Abstract
The Australian freshwater crocodile (Crocodylus johnstoni) is endemic to the northern mainland tropics of Australia and is widespread across the Kimberley region in the northwest Australia. Currently, there is limited understanding of the genetic structure and diversity of these populations, which impacts on our ability to evaluate the conservation status of the species. Population genetic analyses of 173 freshwater crocodiles from the Ord River, Fitzroy River, and Lennard River basins were conducted using single-nucleotide polymorphisms (SNPs). After filtering, 1185 SNPs were retained for downstream population genetic analysis. STRUCTURE and principal component analyses identified three clusters consistent with the three river basins. Population differentiation gave an FST of 0.15 between western and eastern Kimberley and the pairwise FST range was 0.06–0.18 among the three river basins. Assignment tests identified three migration events between the adjacent Fitzroy River and Lennard River basins, which may be explained by possible overland movement across these river basins. The population structure found here indicates that delimitation of management units should be based on river basins with the proximity of adjacent river basins taken into consideration when gene flow exists. Estimates of effective population size showing a low ratio of effective population size to census size in Lake Argyle may raise the concern of future monitoring in this area. Further population genetic studies across the species’ full range are required to better understand the extent of river basins acting as discrete population units, gene flow, population dynamics, and demographic history.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Australian freshwater crocodile (Crocodylus johnstoni) is widely distributed in tropical areas of northern mainland Australia (Webb et al. 1983a, b) where it inhabits freshwater environments including creeks, rivers, lagoons, and pools, and occasionally extends into saline tidal areas (Webb 1985). The ecological role of freshwater crocodiles is yet to be understood (Somaweera et al. in press), but they arguably play a crucial role as prey and predators in freshwater ecosystems. They are also important for ecotourism and in Aboriginal communities where there can be strong cultural and spiritual connections to this species (Delaney et al. 2010). While this species is currently accessed as “Least Concern” in the IUCN Red List (Isberg et al. 2017), there is a growing body of literature showing impacts from cane toads, feral ungulates, land use change, invasive weeds and agrochemicals on local populations of freshwater crocodiles (Yoshikane et al. 2006; Letnic et al. 2008; Somaweera et al. 2019).
Defining discrete population units is crucial for species conservation because it helps delimit management units to best protect genetic diversity (Kolomyjec et al. 2009; Muniz et al. 2018; Kay 2005) and could assist in investigating the relationship between population structure, dispersal and local geographical features (Moritz 1999). Currently, the understanding of the population dynamics of the freshwater crocodile is limited. Mark-recapture studies have reported strong site fidelity among freshwater crocodiles, with movement limited to a small home range (Webb et al. 1983a, b). Some male juveniles showed an extensive movement range (~ 30.3 km) compared with other individuals (~ 1 to 1.9 km) within river systems, and this could relate to mating competition (Tucker et al. 1997). In addition, freshwater crocodiles occasionally move overland in the dry season when they move between drought pools to seek refuge (Lang 1987; Webb et al. 1983a) and can aestivate in suitable terrestrial refugia when these pools dry (Kennett and Christian 1993). However, the scale or distance of movement of this is not documented, making it difficult to evaluate the potential for migration between river basins.
Population genetic and genomic studies have provided insights into gene flow, population structure and genetic diversity of crocodile species (Kay 2005; Muniz et al. 2018; Luck et al. 2012; Versfeld 2016; Russello et al. 2007) but this has been very limited in C. johnstoni. For example, an analysis of 532 unlinked ddRAD loci for Cuvier’s dwarf caiman (Paleosuchus palpebrosus) in Brazil suggested very high population differentiation among four river basins, and four management units consistent with the river basins were proposed (Muniz et al. 2018). Using short tandem repeat (STR) markers, Nile crocodiles inhabiting the Kunene and Okavango Rivers showed some population differentiation, and each river basin population was suggested to be a single management unit (Versfeld 2016). For saltwater crocodiles (C. porosus) in the Kimberley region, Western Australia, management units were suggested at the level of river basin because of moderate genetic differentiation and a 10% first-generation migration rate (Kay 2005). Another saltwater crocodile genetic study based on 8312 SNPs genotype data analyzed the genetic structure, genetic diversity, translocation and homing ability of C. porosus in the Northern Territory, Australia (Fukuda et al. 2019). However, there is no genetic understanding of freshwater crocodile populations including those from the Kimberley region, a region of high ecological importance (Pepper and Keogh 2014). Freshwater crocodiles have established large populations in the Kimberley region, where they inhabit the Ord River basin, including Lake Kununurra and Lake Argyle in eastern Kimberley and the Fitzroy River and Lennard River basins in western Kimberley (McNamara and Wyre 1993). These three river basins are independent river systems without water path connections, with a minimum distance apart of about 50 km (Department of Water 2008). Since the freshwater crocodile is characterized by limited overland movement ability, this may be driving some level of population differentiation between river basins, a hypothesis that is yet to be examined. To address this, we investigated the genetic diversity and differentiation among three river basins from the Kimberly region in northern Western Australia, using single-nucleotide polymorphisms (SNPs) generated by double-digest restriction site-associated DNA sequencing (RADseq).
