Abstract
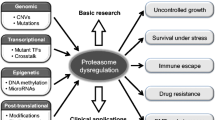
Proteasomal protein degradation is one of the major regulatory mechanisms in the cell. Aberrant proteasome activity is directly related to the pathogenesis of many human diseases including cancers. How proteasome homeostasis is controlled is a fundamental question toward our understanding of proteasome dysregulation in cancer cells. The recent discovery of the Rpn4-proteasome negative feedback circuit provides mechanistic insight into the regulation of proteasome gene expression. This finding also has important implications in cancer therapy that uses small molecule inhibitors to target the proteasome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 The 26S proteasome
The 26S proteasome is the major cellular protease responsible for degradation of regulatory and abnormal proteins [1–4]. It is involved in many cellular processes including but not limited to cell cycle control, transcription, DNA repair, apoptosis, quality control, and antigen presentation. The 26S proteasome is a 2.5-MDa protein complex consisting of a 20S core particle (CP or 20S proteasome) and one or two 19S regulatory particles (RP, also known as 19S proteasome or PA700). The RP is attached to one or both ends of the CP. Crystal structures of the CP from archaeal, actinobacterial species, and several eukaryotes have been solved [5–8]. The CP is a barrel-shaped structure of a stack of four seven-subunit rings in an α7β7β7α7 configuration (Fig. 1). Both exterior rings contain one set of seven different α subunits; and both interior rings contain one set of seven different β subunits. The CP performs three types of catalytic activities inside its chamber: chymotrypsin-like, trypsin-like, and caspase-like activities, which are provided by β5, β2, and β1 subunits. In immune responsive cells, the constitutively expressed β1, β2, and β5 subunits are replaced by three induced β subunits (β1i, β2i, and β5i) to form the immunoproteasome that has higher chymotrypsin-like and trypsin-like activities known to be favorable for antigen processing [9–11]. β5i is replaced by another proteolytic active subunit β5t in cortical thymic epithelial cells, forming the so-called thymoproteasome [12].
Composition of the 26S proteasome. The 26S proteasome consists of the 20S proteasome (CP) and the 19S regulatory particle (RP). The CP is formed by four stacked rings: two outside α-rings and two inner β-rings. Each ring has seven different subunits. The RP is divided into two subcomplexes: the lid and the base. The subunits forming the lid and the base are shown. Rpn10 stabilizes the connection between the lid and the base
Protein substrates enter the CP chamber through a pore or gate at the center of the α subunit ring. The gate is closed in a free CP by interactions among the N-termini of the α subunits, blocking substrate entry into the proteolytic chamber [13]. In the 26S proteasome, the RP acts as an activator to facilitate the opening of the gate by interacting with the α subunits [13–15]. The RP complex can be divided into two subcomplexes called the base and the lid (Fig. 1). The base is in contact with the CP, consisting of six AAA(+)-type ATPases (Rpt1–6) and three non-ATPase subunits (Rpn1, Rpn2, and Rpn13). The primary functions of the base include substrate recruiting, unfolding, and translocation. The lid is formed by at least nine non-ATPase subunits (Rpn3, Rpn5-9, Rpn11, Rpn12, and Rpn15/Sem1). The Rpn11 subunit possesses a deubiquitylating activity, which is required for degradation of polyubiquitylated substrates. The connection between the lid and the base is stabilized by the Rpn10 subunit. Recent studies have shown that a number of ubiquitylating and deubiquitylating enzymes are associated with the 26S proteasome [16–19], suggesting that the 26S proteasome is not only a “garbage disposal” but also plays an active role in ubiquitin chain remodeling. Clearly, the structure and activity of the proteasome system are highly regulated.
2 The discovery of the Rpn4-proteasome negative feedback circuit
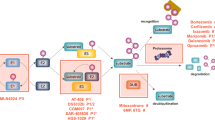
The 26S proteasome is composed of 33 distinct subunits each encoded by a different gene. It was a difficult task to understand how the proteasome genes, in such a large number, are regulated. The central question was whether the proteasome genes are coordinately regulated by a master transcription factor or if they are activated by different transcription factors. Early biochemical studies have shown that the proteasome subunits are nearly stoichiometrically present in Saccharomyces cerevisiae, suggesting that the proteasome genes are coordinately regulated [20, 21]. Recent studies by Mannhaupt et al. and our own demonstrated that Rpn4 encoded by the RPN4 gene (also named SON1 and UFD5) functions as a transcription activator of the proteasome genes [22–24]. An Rpn4 binding site, a 9-bp motif known as proteasome-associated control element (PACE), is located in the promoters of the proteasome genes. Deletion of the PACE motif from one of the proteasome genes markedly reduces the level of assembled/active proteasome in the cell [25]. Interestingly, Rpn4 is an extremely short-lived protein (t 1/2 ≤ 2 min) and is degraded by the proteasome [23]. Moreover, stabilization of Rpn4 by inhibition of proteasome activity results in the upregulation of the proteasome genes [24, 26]. Together, these observations led to the formulation of a model in which proteasome homeostasis is regulated by a negative feedback circuit (Fig. 2). On the one hand, Rpn4 induces the proteasome genes; on the other hand, Rpn4 is rapidly degraded by the assembled/active proteasome.
