Abstract
For several centuries, game management has involved translocations of non-native individuals of many species to reinforce local native populations. However, there are few quantitative studies of potentially negative effects on population viability as expected when taxa with different local adaptations hybridise. The European red deer has been subject to particularly many translocations. Around 1900, a total of 17 red deer of Hungarian (Cervus elaphus hippelaphus) and German (C. e. germanicus) origin were introduced onto the island of Otterøya in Norway where few native red deer (C. e. atlanticus) remained (n ~ 13). To assess interbreeding, the present stock on Otterøya and the indigenous Norwegian and Hungarian populations were characterised in 14 microsatellite loci and in the control region of mtDNA. An intermediate level of genetic variation in the Otterøya population and the presence of population specific alleles from both the indigenous Norwegian and the Hungarian population demonstrate that the introduced red deer interbred with the native. Even distributions of one indigenous and one non-indigenous mtDNA haplotype in the Otterøya population and two point estimates of admixture indicate similar genetic contributions from the two parental populations into the hybrid stock. Low numbers of migrants identified with Bayesian assignment tests demonstrate low recent gene flow from Otterøya into the Norwegian mainland population. The Otterøya hybrid stock has grown vastly in numbers during recent decades, suggesting a high population viability. We observed that the body mass of red deer on Otterøya was similar or greater than in adjacent indigenous Norwegian stocks, indicating that population performance has not been reduced in the hybrid stock and that gene flow probably has not had any negative effects.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Species are distributed along environmental gradients (Begon et al. 1996), and gene frequencies may change locally when adaptations develop in populations by natural selection (Endler 1992; Strickberger 1996). However, locally adapted populations admix to an increasing degree because of range shifts in many species associated with the present use of land and recent climatic changes, especially in temperate areas (IPCC 2001, 2007). In addition, hybridisation rates have increased worldwide because of human-mediated translocations and habitat modifications, causing the extinction of native species, subspecies and locally adapted populations (Allendorf et al. 2001). When genetically different taxa hybridise, local adaptations may be lost from the native taxa by introgression of non-indigenous alleles and loss of local alleles and co-adapted gene complexes (Rhymer and Simberloff 1996; Barton 2001; Burke and Arnold 2001). Gene flow between populations in different environments may therefore constrain local adaptation and lower the short-term fitness of native populations (Storfer 1999). Alternatively, increasing levels of genetic variation from an isolate break (Hartl and Clark 1997) may have positive consequences for population viability through heterosis effects or reduced inbreeding depression (Frankham 1995; Coulson et al. 1998), depending on the genetic divergence of the hybridising taxa (Allendorf et al. 2001; Freeland 2005).
During recent centuries, game management has involved translocations of non-native individuals of many species into former habitats or native populations (Griffith et al. 1989; Hartl 1991; Wolf et al. 1996; Fischer and Lindenmayer 2000; DeYoung et al. 2003; Kruckenhauser and Pinsker 2004). In Europe translocations have been especially common among red deer (Cervus elaphus) populations to re-establish or reinforce local populations and avoid local extinction (Strandgaard and Simonsen 1993; Hartl et al. 1995; Zachos et al. 2003) or to transfer desirable traits for trophy hunters (Hartl et al. 2003). Many of these populations are morphologically different and have been described as separate subspecies (Lønnberg 1906; Whitehead 1972; Whitehead 1993). Many are genetically differentiated (Gyllensten et al. 1983; Kuehn et al. 2003), and even though some argue for one common European subspecies (Groves and Grubb 1987; Polziehn and Strobeck 2002), western and eastern (including Hungarian) European red deer form separate mtDNA haplogroup lineages (Ludt et al. 2004; Skog et al. 2009). The impact of such translocations should be evaluated, considering the growing knowledge of ungulate population genetics and phylogeography (Randi 2005).
The red deer have existed in Norway at least since the sub-boreal period (Ahlèn 1965) and low levels of genetic variation documented by allozyme and microsatellite analyses suggest long-time isolation and previous bottlenecks (Gyllensten et al. 1983; Røed 1998; Haanes et al. 2010). Historically, red deer were distributed across most of southern Norway (Friis 1874; Collett 1877) but after 1750 a major decline limited the population to only a few locations along the west coast (Collett 1909; Ingebrigtsen 1924). To our knowledge only one translocation of non-indigenous red deer into the Norwegian population has occurred in recent times. On the island Otterøya in the northern range of its distribution, the local red deer stock was almost extirpated in 1898, counting only 12–14 individuals including three or four stags (Collett 1909; Collett 1912). To avoid local extinction, 17 captive red deer of a cross breed between the German red deer and the plain—living Hungarian red deer (C. e. hippelaphus), including at least one stag, were translocated from the royal enclosure in Thüringen in central Germany to Otterøya and into the native Norwegian subspecies (C. e. atlanticus) from 1900 to 1903 (Die-Woche 1902; Collett 1909; Finsberg 1934). Among the potential parental origins for the Otterøya stock, the Norwegian, German and Hungarian red deer populations lie at very different latitudes (northern boundaries at 64.5, 54 and 48 N), and offer quite different climates and habitats (Mysterud et al. 2002; Welzholz et al. 2005; Matrai et al. 2004). Further, the western and eastern European red deer lineages are separated by the barriers provided by the Alps and the Carpathians (Hartl et al. 1995; Ludt et al. 2004; Skog et al. 2009), and the introduced German/Hungarian red deer are therefore presumably a mix of western and eastern lineage. Ten years after, the Otterøya population counted 100 individuals (Collett 1909) and since the 1930’s culls have increased considerably to an annual cull of 319 in 2006. Both Hungarian and German red deer are reported to have larger body and antler sizes than the Norwegian subspecies (Lønnberg 1906; Collett 1909; Haigh and Hudson 1993).
We have quantified the level of gene flow between native and introduced continental red deer originating from each of the two major lineages in the European red deer. The success of the German/Hungarian cross was evaluated from the genetic impact on the Otterøya population, and recent gene flow into the mainland population was estimated. We have also assessed the performance of red deer from the mixed stock on the island Otterøya compared with pure stocks from both the mainland and another island (Hitra) in the region. We used body mass as a proxy for performance, which for Norwegian red deer is closely correlated with age of first reproduction (Langvatn et al. 2004) and survival during the first critical winter (Loison et al. 1999).
