Abstract
Biobanking of molecularly characterized colorectal cancer stem cells (CSCs) generated from individual patients and growing as spheroids in defined serum-free media offer a fast, feasible, and multi-level approach for the screening of targeted therapies and drug resistance molecular studies. By combining in vitro and in vivo analyses of cetuximab efficacy with genetic data on an ongoing collection of stem cell-enriched spheroids, we describe the identification and preliminary characterization of microsatellite stable (MSS) CSCs that, despite the presence of the KRAS (G12D) mutation, display epidermal growth factor (EGF)-dependent growth and are strongly inhibited by anti-EGF-receptor (EGFR) treatment. In parallel, we detected an increased resistance to anti-EGFR therapy of microsatellite instable (MSI) CSC lines irrespective of KRAS mutational status. MSI CSC lines carried mutations in genes coding for proteins with a role in RAS and calcium signaling, highlighting the role of a genomically unstable context in determining anti-EGFR resistance. Altogether, these results argue for a multifactorial origin of anti-EGFR resistance that emerges as the effect of multiple events targeting direct and indirect regulators of the EGFR pathway. An improved understanding of key molecular determinants of sensitivity/resistance to EGFR inhibition will be instrumental to optimize the clinical efficacy of anti-EGFR agents, representing a further step towards personalized treatments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
We previously reported on a recently established biobank of colorectal CSCs isolated from fresh patient’s tissues and expanded in serum-free medium as spheroids (De Angelis et al. 2016). Newly established cultures are routinely validated by (i) patient-matching verification through STR (short tandem repeats) analysis, (ii) assessment of CSC capability to originate phenocopies of the original patient’s tumor upon xenografting into NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (NSG) mice, and (iii) analysis of expression of stem cell markers. CSCs are then molecularly characterized by whole exome sequencing (WES) and banked for future uses. Since our procedure of CSC generation reaches 40–50% of efficiency of CSC isolation/primary tissues processed, the biobank is highly representative of patient tumor diversity in terms of genetic landscape and drug sensitivity. Therefore, it offers a powerful tool both for the screening of anticancer drugs in vitro and in vivo, and for the analysis of the molecular events linked to sensitivity versus resistance. In comparison to other methods of CSC collection recently developed, such as organoid biobanks (Fujii et al. 2016; van de Wetering et al. 2015) and patient-derived xenograft (PDX) (Bertotti et al. 2011; Gao et al. 2015; Julien et al. 2012), spheroid cultures can be easily expanded to high cell yield, giving the possibility to fast and affordable tests. Noteworthy, since selective media formulations specifically support the isolation and the expansion of cancer spheroids, cultures are devoid of supporting cells, so that cancer cell-intrinsic events may be specifically identified and analyzed.
By routine analysis of the sensitivity of newly generated CSCs to the anti-estimated glomerular filtration rate (EGFR) MoAb cetuximab, we have identified a small group of CSCs carrying a missense mutation in KRAS (G12D) that are strongly dependent on EGF for proliferation, and can be targeted efficiently by treatment with anti-EGFR MoAbs. Here we report preliminary characterization of these lines, as an example of multiple approach to drug resistance analyses in CSCs.
Material and methods
CSC culture
CSCs were generated from individual patient’s tissues (Table S1) as previously reported (De Angelis et al. 2016), and cultured using an optimized medium (CSC medium, see below) in ultra-low attachment tissue culture flasks (Corning Costar, Cambridge, MA), in humidified atmosphere at 37 °C, and 5% CO2 as previously described in (De Angelis et al. 2018). The CSC medium consists of Advanced DMEM F12 added with 10% Hormone Mix 10× (see below), 100 units/mL of penicillin, 100 μg/mL of streptomycin, 0.29 mg/mL glutamine, 6 g/L glucose, 5 mM HEPES, 3.6 g/L, NaHCO3, 4 mg/L (all from ThermoFisher Scientific, Waltham, MA, USA), 0.1%, BSA, (≥ 700 U/L) heparin, 10 mM nicotinamide (all from Sigma-Aldrich, St. Louis, MO, USA), 20 ng/mL human recombinant EGF, and 10 ng/mL human recombinant bFGF (both from PeproTech, London, UK). Hormone Mix (10×) composition is Advanced DMEM F12 added with 1 g/L apo-transferrin, 250 mg/L insulin, 161 mg/L putrescine, 52 μg/L sodium selenite, 62 μg/L progesterone, and 10 mM HEPES (all from Sigma-Aldrich). Unless otherwise indicated, supplements and reagents were purchased from Sigma-Aldrich, St. Louis, MO, USA.
