Abstract
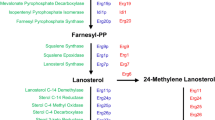
Research on the ergosterol biosynthetic pathway in fungi has focused on the identification of the specific sterol structure required for normal membrane structure and function and for completion of the cell cycle. The pathway and its end product are also the targets for a number of antifungal drugs. Identification of essential steps in ergo-sterol biosynthesis could provide new targets for the development of novel therapeutic agents. Nine of the eleven genes in the portion of the pathway committed exclusively to ergosterol biosynthesis have been cloned, and their essentiality for aerobic growth has been determined. The first three genes;ERG9 (squalene synthase),ERG1 (squalene epoxidase), andERG7 (lanosterol synthase), have been cloned and found to be essential for aerobic viability since their absence would result in the cell being unable to synthesize a sterol molecule. The remaining eight genes encode enzymes which metabolize the first sterol, lanosterol, to ultimately form ergosterol. The two earliest genes,ERG11 (lanosterol demethylase) andERG24 (C-14 reductase), have been cloned and found to be essential for aerobic growth but are suppressed by mutations in the C-5 desaturase (ERG3) gene andfen1 andfen2 mutations, respectively. The remaining cloned genes,ERG6 (C-24 methylase),ERG2 (D8Æ7 isomerase),ERG3 (C-5 desaturase), andERG4 (C-24(28) reductase), have been found to be nonessential. The remaining genes not yet cloned are the C-4 demethylase and the C-22 desaturase (ERG5).
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- PCR:
-
polymerase chain reaction
- SBI:
-
sterol biosynthesis inhibitor
References
Goldstein, J.L., and Brown, M.S. (1990)Nature 243, 425–430.
Lees, N.D., Bard, M., Kemple, M.D., Haak, R.A., and Woods, R.A. (1979)Biochim. Biophys. Acta 553, 469–475.
Lees, N.D., Kleinhans, F.W., Broughton, M.C., Pennington, P.A., Picker, V.A., and Bard, M. (1989)Steroids 53, 567–578.
Bard, M., Lees, N.D., Burrows, L.S., and Kleinhans, F.W. (1978)J. Bacteriol. 135, 1146–1148.
Kleinhans, F.W., Lees, N.D., Bard, M., Haak, R.A., and Woods, R.A. (1979)Chem. Phys. Lipids 23, 143–154.
Rottem, S., Yaskoav, J., Neeman, Z., and Razin, S. (1973)Biochim. Biophys. Acta 323, 495–508.
Lees, N.D., Lofton, S.L., Woods, R.A., and Bard, M. (1980)J. Gen. Microbiol. 118, 209–214.
Dahl, C., Biemann, H.P., and Dahl, J. (1987)Proc. Natl. Acad. Sci. USA 84, 4012–4016.
Quesney-Huneeus, V., Wiley, M.H., and Siperstein, M.D. (1979)Proc. Natl. Acad. Sci. USA 76, 5056–5060.
Quesney-Huneeus, V., Galick, H.A., Siperstein, M.D., Erikson, S.K., Spencer, J.A., and Nelson, J.A. (1983)J. Biol. Chem. 258, 378–385.
Georgopapadakou, N.H., and Walsh, T.J. (1994)Science 264, 371–373.
Travis, J. (1994)Science 264, 360–362.
Davies, J. (1994)Science 264, 375–382.
Klein, R.S., Harris, C.A., Small, C.B., Moll, B., Lesser, M., and Friedland, G.H. (1994)New Engl. J. Med. 311, 554–556.
Beyer, J., Schwartz, S., Heinemann, V., and Siegert, W. (1994)Antimicrob. Agents Chemother. 38, 911–917.
Jolidon, S., Polack, A.M., Guerry, P., and Hartman, P.G. (1989)Biochem. Soc. Trans. 18, 47–49.
Marcireau, C., Guilloton, M., and Karst, F. (1990)Antimicrob. Agents Chemother. 34, 989–993.
Hartman, G., and Polak, A. (1993)Clin. Dermatol. 7, 27–36.
Vanden Bossche, H., Willemsens, G., Cools, W., Cornelissen, F., Lauwers, W.F., and Van Cutsem, J. (1980)Antimicrob. Agents Chemother. 17, 922–928.
Lees, N.D., Broughton, M.C., Sanglard, P., and Bard, M. (1990)Antimicrob. Agents Chemother. 34, 831–836.
Kerridge, D. (1975)Postgrad. Med. J. 55, 653–656.
McCasker, J.H., Clemons, K.V., Stevens, D.A., and Davis, R.W. (1994)Genetics 136, 1261–1269.
Fegueur, M., Richard, L., Charles, N.D., and Karst, F. (1991)Curr. Genet. 20, 365–372.
Jennings, S.M., Tsay, Y.H., Fisch, T.M., and Robinson, G.W. (1991)Proc. Natl. Acad. Sci. USA 88, 6038–6042.
Robinson, G.W., Tsay, Y.H., Kienzle, B.K., Smith-Monroy, C.A., and Bishop, R.W. (1993)Mol. Cell Biol. 13, 2706–2717.