Materials and methods
Study area
The island of Otterøya (64.5°N and 11.3°E, 143 km2) is separated from the mainland by 1–2 km wide sounds that narrows to a few hundred metres in some places. Annual mean precipitation is 1,440 mm and annual mean temperature is 3.7°C. A large area is situated above the tree line (~400 m) and an area of 98.5 km2 qualifies as suitable red deer habitat. Boreal rainforest characterised by Norway spruce (Picea abies) and Scots pine (Pinus Sylvestris) dominates and cultivated areas are scattered (~4 km2). The island constitutes winter pasture for 600 reindeer and has scarce stocks of roe deer (Capreolus capreolus) and moose (Alces alces).
Sampling and genetic analyses
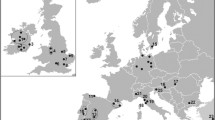
Between 2001 and 2003 muscle tissue was sampled from 20 Hungarian red deer, 40 red deer from Otterøya and 136 red deer from adjacent (mainly mainland) areas in Norway. Since most animals were shot by local hunters, few reliable body mass estimates were available. We were unfortunately not able to attain any samples from Germany. The German population has been subject to bottlenecks and translocations (e.g. Kuehn et al. 2003; Zachos et al. 2007) and to obtain contemporary samples representative of the original indigenous German part of the introduced cross breed may not be possible. In Hungary, wild red deer were sampled in the north at Valkó close to Gödöllö (n = 6), and in the south at Hajósszentgyörgy close to the Gemenc forest (n = 7) and at Vajszló south of the city Pécs (n = 7, Fig. 1). The Valkó population is spatially separate from the two other sampling sites but 40 years ago several individuals were translocated to this area from the south (pers. comm. Làszló Szemethy, St. Stephen University, Hungary). Indigenous Norwegian red deer were sampled for genetic analyses in six municipalities adjacent to Otterøya (Fig. 1), which are 83–236 km (mean = 151; SE = 22) from Otterøya. The genetic variation of these Norwegian municipalities, except P4-1721, has been previously described (Haanes et al. 2010).
Genomic DNA was isolated from whole blood and muscle tissue (Qiagen DNeasy KIT). We selected 14 polymorphic microsatellite loci that show Mendelian heredity in Norwegian red deer (Haanes et al. 2005). These were CSSM03 (Moore et al. 1994), OarCP26 (Ede et al. 1995), RT5 (Wilson et al. 1997), SRCRSP10 (Bhebhe et al. 1994), NVHRT73 and NVHRT48 (Røed and Midthjell 1998), McM58 (Hulme et al. 1994), OarFCB193 and OarFCB304 (Buchanan and Crawford 1993), BM5004, BM888, BMC1009, BM4208 and BM4107 (Bishop et al. 1994). They were amplified on a GeneAmp PCR System 9600 (Applied Biosystems) in 10 μl reaction mixtures with 30–60 ng of genomic template DNA, 2 pmol of each primer, 50 mM KCl, 1.5 mM MgCl2, 10 mM Tris–HCl, 0.2 mM dNTP, and 0.5 U of AmpliTaq DNA polymerase (Applied Biosystems). After denaturation at 94°C for 5 min, 30 cycles of amplification with 1 min at 95°C, 30 s at 55°C and 1 min at 72°C were followed by 10 min extention at 72°C. The PCR products were then separated by size with capillary electrophoresis (ABI310) and electromorphs genotyped with GENOTYPER1.1.1 (both Applied Biosystems).
For a subsample of each population (n = 18 and n = 16 and from Hungary and Otterøya, respectively, and n = 17 from Norway, including two areas south of the sample range) a 463 base pair region of the mitochondrial D-loop adjacent to the tRNA pro gene was amplified using the primers 5′-AATAGCCCCACTATCAGCACCC (L15394) and 5′-TATGGCCCTGAAGTAAGAACCAG (H15947) (c.f. Flagstad and Roed 2003). Thirty-five cycles of amplification with 30 s at 94°C, 30 s at 60°C and 45 s at 72°C were preceded by a 2 min pre-denaturation step and followed by a final 7 min extension. Amplifications were performed in 10 μl volumes containing 1.5 mM MgCl2, 200 μM of each dNTP, 4 pmol of each primer and 0.5 units of AmpliTaq DNA polymerase (Applied Biosystems). PCR products were purified using ExoZapitTM (Amersham Biosciences). Sequencing was performed using BigDye terminator cycle sequencing chemistry on an ABI 3100 instrument, and sequences aligned manually using SeqScape version 2.0 (Applied Biosystems). The sequences have been reported to Genbank (accession nos. GU369821–GU369871).
Statistical analyses of genetic data
Random mating within populations was assessed by exact tests of Hardy–Weinberg equilibrium across the 14 microsatellite loci using GENEPOP 3.4 with default settings (Raymond and Rousset 1995). Significance levels were sequentially Bonferroni adjusted for repeated tests (Rice 1989). To investigate genetic variation we calculated the number of private alleles, allele richness (El Mousadik and Petit 1996) and gene diversity (Nei 1987) in each population across loci using FSTAT 2.9.3 (Goudet 2001). Genetic variation in the control region of mtDNA was expressed as haplotype number, haplotype diversity (h) and nucleotide diversity (Π), calculated using ARLEQUIN 2.000 (Schneider et al. 2000). The observed sequences were blasted in Genbank and compared with sequences from red deer sampled in Germany (DQ520209, DQ520210, DQ520214 and DQ520226; Skog et al. 2009) and Hungary (EU4367XX–EU4368XX; personal communication, Dr. Zsolt Pádár, Institute of Forensic Sciences, Ministry of Justice and Law Enforcement, Hungary), and a neighbour joining tree was built using MEGA 4.0 (Tamura et al. 2007).