In addition to the proteasome genes, Rpn4 controls numerous non-proteasome genes involved in protein ubiquitylation, DNA repair, unfolded protein response, and other cellular processes [22, 27–40]. Interestingly, the RPN4 gene itself is regulated by a wide range of signals. For instance, the RPN4 promoter carries response elements for heat shock transcription factor (Hsf1), multidrug resistance-related transcription factors (Pdr1 and Pdr3), and Yap1, a transcription factor that plays an important role in response to oxidation, toxic metals, and DNA damaging agents [28–31]. It has become clear that environmental stressors activate these transcription factors to induce RPN4 expression. These observations suggest that Rpn4 acts as a major mediator in a stress response network. The Rpn4-proteasome feedback circuit appears to be the core operator in this network. Not only does it regulate proteasome homeostasis but it also gauges the induction of other Rpn4 target genes by keeping the Rpn4 protein level in check. In fact, recent studies have demonstrated that inhibition of Rpn4 degradation severely reduces cell viability under stressed conditions [41, 42].
3 Feedback regulation of proteasome gene expression in higher eukaryotes
A question naturally arising from the discovery of the Rpn4-proteasome negative feedback circuit is whether the proteasome genes are regulated by an Rpn4-like transcription factor in higher eukaryotes. Bioinformatics analyses conducted by several groups revealed that Rpn4 sequence homologs and conserved PACE motifs exist only in Hemiascomycetes, especially among the Saccharomyces “sensu stricto” species [43, 44]. However, the proteasome genes in higher eukaryotes are also coordinately regulated by a feedback mechanism [45–53]. For example, knockdown of individual proteasome subunits by RNAi results in the upregulation of non-targeted subunits in Drosophila cells; and the feedback response is dependent on the 5′-untranslated regions of proteasome genes [47, 48]. In addition, suppression of the proteasome activity by proteasome inhibitors induces upregulation of proteasome genes in mammalian cells [45, 49, 51–53]. These experimental observations strongly suggest that there is a functional homolog of Rpn4 in higher eukaryotes.
Recent studies have suggested that nuclear factor erythroid-derived 2-related factor 2 (Nrf2) may be the sought Rpn4 counterpart in mammalian cells [51, 54–57]. Nrf2 is a cap “n” collar-basic leucine zipper transcription factor that activates its target genes through a common cis-acting element named antioxidant response element (ARE) [58]. Previous studies have shown that Nrf2 is the most potent transcription factor in inducing the expression of antioxidant enzyme genes when the cell is under oxidative and electrophilic stress [59, 60]. Kensler and co-workers found that administration of antioxidant 3H-1,2-dithiole-3-thione, which is known to activate Nrf2, enhances the expression of proteasome genes in the liver of wild-type, but not Nrf2−/− mice [54]. This observation suggests that the proteasome genes are induced by Nrf2 in stress response. The same research group further demonstrated that the Nrf2-dependent induction of proteasome subunit PSMB5 by 3-methycholanthrene requires the AREs of the PSMB5 promoter in murine neuroblastoma Neuro2A cells [55]. Enhanced expression of proteasome genes by Nrf2 was also detected in human fibroblasts and colon cancer cells [56, 57]. Like Rpn4, Nrf2 is very short lived and degraded by the proteasome. The cellular Nrf2 level increases in response to stress because its interaction with Keap1 is disrupted. Keap1 is an adaptor protein required for Nrf2 ubiquitylation by the Cul3-Rbx1 ubiquitin ligase. These features make Nrf2 a perfect candidate for coordinating the feedback expression of the proteasome genes. Indeed, the work by Kraft et al. suggested that feedback induction of proteasome genes in response to proteasome inhibitor MG132 is mediated by Nrf2 in human skin fibroblast cells [51]. In addition to Nrf2, other cap n collar-basic leucine zipper transcription factors may also be involved in the feedback regulation of proteasome genes. A recent study has revealed that Nrf1 but not Nrf2 is required for the feedback induction of proteasome genes in response to proteasome inhibitors in mouse embryonic fibroblasts (MEFs) [53]. It is possible that the transcription factors required for the feedback proteasome gene expression are cell type dependent: The feedback regulation is conducted by Nrf1 in some cell types and by Nrf2 in others.
