Abstract
Understanding the movement ability and the spatial scale(s) of population genetic structure of species can together better ‘tune’ management objectives to prevent potential range contraction and population declines. We studied the Rocky Mountain Sculpin (Cottus sp.), a threatened species in Canada, to demonstrate the utility of using two complementary approaches to assess connectivity of a species. To do so, we used Passive Integrated Transponder (PIT) tags with a stationary tracking array (n = 223) to track movement and genetic data (n = 1,015) from nine microsatellite loci to assess genetic population structure. The PIT tag results indicated that Rocky Mountain Sculpin are sedentary; approximately 50% of individuals only moved a maximum distance of 10 meters (upstream or downstream) over a 5-month period. Genetic analyses indicated that at the spatial scale of our study area (5500 km2), watershed structure (river basins) is the main geographic feature influencing population genetic structure. We used the Bayesian clustering tool STRUCTURE, which suggested four distinct sub-populations of Rocky Mountain Sculpin in Canada. Genetic structure at finer spatial scales (within basins and sub-basins) appears to be influenced by fluvial distance (i.e., geographic distance along a river) and elevation change between sample locations (i.e., isolation-by-distance and isolation-by-environment). Combining movement and genetic analyses provides complimentary evidence of limited dispersal in Rocky Mountain Sculpin and highlights that both approaches together can provide broader insight into connectivity between populations that may ultimately help to aid future management decisions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Improving our understanding of how spatial connectivity and dispersal for species is affected by landscape structure can provide invaluable information for conservation (Hughes et al. 2009; Wagner and Fortin 2013). Spatial connectivity can be evaluated using direct monitoring of dispersal, such as mark and recapture techniques, or inferred using population genetic methods that assess the scale and strength of gene flow among populations (Broquet and Petit 2009; Lowe and Allendorf 2010). By comparison, direct monitoring studies provide insight into demographic connectivity at generally short-term and fine spatial scales, whereas population genetic studies offer insight into connectivity of long-term and broad (i.e., regional) spatial scales (Fraser et al. 2001; Balloux and Lugon-Moulin 2002). Thus, genetic data can impart information regarding the dispersal of successfully reproductive individuals and how this may vary over long time periods, where direct monitoring cannot. Together they provide a complete picture of both local rates of demographic connectivity and how those rates may vary across broader spatial and temporal scales (Lowe and Allendorf 2010). The characterization of connectivity is a pressing conservation issue as many studies predict significant changes in land-use and climate (Dudgeon et al. 2006; McCauley et al. 2015). These changes are expected to have direct impacts on our capacity to ensure population persistence through the maintenance or improvement of spatial connectivity.
Connectivity of obligate freshwater species in dendritic networks is constrained by the aquatic network configuration (Grant et al. 2007). Specifically, movement and gene flow in riverine networks are affected by species-specific habitat preferences, habitat spatial configuration (amount and fragmentation), season, ontogeny, and network structure (stream hierarchy and flow direction) (Hughes et al. 2009; Selkoe et al. 2016). A clear understanding of the importance of these factors and how they interact can be used to guide management and conservation objectives (Lowe and Allendorf 2010). For example, it is important to determine what aspects of network structure and connectivity affect the likelihood of rescue effects (e.g., after local extirpation or declines) or future migration (e.g., as a result of habitat alterations due to disturbances and climate change) (Hanski 1998). This knowledge can highlight when more active management approaches (e.g., assisted migration) are needed to ensure long-term species persistence (McLachlan et al. 2007). Further, identifying the relevant spatial scale of population genetic structure will help ensure that the management, conservation, and restoration resources are being allocated at the appropriate spatial scales (Murphy et al. 2015).
Here, we investigate movement and population genetic structure of Rocky Mountain Sculpin (Cottus sp.), a newly identified species that is genetically and morphologically distinct from other freshwater sculpin (Cottidae) species (COSEWIC 2010). In Canada, this species is found on both sides of the Continental Divide in the Flathead River drainage in British Columbia (referred to as western populations), and in the St. Mary and Milk river drainages in Alberta (referred to as eastern populations) (COSEWIC 2010; DFO 2013). Recently, the Committee on the Status of Endangered Wildlife in Canada assessed western populations as Special Concern and eastern populations as Threatened (COSEWIC 2010; DFO 2013). Western populations are listed under the federal Species at Risk Act (SARA) as Special Concern because of their restricted distribution in Canada (only 270 km2), which is thought to make the population vulnerable to anthropogenic activities within the watershed (e.g., forestry practices, road building, exploratory drilling) (COSEWIC 2010). The eastern populations are impacted by flow alterations brought about by drought and the augmentation of water flow within the system alongside anthropogenic activities (DFO 2013).
Currently, Rocky Mountain Sculpin populations are managed as two separate units, eastern and western. However, due to limited adult dispersal, nesting during spawning, and the structure of the dendritic network, there may be further population substructure within these management units. Failure to recognize this substructure could lead to the potential loss of uncharacterized genetic diversity, and may lead to unreliable estimates of dispersal capacity, and genetic connectivity.
Members of the family Cottidae are generally sedentary during their adult life (Petty and Grossman 2004; Schwalb et al. 2011; Radinger and Wolter 2014). Previous mark-recapture studies on Rocky Mountain Sculpin have found that annual dispersal may be limited to a maximum movement of <200 m (Bailey 1952; McCleave 1964). Less is known about larval and juvenile dispersal. However, larvae are quite small (<10 mm), which may result in passive dispersal through downstream drift following emergence from their natal nest sites (Bailey 1952; McCleave 1964; Watkinson and Boguski 2013). Thus gene flow, if present, may be more likely to occur during the larval or juvenile stages of development. If this is the case we would expect that headwaters within each population may be genetically distinct and not as mixed as confluences downstream and that population genetic structure may be described using Isolation-by-Distance (IBD) or Isolation-by-Environment (IBE) models (McRae et al. 2008; Spear et al. 2010; Murphy et al. 2015). Additionally, the potential for dispersal of Rocky Mountain Sculpin among the systems that comprise the eastern population (Lee Creek, St. Mary, North Milk and Milkrivers) is unclear. The addition of a diversion between the St. Mary River and North Milk River, a tributary of the Milk River, within their headwaters (Fig. 1), referred to as the St. Mary Canal, may or may not maintain separation between the river basins, as the structure of the network may still act as a barrier between these rivers. A combination of direct monitoring through mark-recapture and genetic population analysis can together provide insight into the mechanisms that may be driving connectivity within this system.