Animal procedures
All animal procedures were performed according to the Italian national animal experimentation guidelines (D.L.116/92) upon approval of the experimental protocol by the Italian Ministry of Health’s Animal Experimentation Committee. Four- to 6-week-old female NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (NSG) mice (The Jackson Laboratory, Bar Harbor, ME, https://www.jax.org) were used for experiments. For drug testing, 5 × 105 cells were injected subcutaneously in the flank of 12 replicate mice, in 100 mL 1:1 PBS/Matrigel (BD, Franklin Lakes, NJ, http://www.bd.com). Tumors were measured twice weekly by an external digital caliper, and volumes were calculated using the following formula: π/6 × d2 × D, where d and D represent shorter and longer tumor measurements, respectively. Drug treatments were started after tumor establishment (100–150 mm3). Mice were treated with 10 mg/kg cetuximab (Merck KGaA, Darmstad, Germany, http://www.emdgroup.com) intravenously twice weekly. Control animals were treated with vehicle only.
DNA extraction and MSI analysis
Genomic DNA was obtained from CSCs and patient-matched non-tumoral tissues (Dnasy mini kit, Qiagen, Limburg, Netherlands). Five mononucleotide and two pentanucleotide markers were co-amplified by the MSI analysis system (Promega Madison, WI, USA) according to the manufacturer’s directions. The fluorescent-labeled PCR products obtained from CSCs were separated by capillary electrophoresis using an AB3130 xl sequencer and evaluated with the GeneMapper software (Applied Biosystems/Life technologies). Criteria for definition of MSS and MSI are according to the Bethesda guidelines (Umar et al. 2004).
WES data analysis and validation
Exome enrichment was performed using SeqCap EZ Human Exome Kit v3.0 (NimbleGen, Roche) and the resulting target libraries were sequenced using a HiSeq2000 platform (Illumina). Paired-end reads were aligned to the human genome (UCSC GRCh37/hg19) with the Burrows-Wheeler aligner (BWA v. 0.7.10) (Li and Durbin 2009) and PCR duplicates were removed using Picard’s MarkDuplicates (http://broadinstitute.github.io/picard). The Genome Analysis Toolkit (GATK 3.3) (McKenna et al. 2010) was used for base quality score recalibration. Somatic point mutations were detected using Mutect software v.1.1.6 (Cibulskis et al. 2013), and small indels were identified through a comparison between indels called in individual CSC lines and their matched non-tumoral samples by means of the GATK Haplotype Caller algorithm (DePristo et al. 2011), with the following quality filters: quality score > 100 and quality-by-depth score > 1.5; indels below these thresholds or resulting from four or more reads having ambiguous mapping (this number being greater than 10% of all aligned reads) were discarded. The resulting somatic SNVs and small indels were annotated by SnpEff v3.6 (Cingolani et al. 2012) and dbNSFP2.8 (Liu et al. 2013) and filtered to retain only those located in exons with any effect on the coding sequence, and splice site regions (−3 to +8 with respect to exon-intron junctions). Variant validation and genotyping were performed by Sanger sequencing. Amplicons were directly sequenced using the ABI BigDye Terminator Sequencing kit (Applied Biosystems) and an automated capillary sequencer (ABI 3500, Applied Biosystems). Sequence electropherograms were analyzed using Sequencing Analysis Software v.5.4 (Applied Biosystems).
Western blotting
Cell lysates were obtained from approximately 5 × 105 spheroid cells by incubation of cell pellets in lysis buffer (20 mM Tris-HCl pH 7.2; 200 mM NaCl; 1% NP40, added with protease inhibitor cocktail and phosphatase inhibitor cocktails I and II). Cell lysates from xenograft-derived tissues were obtained from ~50 mg pieces of frozen xenograft lysed in 0.40 mL of lysis buffer (as above). Tissues were homogenized with Pro 200 Kema Keur (Pro Scientific Inc. Oxford) at maximum speed at 4 °C for 30 s. Lysate concentrations were determined by Bradford assay (Bio-Rad Laboratories, Hercules, CA, http://www.bio-rad.com) and equal amounts of proteins were loaded on a gradient precast gel (4–12% or; ThermoFisher), then transferred to nitrocellulose membranes (GE Healthcare, Piscataway, NJ, http://www.gelifesciences.com). Blots were blocked with TBST 5% non-fat dry milk and incubated overnight at 4 °C with primary antibodies, then incubated for 45 min with secondary horseradish peroxidase-conjugated antibodies dissolved in TBST 1% BSA. Chemiluminescent signals were detected with prime or select (ThermoFisher Scientific). Images were taken and analyzed with Bio-Rad ChemiDoc Imagers (Bio-Rad Laboratories). For densitometry quantification, immunoblot signals were acquired with ChemiDocMP (Bio-Rad) and the relative intensity was quantified with Image Lab software. Normalization was performed using β-tubulin as reference. Monoclonal anti-α-tubulin was from Sigma-Aldrich; other antibodies were from Cell Signaling Technology.
