Abstract
Background
Discoidin domain receptors1 (DDR1) is associated with tumor progression, and its dysregulated expression has been observed in many cancers.
Aim
We aim to explore molecular mechanism underlying the role of DDR1 in colorectal cancer development.
Methods
Immunohistochemistry and Western blot were applied to examine the DDR1 expression. Real-time RT-PCR and Western blot were performed to determine the expression of miR-199a-5p and DDR1. Luciferase reporter assay was used to determine whether DDR1 was a target of miR-199a-5p. Effects of miR-199a-5p and DDR1 on colorectal cell proliferation, colony formation, cell cycle progression, invasion and migration were then investigated. Western blot was used to determine the relative signal pathways.
Results
Increased DDR1 and decreased miR-199a-5p expression coexisted in CRC, knockdown of DDR1 or overexpression of miR-199a-5p both resulted in reduced colony formation, invasive and migratory capabilities of human CRC LOVE1 and LOVO cells. It was also found that overexpression of miR-199a-5p led to decreased DDR1, MMP2, N-cadherin and vimentin expression and increased E-cadherin expression in both CRC cell lines. However, down-regulation of miR-199a-5p resulted in the opposite effects. Dual luciferase reporter assay confirmed that miR-199a-5p could directly target DDR1 through binding to its 3′-UTR.
Conclusions
Our findings indicated that up-regulation of DDR1 induced by miR-199a-5p down-regulation may contribute to the development and progression of CRC, and this effect may be associated with increased invasiveness, at least in part, via activating the EMT-related signaling.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer (CRC), one of the most common malignances, is an aggressive cancer associated with high morbidity and mortality (1.2 million new cases and over 600,000 death every year), especially in the case of advanced disease [1]. Although the risk factors of CRC are well characterized, the molecular pathogenesis of this particular tumor type is still not pretty understood. As more and more lives of the patients are taken by CRC, it is urgent to clarify the molecular mechanisms of this cancer and to develop novel and more effective treatment strategies against the malignance.
Receptor tyrosine kinases (RTKs) play an important role in the signal transduction pathways that control cell proliferation and differentiation and are involved in tumorigenesis [2]. The discoidin domain receptor1 (DDR1) belongs to a novel class of receptor tyrosine kinases with a characteristic discoidin homology domain, stalk region, transmembrane region, juxtamembrane region and kinase domain [3]. Recently, overexpression of DDR1 was reported in several human cancers, such as breast [4], liver [5], lung [6] and ovary [7]. The precise mechanism(s) by which these receptors may contribute to oncogenesis are not yet known. However, overexpression of DDR1 has been shown to increase the migration and invasion of hepatoma cells [5, 8], implicating a causal role of DDR1 in promoting tumor progression. Unfortunately, the expression and role of DDR1 in CRC remain unclear.
MicroRNAs (miRNAs) represent an abundant class of small, non-coding, single-stranded RNA molecules with 20–25 nucleotides in length [9] capable of mediating a vast gene regulatory network [10]. MiRNAs can regulate gene expression by direct cleavage of targeted messenger RNAs (mRNAs) or by inhibiting translation through complementarity to targeted mRNAs at the 3′untranslated regions (UTRs) [11]. Many studies have showed that microRNAs (miRNAs) can regulate the expression of various genes closely associated with tumor proliferation, invasion, metastasis and prognosis in CRC pathogenesis [12, 13]. It was reported that miR-199a-5p functioned in hepatocellular carcinoma (HCC). DDR1 may be a direct target gene of miR-199a-5p predicted by publicly available PicTar (4-way), TargetScanS and miRanda algorithms. However, whether the targeted relationship is established in CRC or the consequent functional influence is still unknown.
Therefore, in this study, we determined the expression levels of DDR1 in CRC tissues and cells. Subsequently, the roles of DDR1 and miR-199a-5p in migration and invasion of LOVE1 and LOVO cells were investigated. It is confirmed that DDR1 up-regulation induced by miR-199a-5p down-regulation contributes to malignant progression of CRC. Our results may support DDR1 and miR-199a-5p as novel diagnostic or therapeutic target for CRC.
