Abstract
Purpose
Endocrine therapy is the standard treatment for oestrogen receptor positive (ER+) breast cancer. Despite its efficacy, around half of patients will develop resistance to this treatment and eventually relapse. Identification of effective and reliable biomarkers to predict the efficacy of endocrine therapy is of crucial importance in the management of ER+ breast cancer. Emerging evidence has revealed that the cell division regulator CDC20 exhibits an oncogenic function and plays important roles in tumourigenesis and progression of solid tumours. In this study, we investigated the prognostic and predictive role of CDC20 in early ER+ breast cancer patients.
Methods
The biological and clinical impact of CDC20 expression was assessed in large clinical annotated cohort of ER+ breast cancer with long-term follow-up at the mRNA level, using METABRIC and KM-Plotter datasets, and the protein level using immunohistochemistry on patients presenting at Nottingham. CDC20 expression was correlated with clinico-pathological parameters, molecular subtypes, clinical outcome and efficacy of endocrine therapy.
Results
High CDC20 mRNA expression was associated with poor clinico-pathological parameters including large tumour size and high tumour grade (P < 0.0001) in patients with ER+ breast cancer. High CDC20 mRNA expression was significantly associated with poor patient outcome (P < 0.0001). Importantly, high CDC20 expression was correlated with poor response to endocrine treatment in patients who treated with hormonal therapy only (P < 0.01). In multivariate analysis, CDC20 mRNA was an independent predictor of poor clinical outcome after treatment with endocrine therapy (P = 0.02).
Conclusion
CDC20 is a candidate biomarker for a subgroup of ER+ breast cancer characterised by poor clinical outcome. This study shows that the CDC20 could act as potential predictive biomarker of poor response to endocrine therapy in ER+ breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is a heterogeneous disease with various biological subtypes [1] with the most common form, approximately 75%, of breast cancer being oestrogen receptor positive (ER+) [2, 3]. Endocrine therapy is the main treatment for ER+ tumours, which has vastly improved survival and has reduced mortality [4]. However, a high proportion of patients receiving adjuvant endocrine therapy still experience relapse and become resistant to treatment [5, 6]. It is therefore highly desirable to predict, at an early stage of treatment, which ER+ patients will and will not benefit from endocrine therapy.
Cell division cycle 20 homolog (CDC20) is a spindle assembly checkpoint molecule that required for the anaphase-promoting complex/cyclosome (APC/C) activation during mitosis, leading to initiation of chromatid separation and entrance of cell cycle into anaphase [7, 8]. Defects in CDC20 function may therefore terminate mitotic arrest, which lead to tumourgenesis [9, 10]. Consistent with the notion that CDC20 may function as an oncogene, several studies show overexpression of CDC20 in different types of cancers [9, 11,12,13]. Indeed, its overexpression is suggested as a biomarker of poor outcome in pancreatic [14], colon [15], primary non-small cell lung [16] and ovarian cancer [11].
In terms of breast cancer, two reports have demonstrated that CDC20 is a potential key player in the progression of breast cancer where it is significantly higher in breast cancer cells and high-grade primary tumour tissues [17] and indicates an aggressive course of disease risk [18]. We aimed to investigate the role of CDC20 expression in ER+ tumours. In particular, we assessed whether CDC20 had a role in endocrine resistance which could be used to improve therapy prediction in ER+ breast cancer.
Materials and method
CDC20 mRNA expression
The Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) [19], comprising 1506 ER+ breast cancer, was used as a discovery cohort to analyse and explore the prognostic value of CDC20 mRNA and its role as predictive biomarker of clinical outcome for patients who treated with endocrine therapy, Table 1.
The Kaplan–Meier Plotter-Breast Cancer (KM-Plotter) online dataset [20], was used as a validation cohort for the prognostic and predictive value of CDC20 mRNA expression using 2061 patients with ER+ breast cancer. The prognostic value of CDC20 mRNA expression and association with clinical outcome and clinico-pathological parameters were further validated using the Breast Cancer Gene-Expression Miner v4.0 (bc-GenExMiner v4.0) database [21] which includes 5829 cases of ER+ breast cancer.
