3075.2002
Bruns MA, Stephen JR, Kowalchuk GA, Prosser JI, Paul EA (1999) Comparative diversity of ammonia oxidizer 16S rRNA gene sequences in native, tilled, and successional soils. Appl Environ Microbiol 65:2994–3000
Cao H, Hong Y, Li M, Gu J-D (2012) Community shift of ammonia-oxidizing bacteria along an anthropogenic pollution gradient from the Pearl River Delta to the South China Sea. Appl Microbiol Biotechnol 94:247–259. doi:10.1007/s00253-011-3636-1
Christensen PB, Nielsen LP, Sørensen J, Revsbech NP (1990) Denitrification in nitrate-rich streams: diurnal and seasonal variation related to benthic oxygen metabolism. Limnol Oceanogr 35:640–651
Clément J-C, Pinay G, Marmonier P (2002) Seasonal dynamics of denitrification along topohydrosequences in three different riparian wetlands. J Environ Qual 31:1025–1037
Dang H, Zhang X, Sun J, Li T, Zhang Z, Yang G (2008) Diversity and spatial distribution of sediment ammonia-oxidizing crenarchaeota in response to estuarine and environmental gradients in the Changjiang Estuary and East China Sea. Microbiology 154:2084–2095. doi:10.1099/mic.0.2007/013581-0
Dionisi HM, Layton AC, Harms G, Gregory IR, Robinson KG, Sayler GS (2002) Quantification of Nitrosomonas oligotropha-like ammonia-oxidizing bacteria and Nitrospira spp. from full-scale wastewater treatment plants by competitive PCR. Appl Environ Microbiol 68:245–253
Dorador C, Busekow A, Vila I, Imhoff JF, Witzel K-P (2008) Molecular analysis of enrichment cultures of ammonia oxidizers from the Salar de Huasco, a high altitude saline wetland in northern Chile. Extremophiles 12:405–414
Erguder TH, Boon N, Wittebolle L, Marzorati M, Verstraete W (2009) Environmental factors shaping the ecological niches of ammonia-oxidizing archaea. FEMS Microbiol Rev 33:855–869. doi:10.1111/j.1574-6976.2009.00179.x
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Francis CA, Roberts KJ, Beman JM, Alyson ES, Oakley BB (2005) Ubiquity and diversity of ammonia-oxidizing archaea in water columns and sediments of the ocean. Proc Natl Acad Sci USA 102:14683–14688
Freney JR, Smith CJ, Mosier AR (1992) Effect of a new nitrification inhibitor (wax coated calcium carbide) on transformations and recovery of fertilizer nitrogen by irrigated wheat. Fertil Res 32:1–11
Geets J, Boon N, Verstraete W (2006) Strategies of aerobic ammonia-oxidizing bacteria for coping with nutrient and oxygen fluctuations. FEMS Microbiol Ecol 58:1–13
Herrmann M, Saunders AM, Schramm A (2008) Archaea dominate the ammonia-oxidizing community in the rhizosphere of the freshwater macrophyte Littorella uniflora. Appl Environ Microbiol 74:3279–3283. doi:10.1128/aem.02802-07
Herrmann M, Saunders AM, Schramm A (2009) Effect of lake trophic status and rooted macrophytes on community composition and abundance of ammonia-oxidizing prokaryotes in freshwater sediments. Appl Environ Microbiol 75:3127–3136. doi:10.1128/aem.02806-08
Hollibaugh JT, Bano N, Ducklow HW (2002) Widespread distribution in polar oceans of a 16S rRNA gene sequence with affinity to Nitrosospira-like ammonia-oxidizing bacteria. Appl Environ Microbiol 68:1478–1484
Hou P-F (2009) Status of Longfeng wetland and its protection recommendations. Heilongjiang Agr Sci 1:53–59
Ibekwe AM, Grieve CM, Lyon SR (2003) Characterization of microbial communities and composition in constructed dairy wetland wastewater effluent. Appl Environ Microbiol 69:5060–5069
Jiang H, Dong H, Yu B, Lv G, Deng S, Berzins N, Dai M (2009) Diversity and abundance of ammonia-oxidizing archaea and bacteria in Qinghai Lake, Northwestern China. Geomicrobiol J 26:199–211
Kim O-S, Junier P, Imhoff JF, Witzel K-P (2008) Comparative analysis of ammonia monooxygenase (amoA) genes in the water column and sediment-water interface of two lakes and the Baltic Sea. FEMS Microbiol Ecol 66:367–378