Materials and Methods
Cell Culture
Colorectal cancer cell lines LOVE1, LOVO and human colonic epithelial cell line (HCEC) were purchased from the China Center for Type Culture Collection (CCTCC, Wuhan, China). All cell lines were cultured in DMEM supplemented with 10 % fetal bovine serum (FBS), 100 IU/ml penicillin and 100 μg/ml streptomycin sulfate at 37 °C in a humidified incubator containing 5 % CO2.
Tumor Tissue Samples Preparation
A total of 10 patients diagnosed with CRC in The Affiliated Tumor Hospital of Xiangya Medical School of Central South University were recruited. The experimental protocols were approved by the ethics committee of the hospital. RNA or tissue samples prepared from the tumor tissues and their matched adjacent non-tumor tissues (Normal) were then subjected to quantitative real-time RT-PCR or immunohistochemistry analysis.
Immunohistochemistry
Tissue blocks were incubated with anti-DDR1 (Millipore, USA) or normal rabbit IgG as negative control. Immunostaining was performed using the Moticcam3000 System with diaminobenzidine (Zhongshan Jinqiao, China). Imagepro-Plus software was used to quantify the mean density of DDR1 staining, according to the manufacture’s instruction.
Lentivirus Package and Infection
The lentivirus system (Neuron, Shanghai, China) consisted of the pLKD-CMVG & NR-U6-shRNA vector together with the plasmids containing the imperative elements for virus packaging. Short hairpin RNA (shRNA) targeting DDR1 (5′-GGGAUGGACUCCUGUCUUA-3′) and the scrambled control (5′-GCGCGCTTTGTAGGATTCG-3′) were coloned into the vector, respectively, which were then cotransfected with packaging plasmids into 293T cells to generate individual lentiviruses. After 48 h, the DDR1-RNAi-lentivirus and control lentivirus were harvested and purified. The lentivirus expressing DDR1 shRNA and the control lentivirus were termed Lv-DDR1-shRNA and Lv-Con, respectively. The titer of Lv-DDR1-shRNA was 2.17 × 108 TU/ml, and Lv-Con was 1.07 × 109 TU/ml. 1 × 105/well LOVE1 or LOVO cells were plated in 6-well plates to reach 40–50 % confluency, then, 93 μl Lv-DDR1-shRNA or 20 μl Lv-Con were added to the cells at MOI values of 200 and incubated at 37 °C for 48 h.
Transfection
For the miR-199a-5p functional analysis, cells were transfected with the pre-miR-199a-5p, pre-con, anti-199a-5p or anti-con (Genecopoeia, China) were transfected into LOVE1 and LOVO cell lines using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s recommendations.
Real-Time RT-PCR
Total RNA was extracted from cells with Trizol reagent (Invitrogen) following the manufacturer’s instructions. The relative expression level of miR-199a-5p was determined by quantitative real-time RT-PCR using mirVana™ qRT-PCR microRNA detection kit (Ambion) following the manufacturer’s instructions. Specific primer sets for miR-199a-5p and U6 (used as an internal reference) were obtained from Genecopoeia. Expression of DDR1 mRNA was detected by real-time RT-PCR using the standard SYBR Green RT-PCR Kit (Takara, Otsu, Japan) following the manufacture’s instructions. The specific primer pairs are as follows: DDR1 sense, 5′-CTTCAGCGAAATCTCCTTCATC -3′ and antisense,5′- CCAACACCCTCCGTTCAGCCT -3′; β-actin as an internal control, sense, 5′-AGGGGCCGGACTCGTCATACT-3′ and antisense, 5′-GGCGGCACCACCATGTACCCT-3′. The relative expression of DDR1 mRNA or miR-199a-5p was quantified using the GraphPad Prism 4.0 software (GraphPad Software, San Diego, CA, USA) and 2−ΔΔCt method [14].
Cell Proliferation Assay
Cells in exponential growth were plated at a final concentration of 2 × 103 cells/well in 96-well plates. The viability of the cells was evaluated by MTT assay after 24, 48 and 72 h of seeding. The optical density at 570 nm (OD570) of each well was measured with an ELISA reader (ELX-800 type, Bio-Tek).
