Abstract
Neurodegenerative disorders such as amyotrophic lateral sclerosis (ALS), spinal muscular atrophy (SMA), Parkinson’s, Alzheimer’s, and Huntington’s disease affect a rapidly increasing population worldwide. Although common pathogenic mechanisms have been identified (e.g., protein aggregation or dysfunction, immune response alteration and axonal degeneration), the molecular events underlying timing, dosage, expression, and location of RNA molecules are still not fully elucidated. In particular, the alternative splicing (AS) mechanism is a crucial player in RNA processing and represents a fundamental determinant for brain development, as well as for the physiological functions of neuronal circuits. Although in recent years our knowledge of AS events has increased substantially, deciphering the molecular interconnections between splicing and ALS remains a complex task and still requires considerable efforts. In the present review, we will summarize the current scientific evidence outlining the involvement of AS in the pathogenic processes of ALS. We will also focus on recent insights concerning the tuning of splicing mechanisms by epigenomic and epi-transcriptomic regulation, providing an overview of the available genomic technologies to investigate AS drivers on a genome-wide scale, even at a single-cell level resolution. In the future, gene therapy strategies and RNA-based technologies may be utilized to intercept or modulate the splicing mechanism and produce beneficial effects against ALS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
An increasing number of debilitating human illnesses (cancer, muscular dystrophies, and neurodegenerative disorders) are caused by RNA metabolism defects. One of the pivotal participants in the RNA processing is alternative splicing (AS) whose task is to control and diversify gene expression, monitoring the protein productions of more than 90% of the exon-coding genes (Wang and Cooper 2007). Splicing process abnormalities (i.e., mutations in the intron–exon boundaries or in the exonic/intronic RNA regulatory silencer and enhancer elements) and defects in the spliceosome machinery or in the RNA-binding proteins (RBPs) are known to influence disease pathogenesis, and can represent a direct cause or possible modulators of disease susceptibility and severity (La Cognata et al. 2015; Nissim-Rafinia and Kerem 2005; Vanderweyde et al. 2013; Verma et al. 2018). Despite the considerable role of AS program in all aspects of neuronal development (from neurogenesis to mature synaptic functions) and the remarkable efforts of the scientific community to decipher the complexity of splicing regulation, our knowledge about AS in the ALS context is still not completely elucidated.
Herein, we summarize the AS regulation of ALS-related genes and highlight the contribution of splicing changes in pathology. We also discuss the regulation of splicing mechanisms by epigenomic and epi-transcriptomic events, introducing the new technological advances that enable to investigate AS isoforms at a single-cell level.
RNA-based therapeutic applications are quickly becoming a reality in the treatment of complex diseases (i.e., cancer, ocular and cardiovascular diseases, spinal muscular atrophy, Duchenne’s muscular dystrophy, and Alzheimer’s disease) (Baralle and Buratti 2017; Jyotsana and Heuser 2018; Sardone et al. 2017; Wasser and Herz 2016). Investigating the splicing mechanisms involved in ALS could pave the way to new interesting perspectives for developing novel therapeutic approaches and raising hope for this devastating and still incurable pathology.
The Alternative Splicing Program: Molecular Mechanism and Regulation
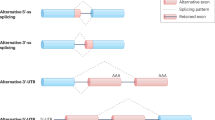
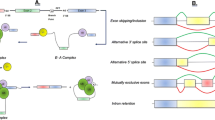
AS is the main mechanism of gene expression regulation that increases transcriptomic and proteomic diversity in eukaryotic cells. It works through five main different events: exon skipping (ES), mutually exclusive exon, alternative 3′ splice site, alternative 5′ splice site, and intron retention (IR) (Tazi et al. 2009).
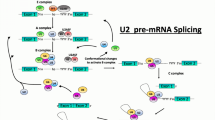
Splicing process is performed by the spliceosome, a dynamic machine able to identify the exon–intron splice sites and to catalyze the cut-and-paste reactions for the intron removal and the exon junction (Matera and Wang 2014). This machinery is composed of a large number of elements, including small nuclear RNA molecules (snRNAs), some of which play a structural role (i.e., the small nuclear ribonucleoproteins, snRNPs), are regulators of the reaction (i.e., SR proteins, rich in Ser and Arg), or work as ATPasi or RNA-helicases (Matera and Wang 2014; Valadkhan 2010).
The splicing process is regulated by auxiliary cis-acting elements, named exonic or intronic splicing enhancers (ESEs or ISEs) and silencers (ESSs or ISSs), which guarantee the correct exon/intron recognition through their binding sites for the SR proteins and operate together with their specific trans-acting RBPs (Fig. 1) (Kapeli et al. 2017; Ram and Ast 2007). Other players of splicing regulation are the hnRNP proteins (heterogeneous nuclear ribonucleoproteins) that promote skipping by binding silencers located in the proximity of the exon–intron junctions (Geuens et al. 2016) (Fig. 1). An example of hnRNPs is the PTB (polypyrimidine tract-binding protein), whose spliced PTBP1 isoform (nPTB1) is able to suppress the neural splicing of specific targets, producing the stop of the neuronal differentiation process (Boutz et al. 2007).
RNA-binding proteins regulating splicing events. Serine and Arginine-rich proteins (SR proteins) are known to bind exonic or intronic splicing enhancer (ESEs/ISEs), while heterogeneous nuclear ribonucleoproteins (hnRNPs) bind intronic or exonic splicing silencers (ISSs/ESSs). The splicing process is enhanced by SR proteins and repressed by hnRNPs. In the blue circles we report a small list of RS proteins and hnRNP while in the red circle we list RBPs that have been associated with various diseases, including neurological diseases. For further details, please refer to the databases of RNA-binding protein specificities (RBPDB, http://rbpdb.ccbr.utoronto.ca/), which collects RNA-binding proteins associated to neurological diseases
Recent discoveries have shown that AS is powerfully conditioned by epigenomic and epi-transcriptomic regulation, based on histone modifications, chromatin structure, and transcription rate changes (Prasad et al. 1999; Zhu et al. 2018). The action of hnRNPs, SR proteins and the activity of kinase/phosphatase enzymes rely on post-translational modifications (phosphorylation and de-phosphorylation reactions) that are essential to promote splicing (Bedford and Richard 2005; Blackwell and Ceman 2012). The advancement in the understanding of splicing regulation by epigenomic/epi-transcriptomic processes will provide new molecular insights into the function of individual RNA modifications.
Alternative Splicing Regulation of ALS Genes
ALS is a neurodegenerative disease characterized by the progressive degeneration of both upper and lower motor neurons (MN) in spinal cord and motor cortex (Brown 1997), arising from the complex interaction of several molecular and cellular phenomena, including oxidative stress, mitochondrial dysfunction, axonal transport alteration, inflammation, excitotoxicity, and protein aggregation (Mendonca et al. 2012; Rothstein 2009).
Several genes are known to play a role in ALS pathogenesis (such as TBK1, SOD1, VCP, TARDBP, FUS, GRN, MAPT, CHCHD10, and TUBA4A) [for an updated review, the reader is referred to (Volk et al. 2018)], and a complete and updated list is accessible in the Online genetic database of amyotrophic lateral sclerosis (ALSoD, http://alsod.iop.kcl.ac.uk/). Among the numerous listed genes, TARDBP, ELP3, ANG, TAF15, and FUS deserve particular attention because they are directly involved in RNA processing (transport control, stability, and translocation) (Baumer et al. 2010; Greenway et al. 2006; Kabashi et al. 2008; Mackenzie and Neumann 2012; Zhao et al. 2018).
A clear relationship between ALS-related genes and splicing factors is revealed by a bioinformatic analysis performed with STRING, a free web resource collecting known and predicted protein–protein interactions (https://string-db.org/) (Fig. 2). To build the network, we screened genes from both ALSoD database and SpliceAid-F database (http://srv00.recas.ba.infn.it/SpliceAidF/), which collect experimentally validated splicing factors (many ALS genes are splicing factors themselves) and related binding sites. Among RNAs or proteins involved in the splicing machinery or regulation, we included those associated with neurodegenerative, neuromuscular, and neurological diseases (i.e., spinal muscular atrophy, frontotemporal dementia, muscular dystrophy, neurofibromatosis type 1, myotonic dystrophy, fragile X syndrome, congenital myasthenic syndrome, and paraneoplastic encephalomyelitis). Figure 2 shows a number of existing potential connections and protein–protein interactions between the elements, strengthening the hypothesis of the AS contribution in ALS pathogenesis.
