Abstract
Chiasmata are usually formed in the distal half of cereal chromosomes. Previous studies showed that the crossover-rich region displays a more active role in homologous recognition at early meiosis than crossover-poor regions in the long arm of rye chromosome 1R, but not in the long arm of chromosome 5R. In order to determine what happens in other chromosomes of rye and wheat, we have partitioned, by wheat–rye translocations of variable-size, the distal fourth part of chromosome arms 1BS and 2BL of wheat and 1RS and 2RL of rye. Synapsis and chiasma formation in chromosome pairs with homologous (wheat–wheat or rye–rye) and homoeologous (wheat–rye) stretches, positioned distally and proximally, respectively, or vice versa, have been studied by rye chromatin labelling using fluorescence in situ hybridisation. Chromosome arm partitioning showed that the distal 12 % of 1BS form one crossover in 50 % of the cells, while the distal 6.7 % of 2RL and the distal 10.5 % of 2BL account for 94 % and 81 % of chiasmata formed in these arms. Distal homoeologous segments reduce the frequency of chiasmata and the possibility of interaction between the intercalary/proximal homologous segments. Such a reduction is related to the size of the homoeologous (translocated) segment. The effect on synapsis and chiasma formation was much lower in chromosome constructions with distal homology and proximal homoeology. All of these data support that among wheat and rye chromosomes, recombining regions are more often involved in homologous recognition and pairing than crossover-poor regions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Reciprocal exchanges between homologous chromosomes produced at prophase I of meiosis permit the formation of bivalents, which are key structures for the reduction of the chromosome number after chromosome segregation at anaphase I. In addition, new allele combinations generated by recombination contribute to an increase in genetic variability. Meiotic recombination is the underlying mechanism that facilitates the production, and subsequent selection, of genotypes that improve traits of interest in a wide range of crops and domestic animals. Exploitation of meiotic recombination in breeding programs has, however, serious restrictions imposed by the variable distribution of crossovers along the chromosome. Data on crossover distribution in plants have been obtained either from the construction of genetic maps or from cytological observation of crossovers as chiasmata or as late recombination nodules (Mézard et al. 2007). Some species with large chromosomes, such as Fritillaria meleagris (Darlington 1935; Fogwill 1958) or Allium fistulosum (Jones 1984) show proximal localisation of crossovers, but in many others, such as maize, wheat, barley, or rye, crossovers locate distally (Lukaszewski and Curtis 1993; Künzel et al. 2000; Akhunov et al. 2003; Anderson et al. 2004; Lukaszewski 2008; Higgins et al. 2012).
Deletions or inversions are chromosome aberrations that change the position of segments in the telomere-centromere axis. This feature has been used to establish the role of different chromosome regions on chiasma formation in wheat and rye. Hemizygosis for the loss of long terminal segments show a considerable reduction of chiasma frequency in the truncated chromosome arm in wheat. However, the amount of chiasmate associations at metaphase I is restored to a normal level in deficient chromosome homozygotes (Curtis et al. 1991; Endo et al. 1991; Lukaszewski 1995; Gill et al. 1997; Jones et al. 2002; Qi et al. 2002). The construction of genetic maps, in addition to the frequency of chiasmata at metaphase I, demonstrated that the low chiasma frequency of the region located in the middle of the normal chromosome arms 1BL or 5BL, increased in truncated chromosomes with the distal 25 % or 41 % missing, respectively (Jones et al. 2002; Qi et al. 2002). However, the chiasma rate increase concerned the very distal part of the truncated chromosome pair, but not its proximal region. In contrast, chiasmata are infrequent in the proximal third of the 5RL chromosome arm of rye both in normal homozygotes and homozygotes for truncated chromosomes lacking the distal 70 % (Naranjo et al. 2010). This result indicates that crossover formation in 5RL depends on the DNA sequence, or the chromatin organisation, present in each chromosome region. That it is not the position, but the DNA sequence, or chromatin organisation, normally present in the distal part of chromosome arm 1RL that determines the crossover formation, was demonstrated by the presence of a paracentric inversion covering 90–95 % of this arm. This inversion changes the pattern of chiasma distribution; inversion homozygotes and heterozygotes produce only proximal chiasmata in the inverted arm (Lukaszewski 2008; Valenzuela et al. 2012). The pattern of chiasma localisation is also modified in reverse tandem duplications for wheat chromosome arms 2BS and 4AL. The inversion–duplication places the recombining segments in the middle of the arms, and chiasmata, if present, are always located in this recombining region (Lukaszewski et al. 2012).