Unlike Rpn4, which controls both basal and inducible proteasome expression in yeast [24, 25], neither Nrf1 nor Nrf2 appears to play a major role in basal expression of proteasome genes in mammalian cells. Both Nrf1−/− and Nrf2−/− MEFs and the hepatic cells of Nrf2−/− mice display similar endogenous levels of proteasome subunits as their wild-type counterparts [53, 54]. These observations suggest that there may be two distinct systems to regulate the expression of the proteasome genes in mammalian cells: one for the basal level and the other for the feedback upregulation. Whereas recent studies have provided some clues to understand the feedback regulation mechanism, more work needs to be done to identify the pathway that controls the basal level expression of the proteasome genes in mammalian cells.
4 Implications of feedback regulation of proteasome gene expression in cancer therapy
The proteasome has emerged as a drug target for cancer therapy [61]. The illustration of the feedback regulation of proteasome gene expression has several important implications in cancer therapy that targets the proteasome. First, it provides a clue to understand the cause of proteasome overexpression often detected in cancers [62–66]. It is possible that the feedback induction, which normally occurs only when the proteasome activity is suppressed, may become constitutively active in cancer cells. In support of this hypothesis, Xu et al. demonstrated that the feedback expression of proteasome genes in response to proteasome inhibitors is less inducible in breast cancer cells than in noncancerous breast epithelial cells [49]. Second, the feedback mechanism may contribute to the emergence of bortezomib resistance in cancer therapy. Bortezomib is the only FDA-approved proteasome inhibitor in clinical use. Although this drug has shown promising results in the treatment of multiple myeloma and mantle cell lymphoma, it has limited efficacy in other types of cancers [61, 67]. The bortezomib resistance can be explained by multiple mechanisms. One possible mechanism is the feedback expression of proteasome genes. Bortezomib is a reversible inhibitor and is rapidly cleared from the patients’ blood [61, 68, 69]. The transiently inhibited proteasome activity recovers after drug clearance and drug dissociation from the active site. The recovery is conceivably boosted by new proteasome synthesis induced by the feedback mechanism, thereby reducing the extent and duration of proteasome inhibition. Whereas the compromised efficacy of bortezomib may be strong enough to kill myeloma tumor cells, it may not be sufficient to be effective in other cancers, especially solid tumors. Third, the feedback pathway is a potential target for cancer therapy. An early study by Ju et al. has already raised the concern that the efficacy of proteasome inhibitors may be compromised by the feedback upregulation of proteasome genes [24]. Ju et al. have also suggested that simultaneous targeting of the proteasome active sites and the transcription factor of the proteasome genes would be an efficient regimen for the treatment of cancer. This idea originated from the observation that the depletion of Rpn4 displays a strong synthetic growth defect with impairment of proteasome activity [24]. In support of this prediction, a recent study demonstrated that knockdown of Nrf1 enhances the cytotoxic effect of proteasome inhibitor YU101 in MDA-MB-231 breast cancer and U2OS osteosarcoma cell lines [53]. It will be of interest to examine whether this combined treatment is applicable to other types of cancer cells and whether it has a beneficial effect in clinical trials.
It is worthy of note that the downregulation of one of the proteasome genes by deletion of the Rpn4-binding site from the PRE1 promoter dramatically reduces the assembled proteasome level and proteasome activity [25]. This finding is of clinical relevance because it provides a potentially important approach to reduce the proteasome activity in cancer cells. To date, proteasome inhibitors attacking the catalytic sites of the proteasome are the only tool to reduce the proteasome activity. However, many types of cancers are resistant to proteasome inhibitors by a variety of mechanisms. Knockdown of individual proteasome genes may present a promising alternative to proteasome inhibitors in cancer therapy. In fact, a recent study revealed that knockdown of proteasome subunits by RNA interference impairs cancer cell growth and exhibits a synthetic lethal phenotype when combined with proteasome inhibitors [70].
5 Combined therapy via targeting the pathways that display synthetic lethal phenotypes with the proteasome
Combined therapy is a feasible approach to increase the efficacy of proteasome inhibitors in the treatment of cancers. Unfortunately, current regimens of combined therapy are simply an application of a second known anti-cancer agent together with bortezomib, and have failed to highlight a significant synergistic effect [61, 67]. The key to the success of combined therapy is to target a specific pathway, which is essential for cell growth and survival when the proteasome activity is inhibited. Whereas it is difficult to define such pathways in mammalian cells, recent studies in S. cerevisiae have provided the needed information.