Representation of known and predicted molecular interactions between ALS genes and splicing regulators. A network created with STRING (https://string-db.org/) shows ALS disease-causing or related genes with known or predicted interactions with splicing factors, as reported in the legend. Red circles represent ALS genes, while blue circles represent splicing factors. For the STRING analysis we used a confidence interaction score of 0.400, focusing on specific and meaningful associations
In the following sections, ALS genes (listed in Table 1) will be divided in two classes: we will first focus on the splicing regulation of those genes that have their own specific pathology and contribute to a loss-of-function mechanism in ALS (TARDBP, FUS, and C9ORF72), and then we will discuss other susceptibility or ALS-related genes (Conlon and Manley 2017).
TARDBP
TAR DNA-binding protein 43 (TDP-43), encoded by TARDBP localized on chromosome 1, is one of the components of ubiquitinated protein aggregates found in familial and sporadic ALS patients with different degree of severity (Tsuji et al. 2012). It is an essential splicing factor, since it is a member of the hnRNPs family and thus is involved in the RNA metabolism processes (splicing, transport, and translation) (Buratti and Baralle 2008). TDP-43 is composed of several functional domains: an N-terminal domain (NTD), two-tandem RNA recognition (RRM1-2), and a glycine-rich term prion-like domain located at the C-term (PrLD). This latter is particularly important for AS regulation, since it is involved in mediating the phase transitions underpinning RNP granule assembly (Shorter and Taylor 2013), but on the other hand acts as a pathogenic mutation site rendering TDP-43 prone to misfolding into conformers that accumulate in pathological inclusions (Harrison and Shorter 2017). The absence of PrLD domain prevent aberrant misfolding and toxicity events (Ash et al. 2010; Johnson et al. 2009) and could be an interesting target for an oligonucleotide therapeutic strategy.
Several lines of evidence suggest the crucial role of TDP-43 in splicing regulation and ALS onset, encompassing both mutations in the genetic sequence of TARDBP itself and the AS regulation of specific targets (SMN2, APOAII, CFTR HNRNPA1, POLDIP3, ATG4B, STMN) (Butti and Patten 2018; Deshaies et al. 2018; Torres et al. 2018). About the genetic sequence, two heterozygous missense variants have been described in exon 6 of TARDBP that modulate AS and are likely involved in ALS onset or progression (Van Deerlin et al. 2008),. With regard to targets, TDP-43 depletion interfere with the AS of hnRNPA1 pre-mRNA, determining the inclusion of exon7B and culminating in a longer hnRNAP A1B isoform that is prone to aggregation and cytotoxic (Deshaies et al. 2018). Similarly, downregulation of TDP-43 results in an increase of cryptic sites in ATG4B (autophagy related 4B cysteine peptidase) causing an impairment of autophagy (Torres et al. 2018). Another identified TPD-43 downstream target is POLDIP3 (Polymerase delta-interacting protein 3), which is involved in regulating splicing efficiency (Shiga et al. 2012). POLDIP3 isoform 2 is rarely expressed in healthy tissues and its concentration increases when TDP-43 is depleted, making it a candidate biomarker for TDP-43 dysfunction (Shiga et al. 2012). Furthermore, TDP-43 regulates the splicing of STMN2 (Stathmin-2), necessary for normal growth and axonal regeneration. TDP-43 knockdown leads to a decreased STMN2 expression, probably through splicing in a cryptic exon (Klim et al. 2019).
Further evidence shows that TDP-43 dysfunctions alter the splicing efficiency by modifying U snRNP biogenesis and leading to neuronal death (Yahara et al. 2017). In addition, TDP-43 depletion leads to a co-depletion of U12 snRNA, altering snRNPs assembly and providing a potential role for the minor spliceosome machinery in ALS (Ishihara et al. 2013). Taken together, all these data suggest that TDP-43 depletion leads to the inclusion of different cryptic exons, and the maintenance of a homeostatic level of this protein is crucial for AS regulation.
FET-Proteins: FUS, EWSR and TAF15
FUS (Fused in Sarcoma), EWSR (Ewing sarcoma breakpoint region), and TAF15 (TATA-Box Binding Protein Associated Factor 15) belong to the family of FET-proteins (Mackenzie and Neumann 2012). These proteins are composed of an N-terminal domain rich in Gln, Gly, Ser, and Tyr, a conserved RNA-binding domain (RBD), the Arginine-Glycine-rich (RGG) domain that may influence RNA-binding, and a Cys2–Cys2 zinc finger motif binding nucleic acids (Morohoshi et al. 1998). Similarly to TDP-43, FUS and TAF15 aggregates characterize different neurodegenerative conditions including ALS (Lagier-Tourenne et al. 2010). FET-proteins bind single or double RNA/DNA strands and many experiments suggested their possible implication in transcription, RNA transport and pre-mRNA splicing (Lagier-Tourenne et al. 2010; Wu and Green 1997). Interestingly, FUS is enclosed in spliceosomal complex and interacts with several SR proteins and minor splicing factors, as well as with the U1 snRNP (Butti and Patten 2018; Kapeli et al. 2016; Reber et al. 2016; Shang and Huang 2016). FUS depletion is known to affect the splicing of genes involved in neurogenesis (PPP2R2C), dendritic development (ACTL6B), and action potential transmission in skeletal muscles (SCN8A and SCN4A) (Reber et al. 2016). Moreover, FUS is able to interfere with the AS of genes involved in axonal growth and cytoskeletal organization, including MAPT, NTNG1, NRCAM, and ABLIM1 genes (Butti and Patten 2018).
Genomic FUS mutations determine an alternatively spliced exon 7, inducing a frameshift and the consequent splicing variants degradation (Zhou et al. 2013). Moreover, it is involved in some back-splicing events that regulate the formation of circular RNAs (circRNAs) in murine embryonic stem cell-derived motor neurons, causing a reduction in circRNA expression levels (Errichelli et al. 2017).
Differently from FUS, TAF15 plays a minimal role in AS altering the splicing of few known genes: GPCPD1 (glycerophosphocholine phosphodiesterase 1), KCNMA1 (the alpha-1 gene of the calcium-activated potassium channel subunit), and GRIN1 (N-methyl-d-aspartate receptor subunit NR1) (Kapeli et al. 2016).
Given the essential role that FUS plays in splicing regulation, further studies deserve to be carried out to better understand the consequence of the loss of function of FUS (and of other FET-proteins) on RNA splicing and its potential contribution to ALS pathogenesis.
C9ORF72
Hexanucleotide GGGGCC (G4C2) repeat expansion in C9ORF72 (C9) is the main cause of ALS and Frontotemporal Dementia (FTD). This G4C2 expansion results in a dipeptide repeat protein (DPR), that accumulates in the cerebellum, cortical region and hippocampus of ALS patients (Gijselinck et al. 2012; Liscic 2015).
The differential use of transcription alternative start and termination sites in C9 is known to produce at least three RNA variants, encoding a long protein isoform (called isoform A) of approximately 54 kDa and a short isoform (named isoform B) of about 24 kDa (Barker et al. 2017). Expansion carriers exhibit a reduction in both C9ORF72 mRNAs and protein levels, suggesting that toxicity may be mediated by a loss-of-function mechanism (Barker et al. 2017).
Mis-splicing of the expanded C9 transcript may play a role in C9-mediated toxicity; whereas C9 is able to sequester several members of the hnRNP family (such as hnRNP A1, hnRNP A3, hnRNP H) resulting in altered splicing patterns of their RNA targets (Conlon et al. 2016, 2018; Lee et al. 2013; Mori et al. 2013).
A number of studies described global splicing alterations in C9 expansion carriers. Transcriptome analysis in lymphoblastoid cells and motor neurons of C9-FTD/ALS cases revealed an increased occurrence of splicing errors (most evident among patients with faster disease progression) and an enrichment of upregulated transcripts involved in RNA splicing, thus suggesting that such increased error rate could be a consequence of RBPs sequestration into foci, which in turn would contribute to disease progression and severity (Cooper-Knock et al. 2015). Brain transcriptome profiling analysis showed extensive AS and alternative polyadenylation defects in the cerebellum of C9ALS subjects, involving also ALS-associated genes (e.g., ATXN2 and FUS) (Prudencio et al. 2015). Experiments conducted on lymphoblastoid cell lines revealed an increased number of aberrant splicing events, especially in the ALS patients with a rapid disease progression (Prudencio et al. 2015).
Interestingly, the G-quadruplex structure of C9 is able to sequester the splicing factor SRSF2 (Serine and arginine SF factor 2) (Conlon et al. 2016; Zhang et al. 2015), which in turn helps the binding of U1 snRNP to the 5′ splice site or other factors to the 3′ splice site of target genes (Mure et al. 2018). Very recently, DPRs were found to block spliceosome assembly associating and interfering with U2 snRNP, causing a global splicing alteration in ALS patients (Yin et al. 2017).
Alternative Splicing Regulation of ALS-Related Genes
A substantial number of other ALS-related genes are susceptible to splicing-based regulation. In the next sections, we will focus on SETX, OPTN, NEK1, SPG11, VAPB, DCTN1, CHCHD10, KIF5A, EEAT2, and ADAR2.