The use of fluorescence in situ hybridisation (FISH) has permitted studies of the initiation and progression of synapsis in rye chromosomes added to wheat and the determination of whether there is or not some parallelism between the crossover formation and chromosome pairing and synapsis development. By visualizing the behaviour of two 1RS homologues that are distinct in their subtelomeric heterochromatin, Colas et al. (2008) have shown that chromatin remodelling at the onset of meiosis enables the chromosomes to become competent to pair and recombine efficiently. The synaptic pattern of chromosome arm 1RL in homozygotes for the normal or inverted arm, and in heterozygotes, showed that crossover-rich regions are more efficient in finding the homologous partner and developing synapsis than crossover-poor regions (Valenzuela et al. 2012). However, in the case of the 5RL arm, homologous recognition and synapsis is not affected when only the proximal crossover-poor region is present (Naranjo et al. 2010). The behaviour of 1RL and 5RL indicates differences between chromosomes for the links concerning chromosome pairing, synapsis, and recombination. Complex networks of interactions between these processes have been reported (reviewed by Pawlowski and Cande 2005). Such interactions become even more complex in allopolyploid wheats, which have developed genetic systems that ensure a diploid-like meiotic behaviour at metaphase I (reviewed by Sears 1976) by suppression of chiasmata between homoeologous chromosomes (Dubcovski et al. 1995; Luo et al. 1996). In this paper, we analyse the contribution of specific distal or intercalary segments of chromosome arms 1BS and 2BL of wheat and 1RS and 2RL of rye to homology recognition, synapsis, and chiasma formation. Chromosome segments are defined by various-size translocations between either 1BS and 1RS or 2BL and 2RL. Each wheat–rye translocation is studied in both a homologous and a homoeologous context.
Material and methods
Plant material
All plants studied were derived from the series of wheat–rye translocations involving chromosome arms 1BS and 1RS, or 2BL and 2RL, produced by induced homoeologous recombination in a wheat (Triticum aestivum, cv. Pavon 76, 2n = 6x = 42) background and mapped by Lukaszewski et al. (2005). Five 1BS/1RS translocation lines, T-9, T-8, T-26, T-14, and T-6, carrying proximal 1RS and distal 1BS genetic material were crossed to Pavon 76 wheat to produce heterozygotes called PAT (PAvon Translocation) and to the Robertsonian translocation 1RS.1BL of Pavon 76 to produce heterozygotes called ROT (RObertsonian Translocation). Among the 2BL/2RL translocation lines, two different sets were used: translocations 2BL + 12, 2BL + 19, and 2BL + 15, with proximal 2BL and distal 2RL segments, and translocations 2RL-2, 2RL-12, 2RL-10, and 2RL-19, with proximal 2RL and distal 2BL segments. All 2BL/2RL translocations were crossed to Pavon 76 to produce PAT heterozygotes and to the Pavon 76 Robertsonian translocation line 2BS.2RL to produce ROT heterozygotes. The crosses carried out to obtain the PAT and ROT heterozygotes and their chromosome structure are represented in Fig. 1. For the translocated arms, heterozygotes ROT9, ROT8, ROT26, ROT14, ROT6, ROT2RL2, ROT2RL12, ROT2RL10, and ROT2RL19 bear proximal rye–rye homology and distal wheat–rye homoeology, while heterozygotes ROT2BL12, ROT2BL19, and ROT2BL15 show proximal wheat–rye homoeology and distal rye homologous segments. In contrast, proximal homoeologous segments of wheat and rye and distal homologous segments of wheat are present in heterozygotes PAT9, PAT8, PAT26, PAT14, PAT6, PAT2RL2, PAT2RL12, PAT2RL10, and PAT2RL19, while heterozygotes PAT2BL12, PAT2BL19, and PAT2BL15 bear proximal wheat–wheat homology and distal wheat–rye homoeology. The chromosome constitution of all the heterozygotes studied was verified by FISH analysis of root tips in squashed preparations as described for meiosis. All plants used were grown in a greenhouse. At meiosis, one of the three anthers of each flower was checked to establish the meiotic stage and the other two were fixed in 3:1 ethanol–acetic acid, and stored at 4 °C.
Origin and chromosome constitution of the wheat/rye translocation heterozygotes analysed. a Heterozygotes involving translocations between the short arm of rye chromosome 1R (1RS, red) and the short arm of wheat chromosome 1B (1BS, blue). b, c Heterozygotes involving translocations between the long arm of wheat chromosome 2B (2BL, blue) and the long arm of rye chromosome 2R (2RL, red)
Fluorescence in situ hybridisation
Fixed anthers (or root tips) were digested in a pectolytic enzyme mixture, transferred to a clean slide, and gently squashed as previously described (Maestra et al. 2002). For studying synapsis, the following repeat DNA probes were used: clone pAtT4 containing the Arabidopsis telomeric tandem repeat (Richards and Ausubel 1988), clone pAWRC.1 containing a rye-specific centromere repeat (Franki 2001), clone pSc74 containing a rye-specific 480-bp tandem repeat (Bedbrook et al. 1980; Cuadrado and Schwarzacher 1998), and clone pUCM600 containing a rye-specific repeat from the R173 family (González-Garcia et al. 2011).
The analysis of the position of the rye chromosome segments at prophase I was carried out using probes pAWRC.1, pUCM600, and pSc74 labelled by nick translation with biotin-16-dUTP. Labelling of the telomere probe pAt74 by nick translation with digoxigenin-11-dUTP allowed the identification of different early prophase I substages as previously described (Maestra et al. 2002; Corredor et al. 2007; Naranjo et al. 2010). In summary, telomeres form aggregates at early leptotene, which converge in a tight cluster at the leptotene–zygotene transition. Chromatin undergoes a conformational change that results in chromosome elongation, concomitant with the formation of the bouquet structure. The telomere cluster disorganises at mid-zygotene as synapsis progresses. The bouquet is completely disorganised at late-zygotene and pachytene, stages that can be identified by a different degree of chromatin condensation. Chromosomes arms with rye genetic material were identified at metaphase I and anaphase I using probes pUCM600 and pAWRC.1 labelled with biotin-16-dUTP and probe pSc74 labelled with digoxigenin-16-dUTP.