CDC20 protein expression
CDC20 protein expression was assessed in a series cohort of clinical samples for patients with ER+ (n = 347) using immunohistochemistry (IHC). Patients presented at Nottingham City Hospital between 1989 and 2006. Patient management was uniform and based on tumour characteristics by Nottingham Prognostic Index (NPI) and hormone receptor status. No adjuvant therapy was given to patients with good prognostic NPI score (≤ 3.4), while for patients with poor NPI scores (> 3.4) endocrine therapy was given. Premenopausal patients within the moderate and poor prognostic NPI were given chemotherapy, whereas postmenopausal patients with moderate or poor NPI were candidate for hormonal therapy. None of the patients in this study received neoadjuvant therapy. Clinical history, information on therapy and outcomes and tumour characteristics are prospectively maintained. The clinico-pathological parameters for the cohort series are summarised in Table 1.
Western blotting
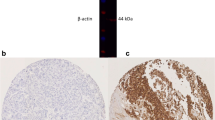
Immunoblotting was performed as previously described [22], the specificity of CDC20 was validated on BT474 human breast cancer cells (American Type Culture Collection; Rockville, MD, USA) using primary CDC20 antibody (HPA039484, Sigma-Aldrich, 1:2000). This showed a specific band for CDC20 protein at the predicted size of 55 kDa (Fig. 1a).
IHC staining and evaluation
Tumour samples were arrayed as previously described [23]. The IHC staining was performed on 4 μm tissue microarrays (TMAs) sections using Novolink polymer detection system (Leica Biosystems, RE7150-K), detailed method was described in previous publication [22]. Sections were incubated, overnight at 4 °C, with the primary CDC20 antibody diluted at 1:500. CDC20 immunoreactivity was assessed using high-resolution digital images (Nanozoomer, Hamamatsu Photonics) and viewing software (Xplore; Philips, UK). Evaluation was based on a semiquantitative assessment using a modified histochemical score (H-score), which includes an assessment of both the intensity and the percentage of stained cells [24]. The staining intensity of invasive tumour cells was scored into four categories 0 (no staining); 1 (weak staining); 2 (moderate staining) and 3 (strong staining). The percentage of each category was estimated, and the H-score calculated. TMA cores were only assessed if tumour burden was > 15%.
Clinical outcome data and events definition
Clinical outcomes including breast cancer specific survival (BCSS) was defined as the time in months from the diagnosis to the date of breast cancer-related death. Recurrence free survival (RFS) was defined as the time in months from diagnosis until developing local or regional recurrence. Distant-metastasis free survival (DMFS) was defined as the time in months from diagnosis until developing distant-metastasis. For the benefit of endocrine therapy, the expression of CDC20 was investigated with clinical outcome on the endocrine-treated cohort only. Secondary outcomes included associations with clinico-pathological parameters.
Statistical analysis
Data analysis was performed using SPSS statistical software (version 25, Chicago, IL, USA). The analysis for this study compared low and high expression of CDC20. The Chi square test was performed for inter-relationships between categorical variables. Spearman’s correlation coefficient was used to examine the association between continuous variables. One-way analysis of variance (ANOVA) with the post hoc Tukey was used for differences between three or more groups. Kaplan–Meier analysis was used to assess the association of CDC20 expression with clinical outcome. Multivariate Cox Regression analysis with adjustment of covariates was used to identify independent prognostic biomarkers. Benjamini–Hochberg procedure for multiple test correction was performed. P value of ≤ 0.05 was considered significant. The dichotomisation of CDC20 mRNA and protein expression into low and high groups was determined using X-Tile (X-Tile Bioinformatics Software, Yale University, version 3.6.1).