Materials and methods
Sample collection and DNA extraction
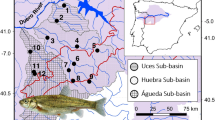
Freshwater crocodile tissue samples were collected from the Fitzroy River (Geikie Gorge and water pools downstream), Lennard River (Windjana Gorge and water pools downstream), and Ord River (Lake Argyle and creeks flowing in) basins, which are known to contain large populations of freshwater crocodiles (Mawson 2004). Hatchling and subadult crocodiles were caught at night using hand nets from boats. Adult crocodiles were caught using nooses (Fitzroy and Ord Rivers) or drift nets swept across drying water pools (Lennard River), the latter during annual mark and recapture surveys by the Department of Biodiversity Conservation and Attraction and Bunuba Rangers. All the captured animals were restrained using methods appropriate for their size (Webb and Messel 1977; Combrink et al. 2012). Dorsal tail scutes were cut using scissors or a sharp pocketknife sterilized in 70% ethanol and dried using a clean tissue. The harvested tissue samples were then stored in 100% ethanol. The animals were released at the site of capture. In addition to the above, two samples from Duck Hole and six samples from Lake Kununurra along the Ord River (Fig. 1, Table 1) were included. In total, 174 samples from five sample sites representing three river basins were used for genomic DNA extraction. DNA was extracted using the DNeasy Blood and Tissue Extraction Kit from Qiagen (Germany) or the phenol–chloroform protocol (Green and Sambrook 2012) Qubit (Life Technologies, USA). The concentration and quality of the extracted DNA was determined using Nanodrop spectroscopy and 1% agarose gels. The samples were then sent to the Australian Genomic Research Facility for dd-RADseq, as described below.
The map shows the sample collecting sites and the numbers from each location. The blue lines represent the river and the dot represent the collecting site of samples, with the specific river basin names labelled in blue and sample size indicated by arrow. The red dot indicates the samples collecting site, with the number of samples collected there
Library preparation and SNP discovery
Library preparation was performed according to the protocols by Peterson et al. (2012). In brief, 200 ng of genomic DNA was double-digested using ecoRI and NlaIII, with the ligation-compatible barcode adapters A and P2 adapter and restriction site overhang. The fragmented DNA was enriched by polymerase chain reaction, and only the tags with ligation of both P2 and A adapters being amplified. The enriched fragments were size-selected using Blue Pippin, which selected 60-bp fragments for sequencing. The prepared libraries were sequenced using the NextSeq platform (Illumina, USA) in four lanes, following the manufacturer’s instructions.
Raw sequences were demultiplexed and assembled using Stacks 1.47 (Catchen et al. 2011, 2013b). Demultiplexing was performed using process RAD-tags, which excluded reads with uncalled bases as well as low-quality data with Phred scores < 20 and barcode and RAD-tags. The resulting reads were analyzed for SNP discovery de novo since a reference genome sequence from freshwater crocodile is not available. We employed UStack with a maximum likelihood statistical model (Catchen et al. 2011), which requires a minimum of two raw reads to create a stack. Cstack allowed a maximum of one mismatch to generate the catalogue and SStack verified individuals with a default parameter. All generated files were used in the population component to link individuals to their respective populations and generate a data matrix stored in a VCF file. The genotype rate and locus depths can cause bias in downstream analysis (Fumagalli 2013; Chattopadhyay et al. 2014; Linck and Battey 2019). Therefore, we filtered the dataset to include only those loci that provided genotypes in at least 50% of all individuals to maximize population structure informativeness parameters (Streicher et al. 2016). The SNP filtering steps were conducted using VCFtools (Danecek et al. 2011). Finally, the filtered VCF file was converted into a compatible format with PGDSpider v. 2.0.8.2 (Lischer and Excoffier 2011) for analysis using GenoDive (Meirmans and Van Tienderen 2004), and STRUCTURE (Evanno et al. 2005).
Population structure and differentiation
The genetic relatedness of individuals was investigated to determine whether the genetic structure was consistent with river basin location. Principal component analysis (PCA) was performed in GenoDive (Meirmans and Van Tienderen 2004) to infer population structure and identify and adjust for ancestral differences among the sampled animals. A covariance matrix was used to calculate eigenvalues of individual principal components (PCs), with 100 replications for significance testing. The two PCs with the highest eigenvalues were plotted. Another PCA analysis were performed by removing Duck Hole and Lake Kununurra samples due to their small sample size.