Viability assay and drug screening
CSC viability upon treatment with cetuximab and/or kinase inhibitors was determined by CellTiter-Glo luminescent cell viability assay (Promega, Madison, WI, http://www.promega.ca) according to the manufacturer’s directions. Briefly, CSCs (2.5–3 × 103 per well) were dissociated with TrypLE Express, seeded in 96-well plates (six replicates per experimental point) in CSC medium, and incubated in a humidified atmosphere at 37 °C, 5% CO2. Luminescence was detected with a DTX880 multimode microplate reader (Beckman Coulter, Brea, CA, https://www.beckmancoulter.com). Cetuximab was used at a concentration of 100 μg/mL; EGF and kinase inhibitors (Selleck Chemicals, Houston, TX, http://www.selleckchem.com) were used at the concentrations indicated in the text.
Statistical analysis
Statistical analyses and non-linear regressions were done by GraphPad Prism v.4.0 for Windows (GraphPad Software, San Diego, CA, www.graphpad.com). Statistical significance of data was evaluated by ANOVA and Bonferroni pairwise post-tests or by unpaired Student’s t test, as specified for each experiment in the legends to figures. Statistical significance up to 0.05 was accepted. P values are indicated as follows: ***p value < 0.001, **p value < 0.01, *p value < 0.05.
Results
KRAS-mutated CSCs are inhibited by cetuximab and EGF deprivation
By routine analysis of newly generated CSCs for cetuximab sensitivity, we identified two lines, CSC28 and CSC26, both carrying a somatically acquired missense substitution in KRAS (G12D) that were highly inhibited by the MoAb and strictly depended on EGF for growth. Figure 1a shows the dose-response to EGF of the KRAS-mutated, cetuximab-sensitive (Mut-S) CSCs, in the absence or in the presence of 100 μg/mL cetuximab. The analysis of representative KRAS wild-type, cetuximab-sensitive (WT-S, CSC1, and CSC11) (Fig. 1b) and KRAS (G12D) mutant, resistant lines (Mut-R, CSC10, and CSC8) (Fig. 1c) is shown for comparison. In both Mut-S CSCs, after 3 days of treatment, cell growth decreased at levels lower than 25% of the maximum, at EGF concentrations of 0.02 μg/mL, and was completely inhibited in the absence of the growth factor. No resistant clones emerged from the cultures, and no residual alive cells were observed after 8 days of culture (not shown). Addition of cetuximab exacerbated the growth inhibition, this likely indicating autocrine production of EGFR ligand(s) by the cells. By comparing the inhibition curves of Mut-S and WT-S CSCs, no major difference was evident in the degree of dependence on EGF. The EC50 values of EGF of the different lines, both in the absence and in the presence of cetuximab (Fig. 1d), did not correlate with the presence or the absence of the mutation. Conversely, two representative mutated CSCs were completely resistant to EGF removal and/or cetuximab treatment. Western blot analysis of the CSCs tested confirmed that phosphorylation level of ERK and AKT decreases in Mut-S CSCs following EGF deprivation, at similar levels to those observed in WT-S CSCs. Conversely, no effect on protein phosphorylation was evident in control Mut-R lines (Fig. 1e–f). To verify these data in vivo, we analyzed the effect of cetuximab on xenografts generated with Mut-S, WT-S, and Mut-R CSCs (Fig. 2a). Cetuximab completely inhibited the growth of the Mut-S CSC, similarly than in the WT-S lines. In contrast, Mut-R CSC xenografts were fully resistant to the drug, altogether confirming the good correlation between cetuximab effects in vitro and on xenografts we had previously shown (De Angelis et al. 2016). Western blot analysis of xenograft cells ex vivo (Fig. 2b–c) confirmed that AKT and ERK phosphorylation is reduced in xenografts of sensitive CSCs, whether WT or mutated. Conversely, Mut-R CSC xenografts showed no variations in phosphorylation levels upon treatment.