Colony Formation Assay
For all groups, 3 ml complete medium containing 150 cells were added to each well of a six-well plate. Plates were incubated at 37 °C, 5 % CO2 for 14 days. After that, cells were gently washed and stained with Giemsa. Colonies containing at least 50 cells were counted.
Cell Cycle Assay
For all groups, 106 cells were collected in 1 × PBS and resuspended in 70 % ethanol to fix overnight at −20 °C. Cells were pelleted at 1,000 rpm for 5 min, washed in 1 × PBS, and then pelleted at1,000 rpm for 5 min. Cells were resuspended in 300 μl propidium iodide staining buffer and incubated for 30 min at room temperature. DNA content analyses were carried out using a flow cytometry (FACSCalibur, Beckman Coulter).
Cell Invasion Assay
The invasive ability of colorectal cancer cells was then studied in 24-well transwell chambers (Chemicon,USA) that has a layer of matrigel. For all groups, 200 μl of cell suspension (1 × 106 cells/ml) was added in triplicate wells. After 24-h incubation, the dye on the membrane was dissolved with 10 % acetic acid, dispensed into 96-well plates (150 μl/well) and the optical density at 570 nm (OD570) of each well was measured with an ELISA reader (ELX-800 type; BioTek).
Cell Migration Assay
Cell migratory capability was estimated using a wound healing assay as described previously [15]. In brief, cells were cultured to confluence. Wounds of approximately 1 mm width were created with a plastic scriber, and cells were washed and incubated in a serum-free medium. After wounding for 24 h, cells were incubated in a medium including 10 % fetal bovine serum. Cultures at 0, 24 and 48 h were fixed and observed under a microscope.
Dual Luciferase Reporter Assay
The 3′-UTR of DDR1 (NM_013993) containing the miR-199a-5p binding sites and its corresponding mutated sequence were cloned into the psi-CHECK2 luciferase reporter vector (Promega) downstream of Renilla luciferase, named DDR1-3′-UTR and DDR1-Mut 3′-UTR, respectively. Using lipofectamine 2000 (Invitrogen), LOVE1 and LOVO cells were co-transfected with the reporter constructs and miR-199a-5p mimics, the miR-199a-5p inhibitor, negative control (NC) or the negative control inhibitor. Luciferase activity was determined after 48 h using the Dual-Glo substrate system (Promega) and a Beckman Coulter LD400 luminometer. Data are presented as the ratio of experimental (Renilla) luciferase to control (Firefly) luciferase.
Western Blotting
Tissues or cells were solubilized in cold RIPA lysis buffer. After that, proteins were separated with 10 % SDS-PAGE and then transferred to a PVDF membrane. Membranes were blocked in 10 % non-fat dried milk in PBST for 3 h and then incubated overnight with specific primary antibodies (Santa Cruz, USA) with β-actin as a control. After incubation with the corresponding secondary antibody, immune complexes were detected using an ECL kit (Biyuntian, China).
Statistical Analysis
Data are expressed as the mean ± SD from at least three separate experiments. Statistical analysis was carried out using SPSS 15.0 software. The difference between two groups was analyzed by the Student’s t test. A value of P < 0.05 was considered to indicate a statistically significant result.
Results
Expression of DDR1 and miR-199a-5p in CRC
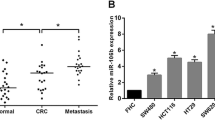
The mRNA levels of miR-199a-5p and DDR1 in clinical CRC tissues or in LOVE1 and LOVO tumor cell lines were determined using quantitative RT-PCR (qRT-PCR). Compared to normal adjacent tissues, the mRNA levels of DDR1 in tumor tissues were significantly higher than normal tissues, while the mRNA level of miR-199a-5p was drawn opposite conclusion (Fig. 1a, b). Meanwhile, the DDR1 protein expression was apparently increased in tumor tissues than in normal tissue when measured with immunohistochemistry (Fig. 1c). These results suggest that DDR1 may play an important role in malignant progression of CRC. Furthermore, the coexistence of DDR1 up-regulation and miR-199a-5p down-regulation in CRC cells implies a potential regulatory correlation between DDR1 and miR-199a-5p.
