Abstract
We investigated the endangered status and taxonomic status of the freshwater crayfish Procambarus ferrugineus, a crayfish species considered for the candidate list of the Endangered Species Act. This species has a narrow distribution from central Arkansas, USA and is codistributed with its presumed sister species, Procambarus liberorum. We sampled extensively throughout the ranges of both primary burrowing species and collected mitochondrial DNA from a hypervariable fragment of the 16S gene from 109 individuals across 22 sites. We also collected data from a variable region of the 12S gene from a subset of the resulting 16S haplotypes. Due to our inability to sample what we considered P. ferrugineus in the field, we included museum specimens from the United States Natural History Museum of both P. ferrugineus and P. liberorum. Analyses of the resulting data suggested that these two species are indeed the same and we therefore synonymize them under the name of priority—P. liberorum. Additionally, our sampling discovered three new cryptic species from southwestern Arkansas all from the genus Procambarus. Nested clade phylogeographic analysis coupled with population genetic analyses suggested that P. liberorum has had three rounds of range expansion throughout the inferred evolutionary history. Using IUCN Red List criteria for conservation assessment, we conclude that the species P. liberorum should be considered stable, but with special concern because of habitat fragmentation and urbanization, small restricted range, and a moderate level of genetic diversity. Procambarus reimeri should be considered endangered due to its limited geographic range and the potential for a decline in suitable habitat. The three potentially newly discovered species should be considered data deficient until more information is obtained on their distributional limits and habitat requirements. Our study highlights the importance of thorough geographic and taxonomic sampling coupled with the utility of collecting data from museum specimens to reach robust taxonomic and conservation conclusions for endangered species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
For better or worse, species are the central currency of much of conservation biology. For example, the International Conservation Union (IUCN) keeps a Red List of endangered species (2001), which is often used to define “hot spots” of global and regional conservation efforts (Myers et al. 2000; Orme et al. 2005). Likewise, many local, regional, and national governments are focused on single species approaches to conservation. For example, most governmental conservation action in the United States is dictated by the implications of the Endangered Species Act (Clegg 1995). Given such emphasis on “the species,” it is critical to evaluate the validity of these taxonomic entities through a variety of means using a variety of ecological, morphological, and molecular data (Paquin and Hedin 2004) coupled with explicit and diagnosable species concepts (Sites and Crandall 1997; Sites and Marshall 2003). In some cases of highly endangered species, however, relevant data may not be available because of the rarity of the species. In such cases, museum specimens become essential in determining species status, historical population structure and levels of gene flow (Thomas et al. 1990; Graham et al. 2004; Wandeler et al. 2007), and even habitat reconstruction and climate change insights (Rowe 2007). Indeed, often such samples can be obtained while preserving the external morphology of the specimens themselves (e.g., Gilbert et al. 2007). We present such a case and demonstrate the utility of combining molecular phylogenetic investigation with museum samples to obtain relevant data on species status for endangered species. We demonstrate this approach by investigating the species status of the freshwater crayfish Procambarus ferrugineus.
Freshwater crayfishes are a highly endangered group of organisms with high alpha biodiversity in the eastern United States (Crandall 1997). With over 360 described species from North America alone (Crandall and Buhay 2008), freshwater crayfish share the dubious distinction of having more species imperiled than not (a distinction held only by one other group—the freshwater mussels) (Conservancy 1996). One such imperiled species is P. ferrugineus.
Originally, Hobbs and Robison (1988) described P. ferrugineus from collections at two sites in Lonoke County in central Arkansas along the Coastal Plain physiographic province. The presumed sister taxon, Procambarus liberorum, was described earlier by Fitzpatrick (1978) from Washington County in northern Arkansas in the Ozark Mountains. Both species were originally placed in the gracilis group of the subgenus Girardiella within the genus Procambarus (Hobbs and Robison 1988).
Procambarusferrugineus was differentiated from P. liberorum by a combination of the following characters: (1) P. liberorum had a prominent angular excision in the basal third of the opposable margin of the dactyl while P. ferrugineus either lacked such a prominent angular excision or had a weak excision in the basal third of the opposable dactyl; (2) the dorsolateral surface of the palm in P. ferrugineus was described as tuberculate while the dorsolateral surface of the palm in P. liberorum was punctuate; and (3) the first form male of P. liberorum was mostly red while the first form male of P. ferrugineus was mostly brown or tan, although inexplicably in the color notes, Hobbs and Robison (1988) described the carapace of the holotype as brick red. However, Hobbs and Robison (1988) noted that P. ferrugineus was similar to P. liberorum in possessing a very narrow areola. Furthermore, as additional specimens were collected by Robison from the intervening areas between the Ozark Mountains and the Coastal Plains in Lonoke County, doubts about the specific distinctiveness of P. ferrugineus were raised. In a letter to Robison–Hobbs suggested that “he had erred in describing P. ferrugineus, as the original morphological characters differentiating the Coastal Plain P. ferrugineus from the mountain dwelling P. liberorum were beginning to blur.”