Results
CDC20 expression in ER+ BC
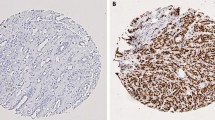
High CDC20 mRNA expression in the METABRIC cohort was observed in 870 cases (58%), where low expression was observed in 636 cases (42%). CDC20 protein expression was localised to the cytoplasm of invasive tumour cells, with expression levels varying from absent to high (H-score range 0–250) (Fig. 1b, c). CDC20 expression was dichotomised into low and high using an H-score of 120 resulting in 85 (25%) cases showing high expression and 262 (75%) cases with low expression.
Association of CDC20 expression with clinic-pathological characteristics in ER+ breast cancer
CDC20 mRNA expression was associated with aggressive clinico-pathological parameters including negative expression of PR, poor Nottingham Prognostic Index (NPI) and high tumour grade (P < 0.0001, Fig. 2a–c) using the METABRIC dataset. The association with PR and NPI was validated using the bc-GenExMiner v4.0 dataset (P < 0.0001; Supplementary Fig. 1A and B). In contrast, CDC20 protein expression showed no statistical significance association with any of the clinico-pathological parameters.
The METABRIC dataset was used to investigate the correlation between CDC20 mRNA and proliferation related-genes. There was positive correlation of CDC20 mRNA with the expression of MKI67, CCNB1, CCNA2 and CCND1 (p < 0.0001), Table 2. These findings were validated using bc-GenExMiner v4.0 dataset (P < 0.0001; Fig. 2d–g).
Clinical significance of CDC20 in patients with ER+ BC
High mRNA CDC20 expression was significantly correlated with poor clinical outcome. Thus, results of METABRIC dataset showed that high expression of CDC20 mRNA was associated with poor RFS (P < 0.0001), DMFS (P < 0.0001) and high risk of death from breast cancer (P < 0.0001) (Fig. 3a–c). To further validate our findings we used the KM-Plotter dataset which showed that high mRNA expression of CDC20 was also associated with poor clinical outcomes including RFS (P < 0.0001), DMFS (P < 0.0001) and BCSS (P < 0.0001) (Supplementary Fig. 2A–C). Furthermore, results from the bc-GenExMiner v4.0 datasets showed that cases with low expression of CDC20 mRNA had favourable clinical outcomes compared to the high expression group, which showed poor clinical outcome (P < 0.0001; Supplementary Fig. 2D).
CDC20 is a predictive biomarker of poor response to endocrine therapy
In patients who received endocrine therapy, tumours with high CDC20 mRNA expression were significantly associated with adverse clinical outcome, of which high risk of recurrence (P = 0.004; Fig. 4a) and distant-metastasis (P = 0.001; Fig. 4b) compared to patients with low CDC20 expression. In term of BCSS, high CDC20 mRNA expression was associated with worse survival and higher risk of death from breast cancer in patients who were treated with endocrine therapy (P < 0.0001, Fig. 4c) compared with CDC20 low expression. These observations were validated in the KM-Plotter dataset where patients who received endocrine therapy with high CDC20 mRNA showed poor benefit form the hormone treatment: RFS (P < 0.0001), DMFS (P < 0.0001) and BCSS (P = 0.01) (Supplementary Fig. 3a–C) compared to the low CDC20 group who had prolonged survival and lower risk of relapse and death from breast cancer.
CDC20 mRNA expression as a predictive biomarker for poor clinical outcome in patients with ER+ breast cancer after endocrine treatment using METABRIC dataset a recurrence, b distant-metastasis and c BCSS. Kaplan–Meier survival plots for patients with ER+ breast cancer after endocrine treatment and only 5 years follow-up d recurrence and e distant-metastasis
The relationship between CDC20 mRNA expression and risk of relapse after receiving 5-year adjuvant endocrine therapy
In patients with five years of follow-up after endocrine treatment, results showed that high CDC20 mRNA expression was associated with high risk of recurrence (P = 0.008, Fig. 4d) and distant-metastasis (P = 0.001, Fig. 4e). The significance of high CDC20 mRNA expression on predicting high risk of recurrence and distant-metastasis on patients who were treated with endocrine therapy alone was validated using KM-Plotter datasets (P < 0.0001; Supplementary Fig. 3D and E).