SETX
Senataxin (SETX) encodes for an RNA-binding protein with a highly conserved helicase domain involved in regulation of RNA transcription (Bennett et al. 2018) and associated with both ALS4 (a form of juvenile ALS) and AOA2 (ataxia with oculomotor apraxia type 2). Sequencing of SETX coding regions and genetic engineering experiments revealed mutations correlated to an exonic cryptic donor site activation and an ESSs creation, resulting in MNs degeneration (Bennett et al. 2018; Tripolszki et al. 2017).
OPTN
Optineurin (OPTN) is localized on chromosome 10p13 and encodes for a protein involved in membrane and vesicle trafficking, transcription activation and cellular morphogenesis. This gene is mainly known to be responsible for hereditary primary open-angle glaucoma (POAG) and ALS, but its mutations were recently described to cause mRNA downregulation and degradation (Maruyama et al. 2010; Toth and Atkin 2018). In particular, an intronic mutation is known to activate a cryptic exonic donor site, resulting in ES of exon 6 (Del Bo et al. 2011); a non-sense mutation causes a stop codon and generates a frameshift due to exon 5 deletion (Maruyama et al. 2010); additional OPTN variants were reported to create either an ESEs or an ESSs (Johnson et al. 2012).
NEK Family
NIMA-related kinases (NEK) protein family are players in several fundamental biological processes, including cell cycle regulation (Moniz et al. 2011). These proteins share a common kinase domain, a basic domain, one or more coiled-coil motifs (CC), and two nuclear export sequences (NES). Eleven mammalian NIMA-related kinases are currently known, and among these, NEK10 is the only that does not contain the N-terminal catalytic domain, while NEK4, 6, and 7 do not have the coiled–coiled motifs. A growing number of NEKs are implicated in DNA damage response, some of which have tissue-specific functions, such as NEK3 and NEK7 in neurons (He et al. 2016; Shi et al. 2016).
A recent whole exome sequencing study suggested an association between NEK1 mutations and ALS, emphasizing a loss-of-function mechanism in 0.8% of ALS patients and highlighting a link between NEK1 and other known ALS genes (SOD1, TBK1, C21orf2) (Brenner et al. 2016; Cirulli et al. 2015; Kenna et al. 2016). To the best of our knowledge, there are currently no evidence correlating NEK1 splicing alterations to ALS, although it is known that the other NIMA-related proteins undergo splicing regulation. Specifically, NEK4 isoform is engaged in mRNA processing mediated by the spliceosome and is localized in nuclear speckles and substrates containing snRNPs (Basei et al. 2015). NEK2 phosphorylates SRSF1 and, therefore, modulates SRSF1’s target genes controlling important AS events (Naro et al. 2014). Moreover, SF phosphorylation and NEK2 silencing negatively affect the splicing (Naro et al. 2014). Given the importance of splicing regulation in components of the NEK family, it is possible that splicing in NEK1 is involved in ALS pathology.
SPG11
SPG11 (Spastic Paraplegia 11), localized on chromosome 15q13-15, encodes for spatacsin protein, which plays a pivotal role in axonal maintenance, synaptic vesicle transport, and autophagy. Spatacsin is essential for neuronal survival and is ubiquitously expressed in the nervous system, prominently in the cerebellum, cerebral cortex and hippocampus. SPG11 mutations are considered causative for both hereditary spastic paraplegia (HSP) and the autosomal recessive juvenile ALS (ARJALS) form (Orlacchio et al. 2010). ARJALS is a rare disease that occurs before the age of 25 years with a slowly progressive course. Interestingly, HSP and ARJALS have many similarities in clinical presentation, molecular genetics, and cellular pathology. This overlap suggests that the same genetic variants may contribute to a common pathogenic pathway. Mutations in SPG11 affect different splice donor region variants (Pippucci et al. 2009; Yu et al. 2016).
VAPB
Vesicle-associated membrane protein (VAMP)—associated protein B (also known as VAPB) plays a role in cellular stress response of the endoplasmatic reticulum (ER) and in unfolded protein response (UPR) (Suzuki et al. 2009; Walker and Atkin 2011). VAPB is ubiquitously expressed in eukaryotic cells and is involved in cellular calcium homeostasis regulation, protein transport, phospholipidic metabolism and viral infections (Kanekura et al. 2009). It is composed of three structural domains: a MSP (major sperm proteins) conserved domain, a central amphipathic helicoidal structure and a C-terminal transmembrane domain. Five VAPB splice variants are known, all expressed in the human nervous system, which accumulate after proteasomal inhibition and contribute to ALS onset (Nachreiner et al. 2010). The identified variants lack specific exons of wt-VAPB: exon 2 (VAPB-2 isoform), exons 4 and 5 (VAPB-4,5 isoform), exon 3 (VAPB-3 isoform), exons 3 and 4 (VAPB-3,4 isoform) (Nachreiner et al. 2010).
DCTN1
DCTN1 (Dynactin subunit 1) is located on chromosome 2p13 and encodes the dynactin protein (a component of the ubiquitous dynactin complex) that binds both microtubules and cytoplasmic dynein. The complex is involved in vesicle retrograde transport processes (endosomes and lysosomes), axonogenesis, ER-Golgi transports, and chromosomes shift. The AS of this gene results in a brain-specific and an ubiquitously expressed isoform (Lazarus et al. 2013). AS of DCTN1 determine both distal hereditary motor neuropathy type VIIB (HMN7B), also known as distal spinal and bulbar muscular atrophy, and neurodegenerative disorders such as ALS (LaMonte et al. 2002). Different studies reported DCTN1 missense mutations and splicing changes in familial and sporadic ALS patients, mainly in Caucasians (Couthouis et al. 2014; Munch et al. 2004, 2005). Recently, a reduction of DCTN1 mRNA was reported in sALS motor cortex and spinal cord samples (Kuzma-Kozakiewicz et al. 2013).
CHCHD10
CHCHD10 (Coiled-Coil-Helix-Coiled-Coil-Helix Domain Containing 10) encodes a mitochondrial protein, localized in the intermembrane space and involved in mitochondrial organization maintenance. Mutations of this gene cause ALS2 and FTD. A recent study on fALS patients revealed a heterozygous missense variation that actives an acceptor cryptic site (Teyssou et al. 2016). Moreover, two mutations that cause about 50% reduction in CHCHD10 protein levels and contribute to motoneuronal disease were identified (Brockmann et al. 2018).
KIF5A
KIF5A (Kinesin Family Member 5A) encodes a protein of the kinesin family, principally expressed in neurons and involved in axonal and organelle transport. Heterozygous missense mutations in KIF5A cause monogenic spastic paraplegia (HSP10) and Charcot-Marie-Tooth type 2 (CMT2), while frameshift mutations are known to cause neurodevelopmental syndromes (Nicolas et al. 2018). Recently, two studies described the association between the increase of splice site mutations in the KIF5A C-term region and fALS (Brenner et al. 2018; Nicolas et al. 2018), definitively demonstrating that KIF5A mutations interfere with axonal transport and contribute to motor neuron degeneration.
ANXA11
Annexin A11 (ANXA11) is located on chromosome 10q22.3 and encodes for a member of the annexin family, a group of cellular proteins working in calcium-dependent modality (Tawani and Kumar 2015). It is involved in vesicle trafficking, exocytosis, endocytosis, signal transduction, cytokinesis, and apoptosis. ANXA11 is susceptible to phosphorylation in the amino-terminal region (Furge et al. 1999) and a variety of interactions influence its subcellular localization. Annexin A11 has been associated with cancer, autoimmune disorders (such as systemic lupus erythematosus), and multisystem autoimmune disease (sarcoidosis) (Hofmann et al. 2008).
ANXA11 has been recently implicated in ALS (Smith et al. 2017). In particular, the p.D40G mutation was associated to annexin A11-positive protein aggregates in spinal cord motor neurons and hippocampal neuronal axons. ANXA11 is one of the genes most frequently mutated in sALS Chinese patients, where a splice site mutation in exon 6 causes a partial deletions in amino-acidic protein sequence (Zhang et al. 2018).
EAAT2
Oxidative stress is one of the most important mechanisms responsible for motor neuron degeneration (Rothstein 2009). EAAT2 (Excitatory aminoacid transporter 2) is involved in 90% of glutamate reuptake and undergoes RNA defects: exon 9 skipping and intron 7 retention. Splicing defects observed in EAAT2 mRNA in motor cortex and spinal cords of sALS patients were associated to alterations in spliceosomal components (Grabowski 1998; Lin et al. 1998). More recently, reduced EEAT2 activity was linked to aberrant EAAT2 mRNA transcripts as a consequence of abnormal RNA splicing (Bristol and Rothstein 1996).
