Abstract
Bone metastasis accounts for the vast majority of breast cancer (BC) metastases, and is related to a high rate of morbidity and mortality. A number of seminal studies have uncovered gene expression signatures involved in BC development and bone metastasis; each of them points at a distinct step of the ‘invasion-metastasis cascade’. In this review, we provide most recently discovered functions of sets of genes that are selected from widely accepted gene signatures that are implicate in BC progression and bone metastasis. We propose a possible sequential pattern of gene expression that may lead a benign primary breast tumor to get aggressiveness and progress toward bone metastasis. A panel of genes which primarily deal with features like DNA replication, survival, proliferation, then, angiogenesis, migration, and invasion has been identified. TGF-β, FGF, NFκB, WNT, PI3K, and JAK-STAT signaling pathways, as the key pathways involved in breast cancer development and metastasis, are evidently regulated by several genes in all three signatures. Epithelial to mesenchymal transition that is also an important mechanism in cancer stem cell generation and metastasis is evidently regulated by these genes. This review provides a comprehensive insight regarding breast cancer bone metastasis that may lead to a better understanding of the disease and take step toward better treatments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Metastasis, a complex multistep event
Breast cancer (BC) metastasis is the leading cause of cancer related deaths among women (metastasis; the spread of tumor cells from primary site to distant organs) [1, 2]. More than 70 % of BC patients in advanced stage develop bone metastases which are related to high rate of morbidity and mortality [3, 4]. The vast majority of researchers have focused to identify the underlying molecular and cellular mechanisms of this complex phenomenon. Developing gene expression signatures, involved in certain stages of BC metastasis, are among seminal discoveries that may let to understand the basics involved in the ‘invasion-metastasis cascade’, and consequently the generation of effective therapies against metastasis.
‘Invasion-metastasis cascade’ represents a multistep process that consists a set of sequential events, in which tumor cells invade from primary site, disseminate through circulation, and reconstitute secondary tumors at distant tissue(s) [5]. It has been investigated that breast tumor cells encounter a plenty of changes when disseminating from primary to distant sites, including changes in gene expression pattern [6], and in the states of stemness [7], that define clonal evolution and cancer stem cell (CSC) theories, respectively. Genetic alterations and subsequent shift in cellular events and states must be considered in the context of different microenvironments in different steps of metastasis in order to uncover mysteries of the complex process of metastasis cascade.
Clonal evolution and CSC theories are two models for tumor heterogeneity, cancer development, and metastasis that are widely accepted [8, 9]. Clonal evolution theory indicates that different lineages of cancer cells are developed during multistep genetic and epigenetic alterations, which lead to acquiring different features required for tumor development and metastasis [10]. On the other hand, CSCs, which have features of self-renewal, tumorigenesis, multilineage differentiation, motility, invasiveness and apoptosis resistance, are believed to be required for the development and maintenance of several forms of human cancers, including BC. Based on the CSC theory, tumor cells are not naturally alike, based on the state of stemness, and are organized in a hierarchical pattern in which CSCs are considered to be at the top of the apex [11–13]. It is believed that both models of clonal evolution and CSC can be applied in cancer development and metastasis, and that tumors heterogeneity can be generated from both of them [8].
The pattern of gene expression is likely the most critical determinant of CSC state. A quite sophisticated program that leads to expression of a group of genes, and simultaneously suppression of others, directs all features of a tumor cell in a precise time. Gene expression signatures have been developed since more than a decade for a variety of diseases like cancer metastasis. Gene signatures that have been developed for BC metastasis are ostensibly behind-the-scene forces of cellular events. In this study, we provide most recently discovered functions of sets of genes that are selected from seminal widely accepted gene signatures. We then propose a possible sequential pattern of gene expression that may lead to a benign primary breast tumor to get aggressiveness and progress toward bone metastasis. We also discuss most prominent molecular mechanisms involved in BC bone metastasis.
Gene signatures associated to breast cancer progression and bone metastasis
A number of seminal studies have uncovered gene expression signatures involved in BC development and metastasis; each of them points at a distinct step of the ‘invasion-metastasis cascade’ (Tables 1, 2, 3, 4) [14–16]. Today, these findings have entered to the diagnosis as predictors of disease outcome in BC patients [17]. Particularly, such discoveries have heralded the new era of personalized medicine, while predicting the clinical outcome of patients based on a set of distinct gene expression patterns [18]. Although improving, our understanding of the exact molecular and, most importantly, cellular mechanisms of BC metastasis is poor, and therefore reliable treatments are lacking. Analysis of data resulting from high throughput genome wide assays, and translation of the molecular pattern to cellular mechanisms/pathways may provide novel perspective to understand the complex nature of metastasis, and subsequently develop new therapeutic strategies. Among studies that have provided gene signatures for BC progression and bone metastasis, the ones by van’t Veer et al., Smid et al., and Kang et al. are of most seminal and widely accepted.
van’t Veer’s signature
Several studies regarding gene expression pattern in BC have been developed since more than a decade (Table 1). Earliest ones [19–23] were not quite sufficient to be utilized for predictive and therapeutic purposes. This may be because of the inconsistency in different studies (e.g., different kinds of primary tumors), and abundance of heterogeneity in tumors. The first highly applicable reported gene signature, by van’t Veer and colleagues (van’t Veer’s signature), have established a 70-gene prognosis profile from primary tumors of young BC patients. (Table 2 comprises a panel of selected genes from van’t Veer’s signature, based on their significant expression in most poor prognosed patients, and the status of being well studied). They identified a set of genes strongly predicting distant metastasis in patients who were lymph node negative, called poor prognosis signature [14]. Their finding uncovered a pattern of gene expression required for primary tumor cells to become invasive; capable to evade from primary site.
van’t Veer’s signature comprises genes associated to a well-orchestrated program for the regulation of different features required for primary tumors to grow and escape from primary site, including cell cycle, DNA replication, proliferation, tumorigenesis, survival, angiogenesis, migration, and invasion (Table 2). In particular, among those, CCNB2, CCNE2, MCM6, TSPYL5, NUSAP1, CMC2, ECT2, ORC6, DTL, PRC1, MELK, EGLN1, SLC2A3, RAB6B, ESM1, RAD21, CDC25B, CDK16, CENPA, PGK1, MAD2L1, CKS2, BUB1, FGF18, WISP1, and IGFBP5 are known regulators of cell cycle, DNA replication, proliferation, and survival [24–26]. Upregulation of these genes can be considered as the first requirements of primary tumor for its growth in order to be prepared for dissemination. Afterward, ECT2, EGLN1, ESM1, FLT1, EXT1, DIAPH3, EXOC7, NMU, CDC42BPA, VEGF, MMP9, FGF18, WISP1, and TGF-β3, are well-known pivotal elements that participate in angiogenesis, migration, and invasion. Tumor cells which express these genes seem to be adept for invasion from primary site (please see Table 2 for detail functions of gene products).
A variety of key molecular mechanisms are regulated by van’t Veer’s signature gene products. Remarkably, TGF-β signaling pathway likely plays important roles in the regulation of cell cycle, proliferation, induction of EMT, CSCs, and MMPs [27–30] (Table 2). Furthermore, NFκB signaling is also involved in this level of tumor progression taking part in tumor growth and metastasis by the mediation of ESM1, FGF18, and WISP1 [31–33]. In this step, the transcription factor MYC, which is well known for its association to breast tumor proliferation [34, 35], may also play important roles as is shown to be regulated by at least two of van’t Veer’s signature genes including RAD21 and CDC25B [36, 37]. Notably, ‘epithelial to mesenchymal transition’ (EMT), a process that is shown to be essential for tumor dissemination and metastasis of breast carcinomas [38], is enhanced through at least two of genes in this signature including MMP9 and VEGF. These two factors play determining roles in preparing a hospitable microenvironment in which emitted signals trigger EMT in order to induce/maintain CSCs [39–42]. On the other hand, two of genes in this panel may tend to trigger primary tumor cells to metastasize to bone, including FLT1 and PGK1 [43, 44]. PGK1 increases the expression of CXCR4, which is one of the most important bone metastasis factors [45]. FLT1 also provide a premetastatic niche in bone and direct bone metastasis of BC [43]. Together, above information suggest the involvement of key regulators like TGF-β, NFκB, and MYC, as well as the process of EMT in first steps of BC metastasis within the primary tumor.
Biological functions of genes in van’t Veer’s signature define the hallmarks of cancer. Tian and colleagues have recently shown that van’t Veer’s signature gene products functionally meet all the six hallmarks of cancer defined by Hanahan and Weinberg, including sustained proliferation, anti-growth signaling evasion, cell death resistance, immortality, angiogenesis, and invasion/metastasis [46]. They identified interconnected networks and showed that these genes are regulated by key tumorigenic factors like TP53, RB1, MYC, JUN and CDKN2A [47]. Interestingly, van’t Veer’s signature may also reflect the two additional hallmarks of next generation, including reprogramming of energy metabolism and evading immune destruction [1].
Adjustments of energy metabolism in order to fuel cell growth and division, is of most important features that lead to uncontrolled proliferation in neoplasms. Glycolytic fueling has been shown to be one of the most essential mechanisms in the reprogramming of energy, and associated with activated oncogenes like RAS, MYC, mutant tumor suppressors like TP53, certain signaling pathways like PI3K/Akt/PTEN, and hypoxia inducible factor 1(HIF1) [48, 49]. Now, as Tian and colleagues showed, and also from functions of certain genes including TSPYL5, CMC2, CDC25B, EGLN1, SLC2A3, RAB6B, and TGF-β3 (see details of functions in Table 2), van’t Veer’s signature likely associates with the seventh hallmark of cancer, reprogramming of energy metabolism. As for the eighth hallmark, evading immune destruction, it has been shown that tumors that produce transforming growth factor (TGF)-β escape from immune surveillance, mainly by selective and direct suppression of the T cell cytotoxic gene responses [50]. Intriguingly, TGF-β signaling is undeniably of key factors in van’t Veer’s signature. Together, it seems that functions of gene products of van’t Veer’s signature also meet the two additional next generation hallmarks of cancer, in addition to the first six ones.