However, CDC20 protein expression showed no prognostic association with clinical outcome on the whole cohort of patients with ER+ breast cancer, or the impact of endocrine therapy on patient survival, recurrence or distant-metastasis (P > 0.05; Fig. 5a–f).
CDC20 protein and patient outcome in ER+ breast cancer using Nottingham cohort: a recurrence, b distant-metastasis and c BCSS. CDC20 protein expression and clinical outcome in patients with ER+ breast cancer who were treated with endocrine therapy only using Nottingham cohort d recurrence, e distant-metastasis and f BCSS
CDC20 is an independent prognostic marker in ER+ breast cancer
High CDC20 mRNA expression was independent of tumour size, nodal stage and tumour grade in predicting a higher risk of recurrence (P = 0.005), distant-metastasis (P = 0.005) and death from breast cancer (P = 0.0005), Table 3. In those patients treated with endocrine therapy only, CDC20 mRNA expression was an independent prognostic marker of tumour size, grade and nodal stage in predicting the risk of BCSS (P = 0.02), Table 4.
Discussion
Breast cancer is a heterogeneous disease with various biological subtypes [1] with the most common form being ER+/luminal tumours [2, 3]. This subtype remains heterogeneous in terms of recurrence, mortality rates, disease prognosis and response to treatment [3] despite attempts to biologically split them into luminal A and luminal B. Endocrine therapy, especially tamoxifen, still the main treatment for patients with ER+ breast cancer. Although sustained treatment with tamoxifen can successfully reduce postoperative recurrence and mortality rate, 30 to 50% of these patients will develop resistance and later relapse [25]. Therefore, there is still a need for a more precise method for stratifying patients based on their prognosis and response to endocrine therapy.
CDC20 has a key role in activating the APC/C to initiate anaphase and late mitosis exit in the cell cycle [8, 26]. CDC20 has also been shown to be a promising prognostic marker for a variety of tumours; including pancreatic [14, 27], colorectal [15], lung [16] and breast cancer [18]. Here, we focused on the role of CDC20 in ER+ breast cancer and especially in patients who were treated with only endocrine therapy. Our findings revealed that CDC20 is highly expressed in the more aggressive and highly proliferative ER+ tumours, and implicated in resistance to endocrine therapy. Indeed, our findings showed a significant association between high CDC20 mRNA expression and the poor prognostic clinico-pathological features within ER+ breast cancer. Despite our observations that CDC20 protein was not prognostic, it is likely that CDC20 expression plays a vital role in ER+ breast cancer progression.
Proliferation has a key role in the clinical behaviour of breast cancer and correlates strongly with poor clinical outcome and drug resistance. In addition to ER and PR, markers of proliferation seem to influence biological and clinical behaviour of ER+ breast cancer [19]. In light of this, our findings showed that high CDC20 mRNA expression was positively correlated with proliferation-associated genes, including MKI67, CCNB1, CCNA2 and CCND1. This supports the results of previous studies, which reported that knockdown of CDC20 decreased cell proliferation and induced G2/M cell cycle arrest in hepatocellular carcinoma cells [28], and pancreatic tumours [27]. Altogether, these data suggest that CDC20 is implicated in the proliferation of ER+ breast cancer which leads to tumourigenesis and aggressiveness phenotype.
Despite recent efforts to develop new breast cancer biomarkers, only ER and PR measurements are used currently both for clinical diagnosis to classify breast cancer patients and as a guide to endocrine therapy [29]. Multigene signatures, including Oncotype DX, Mammaprint, Prosigna, Breast Cancer Index and EndoPredict, can be valuable as additional prognostic tools with regard to recurrence and the stratification of risk, but so far studies have not validated their value in predicting benefit from endocrine therapy [30]. Therefore, the identification of predictive biomarkers for endocrine therapy efficacy in addition to ER and PR status is of urgent need to stratify patients with ER+ breast cancer for targeted therapy. For this purpose, a key aim of this study was to assess the predictive value of CDC20 mRNA and protein expression as a clinical marker of benefit from endocrine therapy in ER+ breast cancer. Our clinical data found a significant unfavourable effect of CDC20 mRNA expression in patients treated with endocrine therapy. These findings lead to suggestion that assessment of CDC20 mRNA expression prior to adjuvant treatment could predict patents who are highly to resist the endocrine therapy and eventually relapse.