KRAS-mutated CSCs are inhibited by EGF deprival and by cetuximab in vitro. a–c EGF dose-response in the absence (blue) or in the presence (red) of cetuximab (100 μg/mL), on Mut-S, WT-S, and Mut-R CSCs. Mutational status is indicated for each CSC. Spheroid culture viability was assessed at 72 h by CellTiter-Glo Luminescent Cell Viability Assay as described in the “Material and methods,” and is expressed as the percentage of maximal viability. Data represent the average of three independent experiments ± SEM. d LogEC50 for the curves in a–c. Calculations and non-linear regressions were done by GraphPad Prism software. The equation of fitting curve is \( \mathrm{Y}=\mathrm{Bottom}+\frac{\left(\mathrm{Top}\hbox{--} \mathrm{Bottom}\right)}{1+{10}^{\mathrm{LogEC}50\hbox{--} \mathrm{X}}} \). e Representative Western blot analysis for relevant downstream EGFR intermediates on representative Mut-S, WT-S, and Mut-R CSCs following treatments as in a–c. f Densitometry quantification of Western blot, mean ± SEM, n = 3. Statistical significance was calculated by two-tailed Student’s t test; ***p value < 0.001, **p value < 0.01, *p value < 0.05
KRAS-mutated CSCs are inhibited by cetuximab in vivo. a Xenografts of Mut-S, WT-S, and Mut-R CSCs were generated by subcutaneous injection as in “Material and methods,” and after tumor establishment mice were treated with 10 mg/kg cetuximab intravenous twice a week. Mutational status is indicated for each CSC. Data represent the average of six to ten tumors/group ± SEM. Representative experiment is shown. Statistical significance was calculated by two-tailed student’s t test; ***p value < 0.001, **p value < 0.01, *p value < 0.05. b Representative Western blot analysis for relevant downstream EGFR intermediates on xenografts as in a. Cells from six to eight tumors were pooled for analysis. c Densitometry quantification of Western blot ± SEM, n = 3. Statistical significance was calculated by two-tailed student’s t test; ***p value < 0.001, **p value < 0.01, *p value < 0.05
KRAS-mutated CSCs respond to inhibitors of EGFR downstream pathways
Several reports indicate that resistance of KRAS-mutated cancer cells to EGFR inhibition can be circumvented by combining the MoAb with inhibitors of different downstream molecules of the EGFR pathway, including MEK, PI3K, AKT, mTOR, or JAK (Allen et al. 2013; Cheng et al. 2015; Hong et al. 2016; Napolitano et al. 2015; Troiani et al. 2014; Zhang et al. 2015). In order to verify this point in our model, and more in general to investigate whether Mut-S, WT-S, and/or Mut-R CSCs would respond differently to pathway-specific inhibitors, we screened a panel of inhibitors directed to the main EGFR-related pathways (Table S2 and Fig. 3) on representative lines. The inhibitors were tested at 500 nM, in the presence or the absence of cetuximab. Exogenous EGF concentration was 2 ng/mL EGF, chosen to allow the growth of EGF-dependent CSC, without turning off the effect of cetuximab. As shown in Fig. 4, no major qualitative difference in the response to the different classes of targeted agents was evident, among Mut/S (4a), WT/S (4b), and Mut/R CSCs (4c). In fact, three classes of inhibitors proved the most active on all the CSCs tested, regardless of their mutational status: the inhibitors of MEK, the multiple inhibitor of mTOR/PI3K PF05212384, and the multiple inhibitor of mTOR/PI3K/AKT PF0469152. However, several agents directed against AKT, PI3K, mTOR, JAK, or PLK showed an additive effect with cetuximab on both Mut/S and WT/S, but not on Mut/R CSCs. To detail this point, we performed dose-response curves for three representative inhibitors, i.e., PF0469152 (multiple inhibitor of AKT/PI3K/mTOR), TG101348 (directed to JAK2), and AS703026 (directed to MEK) in the presence or in the absence of 100 μg/mL cetuximab, on Mut/S, Mut-R, and WT-S CSCs. The results are shown in Fig. 5 and Table S3. Cetuximab displayed an additive effect in combination with PF0469152, targeted to AKT/PI3K/mTOR (Fig. 5a, d), or with the JAK2 inhibitor TG101348 (Fig. 5b, e), on both Mut-S and WT-S CSCs. An additive effect with MEK inhibition was less evident (Fig. 5c, f). Conversely, cetuximab displayed no combined effect with any of the inhibitors tested, on the two Mut-R CSCs analyzed (Fig. 5h, i, g). Taken together, these data confirm that different subgroups of KRAS (G12D)-mutated CSCs can be distinguished: one in which the mutation parallels complete unresponsiveness to cetuximab, either alone or in combination with the inhibitors of the major EGFR pathway branches. However, a second subgroup is exemplified by CSC26 and CSC28, in which despite the presence of KRAS (G12D), cetuximab is still active on the EGFR signaling pathway, and its action is additive with inhibition of the AKT/PI3K/MTOR and the JAK2 pathways.