Smid’s signature
Focusing on bone metastasis, Smid and colleagues have established a panel of genes in BC patients that are implicated to bone relapse (Smid’s signature). They analyzed primary tumors of lymph node negative BC patients who, subsequently, had developed metastases. A set of 69 genes was identified to be differentially expressed in patients who had experienced bone metastasis versus patients with metastasis to other sites. (Table 3 comprises a panel of selected genes from Smid’s signature that are significantly overexpressed and have been better studied). Notably, they developed classifier of tumors that metastasize to bone that was applicable in clinic [16].
Smid’s signature provides a pattern of gene expression in primary tumors that obligates them to metastasize to bone. Genes in Smid’s signature participate in essential features of metastasis including tumor growth, proliferation, survival, angiogenesis, migration, and invasion (Table 3). Among those, TFF1, TFF3, AGR2, NAT1, CRIP1, TSPAN1, FGFR3, CEACAM6, and TMSB15A may be categorized to play roles in cell growth/proliferation, and survival. Importantly, a number of genes in this signature take part in angiogenesis that include TFF1, TFF3, FGFR3, and FGFBP1. Thereupon, the vast majority of the genes in this panel which have seminal roles in migration and invasion are reported to be TFF1, TFF3, AGR2, NAT1, RND1, TSPAN1, FGFR3, CEACAM6, KRT16, FGFBP1, FOXO3A, KRT6B, and SNAI1. Interestingly, TFF1, TFF3, and FGFR3 are present in all above categories, and seemingly play pivotal roles in the process of BC metastasis to bone.
Well-characterized crucial molecular mechanisms are controlled by Smid’s signature gene products. Trefoil factor-1 (TFF1), that was the most differentially expressed gene associated to bone metastasis in Smid’s signature, is ascertained to play roles in cell survival, anchorage-independent growth, angiogenesis, migration, and invasion in breast (and also other) tumor cells [51–54]. TFF3 induces angiogenesis by the regulation of VEGF, under the control of hypoxia [55]. AGR2, NAT1, FGFR3, and TSPAN1 have shown to play roles in tumor growth/proliferation/survival and/or migration/invasion. CRIP1, CEACAM6, and TMSB15A regulate tumor growth/proliferation and survival. RND1, KRT16, KRT6B, FOXO3A, and SNAI1 induce migration/invasion. It should be noted that, SNAI1 (also SNAIL) has determined to play pivotal roles as a master regulator of EMT [38, 56–58]. On the contrary, SCUBE2, and FOXO3A have generally shown to encode suppressors of tumor growth/proliferation, inducers of apoptosis, and repressor of angiogenesis. SCUBE2, and FOXO3A, although, are distinguished to play roles in line with induction of invasiveness, and metastasis, by regulation of Hedgehog signaling and matrix metalloproteinases, respectively [59–62] (Table 3).
Critical signaling pathways are associated with the genes in the Smid’s signature. Apparently, fibroblast growth factor (FGF) signaling takes fundamental parts in this circuit, as FGFR3 and FGFBP1 that are directly linked to this pathway are two important overexpressed genes in this panel. ERK signaling is likely involved through activation by FGFBP1 [59]. In addition, epidermal growth factor (EGF) and Janus kinase-signal transducers and activators of transcription (JAK-STAT) signaling pathways are demonstrated to be enhanced by TOM1L1 [63]. Together, from the above mentioned findings, it seems that the genes in Smid’s signature direct a complex program in which tumor cells proliferate and survive, then further induce and maintain angiogenesis, and finally enhance migration and invasion. Passing these two steps of gene expression (van’t Veer’s and Smid’s signatures), tumor cells are capable to invade, intravasate, and survive in circulation, heading to bone. These two sets of genes may be regulated in an overlapped, or in a sequential pattern.
Kang’s signature
Focused on the gene expression pattern of breast tumor cells heading to bone, Kang and colleagues investigated a multigenic program in highly aggressive osteolytic BC metastatic cells (Kang’s signature) (Table 4 comprises a panel of selected genes from Kang’s signature that are significantly overexpressed and have been better studied) [15]. Their work provided a framework for the identification of genes mediating metastasis to different organs. Although Kang’s signature was established in animal model, it has recently been confirmed in human BC patients as well [64]. These genes mostly encode secreted and cell membrane proteins, and are associated to the preparation of a compatible metastatic niche.
Kang’s signature provides a panel of genes that may be categorized into four groups including: angiogenesis, migration/invasion, EMT, and growth/angiogenesis inhibition. Of those, MCAM, PTK7, CTGF, FGF5, and CXCR4 play critical roles in angiogenesis. MCAM, PTK7, RGCC, CTGF, FGF5, ADAMTS1, CXCR4, IL-11, and MMP1 are well-known essential factors for tumor migration/invasion. Notably, a number of key genes in this signature encode important inducers of EMT and CSC features including: MCAM, PTK7, RGCC, CTGF, and CXCR4. RGCC and Smad3 direct the induction of EMT through regulation of SNAIL and SLUG EMT transcription factors [65]. CXCR4 activates several signaling pathways, including AKT [66], a process in which Src plays a critical role [67]. It should be noted that, some genes in Kang’s signature controversially function against tumor growth or angiogenesis (fourth group). Those include FHL1, DUSP1, SOCS2, FST, and ADAMTS1 (see below). SLC4A7 and NCF2, which are not categorized in these groups, play roles in preparing the microenvironment and inhibition of apoptosis, respectively (Table 4). From the above mentioned information, MCAM, PTK7, CTGF, and CXCR4 are categorized into all first three groups, and likely play critical roles in the last steps of BC bone metastasis.
Genes in Kang’s signature function toward controlling principal signaling pathways, and govern a sophisticated signaling network that leads to a successful metastasis. Table 4 demonstrates undeniable deviation in the regulation of key signaling pathways, like TGF-β, WNT, NFκB, FGF, and MAPK, as underpinning functions of genes associated to bone metastasis. Almost half of the genes in this panel, including DUSP1, RGCC, FST, CTGF, CXCR4, IL-11, and MMP-1, are directly linked to the TGF-β signaling [65, 68–76]. FST also increase mTOR signaling via Smad3 [77]. PTK7, FST, CXCR4, and MMP-1 take part in the regulation of WNT signaling, and remarkably, RGCC and CTGF function via NFκB pathway. FGF5 mediate FGF signaling, and enhance MAPK pathway [78]. CTGF also act through ERK and FAK pathways [79]. Importantly, distinct gene expression pattern of Kang’s signature specifically direct disseminated tumor cells to overt bone metastasis. CTGF, ADAMTS1, CXCR4, IL-11, and MMP1 are considered crucial inducers of bone metastasis [80–85]. Together, it seems that for a successful bone metastasis such sophisticated signaling network is required, which is controlled by the power of gene expression regulation.
Breast cancer bone metastasis; a multistep cascade of events
Several studies have reported genes that are associated with BC bone metastasis, from which some prognostic tools are provided in order to obtain best available treatments for individual BC patients. However, a comprehensive understanding of the nature of metastasis is yet to be investigated. Accordingly, due to the short knowledge of this complex phenomenon, well-suited therapies are lacking. In this review, we aimed to use published gene signatures, which have been shown to be significantly linked to progression and metastasis of BC to bone, to unmask a sequential pattern of gene expression that leads to colonization of breast tumors in bone.
Primary breast tumors cells ought to arrange a programmed gene expression pattern for their growth, survival, and invasion. Therefore, programs related to cell cycle progression, proliferation, apoptosis resistant, angiogenesis, invasion, and distant colonization need to be directed by nucleus. Genes in van’t Veer’s and Smid’s signature seem to provide such well-coordinated program for tumor cells, as both signatures are derived from primary tumors of BC patients, who developed metastasis (in general) and bone metastasis, respectively [14, 16]. We thought that genes in these two signatures likely express in a time period that primary tumor: first get committed to metastasize, and second committed to metastasize to bone. Then, as is obvious in Tables 2 and 3, primary tumor first get ready for metastasis, and then get features of invasion, and intravasation. Tumor cells, then, acquire bone specific directing features of homing, extravasation, micro-colonization, and eventually macro-colonization (metastatic colonization) from the genes in Kang’s signature (Table 4) (Fig. 1). This is likely because Kang’s signature is primarily derived from metastatic breast tumor cells from bone lesions [15]. Importantly, key genes in Kang’s signature including MMP-1, CXCR4, FGF5 and CTGF, which was initially determined from study on metastatic MDA-MB-231 cells and mouse model, have been recently confirmed in patients with BC and prostate cancer [64]. Figure 2 also shows deviation toward particular functions in each signature. In van’t Veer’s signature almost two out of third number of genes (24 out of 35) are in the category of proliferation. In Smid’s signature 13 out of 17 genes are in the category of migration. In Kang’s signature 9 out of 16 genes are involved in functions of migration and invasion. This may show the evolutionarily pattern of tumor progression and metastasis in BC bone metastasis.