In an effort to better understand the conservation status and taxonomic limits of these endemic species, we embarked on a study to sample and compare P. ferrugineus with its presumed sister taxon P. liberorum as well as to other related species found in similar habitats and geographic locations. Procambarus reimeri was thought to be a close relative of both P. liberorum and P. ferrugineus, and this species occurs in just one Arkansas County (Polk Co.) in similar burrow habitats. To determine the distinctiveness of these taxa, as well as their respective conservation statuses, we collected specimens from throughout the state of Arkansas and applied molecular genetic approaches for determining species validity and conservation needs. Unfortunately, our attempts to collect P. ferrugineus failed and we were forced to consider the possibility that this taxon was now extinct. However, we also acquired new localities for P. liberorum that encroached upon the distributional area of P. ferrugineus. We also sampled new localities south of the known range of both P. liberorum and P. ferrugineus but these individuals could not be confidently identified as either species or P. reimeri. Thus we suspected that P. liberorum and P. ferrugineus might actually be synonymous and might have a larger distribution than previously thought. We therefore collected museum specimens from the United States National Museum of Natural History (Smithsonian Institution), Washington DC from both species in the hopes of extracting quality DNA with which to compare with our field samples. The role of natural history museums in biodiversity studies has recently been highlighted (Graham et al. 2004) and this study provides a concrete example of the utility of such collections to literally save large sums of monies on would be misguided conservation and search efforts.
Materials and methods
Specimen collection
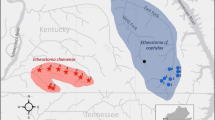
Specimens were collected by excavating burrows and extracting crayfish by hand. Sampling design in species boundary studies is critical (Morando et al. 2003) and includes sampling extensively throughout the species distribution. As P. liberorum is the presumed sister taxon to P. ferrugineus, we sampled extensively throughout its historical distribution, especially in locations close to the limited distribution of P. ferrugineus (Table 1) and in new areas previously not recorded for either species. Sampling locations are shown in Fig. 1 with respect to the individual species distributions. Gill tissue was dissected from each specimen upon collection and stored in 100% EtOH for DNA extraction (see below). The remaining specimen was stored in 70% EtOH as voucher material and deposited in the crayfish collections at either Southern Arkansas University or the Monte L. Bean Life Science Museum at Brigham Young University.
Distribution of Procambarus species included in this study. State boundaries for Arkansas and Oklahoma shown as bold lines, while county boundaries are shown as thin lines. Historical records of P. liberorum and P. ferrugineus are marked as circles with dots in the middle, and historical collections from the Smithsonian Museum (United States National Museum of Natural History) that we obtained genetic data from are circles marked with crosshairs. Open circles depict newly-discovered sites as a result of this study and genetic data was gathered for those samples. Closely-related species are represented as follows: P. reimeri, hexagon; P. sp. nov. 1, square; P. sp. nov. 2, triangle; P. sp. nov. 3, pentagon
DNA extraction, PCR amplification, and sequencing
Genomic DNA was extracted using standard methods and the 16S mtDNA gene was amplified for all individuals (Table 1) during PCR with primers 16sf-cray: GACCGTGCKAAGGTAGCATAATC and 16s-1492r: GGTTACCTTGTTACGACTT (Crandall and Fitzpatrick 1996) which amplify an approximately 500 base pair hypervariable region of the mitochondrial 16S ribosomal gene. The 16S mtDNA is the most variable gene for freshwater crayfishes (Fetzner and Crandall 2001) and has been used extensively and successfully for both population genetic and species diagnosis studies in freshwater crayfish (e.g., Fetzner and Crandall 2003; Buhay and Crandall 2005). The 12S mtDNA gene (Mokady et al. 1999) was also amplified using primers 12sf: 5′ GAAACCAGGATTAGATACCC 3′ and 12sr: 5′ TTTCCCGCGAGCGACGGGCG 3′ from one individual per sampled locality to provide deeper among-species phylogenetic relationships (Table 1). The 12S gene is approximately 400 base pairs and is slightly less variable than 16S (Buhay et al. 2007). Cycle-sequencing reactions were run with purified PCR products and the Big Dye Ready-Reaction kit on a Perkin Elmer Thermocycler. Reactions were cleaned using Millipore plates and then sequenced using an ABI3730XL automated DNA sequencer. There are cases where nuclear and mtDNA sequences can give conflicting signals (e.g., Shaw 2002; Evans et al. 2003), but these tend to be where lineage sorting due to recent radiation is problematic or where there is the potential for hybridization (see Posada and Crandall 2001). Neither of these processes is suspected in this case and the decision was made by the authors to do more thorough sampling across the geographic distribution of the target species rather than half the samples and develop a nuclear marker for those sampled, especially since sampling is critical to the Nested Clade Phylogeographic Analysis.