Schematic representation of EGFR downstream pathway effectors, reporting the activity of the inhibitors tested on CSCs (listed in Table S2)
Targeted pathway inhibition of Mut-S, WT-S, and Mut-R CSCs. Drug screening was performed on dissociated spheroid cells, as described in “Material and methods.” Cell viability was assessed at 72 h by CellTiter-Glo assay and expressed as the percentage of DMSO-treated control cells (black bar). Bars relative to inhibitor-treated cells were assigned different colors according to the pathway inhibited, as specified in the upper right legend. Each compound was tested as a single agent at a 500-nM concentration (colored bars) or in combination with 100 μg/mL cetuximab (white bars). Samples were ordered by their sensitivity to the combination of cetuximab plus targeted inhibitor. Data represent the average of three independent experiments ± SEM. CETU, cetuximab. a Mut-S CSCs. b WT-S CSCs. c Mut-R CSCs
Dose-response of AKT/PI3K/mTOR, JAK2, and MEK inhibitors alone or in combination with cetuximab on Mut-S, WT-S, and Mut-R CSCs. a–i Inhibitor dose-response in the absence (orange, green and blue lines) or in the presence (red lines) of cetuximab (100 μg/mL), on Mut-S, WT-S, and Mut-R CSCs. CSCs mutational status is indicated in the figure. Spheroid culture viability was assessed at 72 h by CellTiter-Glo as described in “Material and methods” and is expressed as the percentage of maximal viability. Data represent the average of three independent experiments ± SEM. a, d, g PF0469152 (multiple inhibitor of AKT/PI3K/mTOR). b, e, h TG101348 (directed to JAK2). c, f, i AS703026 (directed to MEK). Calculations and non-linear regressions were done by GraphPad Prism software. The equation of best fitting curves in a, d, g, c, f, i is \( \mathrm{Y}=\mathrm{Bottom}+\frac{\left(\mathrm{Top}\hbox{--} \mathrm{Bottom}\right)}{1+{10}^{\mathrm{LogEC}50\hbox{--} \mathrm{X}}} \). Equation for best fitting curves in b, e, h is \( \mathrm{Y}=\mathrm{Bottom}+\frac{\left(\mathrm{Top}\hbox{--} \mathrm{Bottom}\right)}{1+{10}^{\left(\mathrm{LogEC}50\hbox{--} \mathrm{X}\right)-\mathrm{HillSlope}}} \). Summary of relevant values for best fitting curves is reported in Table S3
CSC sensitivity to cetuximab correlates with MS status: mutations in GAPs and GEFs recur in cetuximab-resistant MSI CSCs
MSI tumors constitute a distinct subgroup of colorectal cancers, defined by deficiency in the mismatch repair (MMR) machinery. MSI colorectal tumors prevalently occur to the right side intestine, exhibit mucinous histology with abundant lymphocyte infiltration, and have better clinical outcome than microsatellite stable (MSS) tumors (Vilar and Gruber 2010). They also have specific molecular characteristics in terms of mutation landscape (Cortes-Ciriano et al. 2017; Kim et al. 2013) and gene expression profile (Guinney et al. 2015).
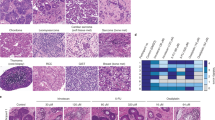
Prompted by the observation that both sensitive CSC26 and CSC28 are MSS, while CSC8 and CSC10 are MSI, we took into examination MS status and cetuximab resistance within a wider panel of CSCs (Table S1 and Fig. S1–S2). As shown in Fig. 6a, average sensitivity of MSS was significantly higher than that of MSI CSCs (average percentage of maximal growth 37.0 ± 7.9 versus 79.2 ± 10.6, t test, p = 0.0075). When analyzed separately (Fig. 6b), both MSI-WT and MSI-Mut CSCs averaged as highly resistant to cetuximab (average viability ~75% of untreated controls), independently from their KRAS mutation status. Conversely, MSS-WT CSCs were highly sensitive to the drug (average viability ~25% of controls). Finally, MSS-Mut CSCs averaged as medium responders (average viability ~50% of controls). MSI lines mutated in BRAF available were all highly resistant to cetuximab (not shown) and were not included in the analysis. A relevant feature of MSI cancer cells is that impairment of the DNA repair machinery determines progressive hypermutation: accordingly, MSI CSCs share high mutation rates (De Angelis et al. 2016; Vilar and Gruber 2010); Fig. S1). Analysis of CSCs mutations in genes involved in cetuximab resistance in CRC, in fact, shows a higher frequency of widespread mutations in MSI versus MSS lines (Fig. 7a and Table S4). To extend this observation to other potentially relevant genes, we examined the pattern of mutations in MSS and MSI CSCs, within panels of genes functionally involved within the KRAS, MAPK, and PI3K-AKT cascades, selected according to the KEGG pathway database (Tanabe and Kanehisa 2012). As shown in Fig. 7b–d and Table S4, specific groups of mutations emerged in MSI cells, recurrently hitting groups of genes structurally and/or functionally related. Such targets included matrix and cell matrix-related genes, i.e., collagens; laminin A, B, and C; integrin A and B; and filamin A, B, and C. Most interestingly, MSI CSCs showed a higher frequency of mutations in genes codifying for proteins regulating RAS function, specifically GTPase-activating proteins (GAPs) including RASAL1, RASAL2, and RASAL3, and NF1, and guanine nucleotide exchange factors (GEFs) such as RASGRF1, SOS, and RAPGEF2. Calcium has a deep impact on RAS signaling, both direct and mediated trough regulators including RASAL and RASGRF1. Of note, mutations affecting several CACNA genes—codifying for voltage-gated calcium channels—were identified in MSI CSCs. By contrast, MSI, cetuximab-highly sensitive CSCs were characterized by lower mutation rates and were scarcely mutated in EGFR-cascade genes. In fact, the most sensitive among WT-S lines, i.e., CSC1, CSC11, CSC12, and CSC27, carried respectively 1, 3, 4, and 5 mutated genes respectively, among the selected panels of functionally relevant genes. Such low mutation incidence was common to Mut-S lines; in fact, CSC26 and CSC28 carried mutations in 5 and 3 genes respectively, including KRAS (Fig. 7).