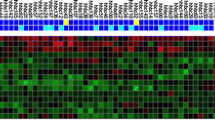
Sequential gene expression pattern, from primary tumor to metastatic colonization. van’t Veer’s, Smid’s, and Kang’s gene signatures shows a sequential pattern of expression that leads to breast cancer progression, metastasis. Genes in van’t Veer’s signature are mostly involved in the regulation of cell cycle, DNA replication, Proliferation, tumorigenesis, and survival, therefor, are essential for primary tumor cells to be prepared for invasion and metastasis. On the other hand some genes in this signature act toward angiogenesis, migration, and invasion. ESM1, ECT2, and EGLN1 are present in both categories and likely play important roles in primary steps of tumor metastasis. Genes in Smid’s signature also play essential roles in cell growth, proliferation, survival, angiogenesis, migration, and invasion. Tumor cells that overexpress genes in these two signatures seem to be able to invade and intravasate from primary site and disseminate through circulation. TFF1, TFF3, and FGFR3 are present in all three categories. Genes in Kang’s signature are mostly associated to angiogenesis, migration/invasion, EMT, and CSC factors. Many genes in this signature have been directly linked to bone metastasis of breast cancer. MCAM, PTK7, and CTGF are common genes in three categories. Genes in Kang’s signature likely act toward homing, extravasation, formation of micrometastasis, and eventually metastatic colonization, in order to end this journey. Abbreviations: EMT epithelial to mesenchymal transition, CSC cancer stem cells
Genes in van’t Veer’s, Smid’s, and Kang’s signatures are each deviated toward distinct functions. Certain numbers of genes in each signature are involved in certain functions categorized into: proliferation (blue), survival (green), angiogenesis (red), migration (yellow), and invasion (brown). Several genes have common functions, which are located in interconnected territories of circles. Red highlighted genes have functions against the corresponding feature (circle). Size of the circles shows the deviation toward that function in each signature. (Color figure online)
A set of well-defined genes govern key molecular pathways, and are the behind-the-scene forces of BC bone metastasis. Genes from van’t Veer’s, Smid’s, and Kang’s signatures control signaling pathways that have been considered as pivotal driving forces of tumor progression and metastasis. TGF-β, FGF, NFκB, WNT, PI3K, and JAK-STAT signaling pathways are induced/enhanced by genes of van’t Veer’s, and Smid’s signatures in primary breast tumor cells. On the other hand, TGF-β, FGF, NFκB, WNT, and PI3K pathways are also induced/enhanced by genes of Kang’s signature in bone colonized tumor cells. In both primary and distant tumors EMT program is induced by these pathways, and also several genes such as SNAI1 and MMP9 in primary tumor, and MCAM, PTK7, CXCR4, RGCC, and CTGF in distant metastatic tumors (Fig. 3) (Tables 2, 3, 4).
Well-orchestrated genes govern key molecular pathways, and are the behind-the-scene forces of breast cancer bone metastasis. Genes in van’t Veer’s, Smid’s, and Kang’s signatures govern a comprehensive signaling network in primary and secondary breast tumors. TGF-β, FGF, JAK-STAT, NFκB, WNT, and PI3K pathways in primary tumor, and TGF-β, FGF, NFκB, and PI3K pathways in secondary tumor are regulated by genes in these three signatures. In the primary tumor the six signaling pathways build a comprehensive signaling network that lead toward tumor growth, proliferation, survival, angiogenesis, migration, and invasion. In primary tumor (up-left), genes from van’t Veer’s, Smid’s signatures and their related signaling molecules are showed. HIF seem to have profound effects in primary tumor development and dissemination. Notably, several genes and signaling cascades induce EMT, and therefor CSC associated features. Intravasated tumor cells form the population of circulating tumor cells that disseminate, home, and extravasate into the secondary organ (bone). Importantly, the majority of differentiated circulating tumor cells (yellow) cannot survive the inhospitable environment while in circulation, and a small proportion of CSCs (red) are able to reach distant sites and form metastasis. In metastatic tumor (down-right), genes from Kang’s signature lead to activation of the five pathways, which build a comprehensive signaling network that governs features like invasion, migration, EMT, and CSC formation. It is important to mention that, tumors in primary and secondary sites can be different or identical regarding their state of differentiation. In most cases secondary tumors are at the same level of differentiation as primary tumor, or even more differentiated. But in some cases, like triple negative breast cancer, both primary and secondary tumors are mostly mesenchymal, and secondary tumors are even more mesenchymal (not shown in this figure). Abbreviations: TGF-β: transforming growth factor-beta, FGF: fibroblast growth factor, JAK-STAT: Janus kinase/signal transducers and activators of transcription, NFκB: nuclear factor kappa B, WNT, and PI3K: phosphatidylinositol 3 kinase, HIF: hypoxia inducible factor, EMT: epithelial to mesenchymal transition, CSC: cancer stem cells. (Color figure online)
Smad-dependent and Smad-independent TGF-β signaling pathways are essential for EMT and BC metastases [86–88]. Smad3 and Smad4 dependent TGF-β signaling have been shown to be indispensable for the induction of EMT and metastasis [89–91]. Smad transcription factors orchestrate overexpression of several important genes involved in EMT and metastasis, including SNAIL, TWIST, and ZEB families of transcription factor coding genes [92]. TGF-β also participates in the activation of several key signaling pathways such as Ras/ERK, and PI3K/Akt, called Smad-independent pathways, and regulates cell growth, survival, cytoskeletal reorganization, migration, and invasion [93]. For instance, MMP9 is shown to be induced by TGF-β-induced Akt-dependent ERK pathway [94]. TGF-β stimulation leads epithelial cells to obtain mesenchymal-like features, and capability of migration, invasion, and dissemination through circulation to distant sites of metastasis, and features of stemness [95].
FGF signaling also acts as an important inducer of EMT and metastasis [96]. FGFs regulate a wide range of biological functions such as proliferation, survival, and migration [97], and likely play pivotal roles in bone metastasis of BC [16]. NFκB is also an important inducer of EMT, and act through direct activation of SNAIL and ZEB family of transcription factors [98, 99]. Interestingly, it is believed that the cooperation of NFκB and TGF-β signaling pathways is critical for EMT and cancer metastasis [100, 101]. Wnt signaling is among the most important pathways involved in the induction of EMT and breast CSCs [86, 102, 103]. Notably, WNT and TGF-β signaling pathways likely induce an mutually reinforcing autocrine signaling network that is indispensable for constant expression of EMT associated transcription factors and CSC niche [87, 104]. PI3K signaling pathway is well-known for its important roles in the induction of EMT and metastasis [105]. Intriguingly, it has been shown that PI3K signaling is essential for autocrine/paracrine TGF-β associated motility, invasiveness, and metastasis [106]. JAK-STAT signaling pathway has an essential regulatory role in growth and proliferation of breast CSCs [107]. JAK-STAT signaling is associated with essential features such as survival, cell cycle regulation, self-sufficiency in growth and metastasis [108, 109]. Importantly, JAK2 also interacts and activates PI3K and RAS signaling molecules [110].
Hypoxia and hypoxia inducible factors (HIFs) seem to have pivotal roles in BC bone metastasis. HIF directly regulates several genes from van’t Veer’s, Smid’s, and Kang’s signatures. TFF3, EGLN1, SNAI1, MMP9, TGFB3, SLC2A3, and CTGF are of genes that are directly regulated by hypoxia. In fact, HIFs are of the essential preliminary factors that trigger gene expression programs that lead to tumor progression and metastasis, and play critical roles in the induction of EMT and stemness state in CSCs [111]. Hypoxia and HIFs are likely essential factors in the regulation of on and off states of EMT between primary and secondary tumors. In primary tumors, localized hypoxia mediates HIFs to be activated, and therefore move toward EMT/CSC induction and metastasis. At the secondary sites, however, with likely no hypoxic environment, lack of hypoxia and other factors lead to the reversion of EMT and CSC features. This phenomenon is essential for metastasis of differentiated carcinomas [87, 112] (for a comprehensive review see Ref. [112]).
It seems that a comprehensive signaling network consisting of TGF-β, FGF, NFκB, WNT, PI3K, and JAK-STAT is indispensable for breast tumor cells to progress to overt bone metastasis. Essential links between key bone metastatic factors, such as vascular cell adhesion molecule 1 (VCAM1), receptor activator of nuclear factor κB ligand (RNAKL), parathyroid-hormone related peptide (PTHrP), and BACH1 with these pathways further confirms this signaling network. VCAM1 Promotes bone metastasis by attracting and tethering osteoclast progenitors that express α4 integrin and facilitating their maturation [113, 114], and induces PI3K-Akt signaling by the mediation of Ezrin [115]. RANKL regulates bone resorption [116], migration [117], invasion [117, 118], bone metastasis [3, 117, 119, 120], and induces tumorigenesis, EMT, stemness [121], and the upregulation of MMP1 [118]. PTHrP can be induced by TGF-β [122], and activates CTGF through protein kinase A/C and ERK pathways [80]. BACH1 is a common regulator of several bone metastasis genes, including MMP1 and CXCR4 [123], induced by TGF-β [124].
Genes play the central role in development and diseases. Controlling the cellular pathways is one of the most critical duties of genes, in which reciprocal feedbacks play essential roles. Gene expression pattern in a given cell likely relates the story of a journey in which the cell is born, grow, proliferate, and/or die. Regulating the cellular behavior is the most critical tasks of gene expression machinery. Malignant behavior in cancers is tightly controlled their by gene expression pattern. Connection of gene signatures discussed in this review may provide novel insight toward better understanding the journey in which tumor cells get features of malignancy and metastasize to distant sites, and therefore providing best fit treatments for any individual cancer.