For the microsatellite data, genetic structure within (Fis) and among (Fst) populations (Weir 1996) was assessed using FSTAT with sequential Bonferroni adjustment. Genetic distances, DA (Nei et al. 1983), among the populations (with Norwegian localities separated) were calculated and a neighbour-joining tree built with 1,000 bootstraps on loci using POPULATIONS (Langella 2002). The tree was visualised using TREEVIEW1.6.6 (Page 1996).
To address the degree of interbreeding between the continental red deer and the native island population at the time of the introduction we used the microsatellite data and estimated the proportionate admixture from two of the parental populations into the Otterøya population using present day Norwegian and Hungarian populations as representatives. We used ADMIX1.0 (Bertorelle and Excoffier 1998) to calculate bootstrap estimates of two admixture estimators (1,000 replicates), the allele frequency based mC (Chakraborty et al. 1992) and the coalescent based mY (Bertorelle and Excoffier 1998). To include the possible affect of genetic drift on the admixture estimates we also estimated admixture using LEA (Chikhi et al. 2001) with 200,000 Monte Carlo Marcov Chain (MCMC) iterations.
To assess the genetic structure of the data set, and to address recent gene flow between the Norwegian mainland and Otterøya, we used the microsatellite data and Bayesian individual assignment as implemented in STRUCTURE2.3 (Pritchard et al. 2000). For each of a different number of genetic clusters (K ∈ [1,11]), a model with uniform priors, an admixture model (α = 1, αmax = 50), correlated allele frequencies (Falush et al. 2003), allowing missing values to be interpreted as null alleles (RECESSIVE ALLELES = 1), 100,000 burnins cycles and 500,000 MCMC iterations was run 10 times. Delta K, which is negatively related to the increasing variance among repeated runs and often increasing posterior probabilities with higher K values, was used to determine the main genetic structure of the data set (Evanno et al. 2005). Dispersers were identified from individuals assigned to clusters other than the one corresponding with where they had been sampled.
Data on red deer body mass
The data on red deer performance derive from a separate data set from the annual Norwegian autumn harvest lasting from 10 September to 15 November in the period 1965 to 2006, for which we do not have genetic data. The date and location (municipality) of harvest together with biological information on sex, body mass and the mandibles for each of 20,161 deer were provided by hunters. Body mass was recorded as dressed mass, which is live mass minus head, skin, viscera, bleedable blood and metapodials, constituting about 58% of live mass and highly correlated with total mass. Using the mandibles, calves and yearlings were aged based on patterns of tooth eruption (Loe et al. 2004). Older animals were aged using annuli in the cementum of the first incisor (Hamlin et al. 2000). The data used are a subset of a larger dataset from the whole of the southwest coast (e.g., Mysterud et al. 2001a, b; Pettorelli et al. 2005). A subset of nine municipalities was selected according to distance from the focal area of Otterøya (Fig. 1). Harvest data may be prone to bias in some cases due to hunter selectivity (see in depth discussion in Mysterud et al. 2008). However, as the tradition for hunting is similar between our focal areas, this is unlikely to be important for the spatial contrast of focus here.
We have good knowledge of the performance of these populations and how they have been affected by density dependence and by climate (the North Atlantic Oscillation (NAO); Mysterud et al. 2001a). As a measure for density, we used the proxy “number of harvested animals” per km2 of red deer habitat (Table S1). Despite this being a crude index, the increase in harvest has been fivefold due to a huge density increase, and over time the measure has shown to correlate well with other direct measures of density (cfr. Mysterud et al. 2007). As an index of climate, we used the station-based winter (Dec-Mar) index of Hurrell (1995).
Statistical analyses of body mass data
We analysed variation in (ln) body mass of red deer with a combination of additive and linear models. Based on previous results, we ran separate models on males and females due to their strongly different life histories (Mysterud et al. 2001b). In addition, it is particularly important to model the age effect correctly, since so much of the variation is found in that parameter. Therefore, and since we are not directly interested in the age effect, we tried both age as a class variable (0, 1, 2, 3, 4, ≥5 years; providing good fit for females, Mysterud et al. 2001b) and modelled the age effect with smoothing splines (for males; Yoccoz et al. 2002), which are very flexible, using the library (mgcv) in (Wood 2006). Similarly, for any growth or decay during the autumn, we used a spline for the “date of harvest” effect to make sure this did not bias our results. We allowed the program to choose the appropriate number of spline knots. Due to age-dependent effort in the rutting of males, we ran this as an interaction between age and date of harvest (cfr. Yoccoz et al. 2002).
Our focal factor is the spatial contrasts between Otterøya and adjacent areas. We therefore used “Treatment” contrasts, i.e., comparing levels of a factor with one specific level—a reference level (the Otterøya stock). However, environmental conditions may not be comparable, so that any spatial difference may not be due to genetic effects, but rather reflect either density or habitat quality. We therefore entered the density index (described above) and the NAO (climate index) to control for annual fluctuations.
Results
Genetic variation
In each investigated population all microsatellite loci were in Hardy–Weinberg equilibrium after sequential Bonferroni adjustment, except BM4208 in the Hungarian population and BM5004 in the Hungarian and Otterøya populations. Across the 14 microsatellite loci, 151 alleles were found in the three investigated populations. Among these, 73 alleles were population specific for either the indigenous Norwegian (7), Otterøya (13) or Hungarian (56) population (Tables 1 and 2). Another 25 microsatellite alleles found in the Otterøya population were shared with only either the Hungarian (16) or the Norwegian (9) population (Table 1), strongly suggesting inheritage from both populations. The Hungarian red deer had by far the highest gene diversity, allele richness and number of private alleles, the Otterøya population was intermediate, while the indigenous Norwegian population was the least genetically variable (Table 2).