Concentrations of DNA probes in the different hybridisation mixes were 5 ng/μl for pAt74, and 10 ng/μl for pAWRC.1, pSc74, and pUCM600. The digoxigenin-labelled probes were detected with 6–8 ng/μl of the FITC-conjugated anti-digoxigenin antibody (Sigma) in 4B (0.5 % blocking reagent in 4× SSC) and biotin-labelled probes with 10–15 ng/μl of the Cy3-conjugated avidin (Sigma) in 4B.
Images of cells were viewed under an Olympus BX60 fluorescence microscope equipped with an Olympus DP70 CCD camera. Images were optimised for brightness and colour using Adobe Photoshop CS4.
Results
The size of the translocated segments
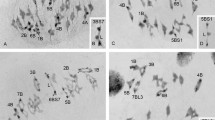
All wheat–rye translocation lines studied in the present work were isolated and identified by Lukaszewski and coworkers (Lukaszewski 2000; Lukaszewski et al. 2003, 2005) as based on the C-banding pattern and the presence/absence of genetic and DNA markers. However, the FISH analysis using genomic DNA as a probe showed limitations in the visualisation of the translocated segment in mitotic chromosomes (Lukaszewski et al. 2005). Among the 1RS/1BS translocations, small wheat-translocated segments with the breakpoint within 7.9 cM of the telomere were masked by the rye chromatin. Because chromosomes are largely elongated at early- and mid-prophase I, we have visualised the translocated segment and identified the break point position in pollen mother cells (PMCs) at pachytene of all wheat–rye translocations studied. In order to establish the translocation breakpoint position, the length of the translocated segment and of the entire rye chromosome arms 1RS and 2RL were measured in cells at pachytene of ROT heterozygotes. In the selected cells, synapsis involving the translocated segment and the entire rye signal was visualised in the same 2D projection (Fig. 2a, b). Meiocytes selected in heterozygotes with proximal rye–rye homology and distal wheat–rye homoeology showed complete or partial subdistal rye–rye synapsis and distal wheat–rye synapsis; the wheat-translocated segment appeared as a terminal chromatin stretch labelled with 4′,6-diamidino-2-phenylindole (DAPI). Cells used in heterozygotes with proximal/intercalary wheat–rye homoeology and distal rye homology showed complete rye-rye synapsis, which involved the entire translocated segment. A number of three to five meiocytes per translocation were measured. The average length of each translocated segment and its fraction relative to the length of the entire rye arm are indicated in Fig. 2c, d. Also indicated are the distal heterochromatin chromomere present in 1RS and the distal and subdistal heterocromatin chromomeres of 2RL. The size of the 1RS/1BS-translocated segments is in the range of 12 % and 18.7 % of the 1RS arm length. All of them were located within the satellite of 1RS (Lukaszewski et al. 2005). The sizes of the 2RL/2BL translocations were between 2.6 % and 11.1 % of the 2RL arm. With the exception of translocation present in ROT2BL15, the breakpoint of 2RL/2BL translocations was located in the interval between the distal and subdistal chromomeres of this arm, which were detected by the pSc74 DNA probe. The translocated segment length measured in ROT heterozygotes also applies to the PAT plants in the positioning of chiasmata (see below).
The size of the translocated segments present in bivalents at pachytene in 1BS/1RS and 2BL/2RL translocation heterozygotes. A1 Nucleus at pachytene of plant ROT2RL19 with the signals of the rye chromatin (red) and telomeres (green). The translocation breakpoint (arrowhead) is inferred from differences in the thickness of the surrounding rye signal. The bright dots correspond to the distal and subdistal chromomeres of 2RL. Synapsis involves the distal homoeologous wheat–rye region (wheat chromatin is labelled with DAPI) and an intercalary homologous segment of rye. A2 The rye chromatin shown in A1. B1 Pachytene nucleus of plant ROT2BL12 showing the translocation breakpoint (arrowhead) and synapsis of the homologous terminal rye segments, the distal chromomere included. B2 The rye chromatin shown in B1. C Size in μm of the 1BS segments (blue) translocated to 1RS (red) in bivalents at pachytene and the corresponding percentage relative to the entire 1RS length. D Same for the 2BL segments (blue) translocated to 2RL and the 2RL segments (red) translocated to 2BL. Green bands on 2RL correspond to the distal and subdistal heterochromatin chromomeres. Scale bar, 10 μm
Synapsis in plants with the ROT chromosome construction
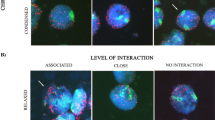
All of these plants carried the translocated chromosome arm accompanied by the rye chromosome arm 1RS or 2RL. The arrangement of the homologous rye segments present in such chromosomes could be established in cells at the leptotene–zygotene transition, mid-zygotene, late-zygotene, and pachytene. According to the degree of homologous rye–rye synapsis, three types of cells were scored in such stages: cells with asynapsis, cells with partial synapsis, and cells with complete synapsis (synapsis degree higher than 90 %). Examples of cells at zygotene are shown in Fig. 3. The distribution of cells with asynapsis, partial synapsis, and complete synapsis in each prophase I substage are represented in Fig. 4. In the two homoeologous groups studied, heterozygotes with proximal rye–rye homology and distal wheat–rye homoeology failed to develop rye–rye synapsis in a variable number of meiocytes. Rye–rye synapsis failure increased in parallel with the size of the wheat-translocated segment. On the other hand, in cells with partial synapsis, the centromere and proximal regions were mainly unsynapsed. This behaviour indicates that the probability of interaction for homologous segments decreases with the distance to the telomere. Accordingly, distal rye homologous segments present in heterozygotes of group 2 with proximal homoeology (ROT2BL12, ROT2BL19, and ROT2BL15) underwent synapsis at zygotene. Synapsis was completed earlier in heterozygotes with smaller-translocated segments.