Expression of DDR1 and miR-199a-5p in CRC. a Relative expression of DDR1 mRNA and miR-199a-5p in human CRC tissue compared to matched adjacent normal colon tissue (Normal), using qRT-PCR. b Relative expression of DDR1 mRNA and miR-199a-5p in human CRC LOVO and LOVE1 cells compared to control (Con) HCEC cells, using qRT-PCR. c Shows the representative results of DDR1 expression (brown particles) in human CRC tissue compared to normal, by immunohistochemical analysis. *P < 0.05 VS Con, in LOVO cells; # P < 0.05 VS Con, in LOVE1 cells
DDR1 Promoting Proliferation, Invasion, and Migration of CRC Cells
To investigate the functions of DDR1 in CRC tumor cells, LOVO and LOVE1cells were transfected with DDR1-shRNA. The mRNA level of DDR1 in both cell lines were significantly decreased (Fig. 2a), indicating that DDR1 was successfully silenced in LOVO and LOVE1 cells. Moreover, it was also found that knockdown of DDR1 reduced cell viablility (Fig. 2b, c) and colony formation (Fig. 2d) in CRC cells. Besides, the reduced DDR1 expression enhanced the cell proportion in G1 phase but reduced the cell proportion in S phase (Fig. 2e, f).That means DDR1 can promote the cell cycle G1/S transition. It suggests that DDR1 promoting proliferation of CRC cells is related to its increasing G1/S transition.
DDR1 promoting proliferation of CRC cells. a DDR1-shRNA significantly decreased DDR1 mRNA expression of DDR1 in LOVO and LOVE1 cells detected by qRT-PCR. Decreased cell viability (b, c) and colony formation (d) in LOVO and LOVE1 cells transfected with DDR1-shRNA. Increased G1 phase population and decreased S phase population in LOVO (e) and LOVE1 (f) cells transfected with DDR1-shRNA. *P < 0.05 VS Control (Con), in LOVO cells; # P < 0.05 VS Control (Con), in LOVE1 cells
Then, it was demonstrated that the invasive and migratory capabilities of LOVO and LOVE1 cells transfected with DDR1-shRNA were decreaced (Fig. 3), suggesting that DDR1 promoting invasion and migration of CRC cells. These results imply that DDR1 plays a promotional role in CRC growth and metastatic progression.
miR-199a-5p Suppressing Proliferation, Invasion, and Migration of CRC Cells
To explore the functional role of miR-199a-5p in CRC, LOVO and LOVE1 cells were transfected with pre-miR-199a-5p. The qRT-PCR result shows that induction of pre-miR-199a-5p significantly increased miR-199a-5p expression in LOVO and LOVE1 cells (Fig. 4a). It was then confirmed that overexpression of miR-199a-5p reduced cell viability (Fig. 4b, c) and colony formation (Fig. 4d) in CRC cells. In addition, the overexpression of miR-199a-5p enhanced the cell proportion in G1 phase but reduced the cell proportion in S phase (Fig. 4e, f). This means miR-199a-5p can inhibit the cell cycle G1/S transition. It suggests that miR-199a-5p suppressing proliferation of CRC cells involves its decreasing G1/S transition.
miR-199a-5p suppressing proliferation of CRC cells. a Pre-miR-199a-5p significantly increased miR-199a-5p expression in LOVO and LOVE1 cells detected by qRT-PCR. Decreased cell viability (b, c) and colony formation (d) in LOVO and LOVE1 cells transfected with pre-miR-199a-5p. Increased G1 phase population and decreased S phase population in LOVO (e) and LOVE1 (f) cells transfected with pre-miR-199a-5p. *P < 0.05 VS Control (Con), in LOVO cells; # P < 0.05 VS Control (Con), in LOVE1 cells
It was then demonstrated that the invasive and migratory capabilities of LOVO and LOVE1 cells transfected with pre-miR-199a-5p were reduced (Fig. 5), indicating that miR-199a-5p inhibiting invasion and migration of CRC cells. These results suggest that miR-199a-5p plays an inhibitor role in CRC growth and metastatic progression, which was completely opposite with DDR1.