References
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674
Siegel R, Naishadham D (2013) Jemal A (2013) Cancer statistics. CA Cancer J Clin 63(1):11–30
Mundy GR (2002) Metastasis to bone: causes, consequences and therapeutic opportunities. Nat Rev Cancer 2(8):584–593
Roodman GD (2004) Mechanisms of bone metastasis. N Engl J Med 350(16):1655–1664
Valastyan S, Weinberg RA (2011) Tumor metastasis: molecular insights and evolving paradigms. Cell 147(2):275–292
Ma XJ et al (2003) Gene expression profiles of human breast cancer progression. Proc Natl Acad Sci USA 100(10):5974–5979
Yu M et al (2013) Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339(6119):580–584
Chaffer CL, Weinberg RA (2011) A perspective on cancer cell metastasis. Science 331(6024):1559–1564
Shackleton M et al (2009) Heterogeneity in cancer: cancer stem cells versus clonal evolution. Cell 138(5):822–829
Greaves M, Maley CC (2012) Clonal evolution in cancer. Nature 481(7381):306–313
Brabletz T et al (2005) Opinion: migrating cancer stem cells - an integrated concept of malignant tumour progression. Nat Rev Cancer 5(9):744–749
Charafe-Jauffret E et al (2009) Breast cancer cell lines contain functional cancer stem cells with metastatic capacity and a distinct molecular signature. Cancer Res 69(4):1302–1313
Jordan CT, Guzman ML, Noble M (2006) Cancer stem cells. N Engl J Med 355(12):1253–1261
van ‘t Veer LJ et al (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415(6871):530–536
Kang Y et al (2003) A multigenic program mediating breast cancer metastasis to bone. Cancer Cell 3(6):537–549
Smid M et al (2006) Genes associated with breast cancer metastatic to bone. J Clin Oncol Off J Am Soc Clin Oncol 24(15):2261–2267
Glas AM et al (2006) Converting a breast cancer microarray signature into a high-throughput diagnostic test. BMC Genom 7:278
van’t Veer LJ, Bernards R (2008) Enabling personalized cancer medicine through analysis of gene-expression patterns. Nature 452(7187):564–570
Perou CM et al (1999) Distinctive gene expression patterns in human mammary epithelial cells and breast cancers. Proc Natl Acad Sci USA 96(16):9212–9217
Perou CM et al (2000) Molecular portraits of human breast tumours. Nature 406(6797):747–752
Zajchowski DA et al (2001) Identification of gene expression profiles that predict the aggressive behavior of breast cancer cells. Cancer Res 61(13):5168–5178
Sorlie T et al (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98(19):10869–10874
West M et al (2001) Predicting the clinical status of human breast cancer by using gene expression profiles. Proc Natl Acad Sci USA 98(20):11462–11467
Grana X, Reddy EP (1995) Cell cycle control in mammalian cells: role of cyclins, cyclin dependent kinases (CDKs), growth suppressor genes and cyclin-dependent kinase inhibitors (CKIs). Oncogene 11(2):211–219
Ohtani K et al (1999) Cell growth-regulated expression of mammalian MCM5 and MCM6 genes mediated by the transcription factor E2F. Oncogene 18(14):2299–2309
Mehdipour P et al (2009) Prognostic implication of CDC25A and cyclin E expression on primary breast cancer patients. Cell Biol Int 33(10):1050–1056
Liu JH et al (1999) Functional association of TGF-beta receptor II with cyclin B. Oncogene 18(1):269–275
Wottawa M et al (2013) Knockdown of prolyl-4-hydroxylase domain 2 inhibits tumor growth of human breast cancer MDA-MB-231 cells by affecting TGF-beta1 processing. Int J Cancer J Int Cancer 132(12):2787–2798
Petrella BL, Armstrong DA, Vincenti MP (2012) Interleukin-1 beta and transforming growth factor-beta 3 cooperate to activate matrix metalloproteinase expression and invasiveness in A549 lung adenocarcinoma cells. Cancer Lett 325(2):220–226
Malanchi I et al (2012) Interactions between cancer stem cells and their niche govern metastatic colonization. Nature 481(7379):85–89
Kang YH et al (2012) ESM-1 regulates cell growth and metastatic process through activation of NF-kappaB in colorectal cancer. Cell Signal 24(10):1940–1949
Sonvilla G et al (2008) FGF18 in colorectal tumour cells: autocrine and paracrine effects. Carcinogenesis 29(1):15–24
Hou CH et al (2011) WISP-1 increases MMP-2 expression and cell motility in human chondrosarcoma cells. Biochem Pharmacol 81(11):1286–1295
Dubik D, Dembinski TC, Shiu RP (1987) Stimulation of c-myc oncogene expression associated with estrogen-induced proliferation of human breast cancer cells. Cancer Res 47 24(Pt 1): 6517-21
Watson PH, Pon RT, Shiu RP (1991) Inhibition of c-myc expression by phosphorothioate antisense oligonucleotide identifies a critical role for c-myc in the growth of human breast cancer. Cancer Res 51(15):3996–4000
McEwan MV, Eccles MR, Horsfield JA (2012) Cohesin is required for activation of MYC by estradiol. PLoS ONE 7(11):e49160
Wu W et al (1998) Overexpression of cdc25A and cdc25B is frequent in primary non-small cell lung cancer but is not associated with overexpression of c-myc. Cancer Res 58(18):4082–4085
Yang J, Weinberg RA (2008) Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev Cell 14(6):818–829
Deryugina EI, Quigley JP (2006) Matrix metalloproteinases and tumor metastasis. Cancer Metastasis Rev 25(1):9–34
Orlichenko LS, Radisky DC (2008) Matrix metalloproteinases stimulate epithelial-mesenchymal transition during tumor development. Clin Exp Metastasis 25(6):593–600
Yang AD et al (2006) Vascular endothelial growth factor receptor-1 activation mediates epithelial to mesenchymal transition in human pancreatic carcinoma cells. Cancer Res 66(1):46–51
Mani SA et al (2008) The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133(4):704–715
Kaplan RN et al (2005) VEGFR1-positive haematopoietic bone marrow progenitors initiate the pre-metastatic niche. Nature 438(7069):820–827
Jung Y et al (2009) Expression of PGK1 by prostate cancer cells induces bone formation. Mol Cancer Res MCR 7(10):1595–1604
Wang J et al (2007) A glycolytic mechanism regulating an angiogenic switch in prostate cancer. Cancer Res 67(1):149–159
Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100(1):57–70
Tian S et al (2010) Biological functions of the genes in the mammaprint breast cancer profile reflect the hallmarks of cancer. Biomark Insights 5:129–138
DeBerardinis RJ et al (2008) The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab 7(1):11–20
Jones RG, Thompson CB (2009) Tumor suppressors and cell metabolism: a recipe for cancer growth. Genes Dev 23(5):537–548
Thomas DA, Massague J (2005) TGF-beta directly targets cytotoxic T cell functions during tumor evasion of immune surveillance. Cancer Cell 8(5):369–380
Buache E et al (2011) Deficiency in trefoil factor 1 (TFF1) increases tumorigenicity of human breast cancer cells and mammary tumor development in TFF1-knockout mice. Oncogene 30(29):3261–3273
Emami S et al (2001) Induction of scattering and cellular invasion by trefoil peptides in src- and RhoA-transformed kidney and colonic epithelial cells. FASEB J: official publication of the Federation of American Societies for Experimental Biology 15(2):351–361
Prest SJ, May FE, Westley BR (2002) The estrogen-regulated protein, TFF1, stimulates migration of human breast cancer cells. FASEB J: official publication of the Federation of American Societies for Experimental Biology 16(6):592–594
Rodrigues S et al (2003) Trefoil peptides as proangiogenic factors in vivo and in vitro: implication of cyclooxygenase-2 and EGF receptor signaling. FASEB J: official publication of the Federation of American Societies for Experimental Biology 17(1):7–16
Guleng B et al (2012) TFF3 mediated induction of VEGF via hypoxia in human gastric cancer SGC-7901 cells. Mol Biol Rep 39(4):4127–4134
Blanco MJ et al (2002) Correlation of Snail expression with histological grade and lymph node status in breast carcinomas. Oncogene 21(20):3241–3246
Nieto MA (2002) The snail superfamily of zinc-finger transcription factors. Nat Rev Mol Cell Biol 3(3):155–166
De Craene B, Berx G (2013) Regulatory networks defining EMT during cancer initiation and progression. Nat Rev Cancer 13(2):97–110
Liu R et al (2009) KLF5 promotes breast cell survival partially through fibroblast growth factor-binding protein 1-pERK-mediated dual specificity MKP-1 protein phosphorylation and stabilization. J Biol Chem 284(25):16791–16798
Harris LG et al (2012) Increased vascularity and spontaneous metastasis of breast cancer by hedgehog signaling mediated upregulation of cyr61. Oncogene 31(28):3370–3380
Harris LG, Samant RS, Shevde LA (2011) Hedgehog signaling: networking to nurture a promalignant tumor microenvironment. Mol Cancer Res MCR 9(9):1165–1174
Storz P et al (2009) FOXO3a promotes tumor cell invasion through the induction of matrix metalloproteinases. Mol Cell Biol 29(18):4906–4917
Lin YC et al (2011) Domain and functional analysis of a novel breast tumor suppressor protein, SCUBE2. J Biol Chem 286(30):27039–27047
Casimiro S et al (2012) Analysis of a bone metastasis gene expression signature in patients with bone metastasis from solid tumors. Clin Exp Metastasis 29(2):155–164
Guo X, Jose PA, Chen SY (2011) Response gene to complement 32 interacts with Smad3 to promote epithelial-mesenchymal transition of human renal tubular cells. Am J Physiol Cell Physiol 300(6):C1415–C1421