Results from recent clinical trials demonstrated that 10 years of endocrine therapy showed improved RFS and overall survival compared with 5 years of endocrine treatment [31, 32]. However, this is at the cost of unnecessary side effects that influence the quality of life for patients [33]. Therefore, it is important to identify a subgroup of patients who are at high risk of relapse and who will not benefit from extended endocrine therapy. Our study demonstrates that for patients with ER+ breast cancer treated with endocrine therapy, high expression of CDC20 mRNA remains a predictive marker for high risk of relapse and death from breast cancer at 5 years follow-up. We suggest that assessment of CDC20 mRNA in clinical practice would be useful to predict patients who would not benefit from endocrine therapy and could spare them these risks and improve quality of life.
Although a previous study has showed the prognostic value of CDC20 protein expression in patients with triple negative breast cancer [18] we found no association with either clinico-pathological parameters or patient clinical outcome in ER+ breast cancer or those who treated with endocrine therapy alone. This is might be explained by using different methods to evaluate the IHC staining.
The level of agreement in our study between results of CDC20 mRNA and protein expression was poor. This discrepancy could be explained by several possibilities, including biological and technical explanations. Biological reasons include differences in post-transcriptional regulation of CDC20 expression or tumour-specific differences in CDC20 mRNA/protein stability, while technical issues may include nature of antibody used in this study [34,35,36]. Such discrepancies between mRNA and protein levels for different reasons have occurred in multiple studies of breast cancer [37, 38].
In summary, our data revealed a positive correlation of high CDC20 expression at the transcriptomic level with poor clinical outcome in patients with ER+ breast cancer. Also, we have provided evidence that CDC20 mRNA expression in ER+ breast cancer is a potentially predictive for selecting patients who might not experience benefits from endocrine therapy. Additional or alternative target therapies could then be given to those who predicted to be resistance to endocrine therapy; this would be a significant shift toward truly individualised medicine. We suggested that CDC20 mRNA expression could be used in clinical either singularly or in combination with other genes as multigene signature to guide the choice of endocrine treatment.
References
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, Zhu SX, Lonning PE, Borresen-Dale AL, Brown PO, Botstein D (2000) Molecular portraits of human breast tumours. Nature 406(6797):747–752. https://doi.org/10.1038/35021093
Rakha EA, El-Sayed ME, Green AR, Paish EC, Powe DG, Gee J, Nicholson RI, Lee AH, Robertson JF, Ellis IO (2007) Biologic and clinical characteristics of breast cancer with single hormone receptor positive phenotype. J Clin Oncol 25(30):4772–4778. https://doi.org/10.1200/jco.2007.12.2747
Dawson SJ, Rueda OM, Aparicio S, Caldas C (2013) A new genome-driven integrated classification of breast cancer and its implications. The EMBO J 32(5):617–628. https://doi.org/10.1038/emboj.2013.19
Musgrove EA, Sutherland RL (2009) Biological determinants of endocrine resistance in breast cancer. Nat Rev Cancer 9(9):631–643. https://doi.org/10.1038/nrc2713