miR-199a-5p suppressing invasion and migration of CRC cells. The invasion (a) and migration (b) decreased in LOVO and LOVE1 cells transfected with pre-miR-199a-5p, using transwell assay and wound healing assay, respectively. *P < 0.05 VS Control (Con), in LOVO cells; # P < 0.05 VS Control (Con), in LOVE1 cells
DDR1 Is a Direct Target of miR-199a-5p
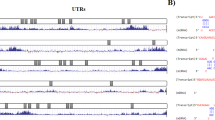
It found that overexpression of miR-199a-5p could reduce the DDR1 expression in LOVO and LOVE1 (Fig. 6a) and miR-199a-5p played completely opposite functional role with DDR1 in both cell lines. To assess, whether DDR1 is a direct target of miR-199a-5p, luciferase reporter assays were applied. We subcloned the DDR1 3′-UTR fragment containing the miR-199-5p binding site and mutated targeting sequence and cloned them into psi-CHECK2 dual luciferase reporter vectors. The miR-199a-5p significantly inhibited the luciferase activity in both LOVO and LOVE1 cells transfected with the DDR1-3′-UTR. However, miR-199a-5p mimics did not suppress the luciferase activity levels in the LOVO and LOVE1 cells transfected with Mut-DDR1-3′-UTR. This confirms that DDR1 is a direct target of miR-199a-5p.
miR-199a-5p targeting DDR1 and its effects on MMP2, N-cadherin, Vimentin and E-cadherin expression. a miR-199a-5p significantly decreased DDR1 mRNA expression in LOVO and LOVE1 cells compared to control (Con), detected by qRT-PCR. Dual luciferase reporter assays were performed to examine the interaction of miR-199a-5p and its targeting sequence in the DDR1 3′-UTR using constructs containing the targeting sequence (DDR1 3′-UTR) and mutated targeting sequence (Mut-DDR1-3′-UTR) in LOVO (b) and LOVE1 (c) cells. Effects of pre-miR-199a-5p and anti-miR-199a-5p on the protein expressions of DDR1, MMP2, N-cadherin Vimentin and E-cadherin in LOVO (d) and LOVE1 (e) cells using western blot analysis. The numbers listed under the lanes represent the relative quantification of each band.*P < 0.05 VS Control (Con), in LOVO cells; # P < 0.05 VS Control (Con), in LOVE1 cells
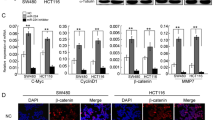
miR-199a-5p/DDR1 Regulation of MMP2, N-Cadherin, Vimentin, and E-Cadherin Expressions
To characterize the molecular mechanism underlying the role of miR-199a-5p/DDR1 in CRC, the protein levels of DDR1, MMP2, N-cadherin, Vimentin and E-cadherin in LOVO and LOVE1 cells transfected with pre-miR-199a-5p or anti-miR-199a-5p were determined using Western blot analysis. The results showed that overexpression of miR-199a-5p significantly decreased DDR1, MMP2, N-cadherin and Vimentin expressions but increased the expression level of E-cadherin in both LOVO and LOVE1 cells. On the contrary, enhanced DDR1 expression mediated by anti-miR-199a-5p could activate MMP-2, N-cadherin and vimentin expression, but reduce E-cadherin expression (Fig. 5d, e). These results indicated that up-regulation of DDR1 induced by reduced miR-199a-5p may contribute to the development and progression of CRC, and this effect may be associated with increased invasiveness, at least in part, via promoting epithelial-to-mesenchymal transition (EMT).
Discussion
DDR1 is a receptor tyrosine kinase that is identified during the search for tyrosine kinase proteins expressed in human malignancies [3, 4]. DDR1, which contains a homology domain to discoidin, is distinct from other members of the large receptor tyrosine kinase and is found to be associated with cell attachment, migration and invasion [16]. Recently, high expression of DDR1 is found in a variety of invasive human cancers, and it is clear that DDR1 is primarily expressed in epithelial cells including lung, liver and prostate [5, 17–20]. For instance, up-regulated DDR1 expression promotes cancer development by enhancing cancer cell survival and invasion, and high DDR1 expression is associated with short hormone resistance interval in prostate carcinoma [20]. Furthermore, high DDR1 expression in non-small cell lung carcinomas implies poor prognosis in terms of disease-free survival [21].