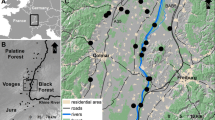
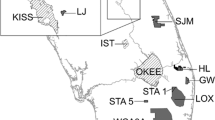
a Sample locations in the Flathead River, Lee Creek, St. Mary River and North Milk River. The star designates the location of the movement array within Lee Creek. Filled black squares denote the locations of the flow stations. b The movement array setup in Lee Creek, Alberta, Canada used to track movement of tagged individuals. The global inset denotes the location of (a)
In this study, we test the hypothesis that due to limited adult dispersal and the potential for downstream drift by larvae or juveniles, sub-populations in each basin and sub-basin within the two managed populations are genetically distinct. We also examine the relative support that exists for IBD and IBE models of population genetic differentiation in the Rocky Mountain Sculpin (Hughes et al. 2009, 2013; Murphy et al. 2015). To address these hypotheses we (1) investigated the movement of adults using Passive Integrated Transponder (PIT) tags, (2) determined the spatial scale of genetic differentiation, and (3) assessed if genetic differentiation is related to fluvial distance (IBD) and/or elevation changes (IBE). In doing so, we demonstrate the utility of combining direct movement and genetic analyses in order to understand the determinants of spatial connectivity in dendritic networks.
Methods
Study area
The study area encompasses a region of southeastern British Columbia and southwestern Alberta, Canada (Fig. 1a). This includes the Flathead River in British Columbia representing the western Rocky Mountain Sculpin population and Lee Creek, St. Mary River and the North Milk River forming the eastern populations, respectively (Fig. 1a). Water flow in each of these systems is sourced from snow melt and precipitation, which contributes to variable seasonal changes in water flow in the Flathead (4.3–90 m3/s), Lee Creek (0.2–10 m3/s), St. Mary (3–90 m3/s), and North Milk (0.7–20 m3/s) (AEP 2015; Canada 2015). From mid-March to mid-October every year, water is diverted from the headwaters of the St. Mary River (Saskatchewan River drainage) to the North Milk River (Missouri River drainage) in Montana, which flows into Alberta and joins the Milk River before flowing back into Montana. This diversion, called the St. Mary Canal, was constructed in 1917 and reconnects two systems which have been separated since the last ice age (Fullerton et al. 2004), and may have permitted the migration of Rocky Mountain Sculpin between the two systems.
Mark-Recapture data
Movement data using mark-recapture methods were collected in Lee Creek, Alberta, Canada from July to October 2013 (Fig. 1b). We used a SmithRoot LR-24 backpack electrofisher (single pass, with block nets downstream) to capture and tag individuals (Total Length ≥ 65 mm; 2+ year olds) with Passive Integrated Transponder (PIT) tags (Fig. 1b). Individuals were anaesthetized in Tricaine Methanesulfonate (TMS; MS-222; 0.2 g L−1) and then a HDX PIT tag (Oregon RFID) was inserted into the peritoneal cavity (Ruetz et al. 2006). The insertion point was then sealed using 3 M Vetbond TM Tissue Adhesive. Individuals were allowed 30 min of recovery prior to being released and were released a maximum of 1 m upstream of the center of our tracking array. The tracking array consisted of 41 transects, spaced 10 meters apart for a total coverage of 400 m of fluvial distance that could be targeted for detections of individual movement (Fig. 1b). The PIT tags were detected using a HDX backpack reader (Oregon RFID), where tag number, location and time stamp are recorded upon detection. To visualize movement data based on the recapture of individuals, we constructed a kernel density model using a Gaussian kernel with a bandwidth of 20 m using the maximum detection distance of movement (the maximum distance moved upstream or downstream from the center of the array). This was conducted using the density function in R software (R Development Core Team 2016). These movement data summarize short-term (~5-month period) and fine spatial scale movements (~400 m), which can compare long-term estimates of gene flow using population genetics.
Genetic survey
One thousand seventeen tissue samples were collected from individual fish in the Flathead River (specifically within two tributaries in the Flathead River drainage), Lee Creek, St. Mary River, and North Milk River during the summers of 2013 and 2014. Sites were chosen prior to field sampling based on available access and distance between sites which ranged from 0.2 to 11 km apart along the rivers (Fig. 1a). We collected pelvic fin clips from individuals that were captured using a SmithRoot LR-24 backpack electrofisher (single pass, with block nets downstream). The objective was to obtain approximately 20–30 individuals at each site (if possible) from 7 to 12 sites in each river/stream (Fig. 1a; Table 1). After individuals were captured, they were allowed to recover for 30 min prior to release. Pelvic fin clips were then preserved immediately in 95% ethanol.