All together, 13 different mtDNA haplotypes were found; two on Otterøya, five in the indigenous Norwegian population and seven in the Hungarian population (Table S2). The nucleotide diversity was intermediate in the Otterøya population while the haplotype diversity was the lowest (Table 2). The two haplotypes in the Otterøya stock were evenly distributed (nA = 9, nB = 7) and both most probably belong to the western mtDNA lineage of European red deer. One of the Otterøya haplotypes was also found in the municipality P3-1563 in the indigenous Norwegian population (nA = 4). It matched a Genbank entry (AF291888), and only differed in one base compared to another indigenous Norwegian haplotype. The other Otterøya haplotype was indigenous to the island, but differed only in two succeeding bases (G–T vs. T–G) from a German sequence (DQ520226) and in two other bases from an observed Norwegian haplotype. Even though it also only differed in two separate base pairs from one Hungarian sequence (EU436831), it differed from all the other Hungarian haplotypes in more than seven different base pairs. Several matches were found in Genbank for one of the other Norwegian haplotypes (e.g. EF636730), as well as for four of the Hungarian haplotypes (e.g.; EU436833, EU436837, EU436838, EU436788). All sampled Hungarian haplotypes except one had from one to three Thymine inserts and grouped out with high bootstrap support (99) in a neighbour-joining tree (not shown), separate from all the haplotypes sampled in North West Europe (including both Otterøya sequences), among which there was no obvious pattern.
The microsatellite data demonstrated limited gene flow and strong genetic structure from significant Fst values between the Otterøya population and both the Hungarian (0.13) and the indigenous Norwegian population (0.19). Similarly, long genetic distances (DA) of 0.40 and 0.33, respectively, showed that the Otterøya population was genetically different and intermediate to both investigated parental populations (Fig. 2). By comparison, a higher Fst value (0.23) and a longer genetic distance (0.48) was found between the Norwegian and Hungarian populations. Within the indigenous Norwegian population much shorter genetic distances (Fig. 2) and lower Fst values (mean = 0.03, SD = 0.02) indicate low genetic structure. An inbreeding coefficient that was not significantly different from zero indicated that there was little genetic structure within the Otterøya population.
The bootstrap estimate of admixture based on microsatellite allele frequencies indicated an even admixture between indigenous Norwegian and Hungarian red deer into the Otterøya population with proportions of mC = 0.55 (SE = 0.06) and mC = 0.45 (SE = 0.06) admixed from the Norwegian and Hungarian populations, respectively. The coalescent based estimator indicated a higher proportion admixed from the Norwegian population with mY = 0.69 (SE = 0.05) and a lower proportion from the Hungarian population with mY = 0.31 (SE = 0.04). The admixture estimate including genetic drift showed more skewed proportions, with 0.85 from the Norwegian (P1) and 0.1 from the Hungarian population (Fig. 3).
In the STRUCTURE analysis, a pronounced higher delta K value for K = 2 suggest that the main genetic structure of the data set is a dichotomy (Table S3), with the Hungarian and Otterøya samples assigned to one cluster and the indigenous Norwegian samples to the other cluster. A slightly higher posterior probability for K = 4 reflects an underlying lower hierarchical structure between; 1) the Otterøya hybrid stock and the Norwegian population, and 2) within the Norwegian population (Table S3 and Fig. 4). The STRUCTURE analyses (K = 2) show a very limited recent gene flow between the Norwegian mainland and Otterøya populations, with all individuals assigned with high probabilities to the cluster corresponding to where they had been sampled, except for three individuals. Two of the individuals sampled on Otterøya were assigned to the Norwegian cluster (0.55 < q < 0.99) while one individual sampled in the Norwegian population was assigned to the Otterøya/German/Hungarian cluster (P = .69). These are most probably first or second generation migrants.
Body mass variation
The models of female (r2 = .853) and male (r2 = .724) body mass variation gave largely similar results concerning the spatial contrast, after controlling for the strong effects of age, date of harvest, density and the NAO (Table 3). Deer from Otterøya were larger than deer from coastal municipalities in Sør-Trøndelag (population P3; municipality no. 1,612, 1,613 and 1,622) including the island of Hitra (population P5), but not larger than all the inland municipalities (municipality no. 1,635 and 1,636). Body mass of red deer from Otterøya was comparable to body mass of those from the mainland municipalities (municipality no. 1,714 and 1,721) in Nord-Trøndelag (population P4). The interaction term between age and municipality could not be entered due to unbalanced data, however, similar differences between municipalities were obtained when only analysing variation for a specific age class (yearlings).
Discussion and conclusions
This study demonstrates that the German/Hungarian red deer introduced onto Otterøya around 1900 interbred with the resident native Norwegian population. Variation in both microsatellite and mtDNA demonstrate that it is a genetic intermediate with heritage from both the native Norwegian and the Hungarian population. Extensive interbreeding between these presumed subspecies (Lønnberg 1906; Whitehead 1972; Whitehead 1993) was evident from the estimators of admixture and from the similar frequencies of the two haplotypes found on Otterøya. However, any negative effects of introgression that may have been expected were not observed in either population growth or body mass. Rather the hybrid stock has grown tremendously in numbers during recent decades and the observed body mass of red deer on Otterøya was similar or larger than those of indigenous Norwegian inland and coastal localities.
Potential limitations and biases on the admixture estimates
For the microsatellite data, the allele frequency-based estimator yielded close to even proportions of admixture from the indigenous Norwegian and Hungarian populations. However, frequency-based estimators are often biased towards even proportions compared to the coalescent-based estimator MY, which also incorporates molecular divergence between alleles and parental populations (Bertorelle and Excoffier 1998; Wang 2003). The coalescence-based estimator yielded skewed proportions of admixture, as could be expected from influence by the unaccounted German part of the introduced red deer cross. Considering that the resident population on Otterøya had a similar size and sex ratio as the group of introduced German/Hungarian red deer (Collett 1909; Ingebrigtsen 1924; Finsberg 1934), the German part would constitute one quarter of admixture with free interbreeding. This is roughly proportionate with and could help explain the deviations from even admixture in the coalescence-based estimator and the genetic drift model (LEA). The private alleles found only on Otterøya probably represent the German contribution but, considering the relatively low sample size, could also be of Hungarian origin. The private alleles and the alleles shared only by Otterøya and Hungarian samples may also explain why these cluster together with K = 2 in the STRUCTURE analysis, since private alleles and large allele frequency differences have substantial effects on the clustering of STRUCTURE (Rosenberg et al. 2001; Musani et al. 2006). The even frequencies of the two mitochondrial haplotypes on Otterøya, which were of Norwegian and non-indigenous origin, provided support for an even admixture. The non-indigenous haplotype was relatively similar to both known German and Hungarian sequences, but since it fell together with the previously described West European haplogroup rather than the East European haplogroup (Ludt et al. 2004; Skog et al. 2009), it most likely represents a German lineage in the introduced cross breed. The even admixture in both mtDNA and microsatellite data further indicates similar contributions from both sexes. In spite of high Fst values and long genetic distances between the parental populations, our data support an even admixture and free interbreeding between the introduced German/Hungarian red deer and the native Norwegian population.