Epstein RJ (2004) The CXCL12-CXCR4 chemotactic pathway as a target of adjuvant breast cancer therapies. Nat Rev Cancer 4(11):901–909
Zhang XH et al (2009) Latent bone metastasis in breast cancer tied to Src-dependent survival signals. Cancer Cell 16(1):67–78
Mikami F et al (2006) The transforming growth factor-beta-Smad3/4 signaling pathway acts as a positive regulator for TLR2 induction by bacteria via a dual mechanism involving functional cooperation with NF-kappaB and MAPK phosphatase 1-dependent negative cross-talk with p38 MAPK. J Biol Chem 281(31):22397–22408
Huang WY et al (2009) RGC-32 mediates transforming growth factor-beta-induced epithelial-mesenchymal transition in human renal proximal tubular cells. J Biol Chem 284(14):9426–9432
Fitzgerald AM et al (2012) The effects of transforming growth factor-beta2 on the expression of follistatin and activin A in normal and glaucomatous human trabecular meshwork cells and tissues. Invest Ophthalmol Vis Sci 53(11):7358–7369
Parada C, et al (2013) CTGF mediates Smad-dependent TGFbeta signaling to regulate mesenchymal cell proliferation during palate development. Mol Cell Biol 33(17):3482–3493
Chu CY, et al (2013) Induction of chemokine receptor CXCR4 expression by transforming growth factor-beta1 in human basal cell carcinoma cells. J Dermatol Sci 72(2):123–133
Bertran E, et al (2013) Overactivation of the TGF-beta pathway confers a mesenchymal-like phenotype and CXCR4-dependent migratory properties to liver tumor cells. Hepatology 58(6):2032–2044
Gupta J et al (2011) TGFbeta-dependent induction of interleukin-11 and interleukin-8 involves SMAD and p38 MAPK pathways in breast tumor models with varied bone metastases potential. Cancer Biol Ther 11(3):311–316
Calon A et al (2012) Dependency of colorectal cancer on a TGF-beta-driven program in stromal cells for metastasis initiation. Cancer Cell 22(5):571–584
Vu TH, Werb Z (2000) Matrix metalloproteinases: effectors of development and normal physiology. Genes Dev 14(17):2123–2133
Winbanks CE et al (2012) Follistatin-mediated skeletal muscle hypertrophy is regulated by Smad3 and mTOR independently of myostatin. J Cell Biol 197(7):997–1008
Kornmann M et al (1997) Fibroblast growth factor-5 stimulates mitogenic signaling and is overexpressed in human pancreatic cancer: evidence for autocrine and paracrine actions. Oncogene 15(12):1417–1424
Tan TW et al (2009) CTGF enhances migration and MMP-13 up-regulation via alphavbeta3 integrin, FAK, ERK, and NF-kappaB-dependent pathway in human chondrosarcoma cells. J Cell Biochem 107(2):345–356
Shimo T et al (2006) Pathogenic role of connective tissue growth factor (CTGF/CCN2) in osteolytic metastasis of breast cancer. J Bone Miner Res Off J Am Soc Bone Miner Res 21(7):1045–1059
Lu X et al (2009) ADAMTS1 and MMP1 proteolytically engage EGF-like ligands in an osteolytic signaling cascade for bone metastasis. Genes Dev 23(16):1882–1894
Muller A et al (2001) Involvement of chemokine receptors in breast cancer metastasis. Nature 410(6824):50–56
McCoy EM et al (2013) IL-11 produced by breast cancer cells augments osteoclastogenesis by sustaining the pool of osteoclast progenitor cells. BMC Cancer 13:16
Gao YB et al (2013) Enhanced production of CTGF and IL-11 from highly metastatic hepatoma cells under hypoxic conditions: an implication of hepatocellular carcinoma metastasis to bone. J Cancer Res Clin Oncol 139(4):669–679
Ren L et al (2013) Bone metastasis from breast cancer involves elevated IL-11 expression and the gp130/STAT3 pathway. Med Oncol 30(3):634
Jing Y et al (2011) Epithelial-mesenchymal transition in tumor microenvironment. Cell Biosci 1:29
Fazilaty H et al (2013) Crosstalk between breast cancer stem cells and metastatic niche: emerging molecular metastasis pathway? Tumour Biol J Int Soc Oncodev Biol Med 34(4):2019–2030
Kalluri R, Weinberg RA (2009) The basics of epithelial-mesenchymal transition. J Clin Investig 119(6):1420–1428
Deckers M et al (2006) The tumor suppressor Smad4 is required for transforming growth factor beta-induced epithelial to mesenchymal transition and bone metastasis of breast cancer cells. Cancer Res 66(4):2202–2209
Kaimori A et al (2007) Transforming growth factor-beta1 induces an epithelial-to-mesenchymal transition state in mouse hepatocytes in vitro. J Biol Chem 282(30):22089–22101
Ashcroft GS et al (1999) Mice lacking Smad3 show accelerated wound healing and an impaired local inflammatory response. Nat Cell Biol 1(5):260–266
Xu J, Lamouille S, Derynck R (2009) TGF-beta-induced epithelial to mesenchymal transition. Cell Res 19(2):156–172
Derynck R, Zhang YE (2003) Smad-dependent and Smad-independent pathways in TGF-beta family signalling. Nature 425(6958):577–584
Byun HJ et al (2006) A splice variant of CD99 increases motility and MMP-9 expression of human breast cancer cells through the AKT-, ERK-, and JNK-dependent AP-1 activation signaling pathways. J Biol Chem 281(46):34833–34847
Dang H et al (2011) Snail1 induces epithelial-to-mesenchymal transition and tumor initiating stem cell characteristics. BMC Cancer 11:396
Ciruna B, Rossant J (2001) FGF signaling regulates mesoderm cell fate specification and morphogenetic movement at the primitive streak. Dev Cell 1(1):37–49
Turner N, Grose R (2010) Fibroblast growth factor signalling: from development to cancer. Nat Rev Cancer 10(2):116–129
Min C et al (2008) NF-kappaB and epithelial to mesenchymal transition of cancer. J Cell Biochem 104(3):733–744
Julien S et al (2007) Activation of NF-kappaB by Akt upregulates Snail expression and induces epithelium mesenchyme transition. Oncogene 26(53):7445–7456
Huber MA et al (2004) NF-kappaB is essential for epithelial-mesenchymal transition and metastasis in a model of breast cancer progression. J Clin Investig 114(4):569–581
Maier HJ et al (2010) NF-kappaB promotes epithelial-mesenchymal transition, migration and invasion of pancreatic carcinoma cells. Cancer Lett 295(2):214–228
Velasco-Velazquez MA et al (2012) Breast cancer stem cells. Int J Biochem Cell Biol 44(4):573–577
Malanchi I et al (2008) Cutaneous cancer stem cell maintenance is dependent on beta-catenin signalling. Nature 452(7187):650–653
Scheel C et al (2011) Paracrine and autocrine signals induce and maintain mesenchymal and stem cell states in the breast. Cell 145(6):926–940
Larue L, Bellacosa A (2005) Epithelial-mesenchymal transition in development and cancer: role of phosphatidylinositol 3′ kinase/AKT pathways. Oncogene 24(50):7443–7454
Muraoka-Cook RS, Dumont N, Arteaga CL (2005) Dual role of transforming growth factor beta in mammary tumorigenesis and metastatic progression. Clin Cancer Res Off J Am Assoc Cancer Res 11(2 Pt 2):937s–943s
Marotta LL et al (2011) The JAK2/STAT3 signaling pathway is required for growth of CD44(+)CD24(-) stem cell-like breast cancer cells in human tumors. J Clin Investig 121(7):2723–2735
Niu G et al (2002) Roles of activated Src and Stat3 signaling in melanoma tumor cell growth. Oncogene 21(46):7001–7010
Bowman T et al (2000) STATs in oncogenesis. Oncogene 19(21):2474–2488
Quintas-Cardama A, Verstovsek S (2013) Molecular pathways: Jak/STAT pathway: mutations, inhibitors, and resistance. Clin Cancer Res Off J Am Assoc Cancer Res 19(8):1933–1940
Lu X, Kang Y (2010) Hypoxia and hypoxia-inducible factors: master regulators of metastasis. Clin Cancer Res Off J Am Assoc Cancer Res 16(24):5928–5935
Brabletz T (2012) To differentiate or not–routes towards metastasis. Nat Rev Cancer 12(6):425–436
Lu X et al (2011) VCAM-1 promotes osteolytic expansion of indolent bone micrometastasis of breast cancer by engaging alpha4beta1-positive osteoclast progenitors. Cancer Cell 20(6):701–714
Chen Q, Massague J (2012) Molecular pathways: VCAM-1 as a potential therapeutic target in metastasis. Clin Cancer Res Off J Am Assoc Cancer Res 18(20):5520–5525
Chen Q, Zhang XH, Massague J (2011) Macrophage binding to receptor VCAM-1 transmits survival signals in breast cancer cells that invade the lungs. Cancer Cell 20(4):538–549
Boyce BF, Xing L (2007) Biology of RANK, RANKL, and osteoprotegerin. Arthr Res Ther 9(Suppl 1):S1
Jones DH et al (2006) Regulation of cancer cell migration and bone metastasis by RANKL. Nature 440(7084):692–696
Casimiro S et al (2013) RANKL/RANK/MMP-1 molecular triad contributes to the metastatic phenotype of breast and prostate cancer cells in vitro. PLoS ONE 8(5):e63153
Park HR et al (2003) Expression of osteoprotegerin and RANK ligand in breast cancer bone metastasis. J Korean Med Sci 18(4):541–546
Peng X et al (2013) Differential expression of the RANKL/RANK/OPG system is associated with bone metastasis in human non-small cell lung cancer. PLoS ONE 8(3):e58361
Palafox M et al (2012) RANK induces epithelial-mesenchymal transition and stemness in human mammary epithelial cells and promotes tumorigenesis and metastasis. Cancer Res 72(11):2879–2888
Yin JJ et al (1999) TGF-beta signaling blockade inhibits PTHrP secretion by breast cancer cells and bone metastases development. J Clin Investig 103(2):197–206
Liang Y et al (2012) Transcriptional network analysis identifies BACH1 as a master regulator of breast cancer bone metastasis. J Biol Chem 287(40):33533–33544
Okita Y et al (2013) Transforming growth factor-beta induces transcription factors MafK and Bach1 to suppress expression of the heme oxygenase-1 gene. J Biol Chem 288(28):20658–20667