Sequence alignment and phylogenetic analyses
Resulting sequences were aligned using BioEdit (Hall 1999). A phylogenetic analysis of the P. liberorum, P. ferrugineus, closely-related species, and outgroup taxa was then performed on the unique 16S haplotypes (determined by TCS, see below) and the combined (since the loci are non-independent) 16S + 12S dataset using Maximum Likelihood (ML) (Felsenstein 1981) with a model of evolution selected from 56 alternatives using the AIC criterion as implemented in the software ModelTest 3.06 (Posada and Crandall 1998; Posada and Buckley 2004). Confidence in the resulting nodes was assessed using the bootstrap approach (Felsenstein 1985) with 1,000 pseudoreplications. ML runs were performed using the software PhyML 2.4.6 (Guindon and Gascuel 2003). Bayesian analyses using MrBayes 3.1.2 (Ronquist and Huelsenbeck 2003) were run for 20 million generations over eight chains with the starting parameters determined by ModelTest. Tracer (Rambaut and Drummond 2003) was used to determine the burnin and a consensus tree was constructed from the remaining trees. Multiple independent identical ML and Bayesian runs were done to ensure convergence on similar results. Nodal support for the Bayesian analyses was assessed using the posterior probability (PP) generated from a consensus tree of the sampled trees past burnin (Huelsenbeck et al. 2001).
Phylogeographic analyses and species diagnoses
Nested Clade Phylogeographic Analysis (NCPA: Templeton 1998a, 2001, 2004) was employed to examine significant associations between genetic variation and geographic location for the P. liberorum and P. ferrugineus individuals (109 individuals from 22 geographic locations). This approach sheds light on historical and contemporary evolutionary patterns and processes, including range expansion, restricted gene flow, and fragmentation. Using the program TCS (Clement et al. 2000) with the 16S dataset, a haplotype network was constructed for the 95% confidence level. The network is a graphic representation of the base pair differences between unique haplotypes. GeoDIS 2.4 (Posada et al. 2000, 2006) was then used to test for significant associations between the haplotypes and their geographic localities using the latitude–longitude coordinates over 5,000 permutations. For the museum specimens used from the Smithsonian, we approximated the coordinates based on the locality descriptions, but for the recent collections, we gathered location information using Global Positioning System (Garmin XL) devices at each site in decimal degrees. The 2005 inference key, available from http://darwin.uvigo/es/software/geodis.html, was used to tease apart contemporary and historical processes. The NCPA is also central to testing both hypotheses of species using the Cohesion Species Concept (Templeton 2001) as well as Evolutionarily Significant Units (Crandall et al. 2000). Here we use Templeton’s Cohesion Species Concept to test the hypothesis that P. liberorum and P. ferrugineus form distinct species (Templeton 1998b, 1999, 2001).
Genetic diversity and demography
The current genetic diversity (θ π ) (Tajima 1983) estimate was obtained from the program DNASP 4.0 (Rozas et al. 2003) using data from the hypervariable region of the 16S gene. Current genetic diversity estimates are based on pairwise nucleotide differences between sequences. Since endangered status assessments use as one of the key criteria the stability of the population size across time, contrasting these estimates provides a robust way of testing for stability in genetic diversity across evolutionary timescales and thereby contributing key insights into the conservation status of the species in question.
We also explored the historical population dynamics of the P. liberorum complex using the Bayesian skyline plot model (Drummond et al. 2005) as implemented in BEASTv1.3 (http://beast.bio.ed.ac.uk/Main_Page). This coalescent-based demographic model uses standard Markov Chain Monte Carlo (MCMC) sampling procedures to estimate a posterior distribution of effective population size through time directly from sequence data under a best-fit substitution model. The hyperparameter m was set to 1/4 of the sequences in each data set. Two independent MCMC analyses 2 × 107 steps long were performed sampling every 1,000th generation, with the burn-in set at 2 × 106 generations. Strict and relaxed clock evolutionary models were tested for consistency across assumptions (Drummond et al. 2006). All the Bayesian MCMC outputs generated by BEAST were analyzed in Tracer v1.3 (Rambaut and Drummond 2003).
Results
Phylogenetic analyses
Our resulting nucleotide sequences were deposited in GenBank under accession numbers EF012312–EF012356 for the 16S fragment and EF012280–EF012311 for the 12S data (Table 1). Phylogenetic relationships among P. liberorum, P. ferrugineus, and their closely-related species were determined using Bayesian and Maximum Likelihood approaches for both the 16S haplotype dataset and the combined 16S and 12S dataset representing one individual from most of the sampled localities (Table 1). ModelTest parameters for the 16S dataset (45 sequences, 490 base pairs) were as follows: GTR + I + G model of evolution (–lnL = 1489.8899, AIC = 2999.7798), base frequencies (A = 0.3461, C = 0.1029, T = 0.3647, G = 0.1863), proportion of invariable sites (I) = 0.5343, gamma distribution shape parameter (G) = 0.7362, Rmat = (0.9033 13.8691 2.8519 0.000 7.1326). ModelTest parameters for the combined 16S + 12S dataset (32 sequences, 878 base pairs) were as follows: TVM + I + G model of evolution (–lnL = 2268.9863, AIC = 4555.9365), base frequencies (A = 0.3744, C = 0.1431, G = 0.1229, T = 0.3596), proportion of invariable sites (I) = 0.6120, gamma distribution shape parameter (G) = –6267), Rmat = (0.5792 7.3728 1.0593 0.0000 7.3728).