Population clustering was further investigated in STRUCTURE (Pritchard et al. 2000) using an admixture model and correlated variance. In order to highlight the genetic structure between western and eastern Kimberley, we further performed a STRUCTURE analysis with combination of the samples from the Lennard and Fitzroy River as the western Kimberley group and the Ord River as the eastern Kimberley group. These analyses were run with K values of 2–6, 5 replications per K, 20,000 burn-in steps, and 20,000 iterations. The K values were selected using the delta K method described by Earl (2012) in Structure Harvester (https://taylor0.biology.ucla.edu/structureHarvester/) and then summarized and drawn using CLUMPP (Jakobsson and Rosenberg 2007) and Distruct (Rosenberg 2004).
Pairwise FST values between the identified populations were calculated using GenoDive (Meirmans and Van Tienderen 2004). We further combined data from the Lennard and Fitzroy River to estimate genetic differentiation between the western Kimberley and the Ord River in the eastern Kimberley. All of the calculations were performed with 10,000 replications to determine significance.
Population assignment test
Assignment tests were used to calculate the probability of an individual belonging to the source population (Rannala and Mountain 1997) using the allele-frequency-based method in GenoDive (Meirmans and Van Tienderen 2004). The likelihood ratio Lhome/Lmax with 100 permutations (significance threshold 0.05) was also calculated (Piry et al. 2004). Misassignments with > 95% probability indicated genotypes that were likely the result of immigration rather than random combinations of the original population.
Genetic diversity and effective population size
We estimated genetic diversity based on the SNP data stored in a VCF file; thus, population statistics only included variable positions. Private alleles, nucleotide diversity (π), expected heterozygosity, observed heterozygosity and inbreeding coefficient (FIS) were measured for each identified population using the POPULATIONS program in Stacks (Catchen et al. 2011, 2013b). We further employed GenoDive to perform Hardy–Weinberg equilibrium with 10,000 permutation to test the significance of FIS (Meirmans and Van Tienderen 2004). To assess the genetic diversity at the genome level, we calculated nucleotide diversity in the all position of RAD-tag in Stacks, to compare genetic diversity to other species. Effective population size was estimated in NeEstimator (Do et al. 2014) using the linkage disequilibrium approach with a minimum allele frequency cutoff of 0.02 and random mating model for the Lake Argyle, Fitzroy River and Lenard River samples (Larson et al. 2014), when Lake Kununurra and Duck Hole were dropped due to the too small sample sizes. We further compared the effective population size to the census size in Lake Argyle (WMI Pty Ltd 2010) and the Fitzroy River (Mawson 2004).
Results
Sequencing data quality and processing
Four lanes of sequencing produced approximately 361 million reads derived from 174 samples. The number of RAD-tags obtained per individual ranged from 58,000 to 210,000, with the tag depth between 2.4–3 (Supplementary Information, Table S1). After de novo assembly and SNP calling, 89,516 SNPs were merged in a data matrix. We summarized the percentage of un-genotyped SNPs across 174 samples and found very high missing rate per loci across individuals and populations (Supplementary Information, Fig S2). We filtered SNP loci by requiring that they be successfully genotyped in 50% of individuals. During data filtering, one individual was eliminated as no loci were available after SNP quality control. Finally, 173 individuals and 1185 SNP loci were retained for the downstream analysis of population genetics.
Population structure and differentiation
Three clusters were identified based on PCA of 1185 SNP loci that are consistent with the three river basins (Fig. 2). Some overlap between the two populations from western Kimberley was observed, as well as a few individuals from Fitzroy River and Duck Hole on the Ord River. The extra PCA analysis excluding Lake Kununurra and Duck Hole show three clear clusters consistent with the river basins, but again some overlap within western Kimberley (Supplementary Material S6). The STRUCTURE analysis detected three genetic clusters (K = 3, Supplementary Material S3, S4), which is consistent with the river basin origins of the samples (Fig. 3). One cluster grouped individuals from Lake Kununurra, Lake Argyle, and Duck Hole, all from the Ord River basin, although some Duck Hole samples showed overlap with the Fitzroy River. The remaining two clusters grouped individuals from the Lennard River or Fitzroy River, although there was some genetic admixture. The STRUCTURE analysis comparing western Kimberley and eastern Kimberley groups shows the same result, which illustrates the clear differentiation between western and eastern Kimberley (Supplementary Material S7).