Group EBCTC (2005) Effects of chemotherapy and hormonal therapy for early breast cancer on recurrence and 15-year survival: an overview of the randomised trials. Lancet 365(9472):1687–1717. https://doi.org/10.1016/S0140-6736(05)66544-0
Early Breast Cancer Trialists’ Collaborative G (2011) Relevance of breast cancer hormone receptors and other factors to the efficacy of adjuvant tamoxifen: patient-level meta-analysis of randomised trials. Lancet 378(9793):771–784. https://doi.org/10.1016/S0140-6736(11)60993-8
Weinstein J (1997) Cell cycle-regulated expression, phosphorylation, and degradation of p55Cdc. A mammalian homolog of CDC20/Fizzy/slp1. J Biol Chem 272(45):28501–28511
Weinstein J, Jacobsen FW, Hsu-Chen J, Wu T, Baum LG (1994) A novel mammalian protein, p55CDC, present in dividing cells is associated with protein kinase activity and has homology to the Saccharomyces cerevisiae cell division cycle proteins Cdc20 and Cdc4. Mol Cell Biol 14(5):3350–3363
Mondal G, Sengupta S, Panda CK, Gollin SM, Saunders WS, Roychoudhury S (2007) Overexpression of Cdc20 leads to impairment of the spindle assembly checkpoint and aneuploidization in oral cancer. Carcinogenesis 28(1):81–92. https://doi.org/10.1093/carcin/bgl100
Wang Z, Wan L, Zhong J, Inuzuka H, Liu P, Sarkar FH, Wei W (2013) Cdc20: a potential novel therapeutic target for cancer treatment. Curr Pharm Des 19(18):3210–3214
Ouellet V, Guyot MC, Le Page C, Filali-Mouhim A, Lussier C, Tonin PN, Provencher DM, Mes-Masson AM (2006) Tissue array analysis of expression microarray candidates identifies markers associated with tumor grade and outcome in serous epithelial ovarian cancer. Int J Cancer 119(3):599–607. https://doi.org/10.1002/ijc.21902
Kim JM, Sohn HY, Yoon SY, Oh JH, Yang JO, Kim JH, Song KS, Rho SM, Yoo HS, Kim YS, Kim JG, Kim NS (2005) Identification of gastric cancer-related genes using a cDNA microarray containing novel expressed sequence tags expressed in gastric cancer cells. Clin Cancer Res 11(2 Pt 1):473–482
Rajkumar T, Sabitha K, Vijayalakshmi N, Shirley S, Bose MV, Gopal G, Selvaluxmy G (2011) Identification and validation of genes involved in cervical tumourigenesis. BMC Cancer 11:80. https://doi.org/10.1186/1471-2407-11-80
Chang DZ, Ma Y, Ji B, Liu Y, Hwu P, Abbruzzese JL, Logsdon C, Wang H (2012) Increased CDC20 expression is associated with pancreatic ductal adenocarcinoma differentiation and progression. J Hematol Oncol 5:15–15. https://doi.org/10.1186/1756-8722-5-15
Wu W-j, Hu K-s, Wang D-s, Zeng Z-l, Zhang D-s, Chen D-l, Bai L, Xu R-h (2013) CDC20 overexpression predicts a poor prognosis for patients with colorectal cancer. J Transl Med 11:142–142. https://doi.org/10.1186/1479-5876-11-142
Kato T, Daigo Y, Aragaki M, Ishikawa K, Sato M, Kaji M (2012) Overexpression of CDC20 predicts poor prognosis in primary non-small cell lung cancer patients. J Surg Oncol 106(4):423–430. https://doi.org/10.1002/jso.23109
Yuan B, Xu Y, Woo JH, Wang Y, Bae YK, Yoon DS, Wersto RP, Tully E, Wilsbach K, Gabrielson E (2006) Increased expression of mitotic checkpoint genes in breast cancer cells with chromosomal instability. Clin Cancer Res 12(2):405–410. https://doi.org/10.1158/1078-0432.ccr-05-0903
Karra H, Repo H, Ahonen I, Löyttyniemi E, Pitkänen R, Lintunen M, Kuopio T, Söderström M, Kronqvist P (2014) Cdc20 and securin overexpression predict short-term breast cancer survival. Br J Cancer 110(12):2905–2913. https://doi.org/10.1038/bjc.2014.252