Arrangement of the rye chromatin (red) and telomeres (green) in cells at zygotene of wheat–rye translocation heterozygotes. A1, 2 Mid-zygotene nucleus of plant ROT8 showing the 1RS homologous segments separated (A2). B1, 2 Late-zygotene nucleus of plant ROT2RL12 with partial synapsis of the 2RL homologous segments (B2). C1, 2 Late-zygotene nucleus of plant ROT14 with complete synapsis of the 1RS homologous segments (C2). The bright dot in A and C and two bright dots in B correspond to the heterochromatic chromomeres of 1RS and 2RL, respectively. Scale bar, 10 μm
Level of synapsis between homologous rye segments in the course of prophase I in wheat/rye translocation heterozygotes arranged in order of increasing length of the translocated segment (LLEZ leptotene–zygotene transition, MZ mid-zygotene, LZ late-zygotene, P pachytene). a 1RS/1BS translocation heterozygotes. b Heterozygotes carrying proximal/intercalary 2RL and distal 2BL segments. c Heterozygotes with proximal/intercalary 2BL and distal 2RL segments. An average number of 93 meiocytes per substage and plant were analysed
The crossover distribution in the arms involved in translocations
The position and frequency of crossovers produced in the arms with the translocated segment were inferred from both the frequency of association at metaphase I and the frequency of recombinant chromosomes at anaphase I. Figure 5 shows the shape of the metaphase I bivalents formed by chromosome pairs involved in translocations 2BL/2RL and the position of the rye signals according to the occurrence or not of one crossover in the homologous region of the two arms. Also indicated is the labelling pattern of the corresponding parental and recombinant chromosomes at anaphase I produced in each case. Examples of cells at metaphase I and anaphase I are shown in Fig. 6. Figure 5a, c is also representative of the chromosome constructions present in plants with translocations 1RS/1BS if the subdistal chromomere is omitted. In cells in metaphase of both ROT and PAT heterozygotes, the arms carrying rye chromatin were associated, in ring bivalents, or unpaired, in rod bivalents with one chiasma between the unlabelled arms of wheat. In cells in anaphase I, homologous chromosomes with distinct parental or recombinant labelling patterns (Fig. 5a, d) were found in heterozygotes carrying proximal homology, either rye–rye or wheat–wheat, and distal homoeology. Recombinant and parental chromosomes showed the same labelling pattern (Fig. 5b, c) in heterozygotes with proximal homoeology and distal homologous segments of wheat or rye. No labelling pattern denoting the occurrence of a crossover between homoeologous segments was observed in chromosomes at anaphase I.
Bivalents at metaphase I and segregating chromosomes at anaphase I with parental (P) or recombinant (R) labelling pattern relative to the translocated segment after the crossovers indicated at pachytene. a Heterozygotes ROT2RL2, ROT2RL12, ROT2RL10, and ROT2RL19. b Heterozygotes ROT2BL12, ROT2BL19, and ROT2BL15. c Heterozygotes PAT2RL2, PAT2RL12, PAT2RL10, and PAT2RL19. d Heterozygotes PAT2BL12, PAT2BL19, and PAT2BL15
Configuration at metaphase I and anaphase I of chromosomes 2BS.2RL, with the normal (N) rye 2RL arm and 2BS.2RL/2BL with the translocated (T) 2RL/2BL arm in the ROT2RL2 heterozygous plant. Chromosome segments of rye (red), the subdistal rye chromomere (green) in T or the distal and subdistal rye chromomeres (green) in N are labelled. a Cell at metaphase I with a labelled ring bivalent. Chiasmate bond between the arms 2RL and 2RL/2BL (arrowheads) impedes their identification. b Rod bivalent showing the arms 2RL and 2RL/2BL separated. The subdistal and distal rye chromomeres are close to one another in N because of chromatin condensation. Although the distal wheat segment of the T arm is masked by the rye chromatin signal, the absence of the distal chromomere is inferred from the smaller size of the chromomeric signal. c Recombinant labelling pattern of the rye chromatin (arrowheads) at anaphase I. d Parental labelling pattern of the N and T arms. Scale bar, 10 μm
The frequency of association at metaphase I for the arms with a translocated segment is indicated in Table 1. This frequency is compared with the frequency of recombinant chromosomes at anaphase I in plants where such chromosomes could be identified. With the exception of plant PAT2BL12, there is a good correspondence between the frequency of association at metaphase I and the frequency of recombination at anaphase I, which supports the idea that all metaphase I associations are chiasmate, and that crossovers occurred in the homologous region of the translocated arms. Accordingly, the frequency of association during metaphase I provided an estimation of the chiasma frequency in plants where recombinant chromosomes and parental chromosomes could not be identified. In both 1RS/1BS and 2BL/2RL translocations, the frequency of chiasmata decreases with the size of the translocated segment in constructions showing subdistal/proximal homology and increases in those with distal homology. The smallest segment of 1BS translocated to 1RS, which represents 12 % of the 1RS arm, forms at least one chiasma in 50 % of meiosis when it is accompanied by 1BS (plant PAT9), while its presence reduces the crossover frequency by almost 40 % when the partner is 1RS (plant ROT9). The largest segment of 1BS translocated to 1RS, with a size of 18.7 % of the 1RS arm, form chiasmata in 71.8 % of PMCs in PAT6, while the intercalary 1RS segment form chiasmata in only 22.1 % of PMCs in plant ROT6. The frequency of chiasmata formed in plants with distal 2BL homology and proximal/subdistal homoeology (plants PAT2RL2, PAT2RL12, PAT2RL10, and PAT2RL19) increases with the translocated segment size from 23.9 % to 81.2 %. In heterozygotes with proximal/subdistal 2RL homology and distal homoeology (plants ROT2RL2, ROT2RL12, ROT2RL10, and RT2RL19), the chiasma frequency decreases from 80.5 % to 60 %. The distal 4.8 % of 2RL present in plant ROT2BL12 forms chiasmata in 70.9 % of PMCs. This frequency increases to 94 % in plant ROT2BL19, which carries a translocated segment occupying 6.7 % of the 2RL arm. In contrast, chiasmata formed in the intercalary wheat segment present in plants PAT2BL12, PAT2BL19, and PAT2BL15 show relatively low values between 29.8 % and 78.7 %.
Discussion
We have quantified the level of synapsis and the frequency of chiasmata in arm pairs composed of both homologous and homoeologous stretches, positioned distally and proximally, respectively, or vice versa. Synapsis of distal homologous segments of chromosome arms 1RS and 2RL is not affected by the presence of intercalary/proximal homoeology, while intercalary/proximal homologous synapsis is conditioned by the size of the distal segment with genetic material of wheat in one chromosome and of rye in the other. This result agrees with previous reports demonstrating that terminal regions of wheat chromosomes 1B and 2B (Corredor et al. 2007) and of the rye chromosome arm 1RL (Lukaszewski 2008; Valenzuela et al. 2012) play an essential role in chromosome interactions leading to recognition, synapsis, and genetic reciprocal exchange of homologous chromosomes. Wheat–rye translocations per se do not restrict the occurrence of at least one chiasma per chromosome arm since, in homozygotes, the translocated arms show frequencies of association at metaphase I that are higher than 95 % (results not shown). Thus, differences between translocation heterozygous plants in the number of chiasmata were caused by the variable size of the translocated segments.
The labelling pattern of recombinant chromosomes at anaphase I denoted the occurrence of at least one crossover in the homologous region. No recombinant chromosome supporting the occurrence of homoeologous recombination was apparent. All plants studied carried the wheat Ph1 homoeologous pairing suppressor that ensures exclusive bivalent formation during metaphase I. In fact, wheat–rye homoeologous recombination is completely suppressed by Ph1 in a mono-2B, mono-2R wheat–rye addition (Corredor et al. 2007). The absence of homoeologous recombination between arms consisting of both homologous and homoeologous segments was reported in plants of wheat heterozygous for introgressed segments of chromosome 1Am from T. monococcum in chromosome 1A of T. aestivum (Luo et al. 1996). No recombination was detected in the homoeologous 1A-1Am segments, irrespective of whether terminally or interstitially positioned in the telomere–centromere axis, whereas the levels of recombination in the juxtaposed homologous 1A–1A segments were normal or close to normal relative to completely homologous 1A chromosomes. However, in a previous report, recombination was significantly reduced in the homologous segment of an arm pair with proximal 1A–1A homology and distal 1A–1Am homoeology (Dubcovski et al. 1995). This is consistent with our result of an inverse relationship between the size of distal homoeologous segments and the number of chiasmata observed during metaphase I and a direct relationship between the size of distal homologous segments and chiasma number. Likewise, the inverse relationships between the level of proximal rye–rye synapsis and the size of the distal homoeologous segment supports that distal homoeology reduces the possibility of encounters between intercalary segments and, therefore, the frequency of homologous crossovers.
With regards to the number and place of chiasmata in chromosome arms of group 2, the distal 11.1 % of 2RL (ROT2BL15) forms at least one chiasma in 93 % of cells, which is a higher frequency than that of 81 % obtained for the distal 10.5 % of 2BL (PAT2RL19). Two translocations of group 2 chromosomes, 2RL − 10, and 2BL + 19, show proximal rye and distal wheat or proximal wheat and distal rye structure, respectively, and contain translocated segments of similar size (4.3 and 4.4 μm in bivalents at pachytene, respectively). These translocations resemble the two products of a reciprocal translocation between 2RL and 2BL. The distal 2RL homologous segments present in plant ROT2BL19 formed more chiasmata (94.2 %) than their 2BL counterparts of plant PAT2RL10 (75 %). On the other hand, the proximal 2RL segments of plant ROT2RL10 also yielded a higher chiasma frequency (73.3 %) than the corresponding 2BL homologous region of plant PAT2BL19 (43.3 %). Thus, the 2RL arm forms more chiasmata in both distal and intercalary regions than 2BL. A genetic map of 2BL involving 22 markers yielded a genetic length of 87.1 cM (Quarrie et al. 2005). This genetic length is somewhat lower than that of 97.3 cM reported in a much higher markers density map of the A and B genomes of tetraploid wheat T. turgidum (Marone et al. 2012). A high-density map or rye lists 2RL at 135 cM (Milczarski et al. 2011). These genetic lengths of 2BL and 2RL are consistent with the higher frequency of chismata of 2RL reported here.