Genotyping
Total genomic DNA was isolated from fin clips using a DNeasy 96 Blood and Tissue Kit (Qiagen) following the manufacturer’s protocol. A total of 15 microsatellite loci were selected from the literature based on their previous use in sculpin population genetic studies (Table S1; Cba from Fiumera et al. 2002; Cco from; Fujishin et al. 2009; Cgo from; Englbrecht et al. 1999 and Cott from; Nolte et al. 2005). For each primer set, the forward primer was labeled with one of 6FAM, TET or HEX fluorescent dyes to allow for genotyping on an Applied Biosystems genetic analyzer. Loci were initially tested using eight individuals from across the species range. These amplification tests were carried out in 15 µL reactions consisting of 1× PCR buffer (10 mM Tris pH8.8, 0.1% Triton X-100, 50mM KCl, 016 mg/mL BSA), 2.5 mM MgCl2, 120 µM each dNTP, 0.17 µM fluorescently labelled forward primer, 0.17 µM reverse primer, 1U Taq DNA polymerase (Engelke et al. 1990) and 2.5 µL (approximately 50 ng) of extracted gDNA. Cycling conditions were 95 °C for 5 min, 35 cycles of 95 °C for 30 s, 57 °C for 90 s, and 72 °C for 30 s, and a final extension at 60 °C for 30 min. Amplification reactions were pooled and loaded on an ABI 3730 DNA analyzer (ThermoFisher Scientific). Fragment sizes were determined relative to GeneScan-500 TAMARA (ThermoFisher Scientific) and scored using Genemapper software. Four loci were removed from further analysis (three loci failed to amplify and one locus was monomorphic).
All individuals were genotyped at the remaining 11 polymorphic microsatellite loci. PCR amplifications were performed in two multiplexed 10 µL reactions (Mix 1 and Mix 2; Table S1) consisting of 1× Type-it Microsatellite PCR Master Mix (Qiagen), 1× primer mix, 0.5× Q-Solution (Qiagen), and 2.5 µL of extracted gDNA. The methods and conditions for all individuals for cycling (except for specific annealing temperatures; Table S1), loading, and sizing were identical to those used for testing.
Genetic data analyses
We could not use all loci and individuals in our analysis. Of the 11 microsatellite loci that were genotyped, only nine were suitable for analysis. Loci were omitted from analysis if: (1) loci had many nontyped individuals (>5), (2) loci were fixed for the mean dominant allele frequency (Excel GenAIEx 6.5), and (3) loci had a mean estimated null allele frequency > 0.1 (ML-NULL) (Kalinowski and Taper 2006; Peakall and Smouse 2012; James et al. 2015). Using the above criteria, we removed two loci from the analysis (Cgo310 and CottES19; Table S1) as they both had many nontyped individuals. Finally, of the 1017 individuals initially sampled, two individuals were removed due to multiple nontyped loci (>5) for a final total of 1015 individuals.
Deviations from Hardy–Weinberg Equilibrium (HWE) were tested for using the hw.test function in the pegas package in R Software (Paradis 2010). Linkage disequilibrium between all loci pairs was tested using FSTAT v2.9.3.2 (Goudet 1995). Standard genetic diversity measures were calculated for each river and site using the Excel GenAIEx 6.5 (Peakall and Smouse 2012) and rarefied allelic richness for each river and site was estimated using the allelic.richness function in the hierfstat package in R software (Goudet and Jombart 2015).
Population genetic structure
We assessed spatial genetic structure by first estimating global F ST and 95% confidence intervals using FSTAT v2.9.3.2 (Goudet 1995). We then calculated unbiased pairwise F ST values (Weir and Cockerham 1984) between each of our sites using the genet.dist function in the hierfstat package in R Software (Goudet and Jombart 2015). Genetic differentiation between all sites was visualized using a Principal Coordinates Analysis (PCoA) of a pair-wise matrix of Fst values using the pco function in the ecodist package in R (Goslee and Urban 2007). Genetic distances (F ST) were double square root-transformed prior to analysis to ensure they were Euclidean for use in the PCoA (Legendre and Legendre 1998; James et al. 2015).
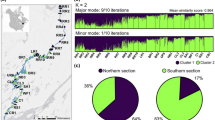
The number of distinct clusters in Rocky Mountain Sculpin was estimated using the software STRUCTURE v2.3.4 (Pritchard et al. 2000). STRUCTURE is a Bayesian fuzzy clustering algorithm that assigns individuals to one or more genetic clusters, to determine the most likely number of clusters (K; or populations) assuming linkage and Hardy–Weinberg equilibrium at each locus. We assumed correlated allele frequencies and admixture among basins in our STRUCTURE model, because a majority of systems (three out of four) were spatially connected allowing for some dispersal and gene flow in our systems (Falush et al. 2003). Our analysis assumed no known priors for K. We assessed values of K from 1 to 10, where for each K we ran 500,000 Monte Carlo Markov Chain (MCMC) generations, after a burn-in of 50,000 generations. Each K had MCMC sampling that was replicated 10 times. We assessed the optimal value of K by reviewing the mean ln probability of K and by using the delta K method (Evanno et al. 2005). We also ran separate STRUCTURE models on each system independently (Flathead, Lee Creek, St. Mary and North Milk) to test if further potential genetic substructure existed beyond the results of overall model, but none was observed (results not shown). Each of the 10 replicates were then summarized using CLUMPP v1.1.2 (Jakobsson and Rosenberg 2007). Finally, we added support to these STRUCTURE-based results by running a similar model, GENELAND, which is a spatial Bayesian fuzzy clustering algorithm (Safner et al. 2011) and provide these results in supplementary material (Fig. S4).
Isolation by distance and isolation by resistance
We tested for the influence of fluvial distance (isolation by distance; IBD) and effective distance based on elevation (isolation by environment; IBE) (McRae et al. 2008; Spear et al. 2010) on genetic differentiation among sample sites. Isolation by distance and resistance were investigated in each basin and sub-basin using a model that compares all pairwise F ST values at sites (Weir and Cockerham 1984) with pairwise fluvial distances and pairwise elevation-based distances, respectively. We only conducted this analysis for eastern sites (Lee Creek, St. Mary River and North Milk River) as there was limited spatial coverage for western samples (Fig. 1a). Fluvial distances were calculated using least-cost paths between sites (Fall et al. 2007) or the shortest path within the river using the gsMPG and shortest.paths functions in the grainscape and igraph packages in R Software (Csardi and Nepusz 2006; Galpern et al. 2014). Elevations for sites were acquired from the Canadian Digital Elevation Model (NRC 2012).