The S. cerevisiae proteasome genes are coordinately activated by Rpn4. Loss of Rpn4 results in decreased expression of proteasome genes and consequently lower proteasome activity, which mimics the effect of proteasome inhibitors. Genome-wide analyses have shown that the deletion of RPN4 exhibits synthetic growth defects with mutations in numerous genes involved in a variety of pathways [71, 72]. Specifically, yeast cells lacking Rpn4 and one of these genes are either inviable or grow much slower than the mutants that carry a single mutation. Since most of these yeast genes have homologs in human cells, it is reasonable to speculate that simultaneous targeting of the proteasome and one of the human homologs may display a synergistic effect against cancer cells. As a proof of concept, Ju et al. explored the relevance of the synthetic growth defect caused by a double deletion of ERG24 and RPN4 to combined therapy [73]. ERG24 encodes the C-14 sterol reductase, which is required for ergosterol biosynthesis in yeast [74]. There are two Erg24 homologs in human cells with C-14 sterol reductase activity, including 3β-hydroxysterol C-14 reductase and lamin B receptor [75, 76]. To test if Egr24 could be a potential target for combined therapy with proteasome inhibitors, Ju et al. took advantage of two recently identified Erg24 inhibitors, dyclonine and alverine citrate [77]. Dyclonine and alverine citrate are the major components of two over-the-counter medicines. Dyclonine is an oral anesthetic found in throat lozenges, whereas alverine citrate is a commonly used smooth muscle relaxant for the treatment of irritable bowel syndrome. Ju et al. found that dyclonine and alverine citrate substantially enhanced the cytotoxic effect of the proteasome inhibitor MG132 in breast cancer cells [73]. The combined treatment with MG132 and dyclonine or alverine citrate markedly induced apoptosis. This study presents a powerful yeast genetic approach to identification of potential targets for combined therapy with proteasome inhibitors.
6 Concluding remarks
The discovery of feedback regulation of proteasome gene expression provides insight into how proteasome homeostasis is regulated. This finding also raises a concern in using proteasome inhibitors as a single regimen for the treatment of cancers, that is, feedback upregulation of proteasome genes may compromise the efficacy of proteasome inhibitors. This problem can be overcome by combined therapy that targets another gene or pathway to produce synthetic lethal phenotypes with inhibition of the proteasome. It is worthy of note that the proteasome activity can be regulated at different levels. In addition to gene expression, proteasome assembly is an important process to control the cellular proteasome activity. Moreover, post-translational modifications of proteasome subunits such as oxidation, phosphorylation, glycosylation, acetylation, and myristoylation have been reported to alter the proteasome activity [78–81]. Further investigation of the biological significance and mechanisms of these modifications will provide more choices for proteasome-targeting cancer therapy.
References
Voges, D., Zwickl, P., & Baumeister, W. (1999). The 26S proteasome: A molecular machine designed for controlled proteolysis. Annual Review of Biochemistry, 68, 1015–1068.
Finley, D. (2009). Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annual Review of Biochemistry, 78, 477–513.
Glickman, M. H., & Ciechanover, A. (2002). The ubiquitin–proteasome proteolytic pathway: Destruction for the sake of construction. Physiological Reviews, 82, 373–428.
Pickart, C. M., & Cohen, R. E. (2004). Proteasomes and their kins: Proteases in the machine age. Nature Reviews. Molecular Cell Biology, 5, 177–187.
Groll, M., Bochtler, M., Brandstetter, H., Clausen, T., & Huber, R. (2005). Molecular machines for protein degradation. Chembiochem, 6, 222–256.
Nickell, S., Beck, F., Scheres, S. H., Korinek, A., Förster, F., Lasker, K., et al. (2009). Insights into the molecular architecture of the 26S proteasome. Proceedings of the National Academy of Sciences of the United States of America, 106, 11943–11947.
Murata, S., Yashiroda, H., & Tanaka, K. (2009). Molecular mechanisms of proteasome assembly. Nature Reviews. Molecular Cell Biology, 10, 104–115.
Marques, A. J., Palanimurugan, R., Matias, A. C., Ramos, P. C., & Dohmen, R. J. (2009). Catalytic mechanism and assembly of the proteasome. Chemical Reviews, 109, 1509–1536.
Boes, B., Hengel, H., Ruppert, T., Multhaup, G., Koszinowski, U. H., & Kloetzel, P. M. (1994). Interferon stimulation modulates the proteolytic activity and cleavage site preference of 20S mouse proteasomes. The Journal of Experimental Medicine, 179, 901–909.
Gaczynska, M., Rock, K. L., Spies, T., & Goldberg, A. L. (1994). Peptidase activities of proteasomes are differentially regulated by the major histocompatibility complex-encoded genes for LMP2 and LMP7. Proceedings of the National Academy of Sciences of the United States of America, 91, 9213–9217.