ADAR2
The possibility that RNA processing defects play a role in neurodegenerative diseases is also confirmed by the regulation of ADAR (adenosine deaminase acting on RNA), an RNA-regulating protein responsible for binding to double-stranded RNA (dsRNA) and converting (editing) adenosine (A) to inosine (I) (Licht et al. 2016).
Defective ADARs were associated to different human pathologies: cancers, metabolic and neurological diseases (Slotkin and Nishikura 2013). Both ADAR enzymes and splicing elements act on the same substrates, dsRNA, determining the interaction of these two processes. Specifically, editing affects post-transcriptional regulation and introduces or removes splice sites. Several studies have shown that editing machinery affects splicing directly by altering the secondary RNA structures or cis-regulatory sequences, or indirectly by binding competitive regions of the dsRNA and, therefore, preventing the access to the splice machinery (Hsiao et al. 2018; Licht et al. 2016). ADAR isoform 2, in particular, catalyzes the adenosine-inosine conversion at the Q/R site of GluA2 pre-mRNA, the most common RNA editing in higher eukaryotes (Hideyama et al. 2012). In ALS patients, downregulation of ADAR2 and reduction in RNA editing were correlated to FUS-positive cytoplasmic inclusions (Aizawa et al. 2016), suggesting that ADAR2 downregulation could affect motor neuron physiology in ALS.
Genome-Wide Assessment of AS Events in ALS: New Technologies and Future Perspectives
Genomic technologies (splice-sensitive microarray or RNA sequencing) have revolutionized the way transcriptome research is conducted, enabling analysis of the entire span of transcripts in a biological sample (Colombrita et al. 2015; Hu et al. 2013; Ishigaki et al. 2012; La Cognata et al. 2016; Morello et al. 2017; Shiga et al. 2012).
Two main RNAseq applications are currently raising particular interest for dissecting the complexity of splicing regulation: (i) the study of AS events on a large scale, spanning from the classic evaluation of differential expression between samples until the characterization of gene expression dynamics, gene boundaries, translation efficiency or RNA–protein interaction, and (ii) the single-cell-level isoform studies (Arzalluz-Luque and Conesa 2018; Zucca et al. 2019).
Bulk RNAseq studies are the most widespread and have been applied to investigate the dynamic variations of transcriptomes in in vitro differentiated motor neurons obtained from human control and patient-specific VCP mutant-derived iPSCs (Luisier et al. 2018). Surprisingly, these time-resolved RNAseq experiments revealed increased IR events as a dominant feature during the early differentiation phases, and identified SFPQ factor (splicing factor proline and glutamine rich) as the major intron-retaining transcript across diverse ALS-causing mutations (VCP, SOD1 and FUS), proposing SFPQ IR events as a hallmark biomarker of familial and sporadic ALS (Luisier et al. 2018).
Despite genome-wide RNAseq studies are nowadays feasible, they rely on the experimental average gene expression across populations of cells, excluding the possibility to capture cell-to-cell variability and thus motivating the development of single-cell strategy (Arzalluz-Luque and Conesa 2018). Indeed, single-cell level insights are required to fully understand the biology of AS and represent the new challenge in RNAseq applications (Hwang et al. 2018). Each splice isoforms can be differently expressed depending on the particular cell type, showing a dominant (i.e., very highly expressed) isoform and several others with significantly lower expression values. This raise the question about whether the diverse and complex isoform expression landscape constitutes an additional layer of gene expression regulation contributing to ALS etiology, or if it is solely a result of the stochastic functioning of the AS machinery (Hwang et al. 2018).
Splicing Modulation Therapy
Deregulated AS is emerging as an important area for therapeutic intervention. Gene therapy, in particular, represents a promising pharmacological option for patients with diseases of genetic origins and is mainly based on antisense oligonucleotides (ASOs), spliceosome-mediated RNA trans-splicing (SMaRT) or small interfering RNAs (siRNAs) approaches (Arechavala-Gomeza et al. 2014).
Antisense oligonucleotides (ASOs), which are synthetic single-stranded nucleic acids, are able to bind the pre-mRNA intron/exon junctions and modulate splicing acting on enhancers or repressor sequences, determining exon skipping or including alternatively spliced exons (Havens and Hastings 2016; McClorey and Wood 2015). Multiple exon skipping strategies with ASOs have been already used in a variety of neurodegenerative and neuromuscular disorders, including Duchenne muscular dystrophy (DMD), spinal muscular atrophy (SMA), and ALS (Aartsma-Rus et al. 2017; Benoit-Pilven et al. 2018; McCampbell et al. 2018; Niks and Aartsma-Rus 2017; Ren et al. 2017; Sardone et al. 2017; Tosolini and Sleigh 2017). ASOs can be chemically modified, generating bifunctional ASOs with added RNA or peptide motifs, and specifically target mutant RNAs or AS protein recruitment to the transcript (Singh et al. 2018).
ASO’s therapy has already entered into the medical field of neurological disorder. Eteplirsen, a third-generation ASO that hybridizes to exon 51 of DMD pre-mRNA, represents the first FDA-approved ASO for the treatment of Duchenne Muscolar Distrophy. Nusinersen is another FDA-approved ASO-based drug (marketed as Spinraza) used for SMA therapy, and works by promoting the exon skipping of exon 7 in SMN2 gene by blocking intronic splicing silencer N1 (ISS-N1) located immediately downstream of exon 7 (Verma 2018).
With regard to ALS, one of the first ASO-based clinical trials was designed to silence SOD1. Intrathecal administration of the ASO IONIS-SOD1Rx resulted both practical and safe in SOD1 ALS patients during phase I testing (NCT01041222) (Miller et al. 2013). A phase Ib/IIa trial (NCT02623699) is currently in process to further evaluate safety, tolerability, and pharmacokinetics of IONIS-SOD1Rx (McCampbell et al. 2018).
Among the ALS-related genes that have been discussed in the present review, C9ORF72 may represent the best candidate for ASOs therapy. As previously described, C9 accumulates in nuclear foci, conferring toxicity associated with the repeat expansion and, therefore, compromising the pre-mRNA processing (Wojciechowska and Krzyzosiak 2011). Given the encouraging results obtained in splicing modulation therapy in other polyglutamic diseases (CAG repeats), such as spinocerebellar ataxia (SCA 2) and Huntington Disease (HD) (Rindt et al. 2017; Tawani and Kumar 2015), it is not surprising to imagine the use of ASOs and/or other exon skipping approaches to restore the reading code or remove exons containing mutations in ALS. Early testing of ASO-based therapeutics for C9ALS was performed on iPSC-derived neurons and fibroblasts (Butti and Patten 2018; Donnelly et al. 2013; Lagier-Tourenne et al. 2013). ASOs were designed to target the repeat expansion or within surrounding N-terminal regions of the C9 mRNA transcript to degrade the transcript or to block the interaction between the repeat expansion and RNA-binding proteins, resulting in a reduction of RNA foci, dipeptide proteins and restored normal gene expression markers (Butti and Patten 2018; Donnelly et al. 2013; Lagier-Tourenne et al. 2013). Recently, ASOs have been also tested in mouse models expressing the expanded C9 (Butti and Patten 2018). Last year, a randomized, double-blind, placebo controlled phase I clinical trial with an antisense oligonucleotide (BIIB078) targeting C9 has been started. The study will involve 80 patients carrying the pathological expansion (ClinicalTrials.gov Identifier: NCT03626012).
Other studies have analyzed the effects of ASO modifications within the oligonucleotide backbone, sugar and heterocycles in order to improve delivery, potency, and stability to target FUS, demonstrating that affinities of various nucleic acid binding domains depend on chemical modifications and that ASO–protein interactions influence the localization of ASOs themselves (Bailey et al. 2017). These preliminary data suggest that ASO-based therapy can be a powerful way for treating ALS-relate genes although it is clear that therapeutic outcomes will depend on the stage of disease progression and on the time of intervention.
The SMaRT method has been used for cystic fibrosis and HD (Rindt et al. 2017). SMaRT is based on the correction of alterations at post-transcriptional level through the introduction of an exogenous RNA into targeted cells to induce a trans-splicing event between exogenous RNA and target endogenous pre-mRNA (Berger et al. 2016). There are currently no applications of this approach for ALS.
Finally, SiRNAs, 21–25 nucleotide double-stranded RNA molecules, are becoming an important therapeutic tool for different diseases including tumors or metabolic/genetic disorders due to genetic malfunction and deregulated expression. The siRNA selectively targets and silences the gene by inhibiting the expression of the protein (Borna et al. 2015; Ozcan et al. 2015). Several siRNAs were designed in silico to target the glycine-rich region of TARDBP via molecular dynamics and thermo-physical analyses (Bhandare and Ramaswamy 2016). Since siRNAs strategy induces a catalytic process resulting in complete gene knockdown, likely detrimental for the cell, the exon skipping approach may be preferred since it corrects the gene-reading frame.