For Bayesian analyses, starting parameters were number of substitution types (nst) = 6 and rates = invgamma. The first 4,000 trees were discarded as burnin and the consensus tree was estimated using the remaining 36,000 trees. For PhyML runs, starting parameters were determined by ModelTest with the initial tree determined by BIONJ with optimization. The TVM model is not an option in PhyML, therefore, for the combined 16S + 12S dataset, we used the GTR + I + G model using the ModelTest maximum likelihood estimated parameters with the exception of the transition/transversion ratio parameter.
Using the 16S data, P. liberorum and P. ferrugineus appear to be synonymous as they form a monophyletic group with a posterior probability of 89% and ML bootstrap support of 91% (Fig. 2). The sister relationship to P. liberorum/P. ferrugineus was unresolved, but the analyses did reveal the existence of three cryptic taxa, two of which have high bootstrap (BS) and posterior probability (PP) support and the third is represented by a single haplotype (although there were seven individuals with this same distinctive haplotype). Procambarus reimeri, Procambarus curdi, and Procambarus nigrocinctus fell out as closely-related species, while Procambarus tenuis was sister to that unresolved assemblage (Fig. 2).
Phylogenetic tree using 16S haplotypes showing relationships of Procambarus species. Numbers of individuals represented by each haplotype are given in parentheses. Haplotype numbers for P. liberorum and P. ferrugineus match those in the parsimony network. Bayesian posterior probabilities given above nodes >50% and ML bootstrap support for nodes >50% given below. Topology was identical with both Bayesian (–lnL = 1636.969) and ML (–lnL = 1501.168) analyses. Procambarus clarkii, Procambarus acutus, and Procambarus ouachitae were used to root the tree
The combined 16S + 12S gene dataset was analyzed using both ML and Bayesian approaches and revealed some similar trends to the 16S haplotype analyses, but clarified the deeper relationships between some of the Procambarus species. Again, P. liberorum and P. ferrugineus fell out in the same clade with strong support of 98% PP/91% BS (Fig. 3). Procambarus reimeri was recovered as the sister species to P. liberorum/P. ferrugineus with 99% PP/73% BS support. Procambarus sp nov 1 and P. sp nov 2 were also strongly supported clades each with 100% PP/100% BS. The phylogenetic positioning of P. sp nov 3 was unresolved yet the haplotype clearly represents a distinct evolutionary lineage.
Phylogenetic tree using combined 16S and 12S representing one individual for most sampled sites of Procambarus species. Bayesian posterior probabilities given above nodes >50% and ML bootstrap support for nodes >50% given below. Topology was identical with both Bayesian (–lnL = 2308.12) and ML (–lnL = 2268.163) analyses. Procambarus clarkii was used to root the tree
Phylogeographic analyses
A total of 22 haplotypes for P. liberorum/P. ferrugineus were recovered using TCS for 109 individuals collected from 22 localities throughout the range of the species complex (including new records that extend the range). The sister species based on the 16S + 12S phylogenetic analyses, P. reimeri, was used to root the TCS network which was set at the 95% confidence limit (nine mutational steps) (Templeton et al. 1992). Procambarus reimeri most closely connected to haplotype 17 at 11 steps, outside the confidence interval.
The total network cladogram was comprised of 11 one-step clades, 6 two-step clades, and 2 three-step clades (Fig. 4). Only 12 of these clades had both genetic and geographic variation, which was examined using the program GeoDIS (Table 2). Of these clades, only six showed significant variation for which Templeton’s 2005 inference key was used to determine the current and historical processes contributing to the variation (Table 3). Contiguous Range Expansion (CRE) and Restricted Gene Flow with Isolation by Distance (RGF w/ IBD) were the inferred patterns for four clades, while the total cladogram pattern was inconclusive. Clade 3-1 could be explained by either Long Distance Colonization (LDC) followed by fragmentation or Past Fragmentation (PF) followed by Range Expansion (RE), therefore, we performed a mismatch analysis of this clade (see below).
Haplotype network of 16S data and corresponding nesting levels for P. liberorum and P. ferrugineus. Procambarus reimeri was used to root the network. The 95% connection limit was determined to be nine steps. Haplotypes containing only P. liberorum individuals were shaded blue, those containing only P. ferrugineus were shaded yellow, and those haplotypes representing individuals from both P. liberorum and P. ferrugineus were shaded green
Genetic diversity and demography
We performed a mismatch analysis (Rogers and Harpending 1992; Harpending et al. 1998) of clade 3-1 to help determine whether colonization followed by fragmentation or fragmentation followed by range expansion can best explain the genetic-geographic patterns for the clade, as suggested by the inference key (Table 3). Examining deviations from neutrality can also help clarify past demographic events, so we determined Tajima’s D (Tajima 1989) which if significantly negative, can provide information about possible bottlenecks associated with range expansion events, typical of large floods and washouts. An initial θ = 1.25965, final θ = 1000, and the expansion parameter τ = 2μt = 0.57994 was used in DNASP for the mismatch analysis. The resulting plot showed a ragged bimodal distribution (Fig. 5), which is indicative of a population that is not expanding [raggedness = r = 0.2837; P (rexpected < robserved) = 0.86877]. Tajima’s D = −1.40484 (P > 0.10) suggested a constant population size. It appears that colonization followed by fragmentation best explains the pattern of diversity shown in clade 3-1, since we did not detect population expansion.