Principal Component analysis plot of average scores of individuals from five collecting sites. The first and second principle components (PC1 and PC2) are shown, representing the x axis and y axis respectively. The first PCA axis explains 6.797% and the second PCA axis explains 2.319% of the total variation. The first 10 PC axes explain 21.223% of variation. The individual sourced from collecting points were represented by different colour, which indicates in the right. Three circles represent 3 river basins: Orange: Ord River basin, Yellow: Lennard River basin, Blue: Fitzroy River basin. Although some overlap between three circles, the identified three clusters generally concordant to the three river basins indicate the presence of genetic structure among river basins
Population structure inferred by 173 individuals from five collecting sites. Each vertical represents an individual. The collecting sites belonged to the river basin placed below the figure, and three collecting sites from Ord River basin was extra labelled above the Ord River basin. The color proportion represents the probability of assignment of each individual to each cluster. K = 3 was the best fit our result based on DeltaK method (Supplement Material). The collecting site is arranged by the geographical location and place adjacent river basin together. Each River basin represent a clear cluster, but some admixture occurs between Lennard River and Fitzroy River
We further analyzed the pairwise FST among the three rivers. The largest differentiation was between Ord River and Lennard River (FST = 0.177, P < 0.0001), followed by 0.119 between Ord River and Fitzroy River (P < 0.0001). Only a moderate differentiation (FST = 0.061, P < 0.0001) was identified between Lennard River and Fitzroy River. Genetic differentiation between western (combined Lennard and Fitzroy Rivers) and eastern Kimberley (Ord River) was FST = 0.152 (P < 0.0001).
Assignment of individuals
The population assignment test correctly assigned 170 of 173 samples to their source populations. Three individuals (20-I6_GCAT, 94-I6_TAGA, and WG378-I19_GGATTGGC) were identified as unlikely to belong to their sample populations and were therefore identified as immigrants. Based on the three reference populations, two samples (20-I6_GCAT and 94-I6_TAGA) were from the Fitzroy River and assigned to the Lennard River. One sample (WG378-I19_GGATTGGC) was from the Lennard River and assigned to the Fitzroy River.
Genetic diversity and effective population size
Based on the SNP data matrix of variable positions the π value for all three basins combined was 0.101, observed heterozygosity was 0.073, expected heterozygosity was 0.10, and FIS was 0.394 (Table 2). The π value for each basin ranged between 0.09 and 0.101 and was the highest for the Fitzroy River (0.101) and lowest for the Lennard River (0.09). The number of private alleles was higher in the Lennard River (263) than in the Ord River (141) and Fitzroy River (84). All clusters showed observed heterozygosity lower than expected heterozygosity and the Hardy–Weinberg Equilibrium test among three cluster (P < 0.001) support the significant difference between observed Heterozygosity and expected heterozygosity. The Lennard River population had the highest value (FIS of 0.33), followed by the Ord River (FIS of 0.18) and Fitzroy River basins (FIS of 0.14). All of the FIS showing statistical significance (P < 0.01). The estimated effective population size was 5678 (2599.8 to infinite) in the Lennard River, 5064 (1134.6 to infinite) in the Fitzroy River, and 2047 (831 to infinite) in Lake Argyle. All of the effective population size estimates contained a large confidence interval showing an infinite size for the upper limit. The ratio of estimated effective population size to census size Ne/N was 0.389 in the Fitzroy River and 0.082 for Lake Argyle (Table 3). Finally, we compared the nucleotide diversity from all positions of RAD-tag to other species and found that the genetic diversity observed in our study population was low (Table 4).
Discussion
This is the first study identified 1185 SNP loci in freshwater crocodiles, this was higher than the 532 loci used in a study of Cuvier’s dwarf caiman (Muniz et al. 2018), and lower than 8312 SNPs loci used in a study of C. porosus (Fukuda et al. 2019). Some studies compared the effectiveness of microsatellites and SNPs in population genetics. Fernández et al. (2013) found 2–3 SNPs were equivalent to 1 STR marker in getting the same exclusion power. Gärke et al. (2011) found that 29 SSR loci were equivalent to 70 SNPs for effectiveness in PCA-based partitioning. Liu et al. (2005) found on average that the informativeness of microsatellite loci was four to twelve times that of SNPs for population comparisons. Considering various population genetics using microsatellites in crocodilian species used up to 20 loci (e.g. Kay 2005; Davis et al. 2002; Velo-Antón et al. 2014; Russello et al. 2007), the number of SNP loci in this study should be sufficient to inform the population structure. The sampling regime allowed comparisons of populations in the adjacent Fitzroy and Lennard River basins in the western Kimberley, to the Ord River basin in the eastern Kimberley. The pairwise estimates of genetic structure (FST) were 2–3 times greater between the Ord River basin and either the Fitzroy or Lennard River, than between the Fitzroy and Lennard River. This pattern of genetic structure was expected given the geology of the region. In the central Kimberley region, a large plateau comprising the uplands of the Prince Regent Plateau, Gibb Hills and the Karunjie Plateau separates the Ord River basin from the Fitzroy River and Lennard River basins (Pepper and Keogh 2014). Sample locations in the two regions were located a minimum of 400 km distant across extensive rocky terrain that would impede immigration. Three genetic clusters were identified based on PCA and STRUCTURE analysis, which were concordant with those found in the three river basins, which could be an indication that these streams are acting as discrete units. One of the factors that may be driving the genetic structure is the strong site fidelity observed in the species. It has been found that relocated freshwater crocodiles have been recaptured in the original water hole after a year of having been released (Webb et al. 1983c) and movement of natural populations appear to be restricted within river basins (Tucker et al. 1997).