Conclusion
Many proteins implicated in motor neuron and neurological disorders have a physiological function in different RNA processes (mRNA stability, transcription regulation, transport of RNA granules, splicing, miRNA biogenesis, and RNA editing). A large number of diseases related to splicing were documented, but this is probably under-estimated because the splicing mutation effects are often not considered as a primary cause of the diseases. An increasing understanding of splicing regulation will create new treatment strategies for modulating this process in disease contexts, leading to personalized medicine. The overlap between clinical data, etio-pathogenetic mechanisms, and gene therapy strategies may offer novel solutions by creating rigorous guidelines in clinical trials (Ludolph et al. 2010). This kind of holistic approach seems to be now the most promising to advance the therapy of complex and multifactorial diseases.
Abbreviations
- ALS:
-
Amyotrophic lateral sclerosis
- AS:
-
Alternative splicing
- RBPs:
-
RNA-binding proteins
- fALS:
-
Familial amyotrophic lateral sclerosis
- sALS:
-
Sporadic amyotrophic lateral sclerosis
- FTD:
-
Frontotemporal dementia
- ES:
-
Exon skipping
- IR:
-
Intron retention
- C9:
-
C9ORF72
- snRNA:
-
Small nuclear RNA
- snRNPs:
-
Small nuclear ribonucleoproteins
- PPIases:
-
Peptidyl-propyl cis/trans isomerases
- ESEs:
-
Exonic splicing enhancers
- ISEs:
-
Intronic splicing enhancers
- ESSs:
-
Exonic splicing silencers
- ISSs:
-
Intronic splicing silencers
- PrLD:
-
Prion-like domain
- hnRNP:
-
Heterogenous nuclear ribonucleoprotein
- PTB:
-
Polypyrimidine tract-binding protein
- MN:
-
Motoneuron
- SMaRT:
-
Transcription of spliceosome-mediated RNA
- siRNA:
-
Small interfering RNA
References
Aartsma-Rus A et al (2017) Development of exon skipping therapies for duchenne muscular dystrophy: a critical review and a perspective on the outstanding issues. Nucleic Acid Ther 27:251–259. https://doi.org/10.1089/nat.2017.0682
Aizawa H, Hideyama T, Yamashita T, Kimura T, Suzuki N, Aoki M, Kwak S (2016) Deficient RNA-editing enzyme ADAR2 in an amyotrophic lateral sclerosis patient with a FUS(P525L) mutation. J Clin Neurosci 32:128–129. https://doi.org/10.1016/j.jocn.2015.12.039
Arechavala-Gomeza V, Khoo B, Aartsma-Rus A (2014) Splicing modulation therapy in the treatment of genetic diseases. Appl Clin Genet 7:245–252. https://doi.org/10.2147/TACG.S71506
Arzalluz-Luque A, Conesa A (2018) Single-cell RNAseq for the study of isoforms-how is that possible? Genome Biol 19:110. https://doi.org/10.1186/s13059-018-1496-z
Ash PE et al (2010) Neurotoxic effects of TDP-43 overexpression in C. elegans. Hum Mol Genet 19:3206–3218. https://doi.org/10.1093/hmg/ddq230
Bailey JK, Shen W, Liang XH, Crooke ST (2017) Nucleic acid binding proteins affect the subcellular distribution of phosphorothioate antisense oligonucleotides. Nucleic Acids Res 45:10649–10671. https://doi.org/10.1093/nar/gkx709
Baralle D, Buratti E (2017) RNA splicing in human disease and in the clinic. Clin Sci 131:355–368. https://doi.org/10.1042/CS20160211
Barker HV, Niblock M, Lee YB, Shaw CE, Gallo JM (2017) RNA misprocessing in C9orf72-linked neurodegeneration. Front Cell Neurosci 11:195. https://doi.org/10.3389/fncel.2017.00195
Basei FL, Meirelles GV, Righetto GL, Dos Santos Migueleti DL, Smetana JH, Kobarg J (2015) New interaction partners for Nek4.1 and Nek4.2 isoforms: from the DNA damage response to RNA splicing. Proteome Sci 13:11. https://doi.org/10.1186/s12953-015-0065-6
Baumer D, Hilton D, Paine SM, Turner MR, Lowe J, Talbot K, Ansorge O (2010) Juvenile ALS with basophilic inclusions is a FUS proteinopathy with FUS mutations. Neurology 75:611–618. https://doi.org/10.1212/WNL.0b013e3181ed9cde
Bedford MT, Richard S (2005) Arginine methylation an emerging regulator of protein function. Mol Cell 18:263–272. https://doi.org/10.1016/j.molcel.2005.04.003
Bennett CL et al (2018) Senataxin mutations elicit motor neuron degeneration phenotypes and yield TDP-43 mislocalization in ALS4 mice and human patients. Acta Neuropathol 136:425–443. https://doi.org/10.1007/s00401-018-1852-9
Benoit-Pilven C et al (2018) Complementarity of assembly-first and mapping-first approaches for alternative splicing annotation and differential analysis from RNAseq data. Sci Rep 8:4307. https://doi.org/10.1038/s41598-018-21770-7
Berger A, Maire S, Gaillard MC, Sahel JA, Hantraye P, Bemelmans AP (2016) mRNA trans-splicing in gene therapy for genetic diseases. Wiley Interdiscip Rev RNA 7:487–498. https://doi.org/10.1002/wrna.1347
Bhandare VV, Ramaswamy A (2016) Identification of possible siRNA molecules for TDP43 mutants causing amyotrophic lateral sclerosis: in silico design and molecular dynamics study. Comput Biol Chem 61:97–108. https://doi.org/10.1016/j.compbiolchem.2016.01.001
Blackwell E, Ceman S (2012) Arginine methylation of RNA-binding proteins regulates cell function and differentiation. Mol Reprod Dev 79:163–175. https://doi.org/10.1002/mrd.22024
Borna H, Imani S, Iman M, Azimzadeh Jamalkandi S (2015) Therapeutic face of RNAi: in vivo challenges. Exp Opin Biol Ther 15:269–285. https://doi.org/10.1517/14712598.2015.983070
Boutz PL et al (2007) A post-transcriptional regulatory switch in polypyrimidine tract-binding proteins reprograms alternative splicing in developing neurons. Genes Dev 21:1636–1652. https://doi.org/10.1101/gad.1558107
Brenner D et al (2016) NEK1 mutations in familial amyotrophic lateral sclerosis. Brain 139:e28. https://doi.org/10.1093/brain/aww033
Brenner D et al (2018) Hot-spot KIF5A mutations cause familial ALS. Brain 141:688–697. https://doi.org/10.1093/brain/awx370
Bristol LA, Rothstein JD (1996) Glutamate transporter gene expression in amyotrophic lateral sclerosis motor cortex. Ann Neurol 39:676–679. https://doi.org/10.1002/ana.410390519
Brockmann SJ et al (2018) CHCHD10 mutations p. R15L and p.G66V cause motoneuron disease by haploinsufficiency. Hum Mol Genet 27:706–715. https://doi.org/10.1093/hmg/ddx436
Brown RH Jr (1997) Amyotrophic lateral sclerosis. Insights from genetics. Arch Neurol 54:1246–1250
Buratti E, Baralle FE (2008) Multiple roles of TDP-43 in gene expression, splicing regulation, and human disease. Front Biosci 13:867–878
Butti Z, Patten SA (2018) RNA dysregulation in amyotrophic lateral sclerosis. Front Genet 9:712. https://doi.org/10.3389/fgene.2018.00712
Cirulli ET et al (2015) Exome sequencing in amyotrophic lateral sclerosis identifies risk genes and pathways. Science 347:1436–1441. https://doi.org/10.1126/science.aaa3650
Colombrita C et al (2015) From transcriptomic to protein level changes in TDP-43 and FUS loss-of-function cell models. Biochem Biophys Acta 1849:1398–1410. https://doi.org/10.1016/j.bbagrm.2015.10.015
Conlon EG, Manley JL (2017) RNA-binding proteins in neurodegeneration: mechanisms in aggregate. Genes Dev 31:1509–1528. https://doi.org/10.1101/gad.304055.117
Conlon EG, Lu L, Sharma A, Yamazaki T, Tang T, Shneider NA, Manley JL (2016) The C9ORF72 GGGGCC expansion forms RNA G-quadruplex inclusions and sequesters hnRNP H to disrupt splicing in ALS brains. eLife. https://doi.org/10.7554/elife.17820
Conlon EG et al (2018) Unexpected similarities between C9ORF72 and sporadic forms of ALS/FTD suggest a common disease mechanism. eLife. https://doi.org/10.7554/elife.37754