Performance of the hybrid stock
Hybridisation among subspecies may have different outcomes. It is rare in some species (Hansen 2002; Lorenzen and Siegismund 2004; Storfer et al. 2004), but happen to various degrees in others (Jensen et al. 2005). Hybridisation is common within the red deer species complex (Hartl et al. 1995; Tate et al. 1998; Hartl et al. 2003), but negative effects are known between genetically and morphologically different taxa like the wapiti and the red deer (Asher et al. 2005). Examples from other species involve loss of local native genetic variation (Shaw et al. 1992; Latch et al. 2006), loss of native adaptations (Storfer 1999), and reduced fitness (Allendorf et al. 2004). However, intentional translocations of species for conservation and management reasons have been common, and have yielded positive results in many species (Griffith et al. 1989; Fischer and Lindenmayer 2000), for example in the Scandinavian wolf (Vila et al. 2003), wild boar (Vernesi et al. 2003) and the Florida Panther (Hedrick 2001). The Norwegian, German and Hungarian red deer populations are located at different latitudes (northern boundaries at 64.5, 54 and 48°N) and genetic variation indicates that the indigenous Norwegian population has been isolated for a long time (Gyllensten et al. 1983; Røed 1998). It therefore seems reasonable to assume that these populations may have developed different local adaptations. However, the negative effects on population viability that may be expected when taxa with different local adaptations hybridise (Rhymer and Simberloff 1996; Allendorf et al. 2001; Burke and Arnold 2001) were not observed in either the population growth or the body mass of the Otterøya hybrid stock. Even though this natural experiment does not offer an adequate evolutionary time frame, our results support the high phenotypic plasticity suggested for red deer (Geist 1998; Lister 2004), rather than different local adaptations in different environments. Ten years after the introduction the Otterøya population counted 100 individuals (Collett 1909). Since then annual culls have increased considerably, reaching 319 in 2005 and 2006. This follows the general trend of expansion of the Norwegian population (Forchhammer et al. 1998; Langvatn, 1998), which is partly explained by climatic variation (Forchhammer et al. 1998; Mysterud et al. 2001a), changes in human harvesting (Langvatn and Loison 1999) and the altered use of agricultural land (Ahlèn 1965; Mysterud et al. 2002). The increase in population size of the hybrid stock could also reflect positive effects on population viability, as could be expected from heterosis and reduced inbreeding involved with hybridisation of more closely related taxa (Haig 1998; Freeland 2005). The increased level of genetic variation in the small initial founding population on Otterøya may thus have prevented the negative effects of inbreeding and counteracted loss of genetic variation from random genetic drift. In red deer, heterosis effects have been documented as both increased lifetime reproductive success and calf body mass (Coulson et al. 1998; Slate et al. 2000), and could explain the heavier body mass on Otterøya compared to most indigenous localities. However, German and Hungarian red deer have a relatively larger body mass than Norwegian red deer (Lønnberg 1906; Collett 1909; Haigh and Hudson 1993) and may indicate effects from additive genetic variation. On the other hand, much geographic variation in body size is attributable to phenotypic plasticity affected by habitat and nutrition (Lister 1984; Geist 1998), as demonstrated by the huge increase in body size and antlers of west European red deer after translocation to New Zealand (Huxley 1931). Further, red deer body mass is generally strongly negatively related to density (Mysterud et al. 2001b) and the higher density on Otterøya may have obscured differences to the inland localities with apparently similar body mass. These comparisons were difficult because of the lack of adequate data on habitat quality, as density relative to resource levels is expected to determine body mass.
Implications for management
Generally, management is concerned with conservation of local biodiversity and indigenous genetic variation (Rhymer and Simberloff 1996; Storfer 1999; Allendorf et al. 2001). Even though no first-generation migrants from the Otterøya population were detected in the mainland population in 2001 and 2002, we observed very low frequencies of some alleles that were only common in the Hungarian and Otterøya populations. Otterøya is separated from the mainland by only a 200-300 meter wide sound, and these alleles are probably the result of introgression into the mainland population during 30 generations. Considering the recent range shifts of many species (IPCC 2001; IPCC 2007), and the population expansion of the Norwegian red deer last century (Forchhammer et al. 1998; Langvatn 1998), some dispersal from the Otterøya seems very likely. Until the effects of heterosis (positive) on Otterøya have been further addressed, it will be quite challenging to decide how to manage the Otterøya hybrid stock. On the one hand it can be argued that the genetically different hybrid population on Otterøya, with its higher genetic diversity, should be allowed to expand and interbreed with the native mainland population to increase its level of genetic variation. On the other hand, the Norwegian population might be one of the few European populations not affected by translocations, and it is probably better to be safe than sorry by preventing any spreading and interbreeding with the indigenous Norwegian population.