In this study, we found that DDR1 expression was higher in CRC tissues than in normal colon tissue for the first time. These suggested that enhanced DDR1 expression may contribute to CRC progression. It has been confirmed that dysregulation of tumor-associated gene is partly due to dysregulation of its regulatory miRs [22–24]. Itis found that miR-199a-5p can inhibit the expression of DDR1 in HCC [5]. Here, our data showed that miR-199a-5p could significantly suppress DDR1 expression in CRC cells. Several reports support that miR-199a-5p could directly target many cancer associated gens, such as GRP78 in prostate cancers [25], clathrin heavy chain (CHC) in B virus (HBV)-associated HCC [26] and autophagy-associated gene 7 (ATG7) in HCC [27] and may have potential in cancer treatment. These results point out the significant influence of miR-199a-5p on the outcome of some malignances.
In this study, we demonstrated that up-regulated miR-199a-5p could suppress DDR1 expression and result in decreasing migration and invasion of CRC cells when compared with control. Furthermore, we found that up-regulated miR-199a-5p could also inhibit expression of MMP2, N-cadherin and vimentin and induce E-cadherin expression. On the other hand, down-regulated miR-199a-5p leads to decreased MMP2, N-cadherin and vimentin expression and increased E-cadherin expression. Researches indicated that the decline of E-cadherin expression or function and increased expression of other cadherins such as N-cadherin and vimentin is necessary for the increased cell motility that accompanies EMT [19, 28, 29].
EMT is a central differentiation process allowing the remodeling of tissues during early embryogenic and is associated with the promotion of tumor invasion and metastasis [30, 31]. It can be triggered by external signals, such as transforming growth factor (TGF)-b, hepatocyte growth factor (HGF), epidermal growth factor (EGF) and fibroblast growth factor (FGF), originating from outside the cell [32, 33]. In addition, numerous researches indicated that EMT process would be a potent mechanism that enhances the detachment of cancer cells from primary tumors [19]. One interesting feature of cells that underwent EMT is the loss of E-cadherin expression, and decreased E-cadherin expression has been reported to be associated with poor clinical outcome in different kinds of human cancers [19, 34–37]. Therefore, EMT signaling pathway may be good potential candidate for cancer treatment and it is important to understand the molecular mechanisms that drive EMT for metastasis prevention of cancer. According to our data, down-regulation of miR-199a-5p activated DDR1 overexpression and accompanied with E-cadherin loss. These might indicate EMT enhancement in CRC cell. However, inhibiting the DDR1 expression by up-regulation of miR-199a-5p with the phenomena of E-cadherin overexpression might suggest EMT process is prevented in CRC cells. Therefore, we propose that DDR1 may be a signal which could trigger EMT in CRC and finally contribute to the development and progression of CRC, while miR-199a-5p could inactivate DDR1 signaling, turn off EMT signaling in CRC and prevent tumor invasion and metastasis.
In conclusion, this study provides the evidence demonstrating that DDR1 is significantly up-regulated and miR-199a-5p is significantly down-regulated in CRC tissues and cells. And miR-199a-5p directly targets DDR1 in CRC cells. Inhibition of DDR1 by overexpression of miR-199a-5p resulted in decreased cell proliferation, cell cycle S phase population, migration and invasiveness in CRC cells. Moreover, our results also indicate that miR-199a-5p could inactivate DDR1 resulting in inhibiting EMT signaling pathway in CRC cells. Thus, we suggest that miR-199a-5p loss up-regulating DDR1 expression contributes to progression of CRC possibly by promoting EMT process.
Our results support DDR1 and miR-199a-5p as novel diagnostic or therapeutic target for CRC. Although this study provides a possible molecular mechanism for miR-199a-5p mediated DDR1 signaling pathway affecting CRC progression, further studies are still needed to elucidate the details of this process.
References
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011;61:69–90.
Longati P, Comoglio PM, Bardelli A. Receptor tyrosine kinases as therapeutic targets: the model of the MET oncogene. Curr Drug Targets. 2001;2:41–55.