We tested for a significant relationship between: (1) linearized genetic structure (F ST/(1-F ST)) and fluvial distance (IBD); and (2) linearized genetic structure and elevation distance (Rousset 1997). This was conducted using a Procrustes rotation analysis, which compares the similarity between the ordination solutions (i.e., PCoA) of two Euclidean distance matrices, with 10,000 permutations using the protest function in the vegan package in R (Peres-Neto and Jackson 2001; Oksanen et al. 2015). Using this approach we compared the PCoA solutions to the double square-root transformed F ST matrix and the fluvial distance matrix (also verified as Euclidean). We chose to use the Procrustes test because it is shown to be more powerful at comparing distance matrices than other methods (Peres-Neto and Jackson 2001). To determine if IBD and IBE independently affect genetic differentiation, we controlled for variation in both fluvial distance and elevation distance using a partial Procrustes test using the PROTEST function (Peres-Neto and Jackson 2001). Finally, to ascertain what amount of variation in genetic differentiation is explained by both IBD and IBE together (when they are both independent and significant predictors) we conducted a Multivariate Distance Matrix Regression (MRDM) using 10,000 permutations. This was conducted using the MRM function in the ecodist library in R (Goslee and Urban 2007; Lichstein 2007).
Flow regime
We assessed how temporal variation in flow rate (m3/s) influences Rocky Mountain Sculpin connectivity by potentially influencing patterns IBD and IBE. Flow data was retrieved from the Government of Canada national water data archive (Canada 2013), which contains daily flow records throughout the year at gauges in Lee Creek (n = 34,840; 1909–2012), St. Mary River (n = 39,931; 1902–2011), and North Milk River (n = 27,253, 1909–2014; Fig. 1). We then summarized data monthly and tested for differences in overall monthly flow rate between these systems using permuted t-tests using R software (Legendre 2005). As these were only single stations along these waterways, we can only infer that flow rate at these stations reflects system-wide flow. Patterns of hydrological differences between the systems should also support our findings of genetic differentiation (i.e., low flow waterways would have reduced connectivity or gene flow).
Results
Mark-Recapture
We marked 223 individuals with PIT tags and recaptured (or detected) 182 of them for a total recapture rate of 81.6%. The mean maximum detection distance of movement from the point of release across all individuals was zero meters (Fig. 2). Further, the first and third quartiles of movement are 10 meters upstream and downstream of the point of release (Fig. 2). Finally, the largest movements are in a downstream direction at 240 m (n = 2) compared to 170 m (n = 1) upstream direction.
Genetic diversity
Overall, we found a small amount of variation in heterozygosity and the number of alleles among eastern populations from Alberta (Lee Creek, St. Mary River and North Milk River); however, heterozygosity was lower and the number of alleles smaller in western individuals recovered from the Flathead River in British Columbia (Table 1). At the site level we see similar patterns (Table S2); however, there appears to be a gradient in heterozygosity for Lee Creek (Table S2; Fig. S1). Specifically, the headwaters in Lee Creek have a lower heterozygosity than sites located at the confluence with the St. Mary River (Figure S1). Almost all loci were in HWE across populations using Bonferroni corrected p-values (p > 0.05; Table S3). The only exception was Cco15 within the St. Mary River population (Table S3). No significant linkage disequilibrium (p > 0.05) was detected.
Spatial genetic structure
Movement analysis revealed that local demographic connectivity is low; however, such connectivity may be variable over different spatio-temporal scales. Genetic data analysis can provide some insight into whether this is the case. Globally, populations were highly structured showing a significant global F ST value of 0.239 and a 95% confidence interval of 0.175–0.292. Pairwise-F ST values and their associated significance levels revealed spatial patterns of genetic differentiation related to the system where sites are found (Table S4). Visualizing these differences in the principal co-ordinates analysis, the largest difference was found along the first axis of variation (53.7%), between the western (Flathead) and eastern (Lee Creek, St. Mary and North Milk) populations (Fig. 3). Along the second axis of variation (30.2%) Lee Creek and St. Mary River are the most similar, yet distinctly different from the North Milk River (Fig. 3).
We detected similar spatial population structure using STRUCTURE, which we identified that there are likely four genetic clusters; one for western individuals (Flathead River) and three for eastern populations (Lee Creek, St. Mary and North Milk; Fig. 4, S2). The overall STRUCTURE model had a mean ln probability that peaked at K = 4 (Fig. S2A) and the highest delta K value at K = 4 (Fig. S2B). STRUCTURE analysis demonstrated that two of our systems, Flathead River and North Milk River, are quite distinct and assigned each of them to their own unique cluster (Fig. 4). Assignment of individuals in Lee Creek and the St. Mary River was less clear (Fig. 4). In spite of this, when plotting the STRUCTURE population assignments geographically it was revealed that this assignment was systematic (i.e., the confluence between Lee Creek and St. Mary River shared more individuals from both genetic groups compared to the headwaters of each system; Fig. 4). Further, a STRUCTURE model run using only individuals from Lee Creek and St. Mary River had a mean ln probability that was identical at K = 1 and K = 2, suggesting that both genetic clusters are just as likely, providing further weight to these systems being genetically distinct from the other two (Fig. S3).