The genetic diversity estimates for P. liberorum/P. ferrugineus were θ π = 0.00698 ± 0.0004 SD which is a moderate level of diversity (Nei 1987). The BEAST analysis indicates a moderate increase in population size over the last two million years (Fig. 6), consistent with the NCPA analysis suggesting Contiguous Range Expansion at all three nesting levels (clades 1-9, 2-3, and 3-2). The analysis also demonstrates that the species is not in decline. Similar results were obtained for both the strict and relaxed clock approaches.
Discussion
Conservation issues often center around taxonomic issues (the validity and/or distinctiveness of a particular taxon) due to much of conservation legislation and indeed much of the conservation science being focused on species or on evolutionarily significant units (ESU) within a species. Thus we need concrete, testable, and repeatable criteria with which to assess species boundaries, species status, and to define conservation units (like ESUs) within species. Here we have taken a broad sampling approach to examining the species status of a presumed endangered species P. ferrugineus to investigate the evolutionary distinctiveness of this taxon relative to closely-related taxa. Our analyses clearly show that the currently described species of P. liberorum and P. ferrugineus form a monophyletic group and distinct evolutionary lineage relative to other closely related species of Procambarus. However, our recent field sampling did not encounter populations of P. ferrugineus. Therefore, this study required museum specimens to represent the species to distinguish between the hypotheses of synonymy versus the alternative that P. ferrugineus was indeed distinct but could not be found in the field due to recent extinction. Ample museum specimens preserved in a fashion that allowed reasonable DNA isolation (70% EtOH) allowed us to test these hypotheses. We used the Cohesion Species Concept to test the hypothesis that P. liberorum and P. ferrugineus are distinct species. A Cohesion Species is first and foremost an evolutionary lineage concept, that is, to designate two populations as distinct species, they must form distinct evolutionary lineages. Thus, the first null hypothesis to be tested is that the individuals sampled from these two presumed species are derived from a single evolutionary lineage versus the alternative that they form distinct evolutionary lineages. We can test this hypothesis formally using the NCA framework by testing if these prior categories correspond to phylogenetic lineages (e.g., Matos and Schaal 2000). As the NCA defines a nested set of hierarchical categories, we can apply an exact random permutation test to the nested design to test the null hypothesis of no association of these prior taxonomic categories with phylogenetic structure. Indeed, in this case, we fail to reject this null hypothesis. The museum specimens from P. ferrugineus, which included a paratype specimen, represented one distinct haplotype and two shared haplotypes with P. liberorum. Given the shared haplotypes coupled with the distinct haplotype of P. ferrugineus being clearly nested within the P. liberorum clade, we fail to reject the null hypothesis of distinct evolutionary lineages and therefore reject the hypothesis that P. liberorum and P. ferrugineus are distinct species. Thus, we hereby synonymize P. ferrugineus and P. liberorum with P. liberorum being the recognized species name due to priority. This example demonstrates the utility of the NCA in testing taxonomic hypotheses. Indeed, had we found distinct species, the NCA approach also immediately identifies diagnostic molecular characters for differentiating species that could effectively be used in barcode designations (Hebert et al. 2003). Finally, the NCA approach also defines the geographic areas associated with distinct lineages. The potential distributional area of species is an essential piece of information in IUCN and other conservation assessments and NCA provides a robust way of estimating this area.
Our study highlights the importance of museum specimens in conservation science. Because much of conservation biology is taxonomically oriented, when assessing the validity and distinctiveness of taxa, it is essential to have broad sampling throughout each species range. In addition, it is ideal to have material from type specimens or specimens collected from type localities to be sure of proper assignment of nomenclature. Finally, good sampling in terms of genetic markers, specimen localities, and number of specimens, is essential. For this study, we sampled broadly throughout the distribution of the species and even included specimens from beyond the known boundaries of the distribution. This broad sampling resulted in robust conclusions about the distinctiveness of the taxa under question and led to the discovery of previously unknown phylogenetic and taxonomic diversity.
During the course of this study, our sampling revealed three possible new lineages of freshwater crayfish closely related to P. liberorum. All three of these lineages are restricted to the southwestern corner of Arkansas with narrow distributions. The description of these new species is beyond the scope of this paper (as a formal species description is typically written for a very different audience and requires distinct data and presentation), but they will be further sampled for a subsequent publication, possibly including new species descriptions as warranted by the data. We are currently collecting nuclear genetic data as well as morphological data to support the distinct species hypotheses suggested by these results. Yet this result highlights the importance of broad sampling for the inclusion of the range of genetic diversity in species diagnoses.
