Abstract
Purpose
The aim of this study is to explore new salivary biomarkers to discriminate breast cancer patients from healthy controls.
Methods
Saliva samples were collected after 9 h fasting and were immediately stored at − 80 °C. Capillary electrophoresis and liquid chromatography with mass spectrometry were used to quantify hundreds of hydrophilic metabolites. Conventional statistical analyses and artificial intelligence-based methods were used to assess the discrimination abilities of the quantified metabolites. A multiple logistic regression (MLR) model and an alternative decision tree (ADTree)-based machine learning method were used. The generalization abilities of these mathematical models were validated in various computational tests, such as cross-validation and resampling methods.
Results
One hundred sixty-six unstimulated saliva samples were collected from 101 patients with invasive carcinoma of the breast (IC), 23 patients with ductal carcinoma in situ (DCIS), and 42 healthy controls (C). Of the 260 quantified metabolites, polyamines were significantly elevated in the saliva of patients with breast cancer. Spermine showed the highest area under the receiver operating characteristic curves [0.766; 95% confidence interval (CI) 0.671–0.840, P < 0.0001] to discriminate IC from C. In addition to spermine, polyamines and their acetylated forms were elevated in IC only. Two hundred each of two-fold, five-fold, and ten-fold cross-validation using different random values were conducted and the MLR model had slightly better accuracy. The ADTree with an ensemble approach showed higher accuracy (0.912; 95% CI 0.838–0.961, P < 0.0001). These prediction models also included spermine as a predictive factor.
Conclusions
These data indicated that combinations of salivary metabolomics with the ADTree-based machine learning methods show potential for non-invasive screening of breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is one of the most common diseases worldwide. Approximately 2.09 million new cases were diagnosed and 627,000 related deaths occurred globally in 2018 [1]. Although the incidence of breast cancer remains high in the United States and Europe, both incidence and mortality have shown a decreasing trend in these countries [2, 3]. However, in Japan, the incidence of breast cancer has been increasing substantially, and mortality has not shown a decreasing trend [4]. These trends are partially due to differences in receiving rates of screening mammography in these countries. Although screening mammography provides age-specific reductions in breast cancer mortality [5], the receiving rate of mammography in Japan is roughly half of the United States and Europe [6].
Organized screening has reduced breast cancer mortality despite various substantial effects such as overdiagnosis, high cost, radiation exposure, and false positive biopsy recommendation [7,8,9,10]. Saliva, an informative biofluid that reflects systemic disease and enables easy, safe, and cost-effective collection, shows the potential for screening various types of cancers [11,12,13]. In addition to the detection of cancers in the oral cavity [14], various salivary biomarkers have been explored [15].
Various types of novel biomarkers in saliva have been reported for detecting breast cancer, such as epidermal growth factor (EGF), human epidermal growth factor receptor 2 (HER2), vascular endothelial growth factor (VEGF), carcinoembryonic antigen (CEA), cancer antigen 15-3 (CA15-3), and tumor suppressor oncogene protein (p53) [16,17,18,19]. Recent omics technologies, such as transcriptomics, proteomics, and glycoproteomics, can simultaneously quantify hundreds of molecules and patterns to discriminate patients with breast cancer from healthy subjects [20,21,22,23].
Metabolomics is a technology that enables profiling of metabolites and has the potential for screening of breast cancer [24,25,26,27,28]. Because it cannot quantify all metabolites by a single method, a limited number of molecules showing similar chemical properties, e.g., lipids, are profiled. Various analytical approaches are used in sample analysis, including nuclear magnetic resonance imaging and mass spectrometry (MS). Separation techniques, such as capillary electrophoresis (CE) [24] or liquid chromatography (LC) [29], are used before MS depending on the molecules of interest.
Hydrophilic metabolites, such as amino acids and polyamines, can reportedly be used to discriminate patients with breast cancer from healthy controls [24,25,26, 29]. We previously observed the elevation of polyamines in saliva collected from patients with pancreatic cancer [30]. In this study, we conducted comprehensive metabolomics of hydrophilic metabolites and assessed their discrimination abilities using machine learning methods.
Methods
Subjects
This study was a cross-sectional study for exploring breast cancer-specific salivary metabolites. The sample size of this study was the number we could recruit within the study periods. All patients had histologically diagnosed with breast cancer. None had received any prior treatment, including hormone therapy, chemotherapy, molecularly targeted therapy, radiotherapy, surgery, or alternative therapy. Healthy controls were volunteer healthcare workers in our hospital. They had no history of any cancer. Two women in the healthy controls had fibrocystic disease confirmed by needle biopsy.