Könneke M, Bernhard AE, Torre JRdl, Walker CB, Waterbury JB, Stahl DA (2005) Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437:543–546
Koops H-P, Pommerening-Röser A (2001) Distribution and ecophysiology of the nitrifying bacteria emphasizing cultured species. FEMS Microbiol Ecol 37:1–9
Kowalchuk GA, Stephen JR (2001) Ammonia-oxidizing bacteria: a model for molecular microbial ecology. Annu Rev Microbiol 55:485–529. doi:10.1146/annurev.micro.55.1.485
Kowalchuk GA, Stienstra AW, Heilig GHJ, Stephen JR, Woldendorp JW (2000) Molecular analysis of ammonia-oxidising bacteria in soil of successional grasslands of the Drentsche A (The Netherlands). FEMS Microbiol Ecol 31:207–215
Laanbroek HJ, Speksnijder AGCL (2008) Niche separation of ammonia-oxidizing bacteria across a tidal freshwater marsh. Environ Microbiol 10:3017–3025
Lozupone CA, Hamady M, Kelley ST, Knight R (2007) Quantitative and qualitative β diversity measures lead to different insights into factors that structure microbial communities. Appl Environ Microbiol 73:1576–1585
Lüdemann H, Arth I, Liesack W (2000) Spatial changes in the bacterial community structure along a vertical oxygen gradient in flooded paddy soil cores. Appl Environ Microbiol 66:754–762
Mintie A, Heichen R, Cromack K Jr, Myrold D, Bottomley P (2003) Ammonia-oxidizing bacteria along meadow-to-forest transects in the Oregon Cascade Mountains. Appl Environ Microbiol 69:3129–3136
Moin NS, Nelson KA, Bush A, Bernhard AE (2009) Distribution and diversity of archaeal and bacterial ammonia oxidizers in salt marsh sediments. Appl Environ Microbiol 75:7461–7468
Mosier AC, Francis CA (2008) Relative abundance and diversity of ammonia-oxidizing archaea and bacteria in the San Francisco Bay estuary. Environ Microbiol 10:3002–3016
Mullins TD, Britschgi TB, Krest RL, Giovannoni SJ (1995) Genetic comparisons reveal the same unknown bacterial lineages in Atlantic and Pacific bacterioplankton communities. Limnol Oceanogr 40:148–158
Nicolaisen MH, Risgaard-Petersen N, Revsbech NP, Reichardt W, Ramsing NB (2004) Nitrification-denitrification dynamics and community structure of ammonia oxidizing bacteria in a high yield irrigated Philippine rice field. FEMS Microbiol Ecol 49:359–369
Norton JM, Alzerreca JJ, Suwa Y, Klotz MG (2002) Diversity of ammonia monooxygenase operon in autotrophic ammonia-oxidizing bacteria. Arch Microbiol 177:139–149
Ottosen LDM, Risgaard-Petersen N, Nielsen LP (1999) Direct and indirect measurements of nitrification and denitrification in the rhizosphere of aquatic macrophytes. Aquat Microb Ecol 19:81–91
Park H-D, Wells GF, Bae H, Criddle CS, Francis CA (2006) Occurrence of ammonia-oxidizing archaea in wastewater treatment plant bioreactors. Appl Environ Microbiol 72:5643–5647. doi:10.1128/aem.00402-06
Pett-Ridge J, Petersen DG, Nuccio E, Firestone MK (2013) Influence of oxic/anoxic fluctuations on ammonia oxidizers and nitrification potential in a wet tropical soil. FEMS Microbiol Ecol 85:179–194. doi:10.1111/1574-6941.12111
Pitcher A, Villanueva L, Hopmans EC, Schouten S, Reichart GJ, Sinninghe Damsté JS (2011) Niche segregation of ammonia-oxidizing archaea and anammox bacteria in the Arabian Sea oxygen minimum zone. ISME J 5:1896–1904
Pommerening-Röser A, Rath G, Koops HP (1996) Phylogenetic diversity within the genus Nitrosomonas. Syst Appl Microbiol 19:344–351
Prosser JI, Nicol GW (2008) Relative contributions of archaea and bacteria to aerobic ammonia oxidation in the environment. Environ Microbiol 10:2931–2941