CSC sensitivity to cetuximab correlates with MS status and low frequency of mutations in functionally relevant genes. a Average inhibition of MSS versus MSI CSCs. Viability is expressed as percentage of untreated controls. Error bars represent SEM from the mean. Statistical significance was calculated by two-tailed student’s t test. b Average inhibition of KRAS MSS-WT, MSS-MUT, MSI-WT, and MSI-MUT CSCs. Error bars represent SEM from the mean. Statistical significance was calculated by one-way ANOVA with Bonferroni pairwise post-test. ***p value < 0.001, **p value < 0.01, *p value < 0.05
WES-based mutation profiling of CSCs analyzed in Fig. 6. a Functionally relevant intragenic lesions in individual CSC lines. b–d WES-based mutation profiling for recurrent genes, within KEGG pathway database panels. b KRAS. c MAPK. d PI3K-AKT
Discussion
By preliminary analysis of a panel of patient-derived CSCs, we have shown that a small subpopulation of KRAS (G12D)-mutated CSCs strictly depends on EGF for growth and is highly sensitive to the anti-EGFR MoAb cetuximab. These cells respond to targeted inhibitors of several intermediates of the EGFR signaling cascade, including AKT, PI3K, mTOR, JAK, or PLK, with an additive effect upon cetuximab co-treatment. In particular, we have detailed the effect of the combination of cetuximab/PF0469152 (directed against AKT/PI3K/mTOR), and cetuximab/TG101348 (directed to JAK2). Both drug combinations exert an additive inhibition on WT-S and Mut-S, but not on representative Mut-R CSCs, altogether indicating a different degree of dysregulation of the EGFR downstream pathway among different groups of mutated CSCs.
We have shown that Mut-S CSCs are characterized by very low frequency of mutations in the EGFR downstream genes, similarly to cetuximab-highly responsive, WT-S lines. In fact, we have also shown that CSC MSI status correlates with high cetuximab resistance, independently from the cell’s mutational status in KRAS. Conversely, MSS, either KRAS WT or mutated, includes CSCs medium-highly responsive to the drug. Mutations in KRAS are clinically associated with resistance to anti-EGFR treatment (De Roock et al. 2010); therefore, determination of KRAS status is now recommended in metastatic CRC patients before starting anti-EGFR treatment. It is generally accepted that constitutively activated, mutated KRAS allows the cell to bypass upstream inhibition of EGFR. However, organoids and PDX generated with KRAS-mutated tumors that are inhibited by cetuximab have been previously reported (Fujii et al. 2016; Isella et al. 2017; Julien et al. 2012; Schutte et al. 2017). Such discrepancy with clinical data may be explained by the absence of stromal cells in these systems: recent studies (Isella et al. 2017; Schutte et al. 2017) demonstrated wide differences in gene expression patterns among patient’s and PDX-developed tumors, pointing to a prominent role of stroma in tumor determination. Stroma-secreted factors such as HGF, TGFβ, AREG, and EREG strongly influence tumor sensitivity to anti-EGFR treatment (Luraghi et al. 2014; Mutsaers et al. 2009; Takahashi et al. 2014). In this frame, serum-free grown spheroids offer a clear-cut approach to the analysis of tumor cell-restricted phenomena. Alternatively or in addition, complete dysregulation of EGFR signaling in KRAS (G12D) CSCs may be linked to a context of other accumulating mutations. Indeed, recent work showed that in colorectal organoids engineered by CRISPR-Cas9 with oncogenic APC, SMAD4, and TP53, the addition of KRAS (G12D) did not induce per se full activation of EGFR signaling, since mutated cells still required EGF for growth (Matano et al. 2015). Among others, aberrant functionality of the KRAS regulators GAPs and GEFS contribute to oncogenesis in some contexts (Vigil et al. 2010), and may cooperate to determine overall CSC dependence on EGF signaling. Indeed, a feedback loop has been recently described, by which the RASGRP-1 GAP turns off oncogenic KRAS, in response to EGFR activation (Depeille et al. 2015). In this frame, it is particularly interesting that MSI CSCs analyzed in this study carry recurrent mutations in several RAS, GAPs, and GEFs. Calcium flux also holds a relevant role in the modulation of GAPs, GEFS, and RAS itself (Cullen and Lockyer 2002): We found that several isoforms of the calcium channel genes are also recurrently mutated in MSI. Altogether, these data strengthen the hypothesis that KRAS (G12D) may not be sufficient per se to confer resistance to CSCs, but other factors may be required to fully deregulate EGFR pathway. Among these, mutations hitting GAFs, GEFs, and/or calcium flux may hold a relevant role. Further research based on the combination of genomic and functional approaches is needed for a complete molecular dissection of colorectal CSC resistance to anti-EGFR targeted agents. The increased availability of different model systems should be exploited in order to achieve both a comprehensive picture of tumor complexity and an improved clinical application of individualized therapies.