Among group 1 chromosomes, homoeology for the distal 13.3 % (ROT8) or 18.7 % (ROT6) of the arms 1RS and 1RS/1BS reduces the chiasma frequency to 50 % and 22 %, respectively; the corresponding distal homologous segments of the arms 1RS/1BS and1BS (plants PAT8 and PAT6) form chiasmata with frequencies of 47.7 % and 71.8 %, respectively. A distal segment with a given length form less chiasmata in 1BS than in 2BS, and the same seems to be true for 1RS and 2RL. This is in agreement with differences in the genetic lengths, 78.4 and 97.3 cM for 1BS and 2BL, respectively (Marone et al. 2012) and 96 and 135 cM for 1RS and 2RL, respectively (Milczarski et al. 2011). Differences in chiasma rates and distribution between 1S and 2L can be explained in part by differences in length. Short arms of group 1 have a tendency to form single crossovers, while long arms of group 2 often form two. To accommodate the second crossover, the first one should display a narrower distribution and be more distal than when only one crossover is formed. Such a situation has been reported in grasshoppers (Henderson 1963; Southern 1967; Fox 1973).
Partitioning of chromosome arms 2RL and 2BL using wheat–rye translocations has revealed that the distal 6.7 % of 2RL (plant ROT2BL19) and the distal 10.5 % of 2BL (plant PAT2RL19) account for 94 % and 81 %, respectively, of chiasmata formed in these arms. Likewise, 70 % of crossovers concentrate in the 14.4 % distal region of 1BS (plant PAT26), and most likely the same occurs in 1RS. Reduction of chiasma frequency in the 1RS and 2RL arms by homoeology in the crossover-rich region is accompanied by synapsis failure in intercalary proximal regions. In the 1RL arm, 70 % of crossovers occur also in the distal 10 % of this arm, but a paracentric inversion involving the recombinogenic region reduces both the level of synapsis and chiasma frequency (Valenzuela et al. 2012). The arms 1RS, 1RL, and 2RL behave differently than 5RL. Homozygosis for the loss of the distal 70 % of 5RL causes a strong reduction of chiasmata while synapsis is not affected; in 5RL/del5RL heterozygotes no chiasma was observed, but the proximal homologous segment completed synapsis in 70 % of PMCs. Poor-crossover regions of 5RL seem to have a higher capability to synapse than those of 1RS, 1RL, and 2RL. Although synapsis could not be analysed in 1BS and 2BL, their variation in chiasma frequency suggests a behaviour similar to that of 1RS and 2RL.
Gene density along the cereal chromosomes is heterogeneous with a majority of genes clustering in relatively gene-rich recombinogenic distal regions; however, a substantial proportion of the gene complement is outside of these regions (Sandhu and Gill 2002; Mayer et al. 2011). In wheat and barley, more than 30 % of the genes are located in recombinationally cold chromosomal regions (Künzel and Waugh 2002; Erayman et al. 2004; Mayer et al. 2011). Although the distal localisation of crossovers may be a limiting factor in the production by homologous recombination of allele combinations, which can be used in crop improvement, hyperrecombination provoked by the fancm mutation in Arabidopsis (Crismani et al. 2012) suggests the possibility of increasing chiasma frequency in crossover-poor regions. Other strategies of genetic improvement of crop plants may be focused on the transfer of disease resistances and other novel traits present in wild relatives. Such strategies often involve the genetic manipulation of homoeologous recombination. A number of transfers in wheat were carried out in the absence of the homoeologous pairing suppressor effect of Ph1 (Riley et al. 1968; Sears 1977; Knott 1989). Although no homoeologous pairing suppressor gene has been found in tomato, homoeologous recombination is also reduced in Solanum lycopersicoides introgression lines of cultivated tomato relative to the interspecific hybrid (Canady et al. 2006). Homoeologous recombination increases with the size of the introgressed segments, which suggests that exchange between homeologous chromosomes is antagonized by homologous interactions. However, in a given chromosome segment, homoeologous recombination can be increased by suppression of the effect of genes, such as MSH2 and MSH7, involved in the mismatch DNA repair system (Tam et al. 2011). This finding may facilitate introgression in cultivated tomato of very short chromosome segments from wild relatives, which carry useful, but not undesirable, genes.