Purkhold U, Pommerening-Röser A, Juretschko S, Schmid MC, Koops H-P, Wagner M (2000) Phylogeny of all recognized species of ammonia oxidizers based on comparative 16S rRNA and amoA sequence analysis: implications for molecular diversity surveys. Appl Environ Microbiol 66:5368–5382. doi:10.1128/aem.66.12.5368-5382.2000
Purkhold U, Wagner M, Timmermann G, Pommerening-Röser A, Koops H-P (2003) 16S rRNA and amoA-based phylogeny of 12 novel betaproteobacterial ammonia-oxidizing isolates: extension of the dataset and proposal of a new lineage within the nitrosomonads. Int J Syst Evol Microbiol 53:1485–1494
Reigstad LJ, Richter A, Daims H, Urich T, Schwark L, Schleper C (2008) Nitrification in terrestrial hot springs of Iceland and Kamchatka. FEMS Microbiol Ecol 64:167–174
Riis T, Sand-Jensen K (1998) Development of vegetation and environmental conditions in an oligotrophic Danish lake over 40 years. Freshwat Biol 40:123–134
Ruiz-Rueda O, Hallin S, Baneras L (2009) Structure and function of denitrifying and nitrifying bacterial communities in relation to the plant species in a constructed wetland. FEMS Microbiol Ecol 67:308–319
Santoro AE, Casciotti KL (2011) Enrichment and characterization of ammonia-oxidizing archaea from the open ocean: phylogeny, physiology and stable isotope fractionation. ISME J 5:1796–1808. doi:10.1038/ismej.2011.58
Schloss PD, Handelsman J (2005) Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71:1501–1506. doi:10.1128/aem.71.3.1501-1506.2005
Schloss PD et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Schramm A, de Beer D, Wagner M, Amann R (1998) Identification and activities in situ of Nitrosospiraand nitrospira spp. as dominant populations in a nitrifying fluidized bed reactor. Appl Environ Microbiol 64:3480–3485
Stehr G, Böttcher B, Dittberner P, Rath G, Koops H-P (1995) The ammonia-oxidizing nitrifying population of the River Elbe estuary. FEMS Microbiol Ecol 17:177–186
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. doi:10.1093/molbev/msr121
Taylor AE, Zeglin LH, Wanzek TA, Myrold DD, Bottomley PJ (2012) Dynamics of ammonia-oxidizing archaea and bacteria populations and contributions to soil nitrification potentials. ISME J 6:2024–2032
Tourna M et al (2011) Nitrososphaera viennensis, an ammonia oxidizing archaeon from soil. Proc Natl Acad Sci USA 108:8420–8425. doi:10.1073/pnas.1013488108
Treusch AH, Leininger S, Kletzin A, Schuster SC, Klenk HP, Schleper C (2005) Novel genes for nitrite reductase and Amo-related proteins indicate a role of uncultivated mesophilic crenarchaeota in nitrogen cycling. Environ Microbiol 7:1985–1995
Venter JC et al (2004) Environmental genome shotgun sequencing of the Sargasso Sea. Science 304:66–74. doi:10.1126/science.1093857
Verhoeven JTA, Meuleman AFM (1999) Wetlands for wastewater treatment: opportunities and limitations. Ecol Eng 12:5–12. doi:10.1016/s0925-8574(98)00050-0
Wagner M, Rath G, Amann R, Koops H-P, Schleifer K-H (1995) In situ identification of ammonia-oxidizing bacteria. Syst Appl Microbiol 18:251–264
Walker CB et al (2010) Nitrosopumilus maritimus genome reveals unique mechanisms for nitrification and autotrophy in globally distributed marine crenarchaea. Proc Natl Acad Sci USA 107:8818–8823. doi:10.1073/pnas.0913533107
Wang Y-F, Gu J-D (2013) Higher diversity of ammonia/ammonium-oxidizing prokaryotes in constructed freshwater wetland than natural coastal marine wetland. Appl Microbiol Biotechnol 97:7015–7033. doi:10.1007/s00253-012-4430-4
Wang Y-F, Gu J-D (2014) Effects of allylthiourea, salinity and pH on ammonia/ammonium-oxidizing prokaryotes in mangrove sediment incubated in laboratory microcosms. Appl Microbiol Biotechnol 98:3257–3274. doi:10.1007/s00253-013-5399-3
Wang Y, Ke X, Wu L, Lu Y (2009) Community composition of ammonia-oxidizing bacteria and archaea in rice field soil as affected by nitrogen fertilization. Syst Appl Microbiol 32:27–36