References
Ahlèn I (1965) Studies on the red deer, Cervus elaphus L. Scandinavia. III. Ecological investigations. Viltrevy 3:177–376
Allendorf FW, Leary RF, Spruell P, Wenburg JK (2001) The problems with hybrids: setting conservation guidelines. TREE 16:613–622
Allendorf FW, Leary RF, Hitt NP, Knudsen KL, Lundquist LL, Spruell P (2004) Intercrosses and the US Endangered Species Act: should hybridized populations be included as Westslope cutthroat trout? Cons Biol 18:203–1213
Asher GW, Archer JA, Scott IC, O’Neill KT, Ward J, Littlejohn RP (2005) Reproductive performance of pubertal red deer (Cervus elaphus) hinds: effects of genetic introgression of wapiti subspecies on pregnancy rates at 18 months of age. Anim Reprod Sci 90:287–306
Barton NH (2001) The role of hybridization in evolution. Mol Ecol 10:551–568
Begon M, Harper JL, Townsend CR (1996) Ecology, individuals, populations and communities, 3rd edn. Blackwell Science, Oxford
Bertorelle G, Excoffier L (1998) Inferring admixture proportions from molecular data. Mol Biol Evol 15:1298–1311
Bhebhe E, Kogi J, Holder DA et al (1994) Caprine microsatellite dinucleotide repeat polymorphism at the SR-CRSP-6, SR-CRSP-7, SR-CRSP-8, SR-CRSP-9 and SR-CRSP-10. Anim Genet 25:203
Bishop MD, Kappes SM, Keele JW et al (1994) A genetic linkage map for cattle. Genetics 136:619–639
Buchanan FC, Crawford AM (1993) Ovine microsatellites at the OarFCB11, OarFCB128, OarFCB193, OarFCB226 and OarFCB304 loci. Anim Genet 24:145
Burke JM, Arnold ML (2001) Genetics and the fitness of hybrids. Ann Rev Gen 35:31–52
Chakraborty R, Kamboh MI, Nwankwo M, Ferrell RE (1992) Caucasian genes in American blacks: new data. Am J Hum Genet 50:145–155
Chikhi L, Bruford MW, Beaumont MA (2001) Estimation of admixture proportions: a likelihood-based approach using Markov chain Monte Carlo. Genetics 158:1347–1362
Collett R (1877) Bemærkninger til Norges pattedyrsfauna (in Norwegian). Nyt Magazin for Naturvidenskaberne 22:93–133
Collett R (1909) Hjorten i Norge (Cervus elaphus atlanticus), nogle biologiske meddelelser (in Norwegian). Bergens museums Aarbok 6:9–31
Collett R (1912) Norges pattedyr (in Norwegian). H. Aschehoug and Co, Kristiania
Coulson TN, Pemberton JM, Albon SD et al (1998) Microsatellites reveal heterosis in red deer. Proc R Soc Lond B 265:489–495
DeYoung RW, Demarais S, Honeycutt RL et al (2003) Genetic consequences of white-tailed deer (Odocoileus virginianus) restoration in Mississippi. Mol Ecol 12:3237–3252
Die-Woche (1902) Rochwildtransport nach Norwegen (in German), Berlin, pp 1111–1113
Ede AJ, Pierson CA, Crawford AM (1995) Ovine microsatellites at the OarCP9, OarCP16, OarCP20, OarCP21, OarCP23 and OarCP26 loci. Anim Genet 25:129–130
El Mousadik A, Petit RJ (1996) High level of genetic differentiation for allelic richness among populations of the argan tree (Argania spinosa L. Skeels) endemic to Morocco. Theor Appl Genet 92:832–839
Endler JA (1992) Genetic heterogeneity and ecology. British Ecological Society 33:315–332
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Finsberg O (1934) Verdens nordligste hjortestamme (in Norwegian). NJFF tidskrift 63:104–164
Fischer J, Lindenmayer DB (2000) An assessment of the published results of animal relocations. Biol Cons 96:1–11
Flagstad O, Roed KH (2003) Refugial origins of reindeer (Rangifer tarandus L.) inferred from mitochondrial DNA sequences. Evolution 57:658–670
Forchhammer MC, Stenseth NC, Post E, Langvatn R (1998) Population dynamics of Norwegian red deer: density-dependence and climatic variation. Proc R Soc Lond B 265:341–350
Frankham R (1995) Conservation genetics. Ann Rev Genet 29:305–327
Freeland JR (2005) Molecular Ecology. Wiley, Chichester
Friis JA (1874) Tilfjelds i ferierne (in Norwegian). Cammermeyer, Christiania
Geist V (1998) Deer of the world: their evolution, behaviour, and ecology. Swan Hill Press, UK
Goudet J (2001) FSTAT, a program to estimate and test gene diversities and fixation indices. http://www.unil.ch/izea/softwares/fstat.html
Griffith B, Scott JM, Carpenter JW, Reed C (1989) Translocation as a species conservation tool—status and strategy. Science 245:477–480
Groves CP, Grubb P (1987) Relationships of living deer. In: Wemmer CM (ed) Proceedings of Biology and management of the Cervidae. Smithsonian Institution Press, Washington
Gyllensten U, Ryman N, Reuterwall C, Dratch P (1983) Genetic differentiation in four European subspecies of red deer (Cervus elaphus L.). Heredity 51:561–580
Haanes H, Rosef O, Veiberg V, Røed KH (2005) Microsatellites with variation and heredity applicable to parentage and population studies of Norwegian red deer (Cervus elaphus atlanticus). Anim Genet 36:454–455
Haanes H, Røed KH, Flagstad Ø, Rosef O (2010) Genetic structure in an expanding cervid population after population reduction. Conserv Genet 11:11–22. doi:10.1007/s10592-008-9781-0
Haig SM (1998) Molecular contributions to conservation. Ecology 79:413–425
Haigh JC, Hudson RJ (1993) Farming wapiti and red deer. Mosby, St. Louis
Hamlin KL, Pac DF, Sime CA, DeSimone RM, Dusek GL (2000) Evaluating the accuracy of ages obtained by two methods for Montana ungulates. J Wildl Manage 64:441–449
Hansen MM (2002) Estimating the long-term effects of stocking domesticated trout into wild brown trout (Salmo trutta) populations: an approach using microsatellite DNA analysis of historical and contemporary samples. Mol Ecol 11:1003–1015
Hartl GB (1991) The influence of game management on allelic variation in large mammals of Central Europe. Supplemento alle Ricerche de biologia della Selvaggina XVIII:95–108