The Interior Highlands which include the Ozark Mountains and the Ouachita Mountains are of considerable biogeographic significance as areas of endemism for both plants and animals (Matthews and Robison 1998). These mountainous regions have provided a safe haven for many forms during geological epochs when most of the rest of the continent was not available for habitation (Robison and Allen 1995). As many as 100–150 species may be endemic to the Interior Highlands (Robison and Smith 1982; Allen 1995). Faunistically, the Ozark Mountains harbor many endemic vertebrates including 14 endemic fish species (Mayden 1985, 1988; Robison 1986; Turner et al. 1996) and five salamander taxa as well as numerous invertebrates such as crayfishes (Williams 1954; Hobbs and Robison 1988; Robison and Allen 1995), stoneflies (Stark et al. 1983; Ernst et al. 1986; Poulton and Stewart 1987), caddisflies (Frazer and Harris 1991; Moulton and Stewart 1996), and other insects (Allen 1990).
Procambarus liberorum appears to have originated in the headwaters of the White River in the Ozark Mountains and then migrated southward through the southern Ozarks through the Arkansas River drainage onto the north flank of the Ouachita Mountains (Arkansas River drainage) and then proceeded eastward through the Arkansas River Valley as far east as Lonoke County in the Coastal Plain province. The Arkansas River floodplain has been hypothesized to be a major barrier to dispersal southward for stream fishes from the Ozarks (Mayden 1985) yielding high endemism in the Ouachita Mountain region. However, this appears not be the case with our burrowing crayfishes which exhibit moderate genetic diversity, expanding population sizes, and extensive gene flow.
There is little in the way of life history information available regarding P. liberorum. This species has not been studied in any depth. Ovigerous females have been found, however, no females carrying eggs have ever been collected (Hobbs and Robison 1988).
Finally, we conclude by providing a conservation assessment of the species of key interest in this study. We used the IUCN (2001) Red List Criteria for endangerment as they provide a globally recognized framework for conservation assessment. At this point, the three potential P. sp nov must be considered data deficient given the lack of information on these populations and lack of formal description. Yet they clearly form distinct evolutionary lineages worthy of conservation and are highly restricted in distribution at the moment. Thus, they are most deserving of future study. Procambarus reimeri is considered endangered based on the IUCN criteria B (geographic range in the form of ); B1 (extent of occurrence estimated to be less than 5,000 km2 and); a—known to exist at no more than five locations and b—continuing decline projected in (i) area of occupancy and (ii) area, extent and/or quality of habitat. The last inference is based on the susceptibility of burrowing crayfish to habitat degradation coupled with the potential for such degradation at the current known locations of this crayfish. Finally, we consider P. liberorum to be stable and not immediately imperiled. However, while this species is more broadly distributed, encroaching urbanization, grassland habitat loss, and deforestation can be detrimental to this species’ terrestrial gene flow routes and dispersal behaviors.
References
Allen RT (1990) Insect endemism in the interior highlands of North America. Fla Entomol 78:539–569
Allen RT (1995) Pedetontus gershneri, a new species of Machilidae from the interior highlands of North America (Insecta: Microcoryphidae). Entomol News 106:195–198
Buhay JE, Crandall KA (2005) Subterranean phylogeography of freshwater crayfishes shows extensive gene flow and surprisingly large population sizes. Mol Ecol 14:4259–4273
Buhay JE, Moni G, Mann N, Crandall KA (2007) Molecular taxonomy in the dark: evolutionary history, phylogeography, and diversity of cave crayfish in the subgenus Aviticambarus, genus Cambarus. Mol Phylogenet Evol 42:435–488
Clegg M (1995) Science and the Endangered Species Act. National Academy Press, Washington
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659
Conservancy TN (1996) TNC priorities for conservation: 1996 annual report card for US plant and animal species. Nature Conservancy, Arlington
Crandall KA (1997) The crayfish component to an endangered aquatic ecosystem of the southeast United States. Freshw Crayfish 11:83–86
Crandall KA, Buhay JE (2008) Global diversity of crayfish (Astacidae, Cambaridae, and Parastacidae-Decapoda) in freshwater. Hydrobiologia 595:295–301
Crandall KA, Fitzpatrick JF Jr (1996) Crayfish molecular systematics: using a combination of procedures to estimate phylogeny. Syst Biol 45:1–26
Crandall KA, Bininda-Emonds ORP, Mace GM, Wayne RK (2000) Considering evolutionary processes in conservation biology. Trends Ecol Evol 15:290–295