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, Graf S, Ha G, Haffari G, Bashashati A, Russell R, McKinney S, Langerod A, Green A, Provenzano E, Wishart G, Pinder S, Watson P, Markowetz F, Murphy L, Ellis I, Purushotham A, Borresen-Dale AL, Brenton JD, Tavare S, Caldas C, Aparicio S (2012) The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486(7403):346–352. https://doi.org/10.1038/nature10983
Gyorffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q, Szallasi Z (2010) An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat 123(3):725–731. https://doi.org/10.1007/s10549-009-0674-9
Jezequel P, Campone M, Gouraud W, Guerin-Charbonnel C, Leux C, Ricolleau G, Campion L (2012) bc-GenExMiner: an easy-to-use online platform for gene prognostic analyses in breast cancer. Breast Cancer Res Treat 131(3):765–775. https://doi.org/10.1007/s10549-011-1457-7
Alfarsi LH, Elansari R, Toss MS, Diez-Rodriguez M, Nolan CC, Ellis IO, Rakha EA, Green AR (2019) Kinesin family member-18A (KIF18A) is a predictive biomarker of poor benefit from endocrine therapy in early ER+ breast cancer. Breast Cancer Res Treat 173(1):93–102. https://doi.org/10.1007/s10549-018-4978-5
Abd El-Rehim DM, Ball G, Pinder SE, Rakha E, Paish C, Robertson JF, Macmillan D, Blamey RW, Ellis IO (2005) High-throughput protein expression analysis using tissue microarray technology of a large well-characterised series identifies biologically distinct classes of breast cancer confirming recent cDNA expression analyses. Int J Cancer 116(3):340–350. https://doi.org/10.1002/ijc.21004
McCarty KS Jr, Miller LS, Cox EB, Konrath J, McCarty KS Sr (1985) Estrogen receptor analyses: Correlation of biochemical and immunohistochemical methods using monoclonal antireceptor antibodies. Arch Pathol Lab Med 109(8):716–721
Clarke R, Tyson JJ, Dixon JM (2015) Endocrine resistance in breast cancer–an overview and update. Molecular and cellular endocrinology 418(Pt 3):220–234. https://doi.org/10.1016/j.mce.2015.09.035
Wasch R, Engelbert D (2005) Anaphase-promoting complex-dependent proteolysis of cell cycle regulators and genomic instability of cancer cells. Oncogene 24(1):1–10. https://doi.org/10.1038/sj.onc.1208017
Taniguchi K, Momiyama N, Ueda M, Matsuyama R, Mori R, Fujii Y, Ichikawa Y, Endo I, Togo S, Shimada H (2008) Targeting of CDC20 via small interfering RNA causes enhancement of the cytotoxicity of chemoradiation. Anticancer Res 28(3a):1559–1563
Li J, Gao JZ, Du JL, Huang ZX, Wei LX (2014) Increased CDC20 expression is associated with development and progression of hepatocellular carcinoma. Int J Oncol 45(4):1547–1555. https://doi.org/10.3892/ijo.2014.2559
Duffy MJ, Harbeck N, Nap M, Molina R, Nicolini A, Senkus E, Cardoso F (2017) Clinical use of biomarkers in breast cancer: updated guidelines from the European Group on Tumor Markers (EGTM). Eur J Cancer 75:284–298. https://doi.org/10.1016/j.ejca.2017.01.017
Alfarsi L, Johnston S, Liu DX, Rakha E, Green A (2018) Current issues with luminal subtype classification in terms of prediction of benefit from endocrine therapy in early breast cancer. Histopathology. https://doi.org/10.1111/his.13523
Gray RG, Rea D, Handley K, Bowden SJ, Perry P, Earl HM, Poole CJ, Bates T, Chetiyawardana S, Dewar JA, Fernando IN, Grieve R, Nicoll J, Rayter Z, Robinson A, Salman A, Yarnold J, Bathers S, Marshall A, Lee M, Group obota C (2013) aTTom: long-term effects of continuing adjuvant tamoxifen to 10 years versus stopping at 5 years in 6,953 women with early breast cancer. J Clin Oncol 31(18_suppl):5–5. https://doi.org/10.1200/jco.2013.31.18_suppl.5