Shubbar E et al (2013) Elevated cyclin B2 expression in invasive breast carcinoma is associated with unfavorable clinical outcome. BMC Cancer 13:1
Caldon CE et al (2012) Cyclin E2 overexpression is associated with endocrine resistance but not insensitivity to CDK2 inhibition in human breast cancer cells. Mol Cancer Ther 11(7):1488–1499
Payton M et al (2002) Deregulation of cyclin E2 expression and associated kinase activity in primary breast tumors. Oncogene 21(55):8529–8534
Epping MT et al (2011) TSPYL5 suppresses p53 levels and function by physical interaction with USP7. Nat Cell Biol 13(1):102–108
Kim EJ et al (2010) TSPYL5 is involved in cell growth and the resistance to radiation in A549 cells via the regulation of p21(WAF1/Cip1) and PTEN/AKT pathway. Biochem Biophys Res Commun 392(3):448–453
Gulzar ZG, McKenney JK, Brooks JD (2013) Increased expression of NuSAP in recurrent prostate cancer is mediated by E2F1. Oncogene 32(1):70–77
Chou HY et al (2011) Phosphorylation of NuSAP by Cdk1 regulates its interaction with microtubules in mitosis. Cell Cycle 10(23):4083–4089
Raemaekers T et al (2003) NuSAP, a novel microtubule-associated protein involved in mitotic spindle organization. J Cell Biol 162(6):1017–1029
Horn D et al (2010) The conserved mitochondrial twin Cx9C protein Cmc2 Is a Cmc1 homologue essential for cytochrome c oxidase biogenesis. J Biol Chem 285(20):15088–15099
Xu J et al (2013) MiR-223/Ect2/p21 signaling regulates osteosarcoma cell cycle progression and proliferation. Biomed Pharmacother 67(5):381–386
Weeks A et al (2012) ECT2 and RASAL2 mediate mesenchymal-amoeboid transition in human astrocytoma cells. Am J Pathol 181(2):662–674
Cook DR et al (2011) The ect2 rho Guanine nucleotide exchange factor is essential for early mouse development and normal cell cytokinesis and migration. Genes Cancer 2(10):932–942
Thomae AW et al (2011) Different roles of the human Orc6 protein in the replication initiation process. Cell Mol Life Sci CMLS 68(22):3741–3756
Ueki T et al (2008) Involvement of elevated expression of multiple cell-cycle regulator, DTL/RAMP (denticleless/RA-regulated nuclear matrix associated protein), in the growth of breast cancer cells. Oncogene 27(43):5672–5683
Pan HW et al (2006) Role of L2DTL, cell cycle-regulated nuclear and centrosome protein, in aggressive hepatocellular carcinoma. Cell Cycle 5(22):2676–2687
Liu CL et al (2007) L2dtl is essential for cell survival and nuclear division in early mouse embryonic development. J Biol Chem 282(2):1109–1118
Mollinari C et al (2002) PRC1 is a microtubule binding and bundling protein essential to maintain the mitotic spindle midzone. J Cell Biol 157(7):1175–1186
Joshi K et al (2013) MELK-dependent FOXM1 phosphorylation is essential for proliferation of glioma stem cells. Stem Cells 31(6):1051–1063
Gu C et al (2013) Tumor-specific activation of the C-JUN/MELK pathway regulates glioma stem cell growth in a p53-dependent manner. Stem Cells 31(5):870–881
Lin ML et al (2007) Involvement of maternal embryonic leucine zipper kinase (MELK) in mammary carcinogenesis through interaction with Bcl-G, a pro-apoptotic member of the Bcl-2 family. Breast Cancer Res BCR 9(1):R17
Nakano I et al (2008) Maternal embryonic leucine zipper kinase is a key regulator of the proliferation of malignant brain tumors, including brain tumor stem cells. J Neurosci Res 86(1):48–60
Peurala E et al (2012) Expressions of individual PHDs associate with good prognostic factors and increased proliferation in breast cancer patients. Breast Cancer Res Treat 133(1):179–188
Metzen E et al (2005) Regulation of the prolyl hydroxylase domain protein 2 (phd2/egln-1) gene: identification of a functional hypoxia-responsive element. Biochem J 387(Pt 3):711–717
Chan DA et al (2009) Tumor vasculature is regulated by PHD2-mediated angiogenesis and bone marrow-derived cell recruitment. Cancer Cell 15(6):527–538
Mak P et al (2013) Estrogen receptor beta sustains epithelial differentiation by regulating prolyl hydroxylase 2 transcription. Proc Natl Acad Sci USA 110(12):4708–4713
Flavahan WA et al (2013) Brain tumor initiating cells adapt to restricted nutrition through preferential glucose uptake. Nat Neurosci 16(10):1373–1382
Mimura I et al (2012) Dynamic change of chromatin conformation in response to hypoxia enhances the expression of GLUT3 (SLC2A3) by cooperative interaction of hypoxia-inducible factor 1 and KDM3A. Mol Cell Biol 32(15):3018–3032
Sureshbabu A et al (2012) IGFBP5 induces cell adhesion, increases cell survival and inhibits cell migration in MCF-7 human breast cancer cells. J Cell Sci 125(Pt 7):1693–1705
Clark GJ, Der CJ (1995) Aberrant function of the Ras signal transduction pathway in human breast cancer. Breast Cancer Res Treat 35(1):133–144
Aitkenhead M et al (2002) Identification of endothelial cell genes expressed in an in vitro model of angiogenesis: induction of ESM-1, (beta)ig-h3, and NrCAM. Microvasc Res 63(2):159–171
Sonoda E et al (2001) Scc1/Rad21/Mcd1 is required for sister chromatid cohesion and kinetochore function in vertebrate cells. Dev Cell 1(6):759–770
Birkenbihl RP, Subramani S (1992) Cloning and characterization of rad21 an essential gene of Schizosaccharomyces pombe involved in DNA double-strand-break repair. Nucleic Acids Res 20(24):6605–6611
Wu G et al (2003) DeltaNp63alpha and TAp63alpha regulate transcription of genes with distinct biological functions in cancer and development. Cancer Res 63(10):2351–2357
Lammer C et al (1998) The cdc25B phosphatase is essential for the G2/M phase transition in human cells. J Cell Sci 111(Pt 16):2445–2453
Li Y et al (2011) ShRNA-targeted centromere protein A inhibits hepatocellular carcinoma growth. PLoS ONE 6(3):e17794
Tomonaga T et al (2003) Overexpression and mistargeting of centromere protein-A in human primary colorectal cancer. Cancer Res 63(13):3511–3516
Shashni B et al (2013) Glycolytic enzymes PGK1 and PKM2 as novel transcriptional targets of PPARgamma in breast cancer pathophysiology. J Drug Target 21(2):161–174
Fang G, Yu H, Kirschner MW (1998) The checkpoint protein MAD2 and the mitotic regulator CDC20 form a ternary complex with the anaphase-promoting complex to control anaphase initiation. Genes Dev 12(12):1871–1883
Sotillo R et al (2007) Mad2 overexpression promotes aneuploidy and tumorigenesis in mice. Cancer Cell 11(1):9–23
Lan Y et al (2008) Aberrant expression of Cks1 and Cks2 contributes to prostate tumorigenesis by promoting proliferation and inhibiting programmed cell death. Int J Cancer J Int Cancer 123(3):543–551
Kang MA et al (2009) Upregulation of the cycline kinase subunit CKS2 increases cell proliferation rate in gastric cancer. J Cancer Res Clin Oncol 135(6):761–769
Johnson VL et al (2004) Bub1 is required for kinetochore localization of BubR1, Cenp-E, Cenp-F and Mad2, and chromosome congression. J Cell Sci 117(Pt 8):1577–1589
Grabsch H et al (2003) Overexpression of the mitotic checkpoint genes BUB1, BUBR1, and BUB3 in gastric cancer: association with tumour cell proliferation. J Pathol 200(1):16–22
Waltenberger J et al (1994) Different signal transduction properties of KDR and Flt1, two receptors for vascular endothelial growth factor. J Biol Chem 269(43):26988–26995
Yoshiji H et al (1996) Expression of vascular endothelial growth factor, its receptor, and other angiogenic factors in human breast cancer. Cancer Res 56(9):2013–2016
Huegel J et al (2013) Perichondrium phenotype and border function are regulated by Ext1 and heparan sulfate in developing long bones: a mechanism likely deranged in Hereditary Multiple Exostoses. Dev Biol 377(1):100–112
Wang Y et al (2013) Involvement of Ext1 and heparanase in migration of mouse FBJ osteosarcoma cells. Mol Cell Biochem 373(1–2):63–72
Hager MH et al (2012) DIAPH3 governs the cellular transition to the amoeboid tumour phenotype. EMBO Mol Med 4(8):743–760
Gupton SL et al (2007) mDia2 regulates actin and focal adhesion dynamics and organization in the lamella for efficient epithelial cell migration. J Cell Sci 120(Pt 19):3475–3487
Block J et al (2008) Filopodia formation induced by active mDia2/Drf3. J Microsc 231(3):506–517
Wilkinson S, Paterson HF, Marshall CJ (2005) Cdc42-MRCK and Rho-ROCK signalling cooperate in myosin phosphorylation and cell invasion. Nat Cell Biol 7(3):255–261
Balasenthil S et al (2011) A migration signature and plasma biomarker panel for pancreatic adenocarcinoma. Cancer Prev Res (Phila) 4(1):137–149
Barkefors I et al (2011) Exocyst complex component 3-like 2 (EXOC3L2) associates with the exocyst complex and mediates directional migration of endothelial cells. J Biol Chem 286(27):24189–24199
Liu J et al (2012) Exo70 stimulates the Arp2/3 complex for lamellipodia formation and directional cell migration. Curr Biol CB 22(16):1510–1515
Wu Y et al (2007) Neuromedin U is regulated by the metastasis suppressor RhoGDI2 and is a novel promoter of tumor formation, lung metastasis and cancer cachexia. Oncogene 26(5):765–773
Ketterer K et al (2009) Neuromedin U is overexpressed in pancreatic cancer and increases invasiveness via the hepatocyte growth factor c-Met pathway. Cancer Lett 277(1):72–81
Ferrara N, Gerber HP, LeCouter J (2003) The biology of VEGF and its receptors. Nat Med 9(6):669–676