Within the western Kimberley, the observed genetic admixture and the identification of three possible migration events between the Lennard River and Fitzroy River basins suggests limited contemporary gene flow between these basins. Although the sample locations in the two collecting site were ~ 110 km apart, overland movement of freshwater crocodiles may have occurred where distances are ~ 50 km through the King Leopold Range. Avenues for temporary connectivity between these rivers may have been facilitated by a number of small stream anabranches found throughout the area between Geikie Gorge (Fitzroy River) and Windjana Gorge (Lennard River) (Department of Water 2008). It may be surmised that during the wet season these creeks, anabranches and temporary water holes may facilitate occasionally suitable habitats and paths for freshwater crocodiles to disperse when the distance apart between river basins is at its minimum. In years of high rainfall due to tropical cyclones (Wende 1997), increased rainfall may have provided aquatic pathways as a stepping-stone for limited dispersal and gene flow between freshwater crocodiles in the Lennard River and Fitzroy River basins. Alternatively, gene flow may have occurred at the mouths of the Lennard River and Fitzroy River, as both rivers empty into King Sound, and during high flood events the entire area can be flooded (Wolanski and Spagnol 2003). However, this scenario would be more difficult since it requires migration distances between the sample’s sites of ~ 500 km river distance.
Low genetic diversity and effective population size
The observed heterozygosity of 1185 SNPs (Ho range 0.077–0.084) were lower than the C. porosus (average Ho = 0.203) based on the 8,312 SNPs (Fukuda et al. 2019). We further compared nucleotide diversity at all position to our species using RADseq and also found the considerably low genetic diversity (Table 4). This evidence support that the genetic diversity of freshwater crocodile in the Kimberley region is low. At present it is not possible to establish a reason for this but some mechanism of genome evolution in crocodilians (Green et al. 2014) may provide a partial explanation. At the genome level, crocodilians show lower genome heterozygosity values and evolutionary mutation rates compared to chicken, turtle or human DNA, which could be related to the longer generation interval and generation overlap, large body size and low evolutionary rate (Green et al. 2014), which may have influenced the genetic diversity in this species.
Estimated effective population sizes for the three rivers ranged from 3480 to 5670, which were not consistent with limited data on population size. Lake Argyle contains the largest C. johnstoni population known and was estimated at over 30,000 non-hatchling individuals in 2010 (WMI Pty Ltd 2010). Limited surveys show that the population at Fitzroy River basin is 13,000 non-hatching (Webb Pty Ltd 1989), which is smaller than that of Ord River basin. Therefore, the expectation of effective population in Lake Argyle would be much higher than Fitzroy River. However, the estimated effective population size of 2047 was much lower than that of the Fitzroy River (5065). An estimated average value of Ne/N in wildlife populations is 0.10–0.11 (Frankham 1995), which is lower than estimated for the Fitzroy River (0.389), but higher than estimated for Lake Argyle (0.068). The interpretation of relatively high effective population size in the Fitzroy River could be related to the genetic admixture between the adjacent Lennard River and Fitzroy River described in the current study. In contrast, the relatively low effective population size in Lake Argyle, needs to be understood for effective management and conservation. The population at the lake arguably has gone through a bottleneck when the Ord River was dammed to create the lake. Currently, the spread of invasive weeds in nesting sites pushes crocodiles to nest in suboptimal substrates causing higher embryo mortality (Somaweera et al. 2019), while selective nest predation pressures by dingoes (Somaweera et al. 2011) and direct mortality through ingestion of invasive cane toads (Somaweera and Shine 2012) continues. While the actual long-term direct and indirect impacts of these interactions are not fully understood yet, they may further influence the effective population size. Continued monitoring of the population is recommended. Future studies with additional genetic markers are required to confirm the effective population size in the Kimberley region given the estimates of Ne had a large confidence intervals. Use of whole genome sequencing, which has been used with other Australian freshwater vertebrate lineages (Martin et al 2018), along with population surveys may provide more genetic markers in shedding the demographical history in Kimberley and determine whether the freshwater crocodiles went through past genetic bottlenecks in this region.