Drummond AJ, Rambaut A, Shapiro B, Pybus OG (2005) Bayesian coalescent inference of past population dynamics from molecular sequences. Mol Biol Evol 22:1185–1192
Drummond AJ, Simon WYH, Matthew JP, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLoS Biol 4:699–710
Ernst MR, Poulton BC, Stewart KW (1986) Neoperla (Plecoptera: Perlidae) of the southern Ozark and Ouachita mountain region, and two new species of Neoperla. Ann Entomol Soc Am 79:645–661
Evans BJ, Supriatna J, Andayani N, Melnick DJ (2003) Diversification of Sulawesi macaque monkeys: decoupled evolution of mitochondrial and autosomal DNA. Evolution 57:1931–1946
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fetzner JW Jr, Crandall KA (2001) Genetic variation. In: Holdich DM (ed) Biology of freshwater crayfish. Blackwell Science, Oxford, pp 291–326
Fetzner JW Jr, Crandall KA (2003) Linear habitats and the nested clade analysis: an empirical evaluation of geographic vs. river distances using an Ozark crayfish (Decapoda: Cambaridae. Evolution 57:2101–2118
Fitzpatrick JF Jr (1978) Systematics of the crawfishes of the Hagenianus group of the genus Procambarus, subgenus Girardiella (Decapoda, Cambaridae). Tulane Stud Zool Bot 20:57–97
Frazer KS, Harris SC (1991) Cladistic analysis of the Ochrotrichia shawnee group (Trichoptera: Hydroptilidae) and description of a new species from the interior highlands of northwestern Arkansas. J Kans Entomol Soc 64:363–371
Gilbert MTP, Moore W, Melchior L, Worobey M (2007) DNA extraction from dry museum beetles without conferring external morphological damage. PLoS One 2:e272
Graham CH, Ferrier S, Huettman F, Moritz C, Peterson AT (2004) New developments in museum-based informatics and applications to biodiversity analysis. Trends Ecol Evol 19:497–503
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phyhlogenies by maximum likelihood. Syst Biol 52:696–704
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Harpending HC, Batzer MA, Gurven M, Jorde LB, Rogers AR (1998) Genetic traces of ancient demography. Proc Natl Acad Sci USA 95:1961–1967
Hebert PDN, Ratnasingham S, deWaard JR (2003) Barcoding animal life: cytochrome c oxidase subunit 1 divergences among closely related species. Proc R Soc London B
Hobbs HH Jr, Robison HW (1988) The crayfish subgenus Girardiella (Decapoda: Cambaridae) in Arkansas, with the descriptions of two new species and a key to the members of the Gracilis group in the genus Procambarus. Proc Biol Soc Wash 101:391–413
Huelsenbeck JP, Ronquist F, Nielsen R, Bollback JP (2001) Bayesian inference of phylogeny and its impact on evolutionary biology. Science 294:2310–2314
IUCN (2001) IUCN Red List Categories: Version 3.1. IUCN Species Survival Commission. Gland, Switzerland
Matos JA, Schaal BA (2000) Chloroplast evolution in the pinus montezumae complex: a coalescent approach to hybridization. Evolution Int J Org Evolution 54:1218–1233
Matthews WJ, Robison HW (1998) Influence of drainage connectivity, drainage area and regional species richness on fishes of the interior highlands of Arkansas. Am Midl Nat 139:1–19
Mayden RL (1985) Biogeography of Ouachita highland fishes. Southw Nat 30:195–211
Mayden RL (1988) Vicariance biogeography, parsimony, and evolution in North American freshwater fishes. Syst Zool 37:329–355
Mokady M, Loya Y, Achituv Y, Geffen E, Graur D, Rozenblatt S, Brickner I (1999) Speciation versus phenotypic plasticity in coral inhabiting barnacles: Darwin’s obervations in an ecological context. J Mol Evol 49:367–375
Morando M, Avila L, Sites JW Jr (2003) Sampling strategies for delimiting species: genes, individuals, and populations in the Liolaemus elongatus-kriegi complex (Squamata: Liolaemidae) in Andean-Patagonian South America. Syst Biol 52:159–185
Moulton SR, Stewart KW (1996) Caddisflies (Trichoptera) of the interior highlands of North America. Mem Am Entomol Inst 56:1–313
Myers N, Mittermeier RA, Mittermeier CG, da Fonseca GAB, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403:853–858
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Orme CD, Davies RG, Burgess M, Eigenbrod F, Pickup N, Olson VA, Webster AJ, Ding TS, Rasmussen PC, Ridgely RS, Stattersfield AJ, Bennett PM, Blackburn TM, Gaston KJ, Owens IP (2005) Global hotspots of species richness are not congruent with endemism or threat. Nature 436:1016–1019
Paquin P, Hedin M (2004) The power and perils of ‘molecular taxonomy’: a case study of eyeless and endangered Cicurina (Araneae: Dictynidae) from Texas caves. Mol Ecol 13:3239–3255
Posada D, Buckley TR (2004) Model selection and model averaging in phylogenetics: Advantages of Akaike Information Criterion and Bayesian approaches over Likelihood Ratio Tests. Syst Biol 53:793–808
Posada D, Crandall KA (1998) Modeltest: Testing the model of DNA substitution. Bioinformatics 14:817–818
Posada D, Crandall KA (2001) Intraspecific gene genealogies: trees grafting into networks. Trends Ecol Evol 16:37–45
Posada D, Crandall KA, Templeton AR (2000) GeoDis: A program for the cladistic nested analysis of the geographical distribution of genetic haplotypes. Mol Ecol 9:487–488