Fine scale genetic structure: isolation by distance and isolation by resistance
Movement in adults is limited (Fig. 2), thus we may predict that our genetic data analysis may also demonstrate patterns of fine scale genetic structure. We found evidence for significant IBD (r = 0.637, p = 0.021; Fig. 5a) and IBE (r = 0.560, p = 0.039; Fig. 5b) within the Lee Creek. Further, using partial Procrustes analysis, we found fluvial distance (IBD; r = 0.799, p = 0.0002) and elevation distance (IBE; r = 0.790, p < 0.0001) to be significant for Lee Creek sites, indicating that IBD and IBE are independent of one another. We also found evidence of significant IBD and IBE in the St. Mary River (r = 0.726, p = 0.019 and r = 0.749, p = 0.014, respectively; Fig. 5c, d). Additionally, partial Procrustes analysis demonstrated non-significance for fluvial distance (IBD) and elevation distance (IBE; r = 0.589, p = 0.329 and r = 0.580, p = 0.577), meaning that these models were not independent of one another. Finally, we found that IBD and IBE in the North Milk River was also non-significant (r = 0.567, p = 0.192 and r = 0.506, p = 0.367, respectively; Fig. 5e, f). Partial Procrustes analysis also demonstrated non-significance for fluvial distance (IBD) and elevation distance (IBE) in the North Milk River (r = 0.585, p = 0.427 and r = 0.549, p = 0.61). As fluvial distance and elevation distance were both significant and independent of each other in Lee Creek, we ran an MRDM and found that a significant amount of variation between fluvial distance and elevation distance explained genetic differentiation between the sites (R 2 = 0.464, p = 0.0015). Finally, we find that overall flow rate between systems was consistently significantly different (p < 0.001) for all pairwise comparisons, where flow rate was the lowest in Lee Creek, followed by the North Milk River, and St. Mary River (Fig. 6).
Isolation by distance (IBD) and isolation by resistance (IBE) plots of genetic distance as pairwise FST/(1 − FST) for a, b Lee Creek, c, d St. Mary River, and e, f North Milk River. Pairwise genetic distance is related to a, c, e pairwise fluvial distances (km) and b, d, f pairwise elevation differences (m). Shown are the correlation coefficients from Procrustes analysis. Asterisk denotes significance at p < 0.05
Mean flow for each month for Lee Creek (1909–2012), St. Mary River (1902–2011), and North Milk Rivers (1909–2014). Locations are shown in Fig. 1a. Grey shaded areas represent 95% confidence intervals
Discussion
We investigated the determinants of movement and gene flow in the threatened Rocky Mountain Sculpin in Canada and demonstrate the utility in combining mark-recapture and population genetic methods to assess connectivity. With the mark-recapture analysis, we confirm that individual Rocky Mountain Sculpin in Canada move very little, where most adult individuals have a displacement of less than 10 meters from their release point over a 5 months period. Further, the largest displacement we find is in the downstream direction. On the other hand, the genetic analysis confirms that the continental divide provides an insurmountable barrier for western and eastern populations. Surprisingly, although three out of the four river systems Rocky Mountain Sculpin were connected during some portion of the year, there are likely four genetically different populations. This lends support to the idea that the population structure of Rocky Mountain Sculpin is related to the basin or sub-basin where they are found and that dispersal between them is limited and may not vary that much over time. Finally, we demonstrate that genetic differentiation in the Rocky Mountain Sculpin is significantly influenced by both fluvial distance and effective distance based on elevation in Lee Creek and the St. Mary River. IBD and IBE also appear to be independent of one another only in Lee Creek, where IBD and IBE explain 46.4% of variation of genetic differentiation within this population. This fine scale genetic structure may be related to differences in flow rate, as low flow may make some reaches unpassable and high flow would make upstream dispersal difficult. Thus, the occurrence of both IBD and IBE could be related to Lee Creek having the lowest flow rates and the St. Mary River having the highest flow rates in the study.
Movement
Understanding the movement, gene flow, and dispersal of fish species is critical to understanding spatial habitat requirements and developing effective conservation plans (Selkoe et al. 2016). Sculpins are thought to be sedentary in their adult phase (Radinger and Wolter 2014) as behaviorally they prefer not to actively swim, but instead occupy the interstitial spaces between unembedded rocks (Bailey 1952; McCleave 1964). This inference is supported by the data presented in this paper, where a majority of individuals do not move more than ten meters from their release point over a 5-month period (Fig. 2). Further, we find the largest movement distances recorded (240 m) were also observed in a downstream direction, suggesting that downstream movement appears to be more likely than upstream movement.
Our results are based on large individuals (Total Length > 65 mm), which are adults (2+ year olds) within our system (COSEWIC 2010). This means that beyond adult dispersal, it is plausible that dispersal during larval or juvenile stages may also contribute to the genetic structure that we observe at coarse and fine spatial scales. For example, the Prickly Sculpin (Cottus asper) has an extended larval stage (30–35 days; the most mobile stage) and high juvenile dispersal rates in coastal populations (Krejsa 1967) that likely contributes to the observed high migration rates (Dennenmoser et al. 2013). As we did not track the movement of these life stages in Rocky Mountain Sculpin and there is a dearth of other studies on movement in this species, it is likely that connectivity may be higher than what we observe in our movement analysis, because we only target adults. Specifically, it may be expected that small fry (<10 mm) may be more prone to shifting downstream than other life stages. Thus, this demonstrates the utility of genetic data analysis, which can provide insight into connectivity at other life stages and over spatio-temporal scales.
Population genetics
Multiple models of spatial population genetic structure in dendritic networks have been proposed including: panmixia, headwater, Death Valley, and stream hierarchy models (Hughes et al. 2009, 2013; Murphy et al. 2015). These models vary with respect to how they consider temporal continuity of flow, species life-history, and dispersal (Murphy et al. 2015). Identifying which models best resemble the spatial population structure of a target species can provide insight for conservation management decisions.
We found that Rocky Mountain Sculpin populations are structured at the river basin and sub-basin level and have four genetic groups associated with unique basins or sub-basins (Figs. 1a, 4). The Flathead and North Milk river populations represent two of these groups. Both of these basins are separated by long-established barriers, such as the Continental Divide for the Flathead River and the North Milk River is found in a completely different drainage (Fullerton et al. 2004). For the most part, we found concordance between these geographical barriers and the identified genetic groups associated with the Flathead and Milk river basins (Fig. 3). On the other hand, the presence of the St. Mary Canal, which connects the St. Mary River to the North Milk River within their headwaters, may suggest that its presence may facilitate dispersal between these two drainage basins. It is interesting that despite this connection being made in 1917, individuals are still genetically different in each system. The stream hierarchy model (Hughes et al. 2013) represents the most plausible model for this scenario, as genetic structure of Rocky Mountain Sculpin reflects the watershed structure of the system (i.e., the highest connectivity is within basins or sub-basins). These results combined with the known adult dispersal limitation, indicate that gene flow likely occurs in a mostly downstream direction during larval or juvenile life stages. These results also confirm that upstream adult dispersal during the months the canal is operational (mid-March to mid-October) is limited (DFO 2013).