Gerhardt H et al (2003) VEGF guides angiogenic sprouting utilizing endothelial tip cell filopodia. J Cell Biol 161(6):1163–1177
Skobe M et al (2001) Induction of tumor lymphangiogenesis by VEGF-C promotes breast cancer metastasis. Nat Med 7(2):192–198
Hiratsuka S et al (2002) MMP9 induction by vascular endothelial growth factor receptor-1 is involved in lung-specific metastasis. Cancer Cell 2(4):289–300
Belotti D et al (2003) Matrix metalloproteinases (MMP9 and MMP2) induce the release of vascular endothelial growth factor (VEGF) by ovarian carcinoma cells: implications for ascites formation. Cancer Res 63(17):5224–5229
Gauglhofer C et al (2011) Up-regulation of the fibroblast growth factor 8 subfamily in human hepatocellular carcinoma for cell survival and neoangiogenesis. Hepatology 53(3):854–864
Wei W et al (2013) FGF18 as a prognostic and therapeutic biomarker in ovarian cancer. J Clin Investig 123(10):4435–4448
Xu L et al (2000) WISP-1 is a Wnt-1- and beta-catenin-responsive oncogene. Genes Dev 14(5):585–595
Su F et al (2002) WISP-1 attenuates p53-mediated apoptosis in response to DNA damage through activation of the Akt kinase. Genes Dev 16(1):46–57
Liu JF et al (2013) CCN4 induces vascular cell adhesion molecule-1 expression in human synovial fibroblasts and promotes monocyte adhesion. Biochim Biophys Acta 1833(5):966–975
Inkson CA et al (2008) TGF-beta1 and WISP-1/CCN-4 can regulate each other’s activity to cooperatively control osteoblast function. J Cell Biochem 104(5):1865–1878
Ono M et al (2013) WISP1/CCN4: a potential target for inhibiting prostate cancer growth and spread to bone. PLoS ONE 8(8):e71709
Nishi H et al (2004) Hypoxia-inducible factor-1 transactivates transforming growth factor-beta3 in trophoblast. Endocrinology 145(9):4113–4118
Medici D, Hay ED, Olsen BR (2008) Snail and Slug promote epithelial-mesenchymal transition through beta-catenin-T-cell factor-4-dependent expression of transforming growth factor-beta3. Mol Biol Cell 19(11):4875–4887
Nguyen AV, Pollard JW (2000) Transforming growth factor beta3 induces cell death during the first stage of mammary gland involution. Development 127(14):3107–3118
Amiry N et al (2009) Trefoil factor-1 (TFF1) enhances oncogenicity of mammary carcinoma cells. Endocrinology 150(10):4473–4483
Katoh M (2003) Trefoil factors and human gastric cancer (review). Int J Mol Med 12(1):3–9
Ahmed AR et al (2012) TFF3 is a normal breast epithelial protein and is associated with differentiated phenotype in early breast cancer but predisposes to invasion and metastasis in advanced disease. Am J Pathol 180(3):904–916
Wang Z, Hao Y, Lowe AW (2008) The adenocarcinoma-associated antigen, AGR2, promotes tumor growth, cell migration, and cellular transformation. Cancer Res 68(2):492–497
Hrstka R et al (2010) The pro-metastatic protein anterior gradient-2 predicts poor prognosis in tamoxifen-treated breast cancers. Oncogene 29(34):4838–4847
Innes HE et al (2006) Significance of the metastasis-inducing protein AGR2 for outcome in hormonally treated breast cancer patients. Br J Cancer 94(7):1057–1065
Park SW et al (2009) The protein disulfide isomerase AGR2 is essential for production of intestinal mucus. Proc Natl Acad Sci USA 106(17):6950–6955
Tiang JM, Butcher NJ, Minchin RF (2010) Small molecule inhibition of arylamine N-acetyltransferase Type I inhibits proliferation and invasiveness of MDA-MB-231 breast cancer cells. Biochem Biophys Res Commun 393(1):95–100
Tiang JM et al (2011) RNAi-mediated knock-down of arylamine N-acetyltransferase-1 expression induces E-cadherin up-regulation and cell–cell contact growth inhibition. PLoS ONE 6(2):e17031
Lanningham-Foster L et al (2002) Overexpression of CRIP in transgenic mice alters cytokine patterns and the immune response. Am J Physiol Endocrinol Metab 282(6):E1197–E1203
Nobes CD et al (1998) A new member of the Rho family, Rnd1, promotes disassembly of actin filament structures and loss of cell adhesion. J Cell Biol 141(1):187–197
Wang GL et al (2012) The effect of NET-1 on the proliferation, migration and endocytosis of the SMMC-7721 HCC cell line. Oncol Rep 27(6):1944–1952
Chen L et al (2010) Suppression of TSPAN1 by RNA interference inhibits proliferation and invasion of colon cancer cells in vitro. Tumori 96(5):744–750
Cheng CJ et al (2009) SCUBE2 suppresses breast tumor cell proliferation and confers a favorable prognosis in invasive breast cancer. Cancer Res 69(8):3634–3641
Tsai MT et al (2009) Isolation and characterization of a secreted, cell-surface glycoprotein SCUBE2 from humans. Biochem J 422(1):119–128
Ilantzis C et al (2002) Deregulated expression of the human tumor marker CEA and CEA family member CEACAM6 disrupts tissue architecture and blocks colonocyte differentiation. Neoplasia 4(2):151–163
Duxbury MS et al (2004) CEACAM6 gene silencing impairs anoikis resistance and in vivo metastatic ability of pancreatic adenocarcinoma cells. Oncogene 23(2):465–473
Elmarghani A, Abuabaid H, Kjellen P (2009) TOM1L is involved in a novel signaling pathway important for the IL-2 production in Jurkat T cells stimulated by CD3/CD28 co-ligation. Mediators Inflamm 2009:416298
Liu NS et al (2009) Participation of Tom1L1 in EGF-stimulated endocytosis of EGF receptor. The EMBO journal 28(22):3485–3499
Hendrix MJ et al (1996) Role of intermediate filaments in migration, invasion and metastasis. Cancer Metastasis Rev 15(4):507–525
Tassi E et al (2001) Enhancement of fibroblast growth factor (FGF) activity by an FGF-binding protein. J Biol Chem 276(43):40247–40253
Abuharbeid S, Czubayko F, Aigner A (2006) The fibroblast growth factor-binding protein FGF-BP. Int J Biochem Cell Biol 38(9):1463–1468
Medema RH et al (2000) AFX-like Forkhead transcription factors mediate cell-cycle regulation by Ras and PKB through p27kip1. Nature 404(6779):782–787
Rena G et al (1999) Phosphorylation of the transcription factor forkhead family member FKHR by protein kinase B. J Biol Chem 274(24):17179–17183
Brunet A et al (1999) Akt promotes cell survival by phosphorylating and inhibiting a Forkhead transcription factor. Cell 96(6):857–868
Hu MC et al (2004) IkappaB kinase promotes tumorigenesis through inhibition of forkhead FOXO3a. Cell 117(2):225–237
Yang JY et al (2008) ERK promotes tumorigenesis by inhibiting FOXO3a via MDM2-mediated degradation. Nat Cell Biol 10(2):138–148
Khatri S et al (2010) FOXO3a regulates glycolysis via transcriptional control of tumor suppressor TSC1. J Biol Chem 285(21):15960–15965
Morelli C et al (2010) Akt2 inhibition enables the forkhead transcription factor FoxO3a to have a repressive role in estrogen receptor alpha transcriptional activity in breast cancer cells. Mol Cell Biol 30(3):857–870
Zou Y et al (2008) Forkhead box transcription factor FOXO3a suppresses estrogen-dependent breast cancer cell proliferation and tumorigenesis. Breast Cancer Res BCR 10(1):R21
Karadedou CT et al (2012) FOXO3a represses VEGF expression through FOXM1-dependent and -independent mechanisms in breast cancer. Oncogene 31(14):1845–1858
Cano A et al (2000) The transcription factor snail controls epithelial-mesenchymal transitions by repressing E-cadherin expression. Nat Cell Biol 2(2):76–83
Vega S et al (2004) Snail blocks the cell cycle and confers resistance to cell death. Genes Dev 18(10):1131–1143
Gu YM et al (2008) Elevated thymosin beta15 expression is associated with progression and metastasis of non-small cell lung cancer. APMIS Acta Pathol Microbiol Immunol Scand 116(6):484–490
Bao L et al (1996) Thymosin beta 15: a novel regulator of tumor cell motility upregulated in metastatic prostate cancer. Nat Med 2(12):1322–1328
Zeng G et al (2012) METCAM/MUC18 augments migration, invasion, and tumorigenicity of human breast cancer SK-BR-3 cells. Gene 492(1):229–238
Zabouo G et al (2009) CD146 expression is associated with a poor prognosis in human breast tumors and with enhanced motility in breast cancer cell lines. Breast Cancer Res BCR 11(1):R1
Jiang T et al (2012) CD146 is a coreceptor for VEGFR-2 in tumor angiogenesis. Blood 120(11):2330–2339
Stalin J et al (2013) Soluble melanoma cell adhesion molecule (sMCAM/sCD146) promotes angiogenic effects on endothelial progenitor cells through angiomotin. J Biol Chem 288(13):8991–9000
Imbert AM et al (2012) CD146 expression in human breast cancer cell lines induces phenotypic and functional changes observed in Epithelial to Mesenchymal Transition. PLoS ONE 7(8):e43752
Zhang X, et al. (2013) MCAM expression is associated with poor prognosis in non-small cell lung cancer. Clin Transl Oncol. Official publication of the Federation of Spanish Oncology Societies and of the National Cancer Institute of Mexico
Chan DN et al (2012) PTK7 marks the first human developmental EMT in vitro. PLoS ONE 7(11):e50432
Yen WW et al (2009) PTK7 is essential for polarized cell motility and convergent extension during mouse gastrulation. Development 136(12):2039–2048
Lu X et al (2004) PTK7/CCK-4 is a novel regulator of planar cell polarity in vertebrates. Nature 430(6995):93–98
Shin WS et al (2008) Soluble PTK7 inhibits tube formation, migration, and invasion of endothelial cells and angiogenesis. Biochem Biophys Res Commun 371(4):793–798
Golubkov VS et al (2010) The Wnt/planar cell polarity protein-tyrosine kinase-7 (PTK7) is a highly efficient proteolytic target of membrane type-1 matrix metalloproteinase: implications in cancer and embryogenesis. J Biol Chem 285(46):35740–35749