Davies C, Pan H, Godwin J, Gray R, Arriagada R, Raina V, Abraham M, Medeiros Alencar VH, Badran A, Bonfill X, Bradbury J, Clarke M, Collins R, Davis SR, Delmestri A, Forbes JF, Haddad P, Hou MF, Inbar M, Khaled H, Kielanowska J, Kwan WH, Mathew BS, Mittra I, Muller B, Nicolucci A, Peralta O, Pernas F, Petruzelka L, Pienkowski T, Radhika R, Rajan B, Rubach MT, Tort S, Urrutia G, Valentini M, Wang Y, Peto R (2013) Long-term effects of continuing adjuvant tamoxifen to 10 years versus stopping at 5 years after diagnosis of oestrogen receptor-positive breast cancer: ATLAS, a randomised trial. Lancet (London, England) 381(9869):805–816. https://doi.org/10.1016/s0140-6736(12)61963-1
van Hellemond IEG, Geurts SME, Tjan-Heijnen VCG (2018) Current status of extended adjuvant endocrine therapy in early stage breast cancer. Curr Treat Options Oncol 19(5):26–26. https://doi.org/10.1007/s11864-018-0541-1
Atkins D, Reiffen KA, Tegtmeier CL, Winther H, Bonato MS, Storkel S (2004) Immunohistochemical detection of EGFR in paraffin-embedded tumor tissues: variation in staining intensity due to choice of fixative and storage time of tissue sections. J Histochem Cytochem 52(7):893–901. https://doi.org/10.1369/jhc.3A6195.2004
Press MF, Hung G, Godolphin W, Slamon DJ (1994) Sensitivity of HER-2/neu antibodies in archival tissue samples: potential source of error in immunohistochemical studies of oncogene expression. Can Res 54(10):2771–2777
Saxby AJ, Nielsen A, Scarlett CJ, Clarkson A, Morey A, Gill A, Smith RC (2005) Assessment of HER-2 status in pancreatic adenocarcinoma: correlation of immunohistochemistry, quantitative real-time RT-PCR, and FISH with aneuploidy and survival. Am J Surg Pathol 29(9):1125–1134
Stark AM, Pfannenschmidt S, Tscheslog H, Maass N, Rosel F, Mehdorn HM, Held-Feindt J (2006) Reduced mRNA and protein expression of BCL-2 versus decreased mRNA and increased protein expression of BAX in breast cancer brain metastases: a real-time PCR and immunohistochemical evaluation. Neurol Res 28(8):787–793. https://doi.org/10.1179/016164106x110364
Dickson BC, Mulligan AM, Zhang H, Lockwood G, O’Malley FP, Egan SE, Reedijk M (2007) High-level JAG1 mRNA and protein predict poor outcome in breast cancer. Mod Pathol 20(6):685–693. https://doi.org/10.1038/modpathol.3800785
Sauerbrei W, Taube SE, McShane LM, Cavenagh MM, Altman DG (2018) Reporting recommendations for tumor marker prognostic studies (REMARK): an Abridged Explanation and Elaboration. J Natl Cancer Inst 110(8):803–811. https://doi.org/10.1093/jnci/djy088
Acknowledgements
We thank the Nottingham Health Science Biobank and Breast Cancer Now Tissue Bank for the provision of tissue samples. We thank the Saudi Arabia Cultural Embassy for funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest that could be perceived as prejudicing the impartiality of the research reported.
Ethical approval
This study was performed according to the REMARK guidelines for tumour prognostic studies [39], and approved by the Nottingham Research Ethics Committee 2 under the title “Development of a molecular genetic classification of breast cancer”.
Informed consent
Informed consent was obtained from the participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Alfarsi, L.H., Ansari, R.E., Craze, M.L. et al. CDC20 expression in oestrogen receptor positive breast cancer predicts poor prognosis and lack of response to endocrine therapy. Breast Cancer Res Treat 178, 535–544 (2019). https://doi.org/10.1007/s10549-019-05420-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-019-05420-8