Abbreviations
- cM:
-
Centimorgan
- DAPI:
-
4′,6-diamidino-2-phenylindole
- FISH:
-
Fluorescence in situ hybridization
- LLEZ:
-
Late leptotene-early zygotene
- MZ:
-
Mid zygotene
- LZ:
-
Late zygotene
- P:
-
Pachytene
- PAT:
-
wheat–rye translocation heterozygotes formed in the cross of 1BL.1RS/1BS translocation homozygotes and the 1BL.1RS Robertsonian translocation line, or 2BS.2RL/2BL and 2BS.2BL/2RL translocation homozygotes and the 2BS.2BL Robersonian translocation line
- PMCs:
-
Pollen mother cells
- ROT:
-
wheat–rye translocation heterozygotes formed in the cross of 1BL.1RS/1BS translocation homozygotes and Pavon 76 wheat, or 2BS.2RL/2BL and 2BS.2BL/2RL translocation homozygotes and Pavon 76 wheat
References
Akhunov ED, Goodyear AW, Geng S, Qi LL, Echalier B et al (2003) The organization and rate of evolution of wheat genomes are correlated with recombination rates along chromosome arms. Genome Res 13:753–763
Anderson LK, Salameh N, Bass HW, Harper LC, Cande WZ et al (2004) Integrating genetic linkage maps with pachytene chromosome structure in maize. Genetics 166:1923–1933
Bedbrook JR, Jones J, O’Dell M, Thompsonand R, Flavell RB (1980) A molecular description of telomeric heterochromatin in Secale species. Cell 19:545–560
Canady MA, Ji Y, Chetelat RT (2006) Homeologous recombination in Solanum lycopersicoides introgression lines of cultivated tomato. Genetics 174:1775–1788
Colas I, Shaw P, Prieto P, Wanous M, Spielmeyer W et al (2008) Effective chromosome pairing requires chromatin remodeling at the onset of meiosis. PNAS 105:6075–6080
Corredor E, Lukaszewski AJ, Pachón P, Allen DC, Naranjo T (2007) Terminal regions of wheat chromosomes select their pairing partners in meiosis. Genetics 177:699–706
Crismani W, Girard C, Froger N, Pradillo M, Santos JL et al (2012) FANCM limits meiotic crossovers. Science 336:1588–1590
Cuadrado A, Schwarzacher T (1998) The chromosomal organization of simple sequence repeats in wheat and rye genomes. Chromosoma 107:587–594
Curtis CA, Lukaszewski AJ, Chrzastek M (1991) Metaphase-I pairing of deficient chromosomes and genetic mapping of deficiency breakpoints in wheat. Genome 34:553–560
Darlington CD (1935) The internal mechanics of the chromosomes II. Prophase paring at meiosis in Fritillaria. Proc Roy Soc Lond B 118:59–73
Dubcovsky J, Luo MC, Dvořák J (1995) Differentiation between homoeologous chromosomes 1A of wheat and 1Am of Triticum monococcum and its recognition by the wheat Ph1 locus. Proc Natl Acad Sci USA 92:6645–6649
Endo TR, Mukai Y, Yamamoto M (1991) Physical mapping of a male-fertility gene of common wheat. Jpn J Genet 66:201–205
Erayman M, Sandhu D, Sidhu D, Dilbirligi M, Baenziger PS et al (2004) Demarcating the gene-rich regions of the wheat genome. Nucleic Acids Res 32:3546–3565
Fogwill M (1958) Differences in crossing over and chromosome size in the sex cells of Lilium and Fritillaria. Chromosoma 9:493–504
Fox DP (1973) The control of chiasma distribution in the locust, Schistocerca gregaria (Forskål). Chromosoma 43:289–328
Franki MG (2001) Identification of Bilby, a diverged centromeric Ty1-copia retrotransposon family from cereal rye (Secale cereale L.). Genome 44:266–274
Gill BS, Gill KS, Friebe B, Endo TR (1997) Expanding genetic maps: reevaluation of the relationship between chiasmata and crossovers. ChromosomToday 12:283–298
González-García M, Cuacos M, González-Sánchez M, Puertas MJ, Vega JM (2011) Painting the rye genome with genome-specific sequences. Genome 54:555–564
Henderson SA (1963) Chiasma distribution at diplotene in a locust Schistocerca gregaria. Heredity 18:173–190
Higgins JD, Perry RM, Barakate A, Ramsay L, Waugh R et al (2012) Spatiotemporal asymmetry of the meiotic program underlies the predominantly distal distribution of meiotic crossovers in barley. Plant Cell 24:4096–4109
Jones GH (1984) The control of chiasma distribution. Symp Soc Exp Biol 38:293–320
Jones LE, Rybka K, Lukaszewski AJ (2002) The effect of a deficiency and a deletion on recombination in chromosome 1BL in wheat. Theor Appl Genet 104:1204–1208
Knott DR (1989) The Wheat Rusts-Breeding for Resistance. Monographs on Theoretical and Applied Genetics 12. Springer Verlag, Berlin
Künzel G, Korzun L, Meister A (2000) Cytologically integrated physical restriction fragment length polymorphism maps for the barley genome based on translocation breakpoints. Genetics 154:397–712
Künzel G, Waugh R (2002) Integration of microsatellite markers into the translocation-based physical RFLP map of barley chromosome 3H. Theor Appl Genet 105:660–665
Lukaszewski AJ (1995) Physical distribution of translocation breakpoints in homoeologous recombinants induced by the absence of the Ph1 gene in wheat and triticale. Theor Appl Genet 90:714–719
Lukaszewski AJ (2000) Manipulation of the 1RS.1BL translocation in wheat by induced homoeologous recombination. Crop Sci 40:216–225