Cooper-Knock J et al (2015) C9ORF72 GGGGCC expanded repeats produce splicing dysregulation which correlates with disease severity in amyotrophic lateral sclerosis. PLoS ONE 10:e0127376. https://doi.org/10.1371/journal.pone.0127376
Couthouis J, Raphael AR, Daneshjou R, Gitler AD (2014) Targeted exon capture and sequencing in sporadic amyotrophic lateral sclerosis. PLoS Genet 10:e1004704. https://doi.org/10.1371/journal.pgen.1004704
Del Bo R et al (2011) Novel optineurin mutations in patients with familial and sporadic amyotrophic lateral sclerosis. J Neurol Neurosurg Psychiatry 82:1239–1243. https://doi.org/10.1136/jnnp.2011.242313
Deshaies JE et al (2018) TDP-43 regulates the alternative splicing of hnRNP A1 to yield an aggregation-prone variant in amyotrophic lateral sclerosis. Brain 141:1320–1333. https://doi.org/10.1093/brain/awy062
Donnelly CJ et al (2013) RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention. Neuron 80:415–428. https://doi.org/10.1016/j.neuron.2013.10.015
Errichelli L et al (2017) FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons. Nat Commun 8:14741. https://doi.org/10.1038/ncomms14741
Furge LL, Chen K, Cohen S (1999) Annexin VII and annexin XI are tyrosine phosphorylated in peroxovanadate-treated dogs and in platelet-derived growth factor-treated rat vascular smooth muscle cells. J Biol Chem 274:33504–33509
Geuens T, Bouhy D, Timmerman V (2016) The hnRNP family: insights into their role in health and disease. Hum Genet 135:851–867. https://doi.org/10.1007/s00439-016-1683-5
Gijselinck I et al (2012) A C9orf72 promoter repeat expansion in a Flanders-Belgian cohort with disorders of the frontotemporal lobar degeneration-amyotrophic lateral sclerosis spectrum: a gene identification study. Lancet Neurol 11:54–65. https://doi.org/10.1016/S1474-4422(11)70261-7
Grabowski PJ (1998) Splicing regulation in neurons: tinkering with cell-specific control. Cell 92:709–712
Greenway MJ et al (2006) ANG mutations segregate with familial and ‘sporadic’ amyotrophic lateral sclerosis. Nat Genet 38:411–413. https://doi.org/10.1038/ng1742
Harrison AF, Shorter J (2017) RNA-binding proteins with prion-like domains in health and disease. Biochem J 474:1417–1438. https://doi.org/10.1042/BCJ20160499
Havens MA, Hastings ML (2016) Splice-switching antisense oligonucleotides as therapeutic drugs. Nucleic Acids Res 44:6549–6563. https://doi.org/10.1093/nar/gkw533
He Y, Zeng MY, Yang D, Motro B, Nunez G (2016) NEK7 is an essential mediator of NLRP3 activation downstream of potassium efflux. Nature 530:354–357. https://doi.org/10.1038/nature16959
Hideyama T, Yamashita T, Aizawa H, Tsuji S, Kakita A, Takahashi H, Kwak S (2012) Profound downregulation of the RNA editing enzyme ADAR2 in ALS spinal motor neurons. Neurobiol Dis 45:1121–1128. https://doi.org/10.1016/j.nbd.2011.12.033
Hofmann S et al (2008) Genome-wide association study identifies ANXA11 as a new susceptibility locus for sarcoidosis. Nat Genet 40:1103–1106. https://doi.org/10.1038/ng.198
Hsiao YE et al (2018) RNA editing in nascent RNA affects pre-mRNA splicing. Genome Res 28:812–823. https://doi.org/10.1101/gr.231209.117
Hu M, Guo Y, Chen H, Duan W, Li C (2013) Exon array analysis of alternative splicing of genes in SOD1G93A transgenic mice. Appl Biochem Biotechnol 170:301–319. https://doi.org/10.1007/s12010-013-0155-9
Hwang B, Lee JH, Bang D (2018) Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp Mol Med 50:96. https://doi.org/10.1038/s12276-018-0071-8
Ishigaki S et al (2012) Position-dependent FUS-RNA interactions regulate alternative splicing events and transcriptions. Sci Rep 2:529. https://doi.org/10.1038/srep00529
Ishihara T et al (2013) Decreased number of Gemini of coiled bodies and U12 snRNA level in amyotrophic lateral sclerosis. Hum Mol Genet 22:4136–4147. https://doi.org/10.1093/hmg/ddt262
Johnson BS, Snead D, Lee JJ, McCaffery JM, Shorter J, Gitler AD (2009) TDP-43 is intrinsically aggregation-prone, and amyotrophic lateral sclerosis-linked mutations accelerate aggregation and increase toxicity. J Biol Chem 284:20329–20339. https://doi.org/10.1074/jbc.M109.010264
Johnson L et al (2012) Screening for OPTN mutations in a cohort of British amyotrophic lateral sclerosis patients. Neurobiol Aging 33(2948):e2915–e2947. https://doi.org/10.1016/j.neurobiolaging.2012.06.023
Jyotsana N, Heuser M (2018) Exploiting differential RNA splicing patterns: a potential new group of therapeutic targets in cancer. Exp Opin Ther Targets 22:107–121. https://doi.org/10.1080/14728222.2018.1417390
Kabashi E et al (2008) TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat Genet 40:572–574. https://doi.org/10.1038/ng.132
Kanekura K, Suzuki H, Aiso S, Matsuoka M (2009) ER stress and unfolded protein response in amyotrophic lateral sclerosis. Mol Neurobiol 39:81–89. https://doi.org/10.1007/s12035-009-8054-3
Kapeli K et al (2016) Distinct and shared functions of ALS-associated proteins TDP-43, FUS and TAF15 revealed by multisystem analyses. Nat Commun 7:12143. https://doi.org/10.1038/ncomms12143
Kapeli K, Martinez FJ, Yeo GW (2017) Genetic mutations in RNA-binding proteins and their roles in ALS. Hum Genet 136:1193–1214. https://doi.org/10.1007/s00439-017-1830-7
Kenna KP et al (2016) NEK1 variants confer susceptibility to amyotrophic lateral sclerosis. Nat Genet 48:1037–1042. https://doi.org/10.1038/ng.3626
Klim JR et al (2019) ALS-implicated protein TDP-43 sustains levels of STMN2, a mediator of motor neuron growth and repair. Nat Neurosci 22:167–179. https://doi.org/10.1038/s41593-018-0300-4
Kuzma-Kozakiewicz M, Chudy A, Kazmierczak B, Dziewulska D, Usarek E, Baranczyk-Kuzma A (2013) Dynactin deficiency in the CNS of humans with sporadic ALS and mice with genetically determined motor neuron degeneration. Neurochem Res. https://doi.org/10.1007/s11064-013-1160-7
La Cognata V, D’Agata V, Cavalcanti F, Cavallaro S (2015) Splicing: is there an alternative contribution to Parkinson’s disease? Neurogenetics. https://doi.org/10.1007/s10048-015-0449-x
La Cognata V, Morello G, Gentile G, D’Agata V, Criscuolo C, Cavalcanti F, Cavallaro S (2016) A customized high-resolution array-comparative genomic hybridization to explore copy number variations in Parkinson’s disease. Neurogenetics 17:233–244. https://doi.org/10.1007/s10048-016-0494-0
Lagier-Tourenne C, Polymenidou M, Cleveland DW (2010) TDP-43 and FUS/TLS: emerging roles in RNA processing and neurodegeneration. Hum Mol Genet 19:R46–R64. https://doi.org/10.1093/hmg/ddq137
Lagier-Tourenne C et al (2013) Targeted degradation of sense and antisense C9orf72 RNA foci as therapy for ALS and frontotemporal degeneration. Proc Natl Acad Sci USA 110:E4530–E4539. https://doi.org/10.1073/pnas.1318835110
LaMonte BH et al (2002) Disruption of dynein/dynactin inhibits axonal transport in motor neurons causing late-onset progressive degeneration. Neuron 34:715–727
Lazarus JE, Moughamian AJ, Tokito MK, Holzbaur EL (2013) Dynactin subunit p150(Glued) is a neuron-specific anti-catastrophe factor. PLoS Biol 11:e1001611. https://doi.org/10.1371/journal.pbio.1001611
Lee YB et al (2013) Hexanucleotide repeats in ALS/FTD form length-dependent RNA foci, sequester RNA binding proteins, and are neurotoxic. Cell Rep 5:1178–1186. https://doi.org/10.1016/j.celrep.2013.10.049
Licht K, Kapoor U, Mayrhofer E, Jantsch MF (2016) Adenosine to inosine editing frequency controlled by splicing efficiency. Nucleic Acids Res 44:6398–6408. https://doi.org/10.1093/nar/gkw325