Alves F, Vogel W, Mossie K, Millauer B, Hofler H, Ullrich A. Distinct structural characteristics of discoidin I subfamily receptor tyrosine kinases and complementary expression in human cancer. Oncogene. 1995;10:609–618.
Johnson JD, Edman JC, Rutter WJ. A receptor tyrosine kinase found in breast carcinoma cells has an extracellular discoidin I-like domain. Proc Natl Acad Sci USA. 1993;90:5677–5681.
Shen Q, Cicinnati VR, Zhang X, et al. Role of microRNA-199a-5p and discoidin domain receptor 1 in human hepatocellular carcinoma invasion. Mol Cancer. 2010;9:227.
Ford CE, Lau SK, Zhu CQ, Andersson T, Tsao MS, Vogel WF. Expression and mutation analysis of the discoidin domain receptors 1 and 2 in non-small cell lung carcinoma. Br J Cancer. 2007;96:808–814.
Laval S, Butler R, Shelling AN, Hanby AM, Poulsom R, Ganesan TS. Isolation and characterization of an epithelial-specific receptor tyrosine kinase from an ovarian cancer cell line. Cell Growth Differ. 1994;5:1173–1183.
Park HS, Kim KR, Lee HJ, et al. Overexpression of discoidin domain receptor 1 increases the migration and invasion of hepatocellular carcinoma cells in association with matrix metalloproteinase. Oncol Rep. 2007;18:1435–1441.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297.
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120:15–20.
Lai EC. Micro RNAs are complementary to 3′ UTR sequence motifs that mediate negative post-transcriptional regulation. Nat Genet. 2002;30:363–364.
Schetter AJ, Harris CC. Alterations of microRNAs contribute to colon carcinogenesis. Semin Oncol. 2011;38:734–742.
Kong Y, Bai PS, Sun H, Nan KJ, Chen NZ, Qi XG. The deoxycholic acid targets miRNA-dependent CAC1 gene expression in multidrug resistance of human colorectal cancer. Int J Biochem Cell Biol. 2012;44:2321–2332.
Arocho A, Chen B, Ladanyi M, Pan Q. Validation of the 2-DeltaDeltaCt calculation as an alternate method of data analysis for quantitative PCR of BCR-ABL P210 transcripts. Diagn Mol Pathol Am J Surg Pathol. 2006;15:56–61.
Saadoun S, Papadopoulos MC, Hara-Chikuma M, Verkman AS. Impairment of angiogenesis and cell migration by targeted aquaporin-1 gene disruption. Nature. 2005;434:786–792.
L’Hote CG, Thomas PH, Ganesan TS. Functional analysis of discoidin domain receptor 1: effect of adhesion on DDR1 phosphorylation. FASEB J. 2002;16:234–236.
Jian ZX, Sun J, Chen W, Jin HS, Zheng JH, Wu YL. Involvement of discoidin domain 1 receptor in recurrence of hepatocellular carcinoma by genome-wide analysis. Med Oncol. 2012;29:3077–3082.
Valencia K, Ormazabal C, Zandueta C, et al. Inhibition of collagen receptor discoidin domain receptor-1 (DDR1) reduces cell survival, homing, and colonization in lung cancer bone metastasis. Clin Cancer Res. 2012;18:969–980.
Miao L, Zhu S, Wang Y, et al. Discoidin domain receptor 1 is associated with poor prognosis of non-small cell lung cancer and promotes cell invasion via epithelial-to-mesenchymal transition. Med Oncol. 2013;30:626.
Shimada K, Nakamura M, Ishida E, et al. Prostate cancer antigen-1 contributes to cell survival and invasion though discoidin receptor 1 in human prostate cancer. Cancer Sci. 2008;99:39–45.
Yang SH, Baek HA, Lee HJ, et al. Discoidin domain receptor 1 is associated with poor prognosis of non-small cell lung carcinomas. Oncol Rep. 2010;24:311–319.
Kadera BE, Li L, Toste PA, et al. MicroRNA-21 in pancreatic ductal adenocarcinoma tumor-associated fibroblasts promotes metastasis. PLoS ONE. 2013;8:e71978.