Lukaszewski AJ (2008) Unexpected behaviour of an inverted rye chromosome arm in wheat. Chromosoma 117:569–578
Lukaszewski AJ, Curtis CA (1993) Physical distribution of recombination in B-genome chromosomes of tetraploid wheat. Theor Appl Genet 86:121–127
Lukaszewski AJ, Kopecky D, Linc G (2012) Inversions of chromosome arms 4AL and 2BS in wheat invert the patterns of chiasma distribution. Chromosoma 121:201–208
Lukaszewski AJ, Lapinski B, Ribka K (2005) Limitations of in situ hybridization with total genomic DNA in routine screening for alien introgressions in wheat. Cytogenet Genome Res 109:373–377
Lukaszewski AJ, Rybka K, Korzun V, Malyshev SV, Lapinski B et al (2003) Genetic and physical mapping of homoeologous recombination points involving wheat chromosome 2B and rye chromosome 2R. Genome 47:36–45
Luo MC, Dubcovsky J, Dvořák J (1996) Recognition of homeology by the wheat Ph1 locus. Genetics 144:1195–1203
Maestra B, De Jong JH, Shepherd K, Naranjo T (2002) Chromosome arrangement and behaviour of two rye telosomes at the onset of meiosis in disomic wheat-5RL addition lines with and without the Ph1 locus. Chromosom Res 10:655–667
Marone D, Laido G, Gadaleta A, Colasuonno P, Ficco DBM et al (2012) A high-density consensus map of A and B wheat genomes. Theor Appl Genet 125:1619–1638
Mayer KF, Martis M, Hedley PE, Šimková H, Liu H et al (2011) Unlocking the barley genome by chromosomal and comparative genomics. Plant Cell 23:1249–1263
Mézard C, Vignard J, Drouaud J, Mercier R (2007) The road to crossovers: plants have their say. Trend Genet 23:91–99
Milczarski P, Bolibok-Bragoszewska H, Myśków B, Stojałowski S, Heller-Uszyńska K et al (2011) A high density consensus map of rye (Secale cereale L.) based on DArT markers. PLoS One 6(12):e28495
Naranjo T, Valenzuela NT, Perera E (2010) Chiasma frequency is region-specific and chromosome conformation-dependent in a rye chromosome added to wheat. Cytogenet Genome Res 129:133–142
Pawlowski WP, Cande WZ (2005) Coordinating the events of the meiotic prophase. Trends Cell Biol 15:664–681
Qi LL, Friebe B, Gill BS (2002) A strategy for enhancing recombination in proximal regions of chromosomes. Chromosom Res 10:645–654
Quarrie SA, Steed A, Calestani C, Semikhodskii A, Lebreton C et al (2005) (2005) A high-density genetic map of hexaploid wheat (Triticum aestivum L.) from the cross Chinese Spring x SQ1 and its use to compare QTLs for grain yield across a range of environments. Theor Appl Genet 110:865–880
Richards EJ, Ausubel SM (1988) Isolation of a higher eukaryothic telomere sequence from Arabidopsis thaliana. Cell 53:127–136
Riley R, Capman V, Johnson R (1968) Introduction of yellow rust resistance of Aegilops comosa into wheat by genetically induced homoeologous pairing. Nature 217:383–384
Sandhu D, Gill KS (2002) Gene-containing regions of wheat and the other grass genomes. Plant Physiol 128:803–811
Sears ER (1976) Genetic control of chromosome pairing in wheat. Annu Rev Genet 10:31–51
Sears ER (1977) Analysis of wheat-Agropyron recombinant chromosomes. In: Sánchez-Monge E, García-Olmedo F (eds). Interspecific Hybridization in Plant Breeding. Proceedings of the Eight Congress of EUCARPIA. Madrid, pp 63–72
Southern DI (1967) Chiasma distribution in Truxaline grasshoppers. Chromosoma 22:164–191
Tam SM, Hays JB, Chetelat RT (2011) Effects of suppressing the DNA mismatch repair system on homeologous recombination in tomato. Theor Appl Genet 123:1445–1458
Stephan W, Langley CH (1998) DNA polymorphism in Lycopersicon and crossing-over per physical length. Genetics 150:1585–1593
Valenzuela NT, Perera E, Naranjo T (2012) Dynamics of rye chromosome 1R regions with high and low crossover frequency in homology search and synapsis development. PLoS One 7(4):e36385. doi:10.1371/journal.pone.0036385
Acknowledgments
We thank A. Lukaszewski for kindly supplying homozygous lines for wheat–rye translocations. This work has been supported by grants: BFU2007-66544 from Dirección General de Investigación, Ministerio de Educación y Ciencia of Spain, and CONICYT for PhD study from the Chile Government.
Ethical standards
The experiments comply with the current laws of the country, Spain, in which they were performed.
Conflict of interests
The authors (Nohelia T. Valenzuela, Esther Perera, and Tomás Naranjo) declare that they have no conflict of interests.
Studies with human or animal subjects
This article does not contain any studies with human or animal subjects performed by the any of the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Hans de Jong.
Rights and permissions
About this article
Cite this article
Valenzuela, N.T., Perera, E. & Naranjo, T. Identifying crossover-rich regions and their effect on meiotic homologous interactions by partitioning chromosome arms of wheat and rye. Chromosome Res 21, 433–445 (2013). https://doi.org/10.1007/s10577-013-9372-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-013-9372-x