This finding is consistent with the findings of Lamphere and Blum (2012) who used a mark-recapture study to demonstrate that movement in the Mottled Sculpin (Cottus bairdi) occurs more in a downstream direction, but that downstream movements may be greater than can be estimated by mark-recapture studies (~500 m maximum) due to limited temporal and spatial coverage of potential movements. Our movement and genetic analysis, strongly supports that Rocky Mountain Sculpin individuals are sedentary and tend to form populations at the basin and sub-basin level with what appears to be predominantly downstream dispersal or gene flow.
Riverscape genetics
The processes that give rise to spatial genetic structure may include Isolation by Distance (IBD), Isolation by Resistance (IBR), and Isolation by Environment (IBE) (Hughes et al. 2009, 2013; Murphy et al. 2015). Isolation by Distance, IBR and IBE processes may also be collinear, as increased elevation change or steeper slopes will also be highly correlated with changes in the environment (e.g., temperature, precipitation, number of road crossings, spring/fall runoffs), this can contribute to connectivity and genetic structure (Wang and Bradburd 2014). Variance in spatial genetic structure as whole will be made up of components of IBD, IBR, IBE, and interactions between each. Thus, IBD, IBR and IBE should also be tested for independence.
We demonstrated that genetic structure is affected by fluvial distance in Lee Creek and St. Mary River (Fig. 5). Specifically, we showed that along the sampled stretch of the Lee Creek (~40 km) and St. Mary River (~40 km) that there is significant IBD, which agrees with our other findings that adult movement is limited and supports groupings identified using STRUCTURE. Additionally, we found significant IBE, related to elevation, in Lee Creek and the St. Mary River. In contrast, we do not find the same result for the North Milk River, where neither IBD nor IBE is significant. There are couple plausible reasons for why connectivity may differ in Lee Creek and St. Mary River compared to the North Milk River. Firstly, this result may be related to the statistical power of our tests; however, the same number of sites are included for both the St. Mary River (n = 9) and North Milk River (n = 9) in this study. Secondly, flow is significantly lower in Lee Creek and significantly higher in the St. Mary River throughout the year compared to the North Milk River (Fig. 6). Reduced or increased flow can reduce connectivity and increase the strength of spatial genetic structure (Meffe and Vrijenhoek 1988; Labbe and Fausch 2000). Thus, it is possible that both lower and higher flow rates in Lee Creek and the St. Mary River, respectively, may cause a greater reduction in gene flow relative to the other systems examined. A logical next step would be to test for possible effects of asymmetrical flow on genetic differentiation, where asymmetrical flow rates could be incorporated into IBR models (McRae et al. 2008). Thirdly, the genetic samples from Lee Creek and St. Mary River were taken from a larger range of fluvial distances (~40 km) compared to the North Milk River (~25 km). Similarly, elevation gradients from Lee Creek and St. Mary River were larger compared to the North Milk River (~300 m for Lee Creek, ~155 m for St. Mary River and ~80 m for North Milk, respectively). It may be possible that we do not detect IBD or IBE in the North Milk River as we do not sample a similar gradient as found in our samples from Lee Creek and the St. Mary River.
Species movement depends on multiple factors, such as habitat configuration, season, ontogeny of the species and flow (Selkoe et al. 2016). All of these factors, either alone or in combination, contribute to patterns of IBD and/or IBE. However, it can be challenging to sample and model them effectively in riverscape environments. For example, in higher latitudes and montane systems it can be difficult to work in the winter season as ice can prevent access to individuals or can easily damage stationary equipment (e.g., tracking stations). There are currently very few studies that investigate movement of fish species in dendritic systems in higher latitudes during the winter season (e.g., Jakober et al. 1998; Heim et al. 2015). Additionally, it can be difficult to determine larval and juvenile movement within rivers and creeks. This is partly related to the size of the organism during this life stage, whereby newly hatched larvae (<10 mm) cannot be tagged or trapped easily in swift water. One solution to these issues may be to use individual based modelling approaches in dendritic systems, which can provide insight into how asymmetrical movement at different time periods, flow rates or life stages might contribute to resulting IBD or IBE that is detected (Landguth et al. 2016).
Conclusions and conservation implications
Rocky Mountain Sculpin are currently managed as two separate units: western and eastern populations (COSEWIC 2010; DFO 2013). The division between these two populations is suitable as they are separated by the Continental Divide and we find that they are genetically the most distinct from one another. However, eastern populations exhibit additional spatial genetic structure between basins, which indicates that connectivity between basins may be low to non-existent. In contrast, there appears to be some movement at the sub-basin scale, as we detect some movement between Lee Creek and the St. Mary River in our STRUCTURE model. This means that at the very least, Rocky Mountain Sculpin should be managed at the basin level to mitigate the reduced probability of successful rescue effects and other potential negative consequences of reduced movement. We also provide evidence that Rocky Mountain Sculpin can be structured genetically at very fine spatial scales, where in Lee Creek and St. Mary River, we detect significant IBD and IBE. All of this stems from the fact that Rocky Mountain Sculpin are sedentary with maximum adult movements within the ~200 m range. This highlights that Rocky Mountain Sculpin may be susceptible to changes in connectivity at very fine spatial scales, such as the addition of new barriers or alterations to flow, as they do not disperse widely. Overall, we demonstrate that combining movement and genetic approaches can provide multiple lines of evidence, cover a long-term temporal scale (movement, days to years; genetics, years to millennia), and ultimately build a broader perspective of the connectivity of a species that can benefit future management decisions.