Sandhu R, Rivenbark AG, Coleman WB. Loss of post-transcriptional regulation of DNMT3b by microRNAs: a possible molecular mechanism for the hypermethylation defect observed in a subset of breast cancer cell lines. Int J Oncol. 2012;41:721–732.
Fang JY, Lu J, Chen YX, Yang L. Effects of DNA methylation on expression of tumor suppressor genes and proto-oncogene in human colon cancer cell lines. World J Gastroenterol. 2003;9:1976–1980.
Su SF, Chang YW, Andreu-Vieyra C, et al. miR-30d, miR-181a and miR-199a-5p cooperatively suppress the endoplasmic reticulum chaperone and signaling regulator GRP78 in cancer. Oncogene. 2013;32:4694–4701.
Wang W, Zhao LJ, Tan YX, Ren H, Qi ZT. Identification of deregulated miRNAs and their targets in hepatitis B virus-associated hepatocellular carcinoma. World J Gastroenterol. 2012;18:5442–5453.
Xu N, Zhang J, Shen C, et al. Cisplatin-induced downregulation of miR-199a-5p increases drug resistance by activating autophagy in HCC cell. Biochem Biophys Res Commun. 2012;423:826–831.
Maeda M, Johnson KR, Wheelock MJ. Cadherin switching: essential for behavioral but not morphological changes during an epithelium-to-mesenchyme transition. J Cell Sci. 2005;118:873–887.
Kang Y, Massague J. Epithelial-mesenchymal transitions: twist in development and metastasis. Cell. 2004;118:277–279.
Thiery JP, Acloque H, Huang RY, Nieto MA. Epithelial-mesenchymal transitions in development and disease. Cell. 2009;139:871–890.
Acloque H, Thiery JP, Nieto MA. The physiology and pathology of the EMT. Meeting on the epithelial-mesenchymal transition. EMBO Rep.. 2008;9:322–326.
Lee JM, Dedhar S, Kalluri R, Thompson EW. The epithelial-mesenchymal transition: new insights in signaling, development, and disease. J Cell Biol. 2006;172:973–981.
Zavadil J, Bottinger EP. TGF-beta and epithelial-to-mesenchymal transitions. Oncogene. 2005;24:5764–5774.
Carico E, Radici M, Losito NS, et al. Expression of E-cadherin and alpha-catenin in T1 N0 laryngeal cancer. Anticancer Res. 2012;32:5245–5249.
Lade-Keller J, Riber-Hansen R, Guldberg P, Schmidt H, Hamilton-Dutoit SJ, Steiniche T. E- to N-cadherin switch in melanoma is associated with decreased expression of phosphatase and tensin homolog and cancer progression. Br J Dermatol. 2013;169:618–628.
van Horssen R, Hollestelle A, Rens JA, Eggermont AM, Schutte M, Ten Hagen TL. E-cadherin promotor methylation and mutation are inversely related to motility capacity of breast cancer cells. Breast Cancer Res Treat. 2012;136:365–377.
Chen X, Wang Y, Xia H, et al. Loss of E-cadherin promotes the growth, invasion and drug resistance of colorectal cancer cells and is associated with liver metastasis. Mol Biol Rep. 2012;39:6707–6714.
Acknowledgments
This work was Supported by the Hunan science and technology planning project of Hunan Province. Grant Number: 2011SK3170.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10620_2014_3136_MOESM1_ESM.tif
Supplementary material 1 DDR1 knockdown of LOVO in LOVE1 cells. DDR1-shRNA significantly decreased DDR1 protein expression of DDR1 in LOVO and LOVE1 cells compared to control (Con) and the cells transfected with scramble shRNA (NC) detected by Western blot (TIFF 6024 kb)
Rights and permissions
About this article
Cite this article
Hu, Y., Liu, J., Jiang, B. et al. MiR-199a-5p Loss Up-Regulated DDR1 Aggravated Colorectal Cancer by Activating Epithelial-to-Mesenchymal Transition Related Signaling. Dig Dis Sci 59, 2163–2172 (2014). https://doi.org/10.1007/s10620-014-3136-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-014-3136-0