Wang J, Dong H, Wang W, Gu J-D (2013a) Reverse-transcriptional gene expression of anammox and ammonia-oxidizing archaea and bacteria in soybean and rice paddy soils of Northeast China. Appl Microbiol Biotechnol. doi:10.1007/s00253-013-5242-x
Wang J, Wang W, Gu J-D (2013b) Community structure and abundance of ammonia-oxidizing archaea and bacteria after conversion from soybean to rice paddy in albic soils of Northeast China. Appl Microbiol Biotechnol. doi:10.1007/s00253-013-5213-2
Wang Y-F, Feng Y-Y, Ma X, Gu J-D (2013c) Seasonal dynamics of ammonia/ammonium-oxidizing prokaryotes in oxic and anoxic wetland sediments of subtropical coastal mangrove. Appl Microbiol Biotechnol 97:7919–7934. doi:10.1007/s00253-012-4510-5
Wang Y-F, Li X-Y, Gu J-D (2014a) Differential responses of ammonia/ammonium-oxidizing prokaryotes in mangrove sediment to amendment of acetate and leaf litter. Appl Microbiol Biotechnol 98:3165–3180. doi:10.1007/s00253-013-5318-7
Wang Y, Zhu G, Song L, Wang S, Yin C (2014b) Manure fertilization alters the population of ammonia-oxidizing bacteria rather than ammonia-oxidizing archaea in a paddy soil. J Basic Microbiol 54:190–197. doi:10.1002/jobm.201200671
Wells GF, Park H-D, Yeung C-H, Eggleston B, Francis CA, Criddle CS (2009) Ammonia-oxidizing communities in a highly aerated full-scale activated sludge bioreactor: betaproteobacterial dynamics and low relative abundance of Crenarchaea. Environ Microbiol 11:2310–2328
Winogradsky S (1890) Investigations on nitrifying organisms. Ann Inst Pasteur 4:213–321
Wuchter C et al (2006) Archaeal nitrification in the ocean. Proc Natl Acad Sci USA 103:12317–12322
Xu C, Zhang G (2009) Biodiversities of Daqing wetland and countermeasures for protection. Chinese Agr Sci Bull 25:215–219
You J, Das A, Dolan EM, Hu Z (2009) Ammonia-oxidizing archaea involved in nitrogen removal. Water Res 43:1801–1809
Zhang L-M, Hu H-W, Shen J-P, He J-Z (2012) Ammonia-oxidizing archaea have more important role than ammonia-oxidizing bacteria in ammonia oxidation of strongly acidic soils. ISME J 6:1032–1045
Zhou Z, Shi X, Zheng Y, Qin Z, Xie D, Li Z, Guo T (2014) Abundance and community structure of ammonia-oxidizing bacteria and archaea in purple soil under long-term fertilization. Europ J Soil Biol 60:24–33
Zuber G, Goldsmith MR, Beratan DN, Wipf P (2005) Assignment of the absolute configuration of [n]-ladderanes by TD-DFT optical rotation calculations. Chirality 17:507–510
Acknowledgments
This research was supported by a studentship from the Graduate School of The University of Hong Kong (KHL), additional financial support of this project was from Environmental Toxicology Education and Research Fund of this laboratory, and National Natural Science Foundation of China (41003031, 41273109, 51378208) and Shanghai Rising-Star Program (12QA1400800), and Fok Ying Tung Education Foundation (141077) (HL). We would like to thank Ms. Jessie Lai and Kelly Lau for support in chemical analysis and Dr. Jing Wang and Mr. Hongshuang Jiao for assistance in field sampling.
Conflict of interest
The authors have no conflict of interest in research results report here.
Author information
Authors and Affiliations
Corresponding author
Additional information
Kwok-Ho Lee and Yong-Feng Wang were contributed to this work equally.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, KH., Wang, YF., Li, H. et al. Niche specificity of ammonia-oxidizing archaeal and bacterial communities in a freshwater wetland receiving municipal wastewater in Daqing, Northeast China. Ecotoxicology 23, 2081–2091 (2014). https://doi.org/10.1007/s10646-014-1334-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10646-014-1334-3