Author contribution summary
MLDA: experimental data generation and assembly. ABr: WES data processing and analysis. FF: experimental data generation. FP: experimental data generation. SV: experimental data generation. AZ: manuscript revision. MT: WES data analysis; MB: conception and design, data assembly and interpretation, manuscript writing.
References
Allen JE, Gallant JN, Dicker DT, Amin S, Irby RB, Sharma AK, et al. The Akt inhibitor ISC-4 synergizes with cetuximab in 5-FU-resistant colon cancer. PLoS One. 2013;8(3):e59380.
Bertotti A, Migliardi G, Galimi F, Sassi F, Torti D, Isella C, et al. A molecularly annotated platform of patient-derived xenografts (“xenopatients”) identifies HER2 as an effective therapeutic target in cetuximab-resistant colorectal cancer. Cancer Discov. 2011;1(6):508–23.
Cheng L, Xia Z, Bian X, Li G, Hu J, Cao Y, et al. Combination of cetuximab and PP242 synergistically suppress the progression of wild-type KRAS colorectal carcinoma. Onco Targets Ther. 2015;8:3185–92.
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol. 2013;31(3):213–9.
Cingolani P, Platts A, Wang le L, Coon M, Nguyen T, Wang L, et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin). 2012;6(2):80–92.
Cortes-Ciriano I, Lee S, Park WY, Kim TM, Park PJ. A molecular portrait of microsatellite instability across multiple cancers. Nat Commun. 2017;8:15180.
Cullen PJ, Lockyer PJ. Integration of calcium and Ras signalling. Nat Rev Mol Cell Biol. 2002;3(5):339–48.
De Angelis ML, Zeuner A, Policicchio E, Russo G, Bruselles A, Signore M, et al. Cancer stem cell-based models of colorectal cancer reveal molecular determinants of therapy resistance. Stem Cells Transl Med. 2016;5(4):511–23.
De Angelis ML, De Maria R, Baiocchi M. How to assess drug resistance in cancer stem cells. Methods Mol Biol. 2018;1692:107–15.
De Roock W, Claes B, Bernasconi D, De Schutter J, Biesmans B, Fountzilas G, et al. Effects of KRAS, BRAF, NRAS, and PIK3CA mutations on the efficacy of cetuximab plus chemotherapy in chemotherapy-refractory metastatic colorectal cancer: a retrospective consortium analysis. Lancet Oncol. 2010;11(8):753–62.
Depeille P, Henricks LM, van de Ven RA, Lemmens E, Wang CY, Matli M, et al. RasGRP1 opposes proliferative EGFR-SOS1-Ras signals and restricts intestinal epithelial cell growth. Nat Cell Biol. 2015;17(6):804–15.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43(5):491–8.
Fujii M, Shimokawa M, Date S, Takano A, Matano M, Nanki K, et al. A colorectal tumor organoid library demonstrates progressive loss of niche factor requirements during tumorigenesis. Cell Stem Cell. 2016;18(6):827–38.
Gao H, Korn JM, Ferretti S, Monahan JE, Wang Y, Singh M, et al. High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nat Med. 2015;21(11):1318–25.
Guinney J, Dienstmann R, Wang X, de Reynies A, Schlicker A, Soneson C, et al. The consensus molecular subtypes of colorectal cancer. Nat Med. 2015;21(11):1350–6.
Hong S, Kim S, Kim HY, Kang M, Jang HH, Lee WS. Targeting the PI3K signaling pathway in KRAS mutant colon cancer. Cancer Med. 2016;5(2):248–55.
Isella C, Brundu F, Bellomo SE, Galimi F, Zanella E, Porporato R, et al. Selective analysis of cancer-cell intrinsic transcriptional traits defines novel clinically relevant subtypes of colorectal cancer. Nat Commun. 2017;8:15107.
Julien S, Merino-Trigo A, Lacroix L, Pocard M, Goere D, Mariani P, et al. Characterization of a large panel of patient-derived tumor xenografts representing the clinical heterogeneity of human colorectal cancer. Clin Cancer Res. 2012;18(19):5314–28.
Kim TM, Laird PW, Park PJ. The landscape of microsatellite instability in colorectal and endometrial cancer genomes. Cell. 2013;155(4):858–68.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25(14):1754–60.