Hartl DL, Clark AG (1997) Principles of population genetics, 3rd edn. Sinauer Associates, Sunderland
Hartl GB, Nadlinger K, Apollonio M et al (1995) Extensive mitochondrial-DNA differentiation among European red deer (Cervus-Elaphus) populations—implications for conservation and management. Z Saugetierkd 60:41–52
Hartl GB, Zachos F, Nadlinger K (2003) Genetic diversity in European red deer (Cervus elaphus L.): anthropogenic influences on natural populations. CR Biol 326:37–42
Hedrick PW (2001) Conservation genetics: where are we now? TREE 16:629–636
Hulme DJ, Silk JP, Redwin JM, Barendse W, Beh KJ (1994) Ten polymorphic ovine microsatellites. Anim Genet 25:434–435
Hurrell JW (1995) Decadal trends in the North Atlantic Oscillation: regional temperatures and precipitation. Science 269:676–679
Huxley JS (1931) The relative size of antlers in deer. Proc Zool Soc Lond 1931:819–864
Ingebrigtsen O (1924) Hjortens utbredelse i Norge (in Norwegian). Bergens Museums Aarbok 1922–1923 Naturvitensk. Række 6:1–58
IPCC (2001) Intergovernmental panel on climate change third assessment report. http://www.ipcc.ch
IPCC (2007) Intergovernmental panel on climate change fourth assessment report. http://www.ipcc.ch
Jensen AB, Palmer KA, Boomsma JJ, Pedersen BV (2005) Varying degrees of Apis mellifera ligustica introgression in protected populations of the black honeybee, Apis mellifera mellifera, in northwest Europe. Mol Ecol 14:93–106
Kruckenhauser L, Pinsker W (2004) Microsatellite variation in autochthonous and introduced populations of the Alpine marmot (Marmota marmota) along a European west-east transect. J Zool Syst Evol Res 42:19–26
Kuehn R, Schroeder W, Pirchner F, Rottmann O (2003) Genetic diversity, gene flow and drift in Bavarian red deer populations (Cervus elaphus). Conserv Genet 4:157–166
Langella O (2002) POPULATIONS v. 1.2.28 (12/5/2002): a free population genetic software. CNRS UPR9034 v. 1.2.30. http://bioinformatics.org/~tryphon/populations/
Langvatn R (1998) Hjortens erobring av Norge (in Norwegian). In: Brox KH (ed) Brennpunkt natur. Tapir, Trondheim, pp 49–71
Langvatn R, Loison A (1999) Consequences of harvesting on age structure, sex ratio and population dynamics of red deer Cervus elaphus in central Norway. Wildl Biol 5:213–223
Langvatn R, Mysterud A, Stenseth NC, Yoccoz NG (2004) Timing and synchrony of ovulation in red deer constrained by short northern summers. Am Nat 163:763–772
Latch EK, Harveson LA, King JS, Hobson MD, Rhodes OE (2006) Assessing hybridization in wildlife populations using molecular markers: a case study in wild turkeys. J Wildl Manage 70:485–492
Lister A (1984) Evolutionary and ecological origins of British Deer. Proc R Soc Edinb B 82:205–229
Lister A (2004) The impact of quaternary ice ages on mammalian evolution. Phil Trans R Soc B 359:221–241
Loe LE, Meisingset EL, Mysterud A, Langvatn R, Stenseth NC (2004) Phenotypic and environmental correlates of tooth eruption in red deer (Cervus elaphus). J Zool 262:83–89
Loison A, Langvatn R, Solberg EJ (1999) Body mass and winter mortality in red deer calves: disentangling sex and climate effects. Ecography 22:20–30
Lønnberg E (1906) On the geographic races of red deer in Scandinavia. Ark Zool 3:1–19
Lorenzen ED, Siegismund HR (2004) No suggestion of hybridization between the vulnerable black-faced impala (Aepyceros melampus petersi) and the common impala (A. m. melampus) in Etosha National Park, Namibia. Mol Ecol 13:3007–3019
Ludt CJ, Schroeder W, Rottmanm O, Kuehn R (2004) Mitochondrial DNA phylogeography of red deer (Cervus elaphus). Mol Phylogenet Evol 31:1064–1083
Matrai K, Szemethy L, Toth P, Katona K, Szekely J (2004) Resource use by red deer in lowland nonnative forests, Hungary. J Wildl Manage 68:879–888
Moore SS, Byrne K, Berger KT et al (1994) Characterization of 65 bovine microsatellites. Mamm genome 5:84–90
Musani SK, Halbert ND, Redden DT, Allison DB, Derr JN (2006) Marker genotypes and population admixture and their association with body weight, height and relative body mass in United States federal bison herds. Genetics 174:775–783
Mysterud A, Stenseth NC, Yoccoz NG, Langvatn R, Steinheim G (2001a) Nonlinear effects of large-scale climatic variability on wild and domestic herbivores. Nature 410:1096–1099
Mysterud A, Yoccoz NG, Stenseth NC, Langvatn R (2001b) Effects of age, sex, and density on body weight of Norwegian red deer: evidence of density-dependent senescence. Proc R Soc Lond B 268:911–919
Mysterud A, Langvatn R, Yoccoz NG, Stenseth NC (2002) Large-scale habitat variability, delayed density effects and red deer populations in Norway. J Anim Ecol 71:569–580
Mysterud A, Meisingset EL, Veiberg V et al (2007) Monitoring population size of red deer: an evaluation of two types of census data from Norway. Wildl Biol 13:285–298
Mysterud A, Bonenfant C, Loe LE, Langvatn R, Yoccoz NG, Stenseth NC (2008) The timing of male reproductive effort relative to female ovulation in a capital breeder. J Anim Ecol 77:469–477
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nei M, Tajima F, Tateno Y (1983) Accuracy of estimated phylogenetic trees from molecular data. II. Gene frequency data. J Mol Evol 19:153–170
Page RDM (1996) TREEVIEW: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Pettorelli N, Mysterud A, Yoccoz NG, Langvatn R, Stenseth NC (2005) Importance of climatological downscaling and plant phenology for red deer in heterogeneous landscapes. Proc R Soc Lond B 272:2357–2364
Polziehn RO, Strobeck C (2002) A phylogenetic comparison of red deer and wapiti using mitochondrial DNA. Mol Phylogenet Evol 22:342–356