Conclusions
This study found strong population differentiation with statistical significance between the western and eastern Kimberley, and possible limited gene flow between the adjacent Lennard River and Fitzroy River basins in the western part of this region. Although the specific factors that may have led to this gene flow are unclear, we propose a scenario to explain this, in which freshwater crocodiles may have occasionally dispersed through temporary aquatic pathways in streams, creeks, and floodplains between these two river basins. We recommend additional surveys involving mark-recapture, tracking and genetic analysis between these two river basins and the King Leopold Range to enhance our understanding. Based on the current study, the river basins appear to act as discrete population units for freshwater crocodiles, and thus the delimitation of management units could be considered at the level of river basin. However, a comprehensive study with more sampling sites, including multiple sites within river basins, such as Duck Hole, is required to identify habitat boundaries. The continued threat from the invasion of cane toads across the Kimberley, and variable results concerning effective population size identified in this study should raise the concerns for continued population monitoring in this ecologically important region to ensure the conservation status of freshwater crocodiles remains of “least concern”.
Data availability
All the data files are available from the National Center for Biotechnology Information (NCBI) Sequence-Read Archive (SRA) database with the accession numbers: SRR959989-SRR9600060.
References
Catchen JM, Amores A, Hohenlohe P, Cresko W, Postlethwait JH (2011) Stacks: building and genotyping loci de novo from short-read sequences. G3 Genes Genom Genet 1(3):171–182
Catchen J, Bassham S, Wilson T, Currey M, O'Brien C, Yeates Q, Cresko WA (2013a) The population structure and recent colonization history of Oregon threespine stickleback determined using restriction-site associated DNA-sequencing. Mol Ecol 22(11):2864–2883
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA (2013b) Stacks: an analysis tool set for population genomics. Mol Ecol 22(11):3124–3140
Chattopadhyay B, Garg KM, Ramakerishnan U (2014) Effect of diversity and missing data on genetic assignment with RAD-Seq markers. BMC Res Notes 7(1):841
Combrink X, Warner J, Hofmeyr M, Govender D, Ferreira S (2012) Standard operating procedure for the monitoring, capture and sampling of Nile Crocodiles (Crocodylus niloticus). South African National Parks, Skukuza, South Africa, unpublished report, 1–14.
Danecek P, Auton A, Abecasis G, Aibers CA, Banks E, Depristo MA et al (2011) The variant call format and vcftools. Bioinformatics 27(15):2156–2158
Davis LM, Glenn TC, Strickland DC et al (2002) Microsatellite DNA analyses support an east-west phylogeographic split of American alligator populations. J Exp Zool 294(4):352–372
Delaney R, Fukuda Y, Neave HM et al (2010) Management Program for the Freshwater Crocodile (Crocodylus Johnstoni) in the Northern Territory of Australia, 2010–2015. Department of Natural Resources, Environment, the Arts and Sport
Department of Water (2008) Rivers of the Kimberley. Water notes for river management. Advisory notes for land managers on river and wetland restoration. Government of Western Australia, Kensington
Dierickx EG, Shultz AJ, Sato F, Hiraoka T (2015) Edwards, SV (2015) Morphological and genomic comparisons of Hawaiian and Japanese B lack-footed A lbatrosses (Phoebastria nigripes) using double digest RAD seq: implications for conservation. Evol Appl 8(7):662–678
Do C, Waples RS, Peel D et al (2014) NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour 14(1):209–214
Earl DA (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4(2):359–361
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14(8):2611–2620
Fernández ME, Goszczynski DE, Lirón Juan P et al (2013) Comparison of the effectiveness of microsatellites and SNP panels for genetic identification, traceability and assessment of parentage in an inbred Angus herd. Genet Mol Biol 36(2):185–191
Frankham R (1995) Effective population size/adult population size ratios in wildlife: a review. Genet Res 66(2):95–107
Fukuda Y, Webb G, Manolis C, Lindner G, Banks S (2019) Translocation, genetic structure and homing ability confirm geographic barriers disrupt saltwater crocodile movement and dispersal. PLoS ONE 14(8):e0205862
Fumagalli M (2013) Assessing the effect of sequencing depth and sample size in population genetics inferences. PLoS ONE 8:e79667
Gärke C, Ytournel F, Bed'hom B, Gut I, Lathrop M, Weigend S, Simianer H (2011) Comparison of SNPs and microsatellites for assessing the genetic structure of chicken populations. Anim Genet 43(4):419–428
Gong X, Davenport ER, Wang D (2019) Clark AG (2019) Lack of spatial and temporal genetic structure of Japanese eel (Anguilla japonica) populations. Conserv Genet 20:1–9
Green MR, Sambrook J (2012) Molecular cloning: a laboratory manual: three-volume set. Cold Spring Harbor Laboratory Press, New York
Green RE, Braun EL, Armstrong J, Earl D, Nguyen N, Hickey G et al (2014) Three crocodilian genomes reveal ancestral patterns of evolution among archosaurs. Science 346(6215):1254449
Isberg S, Balaguer-reina SA, Ross JP (2017) Crocodylus johnstoni, Australian Freshwater Crocodile. The IUCN Red List of Threatened SpeciesTM. https://www.iucnredlist.org/species/46589/3010118. Accessed 26 June 2019.
