Abstract
The genetic classification for the N-genome chromosomes has been developed on the basis of C-banding analysis on the set of Triticum aestivum × Aegilops uniaristata single chromosome addition lines and examination of A. uniaristata (\( {\text{2n = 2}} \times { = 14} \), NN), Aegilops ventricosa (\( {\text{2n = 4}} \times { = 28} \), DDNN) and Aegilops recta (\( {\text{2n = 6}} \times { = 42} \), UUXnXnNN) accessions carrying intergenomic translocations using fluorescence in situ hybridisation with probes for three repetitive DNA sequences as well as the 5S and 45S rDNA families. The N-genome chromosomes of the tetraploid A. ventricosa show significant changes relative to the diploid progenitor species, while those of the hexaploid A. recta are similar to A. uniaristata with regard to the distribution of C-bands, 45S and 5S rDNA loci and hybridisation sites of all the three families of tandem repeats. The possible mechanisms of N-genome evolution are discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Aegilops uniaristata Vis. is a diploid species similar in plant morphology to Aegilops comosa Sm. in Sibth. & Sm., and in most taxonomic systems, they are placed in a common section, Comopyrum (Zhukovsky 1928; Eig 1929). However, based on karyotype analysis, Chennaveeraiah (1960) suggested that A. uniaristata possesses a distinct genome and assigned it the genome formula N. The N-genome is also present in two polyploid Aegilops species, Aegilops ventricosa Tausch. and Aegilops recta (Zhuk.) Chennav. (Matsumoto et al. 1957; Chennaveeraiah 1960; Morris and Sears 1967; Kimber and Tsunewaki 1989). The analysis of meiotic chromosome pairing in interspecific hybrids (Kimber et al. 1983; Yen and Kimber 1992), comparison of chromosome morphologies (Chennaveeraiah 1960) and distribution of highly repetitive DNA probes (Bardsley et al. 1999; Badaeva et al. 2002) showed little modification of the N-genome of A. ventricosa relative to the diploid progenitor species. The investigation of the hexaploid A. recta produced more controversial results. Thus, Chennaveeraiah (1960) found only slight resemblance of the N-genome of A. recta to a set of A. uniaristata chromosomes. Meiotic analysis of A. recta × A. uniaristata (4×) hybrids failed to detect an N-genome in A. recta (Yen and Kimber 1992). However, C-banding and fluorescence in situ hybridisation (FISH) studies revealed close similarity between the third genome of this species and the N-genome of A. uniaristata (Badaeva et al. 2004).
The N-genome chromosomes of A. uniaristata have been previously classified based on plant morphology (Miller et al. 1997), cytological principles (Badaeva et al. 1996a; Friebe et al. 1996) or the results of FISH analysis of Triticum aestivum–A. uniaristata addition lines (Iqbal et al. 2000). These classifications, however, are not wholly consistent in their designation of four acrocentric chromosomes, 2N, 3N, 6N, and 7N, due to significant morphological alterations of these chromosomes as a result of asymmetric pericentric inversions (Iqbal et al. 1998). Strong similarities in the lengths and arm ratios of acrocentric chromosomes hampered the establishment of correspondence between their C-banding and FISH patterns. Therefore, the development of a standard genetic nomenclature for the N-genome chromosomes is necessary in which data on plant morphology, C-banding and FISH patterns will be considered together.
The aim of this study was to develop a standard genetic classification for the N-genome chromosomes on the basis of C-banding analysis of a set of T. aestivum–A. uniaristata addition lines and comparison of the N-genomes of diploid and polyploid Aegilops species using C-banding and FISH with probes for three highly repeated non-coding DNA sequences (pAs1, pSc119.2 and Fat) and the 45S and 5S rDNAs.
Materials and methods
Six T. aestivum L. em Thell. cv. ‘Chinese Spring’–A. uniaristata addition lines maintained at the John Innes Centre, Norwich, UK (as described in (Iqbal et al. 2000)) were studied by C-banding. A. uniaristata, A. ventricosa and A. recta accessions carrying intergenomic translocations previously selected using the C-banding method (Badaeva E.D., unpublished) were used for FISH analysis.