Sun Q, et al. (2013) Overexpression of response gene to complement 32 (RGC32) promotes cell invasion and induces epithelial-mesenchymal transition in lung cancer cells via the NF-kappaB signaling pathway. Tumour Biol J Int Soc Oncodev Biol Med
Sonnylal S et al (2013) Connective tissue growth factor causes EMT-like cell fate changes in vivo and in vitro. J Cell Sci 126(Pt 10):2164–2175
Muratoglu SC, et al. (2013) LRP1 protects the vasculature by regulating levels of connective tissue growth factor and HtrA1. Arterioscler Thromb Vasc Biol
Kondo S et al (2002) Connective tissue growth factor increased by hypoxia may initiate angiogenesis in collaboration with matrix metalloproteinases. Carcinogenesis 23(5):769–776
Lau LF, Lam SC (1999) The CCN family of angiogenic regulators: the integrin connection. Exp Cell Res 248(1):44–57
Chen PS et al (2007) CTGF enhances the motility of breast cancer cells via an integrin-alphavbeta3-ERK1/2-dependent S100A4-upregulated pathway. J Cell Sci 120(Pt 12):2053–2065
Hugo HJ et al (2009) Staurosporine augments EGF-mediated EMT in PMC42-LA cells through actin depolymerisation, focal contact size reduction and Snail1 induction: a model for cross-modulation. BMC Cancer 9:235
Basilico C, Moscatelli D (1992) The FGF family of growth factors and oncogenes. Adv Cancer Res 59:115–165
Giordano FJ et al (1996) Intracoronary gene transfer of fibroblast growth factor-5 increases blood flow and contractile function in an ischemic region of the heart. Nat Med 2(5):534–539
Allerstorfer S et al (2008) FGF5 as an oncogenic factor in human glioblastoma multiforme: autocrine and paracrine activities. Oncogene 27(30):4180–4190
Ricciardelli C et al (2011) The ADAMTS1 protease gene is required for mammary tumor growth and metastasis. Am J Pathol 179(6):3075–3085
Krampert M et al (2005) ADAMTS1 proteinase is up-regulated in wounded skin and regulates migration of fibroblasts and endothelial cells. J Biol Chem 280(25):23844–23852
Esselens C et al (2010) The cleavage of semaphorin 3C induced by ADAMTS1 promotes cell migration. J Biol Chem 285(4):2463–2473
Su SC et al (2008) Molecular profile of endothelial invasion of three-dimensional collagen matrices: insights into angiogenic sprout induction in wound healing. Am J Physiol Cell Physiol 295(5):C1215–C1229
Luque A, Carpizo DR, Iruela-Arispe ML (2003) ADAMTS1/METH1 inhibits endothelial cell proliferation by direct binding and sequestration of VEGF165. J Biol Chem 278(26):23656–23665
Vazquez F et al (1999) METH-1, a human ortholog of ADAMTS-1, and METH-2 are members of a new family of proteins with angio-inhibitory activity. J Biol Chem 274(33):23349–23357
Kucia M et al (2005) Trafficking of normal stem cells and metastasis of cancer stem cells involve similar mechanisms: pivotal role of the SDF-1-CXCR4 axis. Stem Cells 23(7):879–894
Teicher BA, Fricker SP (2010) CXCL12 (SDF-1)/CXCR4 pathway in cancer. Clin Cancer Res Off J Am Assoc Cancer Res 16(11):2927–2931
Wang Z et al (2008) Blockade of SDF-1/CXCR4 signalling inhibits pancreatic cancer progression in vitro via inactivation of canonical Wnt pathway. Br J Cancer 99(10):1695–1703
Mimeault M, Batra SK (2013) Hypoxia-inducing factors as master regulators of stemness properties and altered metabolism of cancer- and metastasis-initiating cells. J Cell Mol Med 17(1):30–54
Conley-Lacomb MK et al (2013) PTEN loss mediated Akt activation promotes prostate tumor growth and metastasis via CXCL12/CXCR4 signaling. Mol Cancer 12(1):85
Guo D, Huang J, Gong J (2012) Bone morphogenetic protein 4 (BMP4) is required for migration and invasion of breast cancer. Mol Cell Biochem 363(1–2):179–190
Onoue T et al (2006) Epithelial-mesenchymal transition induced by the stromal cell-derived factor-1/CXCR4 system in oral squamous cell carcinoma cells. Int J Oncol 29(5):1133–1138
Jung MJ et al (2013) Upregulation of CXCR4 is functionally crucial for maintenance of stemness in drug-resistant non-small cell lung cancer cells. Oncogene 32(2):209–221
Li TM et al (2012) Interleukin-11 increases cell motility and up-regulates intercellular adhesion molecule-1 expression in human chondrosarcoma cells. J Cell Biochem 113(11):3353–3362
Yoshizaki A et al (2006) Expression of interleukin (IL)-11 and IL-11 receptor in human colorectal adenocarcinoma: IL-11 up-regulation of the invasive and proliferative activity of human colorectal carcinoma cells. Int J Oncol 29(4):869–876
Nakayama T et al (2007) Expression of interleukin-11 (IL-11) and IL-11 receptor alpha in human gastric carcinoma and IL-11 upregulates the invasive activity of human gastric carcinoma cells. Int J Oncol 30(4):825–833
Shin SY et al (2012) Transcriptional regulation of the interleukin-11 gene by oncogenic Ras. Carcinogenesis 33(12):2467–2476
Foley CJ et al (2012) Matrix metalloprotease-1a promotes tumorigenesis and metastasis. J Biol Chem 287(29):24330–24338
Gupta GP et al (2007) Mediators of vascular remodelling co-opted for sequential steps in lung metastasis. Nature 446(7137):765–770
Masckauchan TN et al (2006) Wnt5a signaling induces proliferation and survival of endothelial cells in vitro and expression of MMP-1 and Tie-2. Mol Biol Cell 17(12):5163–5172
Reunanen N et al (2002) Activation of p38 alpha MAPK enhances collagenase-1 (matrix metalloproteinase (MMP)-1) and stromelysin-1 (MMP-3) expression by mRNA stabilization. J Biol Chem 277(35):32360–32368
Kim MY et al (2009) Tumor self-seeding by circulating cancer cells. Cell 139(7):1315–1326
Lin J et al (2009) Four and a half LIM domains 1 (FHL1) and receptor interacting protein of 140 kDa (RIP140) interact and cooperate in estrogen signaling. Int J Biochem Cell Biol 41(7):1613–1618
Ding L et al (2009) Human four-and-a-half LIM family members suppress tumor cell growth through a TGF-beta-like signaling pathway. J Clin Investig 119(2):349–361
Lin J et al (2012) FHL family members suppress vascular endothelial growth factor expression through blockade of dimerization of HIF1alpha and HIF1beta. IUBMB Life 64(11):921–930
Tong XK, Hamel E (2007) Transforming growth factor-beta 1 impairs endothelin-1-mediated contraction of brain vessels by inducing mitogen-activated protein (MAP) kinase phosphatase-1 and inhibiting p38 MAP kinase. Mol Pharmacol 72(6):1476–1483
Owens DM, Keyse SM (2007) Differential regulation of MAP kinase signalling by dual-specificity protein phosphatases. Oncogene 26(22):3203–3213
Li M et al (2003) The phosphatase MKP1 is a transcriptional target of p53 involved in cell cycle regulation. J Biol Chem 278(42):41059–41068
Liu YX et al (2008) DUSP1 is controlled by p53 during the cellular response to oxidative stress. Mol Cancer Res MCR 6(4):624–633
Bellou S et al (2009) VEGF autoregulates its proliferative and migratory ERK1/2 and p38 cascades by enhancing the expression of DUSP1 and DUSP5 phosphatases in endothelial cells. Am J Physiol Cell Physiol 297(6):C1477–C1489
Farabegoli F et al (2005) Suppressor of cytokine signalling 2 (SOCS-2) expression in breast carcinoma. J Clin Pathol 58(10):1046–1050
Harris J et al (2006) Socs2 and elf5 mediate prolactin-induced mammary gland development. Mol Endocrinol 20(5):1177–1187
Tannahill GM et al (2005) SOCS2 can enhance interleukin-2 (IL-2) and IL-3 signaling by accelerating SOCS3 degradation. Mol Cell Biol 25(20):9115–9126
Karve TM et al (2012) BRCA1 regulates follistatin function in ovarian cancer and human ovarian surface epithelial cells. PLoS ONE 7(6):e37697
Wordinger RJ et al (2002) Expression of bone morphogenetic proteins (BMP), BMP receptors, and BMP associated proteins in human trabecular meshwork and optic nerve head cells and tissues. Mol Vis 8:241–250
Abe Y et al (2004) Follistatin restricts bone morphogenetic protein (BMP)-2 action on the differentiation of osteoblasts in fetal rat mandibular cells. J Bone Miner Res Off J Am Soc Bone Miner Res 19(8):1302–1307
Fainsod A et al (1997) The dorsalizing and neural inducing gene follistatin is an antagonist of BMP-4. Mech Dev 63(1):39–50
Shimonaka M et al (1991) Follistatin binds to both activin and inhibin through the common subunit. Endocrinology 128(6):3313–3315
Yao HH et al (2004) Follistatin operates downstream of Wnt4 in mammalian ovary organogenesis. Dev Dyn Off Pub Am Assoc Anat 230(2):210–215
Ogino H et al (2008) Follistatin suppresses the production of experimental multiple-organ metastasis by small cell lung cancer cells in natural killer cell-depleted SCID mice. Clin Cancer Res Off J Am Assoc Cancer Res 14(3):660–667
Boedtkjer E et al (2013) Contribution of Na+, HCO3(-)-cotransport to cellular pH control in human breast cancer: a role for the breast cancer susceptibility locus NBCn1 (SLC4A7). Int J Cancer J Int Cancer 132(6):1288–1299
Italiano D et al (2012) Identification of NCF2/p67phox as a novel p53 target gene. Cell Cycle 11(24):4589–4596
Conflict of interest
The authors declare that there is no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fazilaty, H., Mehdipour, P. Genetics of breast cancer bone metastasis: a sequential multistep pattern. Clin Exp Metastasis 31, 595–612 (2014). https://doi.org/10.1007/s10585-014-9642-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10585-014-9642-9