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23(14):1801–1806
Kay WR (2005) Population ecology of Crocodylus porosus (Schneider 1801). In: The Kimberley region of Western Australia. PhD Dissertation, The University of Queensland
Kennett R, Christian K (1993) Aestivation by freshwater crocodiles (Crocodylus johnstoni) occupying a seasonally ephemeral creek in tropical Australia. In: Lunney D, Ayers D (eds) Herpetology in Australia. Royal Zoological Society of New South Wales, Mosman, pp 315–319
Kolomyjec SH, Chong JYT, Blair D, Gongora J, Grant TR, Johnson CN, Moran C (2009) Population genetics of the platypus (Ornithorhynchus anatinus): a fine-scale look at adjacent river systems. Aust J Zool 57(4):225–234
Lang JW (1987) Crocodilian behaviour: implications for management. Wildlife management: crocodiles and alligators, pp 273–294
Larson WA, Seeb LW, Everett MV et al (2014) Genotyping by sequencing resolves shallow population structure to inform conservation of Chinook salmon (Oncorhynchus tshawytscha). Evol Appl 7(3):355–369
Letnic M, Webb JK, Shine R (2008) Invasive cane toads (Bufo marinus) cause mass mortality of freshwater crocodiles (Crocodylus johnstoni) in tropical Australia. Biol Conserv 141(7):1773–1782
Linck E, Battey CJ (2019) Minor allele frequency thresholds strongly affect population structure inference with genomic data sets. Mol Ecol Resour 19(3):639–647
Lischer HE, Excoffier L (2011) PGDSpider: an automated data conversion tool for connecting population genetics and genomics programs. Bioinformatics 28(2):298–299
Liu N, Chen L, Wang S et al (2005) Comparison of single-nucleotide polymorphisms and microsatellites in inference of population structure. BMC Genet 6(Suppl 1):S26
Luck NL, Thomas KC, Morin-Adeline VE et al (2012) Mitochondrial DNA analyses of the saltwater crocodile (Crocodylus porosus) from the Northern Territory of Australia. Aust J Zool 60(1):18–25
Martin HC, Batty EM, Hussin J et al (2018) Insights into platypus population structure and history from whole-genome sequencing. Mol Biol Evol 35(5):1238–1252
Mawson PR (2004) Crocodile management in Western Australia. Proceedings of the 17th working meeting of the crocodile specialist group of the species survival commission of the IUCN. Darwin, Australia, pp 28–37
McNamara KJ, Wyre GJ (1993) The conservation, management and farming of crocodiles in Western Australia. In: Crocodiles. Proceedings of the 2nd regional meeting of the IUCN-SSC crocodile specialist group. IUCN: Gland., 1993.
Meirmans PG, Van Tienderen PH (2004) GENOTYPE and GENODIVE: two programs for the analysis of genetic diversity of asexual organisms. Mol Ecol Resour 4(4):792–794
Moritz C (1999) Conservation units and translocations: strategies for conserving evolutionary processes. Hereditas 130(3):217–228
Muniz F, Campos Z, Rangel SH, Martinez J, Souza B, Dethoisy B, Botero-arias R, Hrbek T, Farias I (2018) Delimitation of evolutionary units in Cuvier’s dwarf caiman, Paleosuchus palpebrosus (Cuvier, 1807): insights from conservation of a broadly distributed species. Conserv Genet 19(3):599–610
Pepper M, Keogh JS (2014) Biogeography of the Kimberley, Western Australia: a review of landscape evolution and biotic response in an ancient refugium. J Biogeogr 41(8):1443–1455
Peterson BK, Weber JN, Kay EH et al (2012) Double digest radseq: an inexpensive method for de novo snp discovery and genotyping in model and non-model species. PLoS ONE 7(5):e37135
Piry S, Alapetite A, Cornuet J-M et al (2004) GENECLASS2: a software for genetic assignment and first-generation migrant detection. J Hered 95(6):536–539
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Pujolar JM, Jacobsen MW, Frydenberg J, Als TD, Larsen PF, Maes GE et al (2013) A resource of genome-wide single-nucleotide polymorphisms generated by RAD tag sequencing in the critically endangered European eel. Mol Ecol Resour 13(4):706–714
Rannala B, Mountain JL (1997) Detecting immigration by using multilocus genotypes. PNAS 94(17):9197–9201
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Resour 4(1):137–138