Lin CL, Bristol LA, Jin L, Dykes-Hoberg M, Crawford T, Clawson L, Rothstein JD (1998) Aberrant RNA processing in a neurodegenerative disease: the cause for absent EAAT2, a glutamate transporter, in amyotrophic lateral sclerosis. Neuron 20:589–602
Liscic RM (2015) Molecular basis of ALS and FTD: implications for translational studies. Arh Hig Rada Toksikol 66:285–290. https://doi.org/10.1515/aiht-2015-66-2679
Ludolph AC et al (2010) Guidelines for preclinical animal research in ALS/MND: a consensus meeting. Amyotroph Lateral Scler 11:38–45. https://doi.org/10.3109/17482960903545334
Luisier R et al (2018) Intron retention and nuclear loss of SFPQ are molecular hallmarks of ALS. Nat Commun. https://doi.org/10.1038/s41467-018-04373-8
Mackenzie IR, Neumann M (2012) FET proteins in frontotemporal dementia and amyotrophic lateral sclerosis. Brain Res 1462:40–43. https://doi.org/10.1016/j.brainres.2011.12.010
Maruyama H et al (2010) Mutations of optineurin in amyotrophic lateral sclerosis. Nature 465:223–226. https://doi.org/10.1038/nature08971
Matera AG, Wang Z (2014) A day in the life of the spliceosome. Nat Rev Mol Cell Biol 15:108–121. https://doi.org/10.1038/nrm3742
McCampbell A et al (2018) Antisense oligonucleotides extend survival and reverse decrement in muscle response in ALS models. J Clin Investig 128:3558–3567. https://doi.org/10.1172/JCI99081
McClorey G, Wood MJ (2015) An overview of the clinical application of antisense oligonucleotides for RNA-targeting therapies. Curr Opin Pharmacol 24:52–58. https://doi.org/10.1016/j.coph.2015.07.005
Mendonca DM et al (2012) Neuroproteomics: an insight into ALS. Neurol Res 34:937–943. https://doi.org/10.1179/1743132812Y.0000000092
Miller TM et al (2013) An antisense oligonucleotide against SOD1 delivered intrathecally for patients with SOD1 familial amyotrophic lateral sclerosis: a phase 1, randomised, first-in-man study. Lancet Neurol 12:435–442. https://doi.org/10.1016/S1474-4422(13)70061-9
Moniz L, Dutt P, Haider N, Stambolic V (2011) Nek family of kinases in cell cycle, checkpoint control and cancer. Cell Div 6:18. https://doi.org/10.1186/1747-1028-6-18
Morello G, Guarnaccia M, Spampinato AG, La Cognata V, D’Agata V, Cavallaro S (2017) Copy number variations in amyotrophic lateral sclerosis: piecing the mosaic tiles together through a systems biology approach. Mol Neurobiol. https://doi.org/10.1007/s12035-017-0393-x
Mori K et al (2013) hnRNP A3 binds to GGGGCC repeats and is a constituent of p62-positive/TDP43-negative inclusions in the hippocampus of patients with C9orf72 mutations. Acta Neuropathol 125:413–423. https://doi.org/10.1007/s00401-013-1088-7
Morohoshi F, Ootsuka Y, Arai K, Ichikawa H, Mitani S, Munakata N, Ohki M (1998) Genomic structure of the human RBP56/hTAFII68 and FUS/TLS genes. Gene 221:191–198
Munch C et al (2004) Point mutations of the p150 subunit of dynactin (DCTN1) gene in ALS. Neurology 63:724–726
Munch C et al (2005) Heterozygous R1101K mutation of the DCTN1 gene in a family with ALS and FTD. Ann Neurol 58:777–780. https://doi.org/10.1002/ana.20631
Mure F, Corbin A, Benbahouche NEH, Bertrand E, Manet E, Gruffat H (2018) The splicing factor SRSF3 is functionally connected to the nuclear RNA exosome for intronless mRNA decay. Sci Rep 8:12901. https://doi.org/10.1038/s41598-018-31078-1
Nachreiner T, Esser M, Tenten V, Troost D, Weis J, Kruttgen A (2010) Novel splice variants of the amyotrophic lateral sclerosis-associated gene VAPB expressed in human tissues. Biochem Biophys Res Commun 394:703–708. https://doi.org/10.1016/j.bbrc.2010.03.055
Naro C, Barbagallo F, Chieffi P, Bourgeois CF, Paronetto MP, Sette C (2014) The centrosomal kinase NEK2 is a novel splicing factor kinase involved in cell survival. Nucleic Acids Res 42:3218–3227. https://doi.org/10.1093/nar/gkt1307
Nicolas A et al (2018) Genome-wide analyses identify KIF5A as a novel ALS. Gene Neuron 97(1268–1283):e1266. https://doi.org/10.1016/j.neuron.2018.02.027
Niks EH, Aartsma-Rus A (2017) Exon skipping: a first in class strategy for Duchenne muscular dystrophy. Exp Opin Biol Ther 17:225–236. https://doi.org/10.1080/14712598.2017.1271872
Nissim-Rafinia M, Kerem B (2005) The splicing machinery is a genetic modifier of disease severity. Trends Genet 21:480–483. https://doi.org/10.1016/j.tig.2005.07.005
Orlacchio A et al (2010) SPATACSIN mutations cause autosomal recessive juvenile amyotrophic lateral sclerosis. Brain 133:591–598. https://doi.org/10.1093/brain/awp325
Ozcan G, Ozpolat B, Coleman RL, Sood AK, Lopez-Berestein G (2015) Preclinical and clinical development of siRNA-based therapeutics. Adv Drug Deliv Rev 87:108–119. https://doi.org/10.1016/j.addr.2015.01.007
Pippucci T et al (2009) Autosomal recessive hereditary spastic paraplegia with thin corpus callosum: a novel mutation in the SPG11 gene and further evidence for genetic heterogeneity. Eur J Neurol 16:121–126. https://doi.org/10.1111/j.1468-1331.2008.02367.x
Prasad J, Colwill K, Pawson T, Manley JL (1999) The protein kinase Clk/Sty directly modulates SR protein activity: both hyper- and hypophosphorylation inhibit splicing. Mol Cell Biol 19:6991–7000
Prudencio M et al (2015) Distinct brain transcriptome profiles in C9orf72-associated and sporadic ALS. Nat Neurosci 18:1175–1182. https://doi.org/10.1038/nn.4065
Ram O, Ast G (2007) SR proteins: a foot on the exon before the transition from intron to exon definition. Trends Genet 23:5–7. https://doi.org/10.1016/j.tig.2006.10.002
Reber S et al (2016) Minor intron splicing is regulated by FUS and affected by ALS-associated FUS mutants. EMBO J 35:1504–1521. https://doi.org/10.15252/embj.201593791
Ren X, Deng R, Wang L, Zhang K, Li J (2017) RNA splicing process analysis for identifying antisense oligonucleotide inhibitors with padlock probe-based isothermal amplification. Chem Sci 8:5692–5698. https://doi.org/10.1039/c7sc01336a
Rindt H, Tom CM, Lorson CL, Mattis VB (2017) Optimization of trans-splicing for Huntington’s disease RNA Therapy. Front Neurosci 11:544. https://doi.org/10.3389/fnins.2017.00544
Rothstein JD (2009) Current hypotheses for the underlying biology of amyotrophic lateral sclerosis. Ann Neurol 65(Suppl 1):S3–S9. https://doi.org/10.1002/ana.21543
Sardone V, Zhou H, Muntoni F, Ferlini A, Falzarano MS (2017) Antisense oligonucleotide-based therapy for neuromuscular disease. Molecules. https://doi.org/10.3390/molecules22040563
Shang Y, Huang EJ (2016) Mechanisms of FUS mutations in familial amyotrophic lateral sclerosis. Brain Res 1647:65–78. https://doi.org/10.1016/j.brainres.2016.03.036
Shi H et al (2016) NLRP3 activation and mitosis are mutually exclusive events coordinated by NEK7, a new inflammasome component. Nat Immunol 17:250–258. https://doi.org/10.1038/ni.3333
Shiga A et al (2012) Alteration of POLDIP3 splicing associated with loss of function of TDP-43 in tissues affected with ALS. PLoS ONE 7:e43120. https://doi.org/10.1371/journal.pone.0043120
Shorter J, Taylor JP (2013) Disease mutations in the prion-like domains of hnRNPA1 and hnRNPA2/B1 introduce potent steric zippers that drive excess RNP granule assembly. Rare Dis 1:e25200. https://doi.org/10.4161/rdis.25200
Singh NN, Luo D, Singh RN (2018) Pre-mRNA splicing modulation by antisense oligonucleotides. Methods Mol Biol 1828:415–437. https://doi.org/10.1007/978-1-4939-8651-4_26
Slotkin W, Nishikura K (2013) Adenosine-to-inosine RNA editing and human disease. Genome Med 5:105. https://doi.org/10.1186/gm508