The C-banding protocol followed that of Badaeva et al. (1994). The chromosomes of A. uniaristata were first identified in the T. aestivum–A. uniaristata addition set and were classified accordingly. Chromosomal preparations and FISH were carried out according to Badaeva et al. (1996a). Five DNA probes were used: pAs1 (Rayburn and Gill 1986), pSc119.2 (Bedbrook et al. 1980), Fat, pTa71 (45S rDNA; Gerlach and Bedbrook 1979) and pTa794 (5S rDNA; Gerlach and Dyer 1980). The probes pAs1, pSc119.2, pTa71, and pTa794 were labelled with biotin or digoxigenin by nick translation, according to the manufacturer’s protocol (Roche, Germany), and the Fat element probe was labelled with fluorescein using PCR, as described earlier (Badaeva et al. 2010). The slides were counterstained with 2 μg ml−1 4′,6-diamidino-2-phenylindole in 1× phosphate-buffered saline, mounted in Vectashield medium (Vector Laboratories, Peterborough, UK) and examined on an Imager D1 microscope (Carl Zeiss, Germany). Metaphase cells were captured with an AxioCam HRm black-and-white camera using AvioVision software, release 4.6. The relative positions of sites generated by different repetitive DNA sequences were determined by subsequent hybridisation of the same metaphase plates with the pTa71+pTa794 and pAs1+pSc119.2 or pAs1+Fat probe combinations, in that order.
Results
The examination of six T. aestivum cv. Chinese Spring–A. uniaristata single chromosome addition lines using the C-banding technique confirms the presence of all common wheat chromosomes and permits classification of six out of the seven chromosomes of the N-genome (Fig. 1). All N-genome chromosomes added to wheat are similar in morphology and C-banding patterns to the respective chromosomes of the diploid A. uniaristata, indicating that the development of addition lines does not induce any gross changes in alien chromosomes. However, a translocation T3B:5B (T3BS·5BS+T3BL·5BL) has been identified in the addition line 1N. Although RFLP analysis had revealed a partial deletion of the short arm of chromosome 2N in the respective addition line (Iqbal et al. 2000), the lack of obvious changes in morphology and C-banding pattern relative to this chromosome in the diploid species suggests that this deletion involves a very small chromosomal region.
Genetic classification of the C-banded N-genome chromosomes in the addition lines (adds), Aegilops uniaristata (PI554418, k-1589, TA2768), Aegilops ventricosa (INRA #7, k-2750, INRA #10) and Aegilops recta (k-1638, k-3265, k-1654). Translocated chromosomes are indicated. The positions of new C-bands on A. ventricosa or A. recta chromosomes are indicated by long arrows; the positions of C-bands that are eliminated from the N-genome chromosomes of A. ventricosa are indicated by short arrows
The genetic nomenclature developed in this study for the C-banded N-genome chromosomes only partially corresponds to previously suggested classifications (Badaeva et al. 1996a; Friebe et al. 1996). This is because earlier systems were based on the similarity of homoeologous chromosomes, but the acrocentric chromosomes of A. uniaristata are substantially modified and differ from wheat chromosomes in both morphology and C-banding patterns.
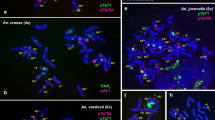
Genetic classification for the N-genome chromosomes labelled by pAs1 and pSc119.2 probes has been suggested by Iqbal et al. (2000) on the basis of FISH analysis of T. aestivum–A. uniaristata addition lines. In addition to these data, the correspondence between C-banding and FISH patterns of individual N-genome chromosomes was verified through analysis of translocation lines. Previously, by means of C-banding, we identified translocations T1N·5N and T3N·5N in A. uniaristata (accessions TA2768 and k-1589); translocations T3D·1N and T5D·5N in A. ventricosa (lines #10 and k-2750), and two other translocations, T6Xn·6N and T2Xn·6N, in A. recta (accession k-3265 and k-1654). These lines were examined here by FISH with five DNA probes (Fig. 2). Combining the C-banding and FISH techniques allowed us to propose a unified system of genetic classification for the N-genome chromosomes, as summarised in Fig. 3. The results obtained allowed us to correct existing classifications of the A. ventricosa 2N and 3N chromosomes, as suggested in our previous paper (Badaeva et al. 2008). The chromosome that was earlier classified as ‘3N’ is actually more similar to A. uniaristata chromosome 2N, whereas A. ventricosa chromosome ‘2N’ is distinct from any of the A. uniaristata chromosomes, although it is a little more similar to the 3N chromosome than the 2N chromosome from the diploid species.