References
AEP (2015) Alberta’s River Basins: River Flows and Levels. Alberta Environment and Parks GoAUhweacabDaDAF, Edmonton
Bailey JE (1952) Life history and ecology of the sculpin Cottus biardi punctulatus in southwestern Montana. Copeia 4:243–255
Balloux F, Lugon-Moulin N (2002) The estimation of population differentiation with microsatellite markers. Mol Ecol 11:155–165
Broquet T, Petit EJ (2009) Molecular estimation of dispersal for ecology and population genetics. Annu Rev Ecol Evol Syst 40:193–216
Canada (2015) Historical Hydrometric Data Search. Government of Canada UhwegcssehshoAoF, 2016.).
Canada WSo (2013) HYDAT Database, Environment Canada. Available from https://www.ec.gc.ca/rhc-wsc/default.asp?lang=En&n=9018B5EC-1. Accessed 27 May 2015
COSEWIC (2010) COSEWIC assessment and status report on the Rocky Mountain Sculpin Cottus sp., Westslope populations, in Canada. Committee on the Status of Endangered Wildlife in Canada. Ottawa. x + 30 pp. (http://www.sararegistry.gc.ca/status/status_e.cfm).
Csardi G, Nepusz T (2006) igraph: The igraph software package for complex network research, InterJournal, Complex Systems. URL: http://igraph.org.
Dennenmoser S, Nolte AW, Vamosi SM, Rogers SM (2013) Conservation genetics of prickly sculpin (Cottus asper) at the periphery of its distribution range in Peace River, Canada. Conserv Genet 14:735–739
DFO (2013) Recovery potential assessment of Rocky Mountain Sculpin (Cottus sp.) eastslope populations in Alberta. Department of Fisheries and Oceans Canadian Science Advisory Secretariat SAR, Canada
Dudgeon D, Arthington AH, Gessner MO, Kawabata ZI, Knowler DJ, Leveque C, Naiman RJ, Prieur-Richard AH, Soto D, Stiassny MLJ, Sullivan CA (2006) Freshwater biodiversity: importance, threats, status and conservation challenges. Biol Rev 81:163–182
Engelke DR, Krikos A, Bruck ME, Ginsburg D (1990) Purification of thermus-aquaticus DNA-Polymerase expressed in Escherichia coli. Anal Biochem 191:396–400
Englbrecht CC, Largiader CR, Hanfling B, Tautz D (1999) Isolation and characterization of polymorphic microsatellite loci in the European bullhead Cottus gobio L-(Osteichthyes) and their applicability to related taxa. Mol Ecol 8:1966–1969
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Fall A, Fortin M-J, Manseau M, O’Brien D (2007) Spatial graphs: Principles and applications for habitat connectivity. Ecosystems 10:448–461
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Fiumera AC, Porter BA, Grossman GD, Avise JC (2002) Intensive genetic assessment of the mating system and reproductive success in a semi-closed population of the mottled sculpin, Cottus bairdi. Mol Ecol 11:2367–2377
Fraser DF, Gilliam JF, Daley MJ, Le AN, Skalski GT (2001) Explaining leptokurtic movement distributions: Intrapopulation variation in boldness and exploration. Am Nat 158:124–135
Fujishin LM, Barker FK, Huff DD, Miller LM (2009) Isolation of 13 polymorphic microsatellite loci for slimy sculpin (Cottus cognatus). Conserv Genet Resour 1:429–432
Fullerton DS, Colton RB, Bush CA, Straub AW (2004) Map showing spatial and temporal relations of mountain and continental glaciations of the northern plains, primarily in northern Montana and northwestern North Dakota. U.S. Department of the Interior, U.S. Geological Survey, Scientific Investigations Map 2843: 4 p. Available through http://pubs.usgs.gov/sim/2004/2843/. Accessed 10 Feb 2016
Galpern P, Rayfield B, Fall A, Manseau M (2014) grainscape: Grains of connectivity and minimum planar graph modelling of landscape connectivity (Windows only). R package version 0.3/r29. https://R-Forge.R-project.org/projects/grainscape/.
Goslee SC, Urban DL (2007) The ecodist package for dissimilarity-based analysis of ecological data. J Stat Softw 22:1–19
Goudet J (1995) FSTAT (Version 1.2): A computer program to calculate F-statistics. J Hered 86:485–486
Goudet J, Jombart T (2015) hierfstat: Estimation and Tests of Hierarchical F-Statistics. R package version 0.04–22. https://CRAN.R-project.org/package=hierfstat.