Posada D, Crandall KA, Templeton AR (2006) Nested clade analysis statistics. Mol Ecol Notes 6:590–593
Poulton BC, Stewart KW (1987) The stoneflies of the Ozark and Ouachita Mountains (Plecoptera). Mem Am Entomol Inst 38:1–116
Rambaut A, Drummond AJ (2003) Tracer: MCMC trace analysis tool. http://www.evolve.zoo.ox.ac.uk. University of Oxford, Oxford
Robison HW (1986) Zoogeography of North American freshwater fishes. In: Hocutt CH, Wiley EO (eds) Zoogeography of North American freshwater fishes. Wiley, New York, pp 267–285
Robison HW, Allen RT (1995) Only in Arkansas: a study of the endemic plants and animals of the state. University of Arkansas Press, Fayetteville
Robison HW, Smith KL (1982) The endemic flora and fauna of Arkansas. Proc Arkansas Acad Sci 36:52–57
Rogers AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol Evol 9:552–569
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Rowe RJ (2007) Legacies of land use adn recent climatic change: the small mammal fauna in the mountains of Utah. Am Nat 170:242–257
Rozas J, Sánchez-DelBarrio JC, Messegyer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Shaw KL (2002) Conflict between nuclear and mitochondrial DNA phylogenies of a recent species radiation: what mtDNA reveals and conceals about modes of speciation in Hawaiian crickets. Proc Natl Acad Sci USA 99:16122–16127
Sites JJ, Marshall J (2003) Delimiting species: a Renaissance issue in systematic biology. Trends Ecol Evol 18:462–470
Sites JW Jr, Crandall KA (1997) Testing species boundaries in biodiversity studies. Cons Biol 11:1289–1297
Stark BP, Stewart KW, Feminella J (1983) New records and descriptions of Alloperla (Plecoptera: Chloroperlidae) from the Ozark-Ouachita region. Entomol News 94:55–59
Tajima F (1983) Evolutionary relationship of DNA sequences in finite populations. Genetics 105:437–460
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Templeton AR (1998a) Nested clade analyses of phylogeographic data: testing hypotheses about gene flow and population history. Mol Ecol 7:381–397
Templeton AR (1998b) Species and speciation: geography, population structure, ecology, and gene trees. In: Howard DJ, Berlocher SH (eds) Endless forms: species and speciation. Oxford University Press, Oxford, pp 32–43
Templeton AR (1999) Using gene trees to infer species from testable null hypotheses: cohesion species in the Spalax ehrenbergi complex. In: Wasser SP (ed) Evolutionary theory and processes: modern perspectives, papers in Honour of Eviatar Nevo. Kluwer Academic Publishers, Dordrecht, pp 171–192
Templeton AR (2001) Using phylogeographic analyses of gene trees to test species status and processes. Mol Ecol 10:779–791
Templeton AR (2004) Statistical phylogeography: methods of evaluating and minimizing inference errors. Mol Ecol 13:789–810
Templeton AR, Crandall KA, Sing CF (1992) A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genetics 132:619–633
Thomas WK, Paabo S, Villablanca F, Wilson A (1990) Spatial and temporal continuity of kangaroo rat populations shown by sequencing mitochondrial DNA from museum specimens. J Mol Evol 31:101–112
Turner TF, Trexler JC, Kuhn DN, Robison HW (1996) Life-history variation and comparative phylogeography of darters (Pisces: Percidae) from the North American central highlands. Evolution 50:2023–2036
Wandeler P, Hoeck PEA, Keller LF (2007) Back to the future: museum specimens in population genetics. Trends Ecol Evol 22:634–642
Williams AB (1954) Speciation and distribution of the crayfishes of the Ozark Plateaus and Ouachita Provinces. Univ Kansas Sci Bull 36:803–918
Acknowledgements
We thank two anonymous reviewers for helpful comments in improving this manuscript. We gratefully acknowledge the travel and field support from the Arkansas Game & Fish Commission and the mentoring grant provided by the Roger and Victoria Sant Endowment for Conservation at Brigham Young University. We thank Savel Daniels, James Finlay, Betty Crump (USDA Forest Service), Brian Wagner (Arkansas Game and Fish Commission), Michael D. Warriner (Arkansas Natural Heritage Commission), Ron Goddard and students at Waldron High School, Michelle McGee and students at Acorn High School, Gene Leeds, Louie Leeds, and Joe Kremers (Clarksville) for their excellent assistance in collecting crayfish for this study. We thank Karen Reed and Rafael Lemaitre for allowing us access to the crayfish collection at the US Natural History Museum and for their assistance during our visits to the Smithsonian. KAC was partially supported by NSF grant EF-0531762.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Crandall, K.A., Robison, H.W. & Buhay, J.E. Avoidance of extinction through nonexistence: the use of museum specimens and molecular genetics to determine the taxonomic status of an endangered freshwater crayfish. Conserv Genet 10, 177–189 (2009). https://doi.org/10.1007/s10592-008-9546-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-008-9546-9