Cardozo, C., & Kohanski, R. A. (1998). Altered properties of the branched chain amino acid-preferring activity contribute to increased cleavages after branched chain residues by the “immunoproteasome”. The Journal of Biological Chemistry, 273, 16764–16770.
Murata, S., Sasaki, K., Kishimoto, T., Niwa, S., Hayashi, H., Takahama, Y., et al. (2007). Regulation of CD8+ T cell development by thymus-specific proteasomes. Science, 316, 1349–1353.
Rabl, J., Smith, D. M., Yu, Y., Chang, S.-C., Goldberg, A. L., & Cheng, Y. (2008). Mechanism of gate opening in the 20S proteasome by the proteasomal ATPases. Molecular Cell, 30, 360–368.
Smith, D. M., Chang, S. C., Park, S., Finley, D., Cheng, Y., & Goldberg, A. L. (2007). Docking of the proteasomal ATPases’ carboxyl termini in the 20S proteasome’s alpha ring opens the gate for substrate entry. Molecular Cell, 27, 731–744.
Tomko, R. J., Jr., Funakoshi, M., Schneider, K., Wang, J., & Hochstrasser, M. (2010). Heterohexameric ring arrangement of the eukaryotic proteasomal ATPases: Implications for proteasome structure and assembly. Molecular Cell, 38, 393–403.
Xie, Y., & Varshavsky, A. (2000). Physical association of ubiquitin ligases and the 26S proteasome. Proceedings of the National Academy of Sciences of the United States of America, 97, 2497–2502.
Xie, Y., & Varshavsky, A. (2002). UFD4 lacking the proteasome-binding region catalyses ubiquitination but is impaired in proteolysis. Nature Cell Biology, 4, 1003–1007.
Crosas, B., Hanna, J., Kirkpatrick, D.S., Zhang, D.P., Tone, Y., Hathaway, N.A., et al. (2006). Ubiquitin chains are remodeled at the proteasome by opposing ubiquitin ligase and deubiquitinating activities. Cell, 127, 1401–1413.
You, J., & Pickart, C. M. (2001). A HECT domain E3 enzyme assembles novel polyubiquitin chains. The Journal of Biological Chemistry, 276, 19871–19878.
Russell, S. J., Steger, K. A., & Johnston, S. A. (1999). Subcellular localization, stoichiometry, and protein levels of the 26S proteasome subunits in yeast. The Journal of Biological Chemistry, 274, 21943–21952.
Glickman, M. H., Rubin, D. M., Fried, V. A., & Finley, D. (1998). The regulatory particle of the Saccharomyces cerevisiae proteasome. Molecular and Cellular Biology, 18, 3149–3162.
Mannhaupt, G., Schnall, R., Karpov, V., Vetter, I., & Feldmann, H. (1999). Rpn4p acts as a transcription factor by binding to PACE, a nonamer box found upstream of 26S proteasomal and other genes in yeast. FEBS Letters, 450, 27–34.
Xie, Y., & Varshavsky, A. (2001). RPN4 is a ligand, substrate, and transcriptional regulator of the 26S proteasome: A negative feedback circuit. Proceedings of the National Academy of Sciences of the United States of America, 98, 3056–3061.
Ju, D., Wang, L., Mao, X., & Xie, Y. (2004). Homeostatic regulation of the proteasome via an Rpn4-dependent feedback circuit. Biochemical and Biophysical Research Communications, 321, 51–57.
Wang, X., Xu, H., Ju, D., & Xie, Y. (2008). Disruption of Rpn4-induced proteasome expression in Saccharomyces cerevisiae reduces cell viability under stressed conditions. Genetics, 180, 1945–1953.
London, M., Keck, B. I., Ramos, P. C., & Dohmen, R. J. (2004). Regulatory mechanisms controlling biogenesis of ubiquitin and the proteasome. FEBS Letters, 567, 259–264.
Jelinsky, S. A., Estep, P., Church, G. M., & Samson, L. D. (2000). Regulatory networks revealed by transcriptional profiling of damaged Saccharomyces cerevisiae cells: Rpn4 links base excision repair with proteasomes. Molecular and Cellular Biology, 20, 8157–8167.
Gasch, A. P., Huang, M., Metzner, S., Botstein, D., Elledge, S. J., & Brown, P. O. (2001). Genomic expression response to DNA-damaging agents and the regulatory role of the yeast ATR homolog Mec1p. Molecular Biology of the Cell, 12, 2987–3003.
Owsianik, G., Balzi, E., & Ghislain, M. (2002). Control of 26S proteasome expression by transcription factors regulating multidrug resistance in Saccharomyces cerevisiae. Molecular Microbiology, 43, 1295–1308.