Grant EHC, Lowe WH, Fagan WF (2007) Living in the branches: population dynamics and ecological processes in dendritic networks. Ecol Lett 10:165–175
Hanski I (1998) Metapopulation dynamics. Nature 396:41–49
Heim KC, Wipfli MS, Whitman MS, Arp CD, Adams J, Falke JA (2015) Seasonal cues of Arctic grayling movement in a small Arctic stream: the importance of surface water connectivity. Environ Biol Fish 99:49–65
Hughes JM, Schmidt DJ, Finn DS (2009) Genes in streams: using DNA to understand the movement of freshwater fauna and their riverine habitat. Bioscience 59:573–583
Hughes JM, Huey JA, Schmidt DJ (2013) Is realised connectivity among populations of aquatic fauna predictable from potential connectivity? Freshw Biol 58:951–966
Jakober MJ, McMahon TE, Thurow RF, Clancy CG (1998) Role of stream ice on fall and winter movements and habitat use by bull trout and cutthroat trout in Montana headwater streams. Trans Am Fish Soc 127:223–235
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
James PMA, Cooke B, Brunet BMT, Lumley LM, Sperling FAH, Fortin MJ, Quinn VS, Sturtevant BR (2015) Life-stage differences in spatial genetic structure in an irruptive forest insect: implications for dispersal and spatial synchrony. Mol Ecol 24:296–309
Kalinowski ST, Taper ML (2006) Maximum likelihood estimation of the frequency of null alleles at microsatellite loci. Conserv Genet 7:991–995
Krejsa RJ (1967) The systematics of the prickly sculpin, Cottus asper Richardson, a polytypic species: part II. Studies on the life history, with especial reference to migration. Pacific Sci 21:414–422
Labbe TR, Fausch KD (2000) Dynamics of intermittent stream habitat regulate persistence of a threatened fish at multiple scales. Ecol Appl 10:1774–1791
Lamphere BA, Blum MJ (2012) Genetic estimates of population structure and dispersal in a benthic stream fish. Ecol Freshw Fish 21:75–86
Landguth EL, Bearlin A, Day CC, Dunham J (2016) CDMetaPOP: an individual-based, eco-evolutionary model for spatially explicit simulation of landscape demogenetics. Methods Ecol Evol. doi:10.1111/2041-210X.12608
Legendre P (2005) Code for t-test for independent samples with permutation test. http://adn.biol.umontreal.ca/~numericalecology/Rcode/
Legendre P, Legendre L (1998) Numerical Ecology, 2nd English Edn. Elsevier, Amsterdam
Lichstein JW (2007) Multiple regression on distance matrices: a multivariate spatial analysis tool. Plant Ecol 188:117–131
Lowe WH, Allendorf FW (2010) What can genetics tell us about population connectivity? Mol Ecol 19:3038–3051
McCauley DJ, Pinsky ML, Palumbi SR, Estes JA, Joyce FH, Warner RR (2015) Marine defaunation: animal loss in the global ocean. Science 347:8
McCleave JD (1964) Movement and population of the mottled sculpin (Cottus bairdi Girard) in a small Montana stream. Copeia 1964:506–513
McLachlan JS, Hellmann JJ, Schwartz MW (2007) A framework for debate of assisted migration in an era of climate change. Conserv Biol 21:297–302
McRae BH, Dickson BG, Keitt TH, Shah VB (2008) Using circuit theory to model connectivity in ecology, evolution, and conservation. Ecology 89:2712–2724
Meffe GK, Vrijenhoek RC (1988) Conservation genetics in the management of desert fishes. Conserv Biol 2:157–169
Murphy AL, Pavlova A, Thompson R, Davis J, Sunnucks P (2015) Swimming through sand: connectivity of aquatic fauna in deserts. Ecol Evol 5:5252–5264
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nolte AW, Stemshorn KC, Tautz D (2005) Direct cloning of microsatellite loci from Cottus gobio through a simplified enrichment procedure. Mol Ecol Notes 5:628–636
NRC (2012) Canadian Digital Elevation Model (CDEM). (ed. Government of Canada NRC, Earth Sciences Sector. http://geogratis.gc.ca/api/en/nrcan-rncan/ess-sst/C40ACFBA-C722-4BE1-862E-146B80BE738E.html. Accessed 20 Jun 2016)
Oksanen J, F. Guillaume Blanchet, R. Kindt, P. Legendre, P. R. Minchin, R. B. O’Hara, G. L. Simpson, P. Solymos, M. H. H. Stevens, Wagner H (2015) vegan: Community Ecology Package. In: R package version version 22–1 http://CRAN.R-project.org/package=vegan.
Paradis E (2010) pegas: an R package for population genetics with an integrated-modular approach. Bioinformatics 26:419–420
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539
Peres-Neto PR, Jackson DA (2001) How well do multivariate data sets match? The advantages of a Procrustean superimposition approach over the Mantel test. Oecologia 129:169–178
Petty JT, Grossman GD (2004) Restricted movement by mottled sculpin (pisces : cottidae) in a southern Appalachian stream. Freshw Biol 49:631–645
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
R Development Core Team (2016) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Radinger J, Wolter C (2014) Patterns and predictors of fish dispersal in rivers. Fish Fish 15:456–473
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Ruetz CR, Earl BM, Kohler SL (2006) Evaluating passive integrated transponder tags for marking mottled sculpins: effects on growth and mortality. Trans Am Fish Soc 135:1456–1461
Safner T, Miller MP, McRae BH, Fortin MJ, Manel S (2011) Comparison of bayesian clustering and edge detection methods for inferring boundaries in landscape genetics. Int J Mol Sci 12:865–889
Schwalb AN, Poos MS, Ackerman JD (2011) Movement of logperch-the obligate host fish for endangered snuffbox mussels: implications for mussel dispersal. Aquat Sci 73:223–231
Selkoe KA, Srcribner KT, Galindo HM (2016) Waterscape genetics - applications of landscape genetics to rivers, lakes, and seas. In:Balknhol N, Cushman SA, Storfer AT, Waits LP (eds) Landscape genetics: Concepts, methods, applications, 1 edn. Wiley, Chichester, pp. 264
Spear SF, Balkenhol N, Fortin MJ, McRae BH, Scribner K (2010) Use of resistance surfaces for landscape genetic studies: considerations for parameterization and analysis. Mol Ecol 19:3576–3591
Wagner HH, Fortin MJ (2013) A conceptual framework for the spatial analysis of landscape genetic data. Conserv Genet 14:253–261
Wang IJ, Bradburd GS (2014) Isolation by environment. Mol Ecol 23:5649–5662
Watkinson DA, Boguski DA (2013) Information in support of a recovery potential assessment of Rocky Mountain Sculpin (Cottus sp.), Eastslope populations, in Alberta. Canadian Science Advisory Secretariat, Ottawa
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evol Int J Org Evol 38:1358–1370
Acknowledgements
The authors would like to acknowledge individuals part of the field sampling team, including: Wesley Donaldson, Elliot Macdonald, Kelly Mulligan, Caitlin Good, Christine Lacho, Troy Adams, Kenton Neufeld, Elashia Young, and Denyse Dawe. Thanks are also due to Fisheries and Oceans Canada Species at Risk group for funding to MP and DW.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ruppert, J.L.W., James, P.M.A., Taylor, E.B. et al. Riverscape genetic structure of a threatened and dispersal limited freshwater species, the Rocky Mountain Sculpin (Cottus sp.). Conserv Genet 18, 925–937 (2017). https://doi.org/10.1007/s10592-017-0938-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-017-0938-6