Localisation of cloned repetitive DNA sequences on the chromosomes of Aegilops uniaristata accessions (a TA2768, d k-1589, g k-3310), Aegilops ventricosa (b ssd32, e k-2750T, h k-2750N) and Aegilops recta (c k-2509, f k-3265, i k-3264) using FISH with the following DNA probe combinations: a pSc119.2 (red) and Fat (green); b, c pAs1 (red) and Fat (green); d, e, f pSc119.2 (red) and pAs1 (green); g, h, i pTa71 (red) and pTa794 (green). The N-genome chromosomes are designated according to our genetic nomenclature; chromosomes involved in translocations are indicated with arrows. Scale bar, 10 μm
An idiogram of the N-genome chromosomes in the karyotypes of Aegilops uniaristata, Aegilops ventricosa and Aegilops recta showing chromosome morphology and the positions of the pAs1 and pSc119.2 repetitive DNA sequences, the sites of pTa71 (45S rDNA) and pTa794 (5S rDNA) probes, distribution of the Fat element and the location of heterochromatin bands as revealed by C-banding. The positions of new bands, C-bands or FISH sites on A. ventricosa chromosomes are indicated by asterisks; the positions of bands or FISH sites that are eliminated from the N-genome chromosomes of A. ventricosa are indicated by arrows
According to our C-banding and FISH analyses, the N-genome of A. recta remains nearly unchanged relative to its diploid donor, as only minor differences are observed in the C-banding and FISH patterns of the 1N and 5N chromosomes. The nucleolar organising regions (NORs) on chromosome 5N of A. recta are suppressed, as evidenced by the lack of secondary constriction and the significant reduction in pTa71 signal strength (Fig. 2i), meaning that in this species, amphiplasty is associated with a partial loss of 45S rRNA gene copies.
At the same time, the N-genome of A. ventricosa is substantially modified compared with A. uniaristata. Changes in the C-banding patterns of the N-genome chromosomes are due to emergence (Fig. 1, indicated by long arrows) or elimination (Fig. 1, indicated by short arrows) of particular C-bands or changes in their sizes (Figs. 1 and 3). The heterochromatin content of chromosomes 3N and 7N is substantially decreased, whereas chromosomes 2N and 4N have become more heterochromatic (Fig. 1). The amount of hybridisation with the pAs1 sequences on chromosomes 1N, 4N, 6N and 7N is reduced, due to both the decreased number of sites and the smaller signal strengths (Figs. 2d, e and 3). The distribution of pSc119.2 or/and Fat sequences on some A. ventricosa chromosomes has also been altered. Chromosome 4N contains only one pSc119.2 site, in its long arm, and a bright cluster of Fat element hybridisation was observed only in the proximal part of its short arm (Fig. 2a, b). A. ventricosa chromosome 1N has lost two of the three minor NORs (Fig. 3h). A minor pTa71 locus is also missing from the long arm of chromosome 7N. At the same time, two unequal, minor NORs appear in the terminal regions of a pair of 3D chromosomes. Such loci have not been observed earlier in Aegilops tauschii (Badaeva et al. 1996b) or A. ventricosa (Bardsley et al. 1999; Badaeva et al. 2002).