This study was conducted according to the Declaration of Helsinki principles. The study protocol was approved by the ethics committees of Keio University (No.20120143), Teikyo University (No.15-047-2), and Kitasato University, Kitasato Institutional Hospital (No.17006). Written, informed consent was obtained from all participants who agreed to serve as saliva donors.
Saliva collection
Saliva was collected as described previously [31]. Subjects were allowed only water after 9:00 p.m. on the day prior to collection. All samples were collected between 9:00 and 11:00 a.m. The subjects were required to brush their teeth without toothpaste on the day collection and could not use lipstick, drink water, smoke, brush their teeth, or exercise intensively 1 h before saliva collection. A polypropylene straw 1.1 cm in diameter was used to assist in collection. Subjects were required to gently gargle with water just before saliva collection. Approximately 400 µL of unstimulated saliva was collected and stored in 50 cc polypropylene tubes on ice to prevent degeneration of salivary metabolites [32]. After collection, saliva samples were immediately stored at − 80 °C.
Saliva preparation and metabolomics analyses
The saliva samples were analyzed by two methods. CE-time-of-flight-MS (TOF–MS) was used for non-targeted analyses of hydrophilic metabolites, and LC-triple quadrupole MS (QQQMS) was used for accurate quantification of polyamines as described previously with slight modifications [32, 33]. Frozen saliva was thawed at 4 °C for approximately 1.5 h and subsequently dissolved using a Vortex mixer at room temperature (Thermo Fisher Scientific, Waltham, MA, USA). Ten microliters of each sample were then used in LC–MS analysis, and the rest in CE-MS analysis.
For LC–MS analysis, saliva was mixed with methanol (90 µL) containing 149.6 mM ammonium hydroxide (1% (v/v) ammonia solution) and 0.9 µM internal standards (d8-spermine, d8-spermidine, d6-N1-acetylspermidine, d3-N1-acetylspermine, d6-N1,N8-diacetylspermidine, d6-N1,N12-diacetylspermine, hypoxanthine-13C,15N, 1,6-diaminohexane, 13C,15N-Arg, 13C,15N-Lys, 13C,15N-Met, 13C,15N-Pro, 13C,15N-Trp, d3-Leu, and d5-Phe). After centrifugation at 15,780×g for 10 min at 4 °C, the supernatant was transferred to a fresh tube and vacuum-dried. The sample was reconstituted with 90% methanol (10 µL) and water (30 µL), and then vortexed and centrifuged at 15,780×g for 10 min at 4 °C. One microliter of supernatant was then injected into the LC–MS.
For CE-MS, saliva was centrifuged through a 5 kDa-cutoff filter (EMD Millipore, Billerica, MA, USA) at 9100×g for at least 2.5 h at 4 °C. The filtrate (45 µL) was transferred to a 1.5 mL Eppendorf tube with 2 mM of internal standards (methionine sulfone, 2-[N-morpholino]-ethanesulfonic acid (MES), d-camphol-10-sulfonic acid, sodium salt, 3-aminopyrrolidine, and trimesate). The instrumentation and measurement conditions used for LC-QQQMS and CE-TOFMS were as described previously [32,33,34].
Processing of raw data was conducted by following the typical data processing flow [35]. LC–MS data were processed using Agilent MassHunter Qualitative Analysis and Quantitative Analysis software, including the MassHunter Optimizer and the Dynamic Multiple Reaction Monitoring Mode (DMRM) software (version B.08.00; Agilent Technologies, Santa Clara, CA, USA). Polyamine concentrations were calculated based on the peak area of corresponding internal standards. CE-MS data were analyzed by MasterHands (Keio University, Tsuruoka, Japan) [24] with noise filtering, subtraction of baselines, peak integration for each sliced electropherogram, estimation of accurate m/z in mass spectrometry, alignment of multiple datasets to generate peak matrices, and identification of each peak by matching m/z values and corrected migration times to corresponding entries in a standard library. Metabolite concentrations in CE-MS were calculated based on the ratio of peak area divided by the area of the internal standards in the samples and standard compound mixtures. Polyamine LC–MS data were used for subsequent analyses since their peaks were redundantly detected by both methods.