Liu X, Jian X, Boerwinkle E. dbNSFP v2.0: a database of human non-synonymous SNVs and their functional predictions and annotations. Hum Mutat. 2013;34(9):E2393–402.
Luraghi P, Reato G, Cipriano E, Sassi F, Orzan F, Bigatto V, et al. MET signaling in colon cancer stem-like cells blunts the therapeutic response to EGFR inhibitors. Cancer Res. 2014;74(6):1857–69.
Matano M, Date S, Shimokawa M, Takano A, Fujii M, Ohta Y, et al. Modeling colorectal cancer using CRISPR-Cas9-mediated engineering of human intestinal organoids. Nat Med. 2015;21(3):256–62.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20(9):1297–303.
Mutsaers AJ, Francia G, Man S, Lee CR, Ebos JM, Wu Y, et al. Dose-dependent increases in circulating TGF-alpha and other EGFR ligands act as pharmacodynamic markers for optimal biological dosing of cetuximab and are tumor independent. Clin Cancer Res. 2009;15(7):2397–405.
Napolitano S, Martini G, Rinaldi B, Martinelli E, Donniacuo M, Berrino L, et al. Primary and acquired resistance of colorectal cancer to anti-EGFR monoclonal antibody can be overcome by combined treatment of regorafenib with cetuximab. Clin Cancer Res. 2015;21(13):2975–83.
Schutte M, Risch T, Abdavi-Azar N, Boehnke K, Schumacher D, Keil M, et al. Molecular dissection of colorectal cancer in pre-clinical models identifies biomarkers predicting sensitivity to EGFR inhibitors. Nat Commun. 2017;8:14262.
Takahashi N, Yamada Y, Furuta K, Honma Y, Iwasa S, Takashima A, et al. Serum levels of hepatocyte growth factor and epiregulin are associated with the prognosis on anti-EGFR antibody treatment in KRAS wild-type metastatic colorectal cancer. Br J Cancer. 2014;110(11):2716–27.
Tanabe, M. & Kanehisa, M. (2012) Using the KEGG database resource. Curr Protoc Bioinformatics, Chapter 1, Unit1 12.
Troiani T, Napolitano S, Vitagliano D, Morgillo F, Capasso A, Sforza V, et al. Primary and acquired resistance of colorectal cancer cells to anti-EGFR antibodies converge on MEK/ERK pathway activation and can be overcome by combined MEK/EGFR inhibition. Clin Cancer Res. 2014;20(14):3775–86.
Umar A, Boland CR, Terdiman JP, Syngal S, de la Chapelle A, Ruschoff J, et al. Revised Bethesda Guidelines for hereditary nonpolyposis colorectal cancer (Lynch syndrome) and microsatellite instability. J Natl Cancer Inst. 2004;96(4):261–8.
van de Wetering M, Francies HE, Francis JM, Bounova G, Iorio F, Pronk A, et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell. 2015;161(4):933–45.
Vigil D, Cherfils J, Rossman KL, Der CJ. Ras superfamily GEFs and GAPs: validated and tractable targets for cancer therapy? Nat Rev Cancer. 2010;10(12):842–57.
Vilar E, Gruber SB. Microsatellite instability in colorectal cancer-the stable evidence. Nat Rev Clin Oncol. 2010;7(3):153–62.
Zhang FQ, Yang WT, Duan SZ, Xia YC, Zhu RY, Chen YB. JAK2 inhibitor TG101348 overcomes erlotinib-resistance in non-small cell lung carcinoma cells with mutated EGF receptor. Oncotarget. 2015;6(16):14329–43.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All animal procedures were performed according to the Italian national animal experimentation guidelines (D.L.116/92) upon approval of the experimental protocol by the Italian Ministry of Health’s Animal Experimentation Committee.
Electronic supplementary material
Supplementary Figure 1
Somatic mutation frequencies of individual CSCs in Fig. 6 (a-b), estimated by WES analysis. (JPEG 142 kb)
Supplementary Figure 2.
Sensitivity to cetuximab in EGF-deprived cultures of individual CSCs in Fig. 6 (a-b). Cell growth is expressed as the percentage of maximal viability occurring in the presence of 20 ng/mL exogenous EGF. Spheroid culture viability was assessed at 72 h by CellTiter-Glo Assay as described in the Methods section. Data represent the average of three independent experiments ± SEM. (JPEG 157 kb)
ESM 1
(XLS 34 kb)
ESM 2
(XLS 43 kb)
ESM 3
(XLSX 11 kb)
ESM 4
(XLSX 27 kb)
Rights and permissions
About this article
Cite this article
De Angelis, M.L., Bruselles, A., Francescangeli, F. et al. Colorectal cancer spheroid biobanks: multi-level approaches to drug sensitivity studies. Cell Biol Toxicol 34, 459–469 (2018). https://doi.org/10.1007/s10565-018-9423-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10565-018-9423-3