Discussion
Development of the genetic classification of A. uniaristata chromosomes on the basis of integrated results from classical (C-banding) and a molecular-cytogenetic (FISH) analyses extends our knowledge of the N-genome organisation and evolution. The N-genome was first recognised by Chennaveeraiah (1960), who found that A. uniaristata has a unique karyotype structure. His observation was further supported by the examination of C-banding and FISH patterns of chromosomes (Teoh and Hutchinson 1983; Teoh et al. 1983; Badaeva et al. 1996a, b; Friebe et al. 1996; Iqbal et al. 2000) and by meiotic (Kimber et al. 1983) and isozyme (Kawahara 2000) analyses. However, despite being genetically distant, the N-genome of A. uniaristata shares some common features with the genomes of other diploid Aegilops species. Its high intensity of pAs1 and Fat probe hybridisation is similar to that of the D-genome of A. tauschii, and they both have one major NOR, located on the group 5 chromosomes. Like A. uniaristata, other species, A. comosa var. comosa and var. subventricosa Boiss. (syn. Aegilops heldreichii; M-genome), belonging to the Comopyrum section carry numerous minor NORs and show distinct patterns of hybridisation with the pAs1 sequence (Badaeva et al. 1996a). A. uniaristata is similar to Aegilops speltoides Tausch. (S-genome), as both have large segments of pericentromeric heterochromatin; however, the labelling patterns for the pSc119.2, pAs1 and Fat probes on the S-genome chromosomes, as well as the number and position of 45S and 5S rDNA loci, differ from those seen in the N-genome (Badaeva et al. 1996a, b, 2010). Other Aegilops species belonging to the Emarginata section (S-genomes), Aegilops umbellulata Zhuk. (U-genome), Aegilops caudata L. (C-genome) and Amblyopyrum muticum (Boiss.) Eig (Syn. Aegilops mutica) are similar to A. uniaristata, in that they have large pSc119.2 sites that are predominantly found in terminal regions. However, the chromosomes of these species have a different morphology and show much poorer labelling with pAs1 and Fat sequences, and their C-bands are located mostly in interstitial chromosome regions. These facts suggest that the N-genome is genetically closer to the M- and D-genomes and derived from a common putative diploid ancestor as a result of chromosomal rearrangements and reorganisation of repetitive DNA sequences.
Construction of the genetic map of A. uniaristata chromosome 3N revealed the important role that asymmetric pericentric inversions played in the evolution of N-genome (Iqbal et al. 2000). This mechanism led to the formation of all four acrocentric chromosomes of A. uniaristata and caused transposition of the 5S rDNA locus from the short arm of chromosome 1N to its long arm. The unusual distribution of the Fat element probe that we observed on some N-genome chromosomes could have occurred as a result of these inversions. As was shown before, Fat sequences tend to be located in proximal regions, and signal intensity decreases toward the distal chromosome ends (Badaeva et al. 2010). Therefore, an asymmetric pericentric inversion with breakpoints located close to the centromere in one arm (heavy fluorescence) and distant from it in the opposite arm (faint fluorescence) will lead to the appearance of a low fluorescing area within a brightly fluorescing region, as is seen on chromosomes 2N and 3N. It is interesting to note that two minor 45S rDNA sites in the long arms of these chromosomes are positioned within the regions with pale fluorescence.
Evolution of the N-genome was also associated with amplification, elimination and re-distribution of the different families of repetitive DNA sequences. The most interesting example is the emergence of pericentromeric heterochromatin bands on A. uniaristata chromosomes, which was probably caused by massive amplification of certain classes of highly repetitive DNA sequences. Large pericentromeric C-bands are also found on the chromosomes of A. speltoides; however, the results of a comparative study of several Aegilops species using a TC22b probe, a non-coding repetitive DNA sequence (Teoh et al. 1983), suggest that the molecular composition of pericentromeric heterochromatin in these species is different. Bardsley et al. (1999) also found that the SSR repeats (AAC)5, (ACG)10 and (CGT)10, which are the major components of C-bands in wheat and many Aegilops species, all hybridise poorly to the N-genome chromosomes. Pericentromeric C-bands on A. uniaristata chromosomes are not formed by the pAs1 sequence, either, as their locations do not coincide. Moreover, the long arm of chromosome 7N, which displayed the highest amount of hybridisation with the pAs1 sequence, contains only a few very faint interstitial C-bands, and conversely, the most heterochromatic chromosome, 4N, possesses only two very small pAs1 sites. Thus, the molecular composition and organisation of heterochromatin in the A. uniaristata genome is distinct from other species and remains to be characterised.