Russello M, Brazaitis P, Gratten J et al (2007) Molecular assessment of the genetic integrity, distinctiveness, and phylogeographic context of saltwater crocodiles (Crocodylus porosus) on Palau. Cons Genet 8(4):777–787
Somaweera R, Shine R (2012) The (non) impact of invasive cane toads on freshwater crocodiles at Lake Argyle in tropical Australia. Anim Conserv 15(2):152–163
Somaweera R, Webb JK, Shine R (2011) It’s a dog-eat-croc world: dingo predation on the nests of freshwater crocodiles in tropical Australia. Ecol Res 26:957–967
Somaweera R, Brien ML, Sonneman T et al (2019) Absence of evidence is not evidence of absence: knowledge shortfalls threaten the effective conservation of freshwater crocodiles. Glob Ecol Conserv 2019:e00773
Somaweera R, Nifong J, Rosenblatt A et al (accepted) The ecological importance of crocodylians: towards evidence-based justification for their conservation. Biol Rev
Streicher JW, Nd SJ, Wiens JJ (2016) How should genes and taxa be sampled for phylogenomic analyses with missing data? An empirical study in Iguanian Lizards. Syst Biol 65(1):128–145
Tucker A, Limpus C, Mccallum H et al (1997) Movements and home ranges of Crocodylus johnstoni in the Lynd River. Qld Wildl Res 24(4):379–396
Velo-Antón G, Godinho R, Campos JC et al (2014) Should i stay or should i go? Dispersal and population structure in small, isolated desert populations of West African crocodiles. PLoS ONE 9(4):e94626
Versfeld WF (2016) Nile crocodile (Crocodylus niloticus) genetic diversity and population structure, within the lower Kunene and Okavango Rivers of northern Namibia. Dissertation. Stellenbosch University
Webb GJW (1985) Survey of a pristine population of freshwater crocodiles in the Liverpool River, Arnhem Land, Australia. Natl Geogr Soc Res Rep 1979:841–852
Webb Pty Ltd (1989). The results of Saltwater Crocodile (Crocodylus porosus) and Freshwater Crocodile (C. johnstoni) surveys in King Sound, Stokes Bay and the Fitzroy River, Western Australia in November 1989. Report to Western Australia Department. Conservation and Land Management.
Webb GJW, Messel H (1977) Crocodile capture techniques. J Wildlife Manage 41:572–575
Webb GJW, Manolis SC, Buckworth R (1983a) Crocodylus johnstoni in the McKinlay River Area N. T. II. Dry-season habitat selection and an estimate of the total population size. Wildlife Res 10(2):373–382
Webb GJW, Manolis SC, Buckworth R et al (1983b) Crocodylus johnstoni in the McKinlay River Area N. T. III. Growth, movement and the population age structure. Wildl Res 10(2):383–401
Webb GJW, Manolis SC, Buckworth R (1983c) Crocodylus johnstoni in the McKinlay River Area N. T. IV. A demonstration of homing. Wildl Res 10(2):403–406
Wende R (1997) Aspects of the fluvial geomorphology of the Eastern Kimberley Plateau, Western Australia. PhD dissertation, University of Wollongong
WMI Pty Ltd (1992–2010). Results of helicopter surveys of crocodiles in Cambridge Gulf, Lake Argyle and Lake Kununurra, Reports to CALM/DEC, 1992–2010.
Wolanski E, Spagnol S (2003) Dynamics of the turbidity maximum in King Sound, tropical Western Australia. Estuar Coast Shelf Sci 56(5–6):877–890
Yoshikane M, Kay WR, Shibata Y et al (2006) Very high concentrations of DDE and toxaphene residues in crocodiles from the Ord river, Western Australia: an investigation into possible endocrine disruption. J Environ Monit 8(6):649
Acknowledgements
We thank Juan Montenegro for providing advice on population genetic analysis to RC. Field surveys by RS were approved by the CSIRO AEC (Nos. 2017–2018 and 2017–2022) and the Department of Biodiversity Conservation and Attractions AEC (approval 2014/13) under the research permits 08-000949-1, 08-000950-1 and SC001387. Staff of Department of Biodiversity Conservation and Attractions (Broome and Kununurra offices), Bunuba Rangers and MG Rangers collaborated with RS during field studies.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cao, R., Somaweera, R., Brittain, K. et al. Genetic structure and diversity of Australian freshwater crocodiles (Crocodylus johnstoni) from the Kimberley, Western Australia. Conserv Genet 21, 421–429 (2020). https://doi.org/10.1007/s10592-020-01259-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-020-01259-5