Smith BN et al (2017) Mutations in the vesicular trafficking protein annexin A11 are associated with amyotrophic lateral sclerosis. Sci Transl Med. https://doi.org/10.1126/scitranslmed.aad9157
Suzuki H, Kanekura K, Levine TP, Kohno K, Olkkonen VM, Aiso S, Matsuoka M (2009) ALS-linked P56S-VAPB, an aggregated loss-of-function mutant of VAPB, predisposes motor neurons to ER stress-related death by inducing aggregation of co-expressed wild-type VAPB. J Neurochem 108:973–985. https://doi.org/10.1111/j.0022-3042.2008.05857.x
Tawani A, Kumar A (2015) structural insights reveal the dynamics of the repeating r(CAG) transcript found in Huntington’s disease (HD) and spinocerebellar ataxias (SCAs). PLoS ONE 10:e0131788. https://doi.org/10.1371/journal.pone.0131788
Tazi J, Bakkour N, Stamm S (2009) Alternative splicing and disease. Biochem Biophys Acta 1792:14–26. https://doi.org/10.1016/j.bbadis.2008.09.017
Teyssou E et al (2016) Genetic analysis of CHCHD10 in French familial amyotrophic lateral sclerosis patients. Neurobiol Aging 42(218):e211–e213. https://doi.org/10.1016/j.neurobiolaging.2016.03.022
Torres P et al (2018) Cryptic exon splicing function of TARDBP interacts with autophagy in nervous tissue. Autophagy 14:1398–1403. https://doi.org/10.1080/15548627.2018.1474311
Tosolini AP, Sleigh JN (2017) Motor neuron gene therapy: lessons from spinal muscular atrophy for amyotrophic lateral sclerosis. Front Mol Neurosci 10:405. https://doi.org/10.3389/fnmol.2017.00405
Toth RP, Atkin JD (2018) Dysfunction of optineurin in amyotrophic lateral sclerosis and glaucoma. Front Immunol 9:1017. https://doi.org/10.3389/fimmu.2018.01017
Tripolszki K et al (2017) High-throughput sequencing revealed a novel SETX mutation in a Hungarian patient with amyotrophic lateral sclerosis. Brain Behav 7:e00669. https://doi.org/10.1002/brb3.669
Tsuji H et al (2012) Molecular analysis and biochemical classification of TDP-43 proteinopathy. Brain 135:3380–3391. https://doi.org/10.1093/brain/aws230
Valadkhan S (2010) Role of the snRNAs in spliceosomal active site. RNA Biol 7:345–353
Van Deerlin VM et al (2008) TARDBP mutations in amyotrophic lateral sclerosis with TDP-43 neuropathology: a genetic and histopathological analysis. Lancet Neurol 7:409–416. https://doi.org/10.1016/S1474-4422(08)70071-1
Vanderweyde T, Youmans K, Liu-Yesucevitz L, Wolozin B (2013) Role of stress granules and RNA-binding proteins in neurodegeneration: a mini-review. Gerontology 59:524–533. https://doi.org/10.1159/000354170
Verma A (2018) Recent advances in antisense oligonucleotide therapy in genetic neuromuscular diseases. Ann Indian Acad Neurol 21:3–8. https://doi.org/10.4103/aian.AIAN_298_17
Verma B, Akinyi MV, Norppa AJ, Frilander MJ (2018) Minor spliceosome and disease. Semin Cell Dev Biol 79:103–112. https://doi.org/10.1016/j.semcdb.2017.09.036
Volk AE, Weishaupt JH, Andersen PM, Ludolph AC, Kubisch C (2018) Current knowledge and recent insights into the genetic basis of amyotrophic lateral sclerosis. Med Genet 30:252–258. https://doi.org/10.1007/s11825-018-0185-3
Walker AK, Atkin JD (2011) Stress signaling from the endoplasmic reticulum: a central player in the pathogenesis of amyotrophic lateral sclerosis. IUBMB Life 63:754–763. https://doi.org/10.1002/iub.520
Wang GS, Cooper TA (2007) Splicing in disease: disruption of the splicing code and the decoding machinery. Nat Rev Genet 8:749–761. https://doi.org/10.1038/nrg2164
Wasser CR, Herz J (2016) Splicing therapeutics for Alzheimer’s disease. EMBO Mol Med 8:308–310. https://doi.org/10.15252/emmm.201506067
Wojciechowska M, Krzyzosiak WJ (2011) Cellular toxicity of expanded RNA repeats: focus on RNA foci. Hum Mol Genet 20:3811–3821. https://doi.org/10.1093/hmg/ddr299
Wu S, Green MR (1997) Identification of a human protein that recognizes the 3′ splice site during the second step of pre-mRNA splicing. EMBO J 16:4421–4432. https://doi.org/10.1093/emboj/16.14.4421
Yahara M, Kitamura A, Kinjo M (2017) U6 snRNA expression prevents toxicity in TDP-43-knockdown cells. PLoS ONE 12:e0187813. https://doi.org/10.1371/journal.pone.0187813
Yin S et al (2017) Evidence that C9ORF72 dipeptide repeat proteins associate with U2 snRNP to cause mis-splicing in ALS/FTD patients. Cell Rep 19:2244–2256. https://doi.org/10.1016/j.celrep.2017.05.056
Yu AC, Chan AY, Au WC, Shen Y, Chan TF, Chan HE (2016) Whole-genome sequencing of two probands with hereditary spastic paraplegia reveals novel splice-donor region variant and known pathogenic variant in SPG11. Cold Spring Harbor Mol Case Stud 2:a001248. https://doi.org/10.1101/mcs.a001248
Zhang J et al (2015) Disease-associated mutation in SRSF2 misregulates splicing by altering RNA-binding affinities. Proc Natl Acad Sci USA 112:E4726–E4734. https://doi.org/10.1073/pnas.1514105112
Zhang K et al (2018) ANXA11 mutations prevail in Chinese ALS patients with and without cognitive dementia. Neurol Genet 4:e237. https://doi.org/10.1212/NXG.0000000000000237
Zhao M, Kim JR, van Bruggen R, Park J (2018) RNA-binding proteins in amyotrophic lateral sclerosis. Mol Cells 41:818–829. https://doi.org/10.14348/molcells.2018.0243
Zhou Y, Liu S, Liu G, Ozturk A, Hicks GG (2013) ALS-associated FUS mutations result in compromised FUS alternative splicing and autoregulation. PLoS Genet 9:e1003895. https://doi.org/10.1371/journal.pgen.1003895
Zhu LY, Zhu YR, Dai DJ, Wang X, Jin HC (2018) Epigenetic regulation of alternative splicing. Am J Cancer Res 8:2346–2358
Zucca S et al (2019) RNA-seq profiling in peripheral blood mononuclear cells of amyotrophic lateral sclerosis patients and controls. Sci Data 6:190006. https://doi.org/10.1038/sdata.2019.6
Acknowledgements
The authors gratefully acknowledge the projects “Un approccio diagnostico multidisciplinare per le malattie neurodegenerative dell’invecchiamento” (DSB.AD009.001) and “Life Analytics, human centric platform per la salute ed il benessere dell’uomo” (DSB.AD008.456), and Angelo Bagalà, Ariangela Belvedere, Benedetto Bruno, Walter Carpino, Tiziana Martire, and Patrizia Rizzuto for their administrative and technical support.
Funding
This work has been supported by Grants: (i) “Un approccio diagnostico multidisciplinare per le malattie neurodegenerative dell’invecchiamento”, DSB.AD009.001; (ii) “Sviluppo ed applicazione di tecnologie biosensoristiche in Genomica” CIP 2014.IT.05.SFOP.014/3/10.4/9.2.10/0008, CUP G67B17000170009, n. 11/2017 “Rafforzare l’Occupabilità nel sistema R&S e la nascita di spin off di ricerca in Sicilia—PROGRAMMA OPERATIVO DEL FONDO SOCIALE EUROPEO REGIONE SICILIA 2014–2020”, and (iii) “Life Analytics, human centric platform per la salute ed il benessere dell’uomo”, DSB.AD008.456.
Author information
Authors and Affiliations
Contributions
BP and VLC reviewed the literature and wrote the manuscript. FLC participated in revising the manuscripts. TS, CU, SA provided critical inputs to the manuscript. SC conceived, directed, and supervised the project. All authors have read and approved the final version of this manuscript, and agreed to be accountable for all aspects of the work and consent for publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Perrone, B., La Cognata, V., Sprovieri, T. et al. Alternative Splicing of ALS Genes: Misregulation and Potential Therapies. Cell Mol Neurobiol 40, 1–14 (2020). https://doi.org/10.1007/s10571-019-00717-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-019-00717-0