Hahn, J.-S., Neef, D. W., & Thiele, D. J. (2006). A stress regulatory network for co-ordinated activation of proteasome expression mediated by yeast heat shock transcription factor. Molecular Microbiology, 60, 240–251.
Haugen, A. C., Kelley, R., Collins, J. B., Tucker, C. J., Deng, C., Afshari, C. A., et al. (2004). Integrating phenotypic and expression profiles to map arsenic-response networks. Genome Biology, 5, R95.
Guo, N., Yu, L., Meng, R., Fan, J., Wang, D., Sun, G., et al. (2008). Global gene expression profile of Saccharomyces cerevisiae induced by dictamnine. Yeast, 25, 631–641.
Salin, H., Fardeau, V., Piccini, E., Lelandais, G., Tanty, V., Lemoine, S., et al. (2008). Structure and properties of transcriptional networks driving selenite stress response in yeasts. BMC Genomics, 9, 333.
Mizukami-Murata, S., Iwahashi, H., Kimura, S., Nojima, K., Sakurai, Y., Saitou, T., et al. (2010). Genome-wide expression changes in Saccharomyces cerevisiae in response to high-LET ionizing radiation. Applied Biochemistry and Biotechnology, 162, 855–870.
Ng, D. T. W., Spear, E. D., & Walter, P. (2000). The unfolded protein response regulates multiple aspects of secretory and membrane protein biogenesis and endoplasmic reticulum quality control. The Journal of Cell Biology, 150, 77–88.
Cai, H., Kauffman, S., Naider, F., & Becker, J. M. (2006). Genomewide screen reveals a wide regulatory network for di/tripeptide utilization in Saccharomyces cerevisiae. Genetics, 172, 1459–1476.
Metzger, M. B., & Michaelis, S. (2009). Analysis of quality control substrates in distinct cellular compartments reveals a unique role for Rpn4p in tolerating misfolded membrane proteins. Molecular Biology of the Cell, 20, 1006–1019.
Hausmann, S., Zheng, S., Costanzo, M., Brost, R. L., Garcin, D., Boone, C., et al. (2008). Genetic and biochemical analysis of yeast and human cap trimethylguanosine synthase: Functional overlap of 2, 2, 7-trimethylguanosine caps, small nuclear ribonucleoprotein components, pre-mRNA splicing factors, and RNA decay pathways. The Journal of Biological Chemistry, 283, 31706–31718.
Teixeira, M. C., Dias, P. J., Simões, T., & Sá-Correia, I. (2008). Yeast adaptation to mancozeb involves the up-regulation of FLR1 under the coordinate control of Yap1, Rpn4, Pdr3, and Yrr1. Biochemical and Biophysical Research Communications, 367, 249–255.
Bosis, E., Salomon, D., Ohayon, O., Sivan, G., Bar-Nun, S., & Rabinovich, E. (2010). Ssz1 restores endoplasmic reticulum-associated protein degradation in cells expressing defective cdc48-ufd1-npl4 complex by upregulating cdc48. Genetics, 184, 695–706.
Wang, X., Xu, H., Ha, S.-W., Ju, D., & Xie, Y. (2010). Proteasomal degradation of Rpn4 in Saccharomyces cerevisiae is critical for cell viability under stressed conditions. Genetics, 184, 335–342.
Ju, D., Wang, X., Ha, S.-W., Fu, J., & Xie, Y. (2010). Inhibition of proteasomal degradation of Rpn4 impairs nonhomologous end-joining repair of DNA double-strand breaks. PLoS ONE, 5, e9877.
Mannhaupt, G., & Feldmann, H. (2007). Genomic evolution of the proteasome system among hemiascomycetous yeasts. Journal of Molecular Evolution, 65, 529–540.
Gasch, A. P., Moses, A. M., Chiang, D. Y., Fraser, H. B., Berardini, M., & Eisen, M. B. (2004). Conservation and evolution of cis-regulatory systems in Ascomycete fungi. PLoS Biology, 2, e398.
Meiners, S., Heyken, D., Weller, A., Ludwig, A., Stangl, K., Kloetzel, P.-M., et al. (2003). Inhibition of proteasome activity induces concerted expression of proteasome genes and de novo formation of Mammalian proteasomes. The Journal of Biological Chemistry, 278, 21517–21525.
Fleming, J. A., Lightcap, E. S., Sadis, S., Thoroddsen, V., Bulawa, C. E., & Blackman, R. K. (2002). Complementary whole-genome technologies reveal the cellular response to proteasome inhibition by PS-341. Proceedings of the National Academy of Sciences of the United States of America, 99, 1461–1466.