Jandrositz, A., Turnowsky, F., and Hogenauer, G. (1991)Gene 107, 155–160.
Kelly, R., Miller, S.M., Lai, M.H., and Kirsch, D.R. (1990)Gene 87, 177–183.
Covey, E.J., Matsuda, S.P.T., and Bartel, B. (1994)Proc. Natl. Acad. Sci. USA 91, 2211–2215.
Covey, E.S., Matsuda, S.P.T., and Bartel, B. (1993)Proc. Natl. Acad. Sci. USA 90, 11628–11632.
Trocha, P.J., Jasne, S.J., and Sprinson, D.B. (1977)Biochemistry 16, 4721–4726.
Taylor, F.R., Rodriquez, F.J., and Parks, L.W. (1983)J. Bacteriol. 156, 64–68.
Kalb, V.F., Loper, J.C., Dey, C.R., Woods, C.W., and Sutter, T.R. (1986)Gene 45, 237–245.
Kalb, V.F., Woods, C.W., Turi, T.G., Dey, C.R., Sutter, T.R., and Loper, J.C. (1987)DNA 6, 529–537.
Bard, M., Lees, N.D., Turi, T., Craft, D., Cofrin, L., Barbuch, R., Koegel, C., and Loper, J.C. (1993)Lipids 28, 963–967.
Watson, P.F., Rose, M.E., Ellis, S.W., England, H., and Kelly, S.L. (1989)Biochem. Biophys. Res. Commun. 164, 1170–1175.
Shimokawa, O., Kato, Y., and Nakayama, H. (1986)J. Med. Vet. Mycol. 24, 327–336.
Shimokawa, O., Kato, Y., Kawano, K., and Nakayama, H. (1989)Biochim. Biophys. Acta 1003, 15–19.
Marcireau, C., Guyonnet, D., and Karst, F. (1992)Curr. Genet. 22, 267–272.
Lorenz, R.T., and Parks, L.W. (1992)DNA Cell Biol. 11, 685–692.
Lai, M.H., Bard, M., Pierson, C.A., Alexander, J.F., Goebl, M., Carter, G.T., and Kirsch, D.R. (1994)Gene 140, 41–49.
Shimanuki, M., Goebl, M., Yanagida, M., and Toda, T. (1992)Mol. Biol. Cell 3, 263–273.
Chen, W., Capieaux, E., Balzi, E., and Goffeau, A. (1991)Yeast 7, 305–308.
Pinto, W.J., and Nes, W.R. (1983)J. Biol. Chem. 258, 4472–4476.
Pinto, W.J., Lozano, R., Sekula, B.C., and Nes, W.R. (1983)Biochem. Biophys. Res. Commun. 112, 47–54.
Gaber, R.F., Copple, D.M., Kennedy, B.K., Vidal, M., and Bard, M. (1989)Mol. Cell Biol. 9, 3447–3456.
Hardwick, K.C., and Pelham, H.R.B. (1994)Yeast 10, 265–269.
Baloch, R.I., Mercer, E.I., Wiggins, T.E., and Baldwin, B.C. (1984)Phytochemistry 23, 2219–2226.
Ashman, W.H., Barbuch, R.J., Ulbright, C.E., Jarrett, H.W., and Bard, M. (1991)Lipids 26, 628–632.
Arthington, B.A., Hoskins, J., Skatrud, P.L., and Bard, M. (1991)Gene 107, 173–174.
Rodriguez, R.J., Low, C., Bottema, C.D.K., and Parks, L.W. (1985)Biochim. Biophys. Acta 387, 336–343.
Rodriguez, R.J., and Parks, L.W. (1983)Arch. Biochem. Biophys. 225, 861–871.
Lorenz, R.T., Casey, W.M., and Parks, L.W. (1989)J. Bacteriol. 171, 6169–6173.
Arthington, B.A., Bennett, L.G., Skatrud, P.L., Guyra, C.J., Barbuch, R.J., Ulbright, C.E., and Bard, M. (1991)Gene 102, 39–44.
Worman, H.J., Evans, C.D., and Blobel, G. (1990)J. Cell Biol. 111, 1535–1542.
Fukishima, H., Grimstead, G.F., and Gaylor, J.F. (1981)J. Biol. Chem. 256, 4822–4826.
Faust, J.R., Trzaskos, J.M., and Gaylor, J.F. (1988) inBiology of Cholesterol (Yeagle, P.L., ed.), CRC Press, Boca Raton.
Paltauf, F., Kohlwein, S.D., and Henry, S.A. (1992) inThe Molecular and Cellular Biology of the Yeast Saccharomyces (Jones, E.W., Pringle, J.R., and Broach, J.R., eds.) Vol. 2, Cold Spring Harbor Laboratory Press, New York.
Author information
Authors and Affiliations
About this article
Cite this article
Lees, N.D., Skaggs, B., Kirsch, D.R. et al. Cloning of the late genes in the ergosterol biosynthetic pathway ofSaccharomyces cerevisiae—A review. Lipids 30, 221–226 (1995). https://doi.org/10.1007/BF02537824
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02537824