The formation of polyploid Aegilops species can induce structural and functional changes in the N-genome, and the extent of these modifications varies between different polyploids. The only obvious change seen in the N-genome of A. recta is the suppression of nucleolar activity on chromosome 5N. In contrast, we found many modifications to N-genome chromosomes in A. ventricosa, including small changes to chromosome morphology, likely due to minor chromosomal rearrangements, heterochromatin re-patterning and an overall decrease in pAs1 probe hybridisation and alterations to its distribution. Also, the elimination of minor NORs on chromosomes 1N and 7N (and the emergence of new loci on the chromosome 3D) and the loss of several pSc119.2 sites are also indicative of structural change. These data show that the level of N-genome modification is higher in A. ventricosa than in A. recta, which is concordant with the results of molecular studies (Dubkovsky and Dvořák 1994; Resta et al. 1996). The analysis of meiotic chromosome pairing in hybrids leads to the opposite conclusion (Yen and Kimber 1992), however. These discrepancies suggest that meiosis in such hybrids is controlled by a very complex genetic system, and the failure of pairing between the orthologous chromosomes is due not only to structural divergence but also to regulatory genes located on chromosomes from the same or different genomes.
Abbreviations
- FISH:
-
Fluorescence in situ hybridisation
- NORs:
-
Nucleolar organising regions
- RFLP:
-
Restriction fragment length polymorphism
- PCR:
-
Polymerase chain reaction
References
Badaeva ED, Badaev NS, Gill BS, Filatenko AA (1994) Intraspecific karyotype divergence in Triticum araraticum (Poaceae). Plant Syst Evol 192:117–145
Badaeva ED, Friebe B, Gill BS (1996a) Genome differentiation in Aegilops. 1. Distribution of highly repetitive DNA sequences on chromosomes of diploid species. Genome 39:293–306
Badaeva ED, Friebe B, Gill BS (1996b) Genome differentiation in Aegilops. 2. Physical mapping of 5S and 18S–26S ribosomal RNA gene families in diploid species. Genome 39:1150–1158
Badaeva ED, Amosova AV, Muravenko OV et al (2002) Genome differentiation in Aegilops. 3. Evolution of the D-genome cluster. Plant Syst Evol 231:163–190
Badaeva ED, Amosova AV, Samatadze TE et al (2004) Genome differentiation in Aegilops. 4. Evolution of the U-genome cluster. Plant Syst Evol 246:45–76
Badaeva E, Dedkova O, Koenig J, Bernard S, Bernard M (2008) Analysis of introgression of Aegilops ventricosa Tausch. genetic material in a common wheat background using C-banding. Theor Appl Genet 117:803–811
Badaeva E, Zoshchuk S, Paux E et al (2010) Fat element—a new marker for chromosome and genome analysis in the Triticeae. Chrom Res 18:697–709
Bardsley D, Cuadrado A, Jack P et al (1999) Chromosome markers in the tetraploid wheat Aegilops ventricosa analysed by in situ hybridization. Theor Appl Genet 99:300–304
Bedbrook RJ, Jones J, O’Dell M, Thompson RJ, Flavell RB (1980) A molecular description of telomeric heterochromatin in Secale species. Cell 19:545–560
Chennaveeraiah MS (1960) Karyomorphologic and cytotaxonomic studies in Aegilops. Acta Hortic Gotob 23:85–186
Dubkovsky J, Dvořák J (1994) Genome origin of Triticum cylindricum, Triticum triunciale, and Triticum ventricosum (Poaceae) inferred from variation in repeated nucleotide sequences: A methodological study. Am J Bot 81:1327–1335
Eig A (1929) Monographisch-kritische Ubersicht der Gattung Aegilops. Beihefte, Berlin