Lundgren, J., Masson, P., Realini, C. A., & Young, P. (2003). Use of RNA interference and complementation to study the function of the Drosophila and human 26S proteasome subunit S13. Molecular and Cellular Biology, 23, 5320–5330.
Wójcik, C., & DeMartino, G. N. (2002). RNA interference of valosin-containing protein (VCP/p97) reveals multiple cellular roles linked to ubiquitin/proteasome-dependent proteolysis. The Journal of Biological Chemistry, 277, 6188–6197.
Xu, H., Ju, D., Jarois, T., & Xie, Y. (2008). Diminished feedback regulation of proteasome expression and resistance to proteasome inhibitors in breast cancer cells. Breast Cancer Research and Treatment, 107, 267–274.
Sato, Y., Sakamoto, K., Sei, M., Ewis, A. A., & Nakahori, Y. (2009). Proteasome subunits are regulated and expressed in comparable concentrations as a functional cluster. Biochemical and Biophysical Research Communications, 378, 795–798.
Kraft, D. C., Deocaris, C. C., Wadhwa, R., & Rattan, S. I. S. (2006). Preincubation with the proteasome inhibitor MG-132 enhances proteasome activity via the Nrf2 transcription factor in aging human skin fibroblasts. Annals of the New York Academy of Sciences, 1067, 420–424.
Lee, C.-S., Tee, L. Y., Warmke, T., Vinjamoori, A., Cai, A., Fagan, A. M., et al. (2004). A proteasomal stress response: Pre-treatment with proteasome inhibitors increases proteasome activity and reduces neuronal vulnerability to oxidative injury. Journal of Neurochemistry, 91, 966–1006.
Radhakrishnan, S. K., Lee, C. S., Young, P., Beskow, A., Chan, J. Y., & Deshaies, R. (2010). Transcription factor Nrf1 mediates the proteasome recovery pathway after proteasome inhibition in mammalian cells. Molecular Cell, 38, 17–28.
Kwak, M. K., Wakabayashi, N., Greenlaw, J. L., Yamamoto, M., & Kensler, T. W. (2003). Antioxidants enhance mammalian proteasome expression through the Keap1-Nrf2 signaling pathway. Molecular and Cellular Biology, 23, 8786–8794.
Kwak, M. K., & Kensler, T. W. (2006). Induction of 26S proteasome subunit PSMB5 by the bifunctional inducer 3-methycholanthrene through the Nrf2-ARE, but not the AhR/Arnt-XRE, pathway. Biochemical and Biophysical Research Communications, 345, 1350–1357.
Arlt, A., Bauer, I., Schafmayer, C., Tepel, J., Müerköster, S. S., Brosch, M., et al. (2009). Increased proteasome subunit protein expression and proteasome activity in colon cancer relate to an enhanced activation of nuclear factor E2-related factor 2 (Nrf2). Oncogene, 28, 3983–3996.
Kapeta, S., Chondrogianni, N., & Gonos, E. S. (2010). Nuclear erythroid factor 2 (Nrf2) mediated proteasome activation delays senescence in human fibroblasts. The Journal of Biological Chemistry, 285, 8171–8184.
Moi, P., Chan, K., Asunis, I., Cao, A., & Kan, Y. W. (1994). Isolation of NF-E2-related factor 2 (Nrf2), a NF-E2-like basic leucine zipper transcriptional activator that binds to the tandem NF-E2/AP1 repeat of the beta-globin locus control region. Proceedings of the National Academy of Sciences of the United States of America, 91, 9926–9930.
Kensler, T. W., Wakabayashi, N., & Biswal, S. (2007). Cell survival responses to environmental stresses via the Keap1-Nrf2-ARE pathway. Annual Review of Pharmacology and Toxicology, 47, 89–116.
Nguyen, T., Nioi, P., & Pickett, C. B. (2009). The Nrf2-antioxidant response element signaling pathway and its activation by oxidative stress. The Journal of Biological Chemistry, 284, 13291–13295.
Orlowski, R. Z., & Kuhn, D. J. (2008). Proteasome inhibitors in cancer therapy: Lessons from the first decade. Clinical Cancer Research, 14, 1649–1657.
Schwartz, A. L., & Ciechanover, A. (1999). The ubiquitin-proteasome pathway and pathogenesis of human diseases. Annual Review of Medicine, 50, 57–74.
Kumatori, A., Tanaka, K., Inamura, N., Sone, S., Ogura, T., Matsumoto, T., et al. (1990). Abnormally high expression of proteasomes in human leukemic cells. Proceedings of the National Academy of Sciences of the United States of America, 87, 7071–7075.
Chen, L., & Madura, K. (2005). Increased proteasome activity, ubiquitin-conjugating enzymes, and eEF1A translation factor detected in breast cancer tissue. Cancer Research, 65, 5599–5606.