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Randi E (2005) Management of wild ungulate populations in Italy: captive-breeding, hybridisation and genetic consequences of translocations. Vet Res Commun 29(Suppl 2):71–75
Raymond M, Rousset F (1995) Genepop version-1.2.—population-genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rhymer JM, Simberloff D (1996) Extinction by hybridization and introgression. Ann Rev Ecol Syst 27:83–109
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Røed KH (1998) Microsatellite variation in Scandinavian Cervidae using primers derived from Bovidae. Hereditas 129:19–25
Røed KH, Midthjell L (1998) Microsatellites in reindeer, Rangifer tarandus, and their use in other cervids. Mol Ecol 7:1773–1778
Rosenberg NA, Burke T, Elo K et al (2001) Empirical evaluation of genetic clustering methods using multilocus genotypes from 20 chicken breeds. Genetics 159:699–713
Schneider S, Roessli D, Excoffier L (2000) Arlequin ver.2.000: a software for population genetics data analysis. Genetics and Biometry Laboratory, University of Geneva, Switzerland
Shaw PW, Carvalho GR, Seghers BH, Magurran AE (1992) Genetic consequences of an artificial introduction of Guppies (Poecilia-Reticulata) in N-Trinidad. Proc R Soc Lond B 248:111–116
Skog A, Zachos FE, Rueness EK et al (2009) Phylogeography of red deer (Cervus elaphus) in Europe. J Biogeogr 36:66–77
Slate J, Kruuk LEB, Marshall TC, Pemberton JM, Clutton-Brock TH (2000) Inbreeding depression influences lifetime breeding success in a wild population of red deer (Cervus elaphus). Proc R Soc Lond B 267:1657–1662
Storfer A (1999) Gene flow and endangered species translocations: a topic revisited. Biol Conserv 87:173–180
Storfer A, Mech SG, Reudink MW, Ziemba RE, Warren J, Collins JP (2004) Evidence for introgression in the endangered Sonora Tiger Salamander, Ambystoma tigrinum stebbinsi (Lowe). Copeia 4:783–796
Strandgaard H, Simonsen V (1993) Genetic differentiation in population of red deer, Cervus elaphus, in Denmark. Hereditas 119:171–177
Strickberger MW (1996) Evolution, 2nd edn. Jones and Bartlett Publishers, Boston
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tate ML, Anderson RM, McEwan KM, Goosen GJ, Pearse AJ (1998) Genetic analysis of farmed red deer hybrids. Acta Vet Hung 46:329–340
Vernesi C, Crestanello B, Pecchioli E, Tartari D, Caramelli D, Hauffe H, Bertorelle G (2003) The genetic impact of demographic decline and reintroduction in the wild boar (Sus scrofa): a microsatellite analysis. Mol Ecol 12:585–595
Vila C, Sundqvist AK, Flagstad O et al (2003) Rescue of a severely bottlenecked wolf (Canis lupus) population by a single immigrant. Proc R Soc Lond B 270:91–97
Wang J (2003) Maximum-likelihood estimation of admixture proportions from genetic data. Genetics 164:747–765
Weir BS (1996) Genetic data analysis II: methods for discrete population genetic data. Sinauer Associates, Sunderland
Welzholz JC, Bürger-Arndt R, Bücking W (2005): Country report—Germany: Working Group 1—Task 1.1. Description of the historical background that has led to the development of particular national Protected Forest Area frameworks. In: Latham J, Frank G, Fahy O, Kirby K, Miller H, Stiven R (eds) COST Action E27, protected forest areas in Europe—analysis and harmonisation (PROFOR): reports of signatory states. Federal Research and Training Centre for Forests, Natural Hazards and Landscape (BFW), Vienna, pp 133–157
Whitehead GK (1972) Deer of the world. Constable, London
Whitehead GK (1993) The Whitehead encyclopaedia of deer. Swan Hill Press, Shrewsbury
Wilson GA, Strobeck C, Wu L, Coffin J (1997) Characterization of microsatellite loci in caribou Rangifer tarandus, and their use in other artiodactyls. Mol Ecol 6:697–699
Wolf CM, Griffith B, Reed C, Temple SA (1996) Avian and mammalian translocations: update and reanalysis of 1987 survey data. Conserv Biol 10:1142–1154
Wood S (2006) Generalized additive models: an introduction with R. Boca Raton. Chapman & Hall, UK
Yoccoz NG, Mysterud A, Langvatn R, Stenseth NC (2002) Age- and density-dependent reproductive effort in male red deer. Proc R Soc Lond B 269:1523–1528
Zachos F, Hartl GB, Apollonio M, Reutershan T (2003) On the phylogeographic origin of the Corsican red deer (Cervus elaphus corsicanus): evidence from microsatellites and mitochondrial DNA. Mamm Biol 68:284–298
Zachos F, Althoff C, Steynitz Y, Eckert I, Hartl G (2007) Genetic analysis of an isolated red deer (Cervus elaphus) population showing signs of inbreeding depression. Eur J Wildl Res 53:61–67
Acknowledgements
For providing samples from Norway we thank the Section for Wildlife Diseases at the Norwegian National Veterinary Institute, M. Pearson, H. Holm, O. Hårstad, Ander Børsstad, S. Aglen and the many hunters that sent us samples. For help sampling Hungarian red deer we acknowledge Professors Làszló Szemethy and Sàndor Csànyi at the Institute for Wildlife Conservation, St Stephens University, Hungary. For aid in the laboratory we are in debt to Liv Midthjell, Turid Vikøren and Astrid Stovner. Associate Professor David Griffiths, Institute of Basic Sciences and Aquatic Medicine at the Norwegian School of Veterinary Sciences, assisted with the native proof reading.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Haanes, H., Røed, K.H., Mysterud, A. et al. Consequences for genetic diversity and population performance of introducing continental red deer into the northern distribution range. Conserv Genet 11, 1653–1665 (2010). https://doi.org/10.1007/s10592-010-0048-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-010-0048-1