Friebe B, Badaeva ED, Hammer K, Gill BS (1996) Standard karyotypes of Aegilops uniaristata, Ae. mutica, Ae. comosa subspecies comosa and heldreichii (Poaceae). Plant Syst Evol 202:199–210
Gerlach WL, Bedbrook JR (1979) Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acid Res 7:1869–1885
Gerlach WL, Dyer TA (1980) Sequence organization of the repeated units in the nucleus of wheat which contains 5S-rRNA genes. Nucleic Acid Res 8:4851–4865
Iqbal N, Reader SM, Caligari PDS, Miller TE (1998) Characterization of Aegilops uniaristata chromosomes using molecular markers. In: Slinkard AE (ed) Proceedings of the 9th International Wheat Genetics Symposium, 2–7 August 1998. Printcrafters Inc, Saskatoon, pp 114–116
Iqbal N, Reader SM, Caligari PDS, Miller TE (2000) Characterization of Aegilops uniaristata chromosomes by comparative DNA marker analysis and repetitive DNA sequence in situ hybridization. Theor Appl Genet 101:1173–1179
Kawahara T (2000) Isozyme variation in species of the section Comopyrum of Aegilops. Genet Res Crop Evol 47:641–645
Kimber G, Tsunewaki K (1989) Genome symbols and plasma types in the wheat group. Ann Wheat Newslett 35:24–26
Kimber G, Pignone D, Sallee PJ (1983) The relationships of the M and Mu genomes of Triticum. Can J Genet Cytol 25:509–512
Matsumoto K, Shimotsuma M, Nezu M (1957) The amphiploid MuMuDD and its hybrid with Aegilops ventricosa. Wheat Inf Serv 5:12–13
Miller TE, Iqbal N, Reader SM et al (1997) A cytogenetic approach to the improvement of aluminium tolerance in wheat. New Phytol 137:93–98
Morris R, Sears ER (1967) The cytogenetics of wheat and its relatives. In: Quisenberry RS, Reitz LP (eds) Wheat and wheat improvement. ASA, Madison, USA, pp 19–87
Rayburn AL, Gill BS (1986) Isolation of a D-genome specific repeated DNA sequence from Aegilops squarrosa. Plant Mol Biol Rep 4:104–109
Resta P, Zhang HB, Dubkovsky J, Dvořák J (1996) The origin of the genomes of Triticum biunciale, T. ovatum, T. neglectum, T. columnare, and T. rectum based on variation in repeated nucleotide sequences. Amer J Bot 83:1556–1565
Teoh SB, Hutchinson J (1983) Interspecific variation in C-banded chromosomes of diploid Aegilops species. Theor Appl Genet 65:31–40
Teoh SB, Hutchinson J, Miller TE (1983) A comparison of the chromosomal distribution of cloned repetitive DNA sequences in different Aegilops species. Heredity 51:635–641
Yen Y, Kimber G (1992) Genomic relationships on N-genome Triticum species. Genome 35:962–966
Zhukovsky PM (1928) A critical-systematical survey of the species of the genus Aegilops L. Bull Appl Bot Genet Pl Breed 18:417–609
Acknowledgements
The authors thank Dr. N.N. Chikida (N.I. Vavilov Institute of Plant Industry VIR, St-Petersburg, Russia), Dr. H. Bockelman (Small Grain Collection, Aberdeen, Idaho, USA) and Dr. M. Feldman (Weizmann Institute of Science, Rehovot, Israel) for providing the materials for this study. This work was supported by a grant from the Russian State Foundation for Basic Research (project nos. 11-04-00126 and 11-04-00214).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Hans de Jong.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(JPEG 5849 kb)
Rights and permissions
About this article
Cite this article
Badaeva, E.D., Dedkova, O.S., Zoshchuk, S.A. et al. Comparative analysis of the N-genome in diploid and polyploid Aegilops species. Chromosome Res 19, 541–548 (2011). https://doi.org/10.1007/s10577-011-9211-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-011-9211-x