Bazzaro, M., Lee, M. K., Zoso, A., Stirling, W. L., Santillan, A., Shih, Ie M, et al. (2006). Ubiquitin-proteasome system stress sensitizes ovarian cancer to proteasome inhibitor-induced apoptosis. Cancer Research, 66, 3754–3763.
Pilarsky, C., Wenzig, M., Specht, T., Saeger, H. D., & Grützmann, R. (2004). Identification and validation of commonly overexpressed genes in solid tumors by comparison of microarray data. Neoplasia, 6, 744–750.
Milano, A., Iaffaioli, R. V., & Caponigro, F. (2007). The proteasome: A worthwhile target for the treatment of solid tumours? European Journal of Cancer, 43, 1125–1133.
Papandreou, C. N., Daliani, D. D., Nix, D., Yang, H., Madden, T., Wang, X., et al. (2004). Phase I trial of the proteasome inhibitor bortezomib in patients with advanced solid tumors with observations in androgen-independent prostate cancer. Journal of Clinical Oncology, 22, 2108–2121.
Schwartz, R., & Davidson, T. (2004). Pharmacology, pharmacokinetics, and practical applications of bortezomib. Oncology (Williston Park), 18(14 Suppl 11), 14–21.
Chen, S., Blank, J. L., Peters, T., Liu, X. J., Rappoli, D. M., Pickard, M. D., et al. (2010). Genome-wide siRNA screen for modulators of cell death induced by proteasome inhibitor bortezomib. Cancer Research, 70, 4318–4326.
Pan, X., Ye, P., Yuan, D. S., Wang, X., Bader, J. S., & Boeke, J. D. (2006). A DNA integrity network in the yeast Saccharomyces cerevisiae. Cell, 124, 1069–1081.
Tong, A. H., Lesage, G., Bader, G. D., Ding, H., Xu, H., Xin, X., et al. (2004). Global mapping of the yeast genetic interaction network. Science, 303, 808–813.
Ju, D., Wang, X., & Xie, Y. (2009). Dyclonine and alverine citrate enhance the cytotoxic effects of proteasome inhibitor MG132 on breast cancer cells. International Journal of Molecular Medicine, 23, 205–209.
Lees, N. D., Skaggs, B., Kirsch, D. R., & Bard, M. (1995). Cloning of the late genes in the ergosterol biosynthetic pathway of Saccharomyces cerevisiae—a review. Lipids, 30, 221–226.
Bennati, A. M., Castelli, M., Fazia, M. A. D., Beccari, T., Caruso, D., Servillo, G., et al. (2006). Sterol dependent regulation of human TM7SF2 gene expression: Role of the encoded 3β-hydroxysterol Δ14-reductase in human cholesterol biosynthesis. Biochimica et Biophysica Acta, 1761, 677–685.
Holmer, L., Pezhman, A., & Worman, H. J. (1998). The human lamin B receptor/sterol reductase multigene family. Genomics, 54, 469–476.
Giaever, G., Flaherty, P., Kumm, J., Proctor, M., Nislow, C., Jaramillo, D. F., et al. (2004). Chemogenomic profiling: Identifying the functional interactions of small molecules in yeast. Proceedings of the National Academy of Sciences of the United States of America, 101, 793–798.
Powell, S. R., Wang, P., Katzeff, H., Shringarpure, R., Teoh, C., Khaliulin, I., et al. (2005). Oxidized and ubiquitinated proteins may predict recovery of postischemic cardiac function: Essential role of the proteasome. Antioxidants Redox Signaling, 7, 538–546.
Zong, C., Gomes, A. V., Drews, O., Li, X., Young, G. W., Berhane, B., et al. (2006). Regulation of murine cardiac 20S proteasomes: Role of associating partners. Circulation Research, 99, 372–380.
Zhang, F., Su, K., Yang, X., Bowe, D. B., Paterson, A. J., & Kudlow, J. E. (2003). O-GlcNAc modification is an endogenous inhibitor of the proteasome. Cell, 115, 715–725.
Kimura, Y., Saeki, Y., Yokosawa, H., Polevoda, B., Sherman, F., & Hirano, H. (2003). N-Terminal modifications of the 19S regulatory particle subunits of the yeast proteasome. Archives of Biochemistry and Biophysics, 409, 341–348.
Acknowledgments
I thank Kristen Gaffney for critically reading the manuscript. This work was supported by a National Science Foundation grant (MCB-0816974).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xie, Y. Feedback regulation of proteasome gene expression and its implications in cancer therapy. Cancer Metastasis Rev 29, 687–693 (2010). https://doi.org/10.1007/s10555-010-9255-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-010-9255-y