Data analysis
Collected data were classified into three groups; invasive carcinoma (IC), ductal carcinoma in situ (DCIS), and controls (C). To use only reliable quantification data, metabolites detected in less than 50% of IC samples were eliminated, and metabolites detected below the quantification limit in more than 20 samples were eliminated. The remaining metabolites were subsequently analyzed. The Mann–Whitney test was used for comparisons between two groups, C versus IC. Q-values were calculated by correcting P-values using a false discovery rate (FDR) considering multiple independent tests. The Kruskal–Wallis test and Dunn’s post test were used for comparisons between three groups.
To assess the predictive ability of metabolite combinations, a multiple logistic regression (MLR) model was developed to differentiate IC from C. Prior to the development of the model, stepwise feature selection was conducted to identify the minimum independent features. The threshold to remove a feature was P = 0.05. To evaluate the generalization ability of the model, k-fold cross-validations (k-CV) were conducted, i.e., the datasets were randomly split into training and validation datasets in (k−1):1 ratio. The model was developed using training data and evaluated by validation data. This process was repeated k times, and generalization ability was calculated based on prediction using validation data. We conducted 200 each of two-fold, five-fold, and ten-fold CV using different random values.
We also utilized an alternative decision tree (ADTree), an improved form of conventional if–then decision tree-based machine learning methods [36]. To enhance prediction accuracy, an ensemble approach was used, i.e., multiple ADTree models were developed, and their predictions were integrated to differentiate IC from C. Three-step analyses were conducted. First, to eliminate the bias in the number of datasets, bias-controlled resampling was conducted, i.e., individual data were randomly selected with redundant selection. Second, an ensemble ADTree was developed using the data from the first step. Among several ensemble methods, we utilized bagging methods, i.e., multiple models were developed based on multiple datasets generated by random resampling. Model parameters, including the number of nodes in a tree (boosting number) and the number of trees (bagging number), were determined by two-fold CV. Third, the development model was used to predict the probability of IC using the original data. To assess generalization ability, bootstrap-like analyses were conducted (called resampling analyses), i.e., individual data were randomly selected with redundant selection, and development and validation of the models were conducted. This process was repeated 200 times with different random values.
JMP Pro (ver. 14.1.0; SAS Institute Inc., Cary, NC, USA), GraphPad Prism (ver. 7.0.3; GraphPad Software, Inc., La Jolla, CA, USA), MeV TM4 (ver. 4.9.0; http://mev.tm4.org), and Weka (ver. 3.6.13; University of Waikato, Hamilton, New Zealand) were used for analyses.
Results
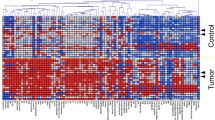
Table 1 summarizes information related to the subjects enrolled in this study. Saliva samples were collected from three groups including C (n = 42), DCIS (n = 23), and IC (n = 101). Benign breast diseases (n = 2) were included in the C group. The IC group included invasive ductal carcinoma of non-specific type (n = 95), mucinous carcinoma (n = 2), invasive lobular carcinoma (n = 2), apocrine carcinoma (n = 1), and invasive micropapillary carcinoma (n = 1). Two hundred sixty metabolites were detected using CE-TOFMS and LC-QQQMS analyses. Of these, 105 were frequently detected in samples collected from patients with breast cancer (≥ 50%) and used for subsequent analyses. Comparisons between C and IC resulted in 31 metabolites showing P-values< 0.05 (Mann–Whitney test); among these, 26 showed Q-values< 0.05 (FDR-corrected P-value). The holistic view of 31 metabolites concentrations is depicted in a heatmap (Fig. 1). Amino acids other than aspartic acid (Asp) had Q-values < 0.05. Polyamines and their acetylated forms also had Q-values < 0.05.
Heatmap showing salivary metabolite concentration. Metabolites with P-values < 0.05 (Mann–Whitney test) in comparisons between C and IC + DCIS were detected. The absolute concentration of each metabolite was divided by the average of those in C. Higher and lower concentrations are indicated in red and blue, respectively. White indicates the averaged concentration in C. Metabolites showing Q-values < 0.05 are highlighted in orange
Figure 2 shows the fold changes of 26 metabolites with Q-values < 0.05 between the IC and C groups. Figure 3 shows comparisons among the quantified concentrations of the top eight-ranked metabolites in Fig. 2 from all 3 groups. Seven metabolites except N1-acetylspermine revealed significant differences (P-value < 0.05, Kruskal–Wallis test with Dunn’s post test) between C and IC and no significant differences between C and DCIS. This finding indicated IC-specific elevation of metabolite concentrations. Additionally, N1-acetylspermine revealed significant difference not only between C and IC but also between DCIS and IC.
Discrimination of IC from C was evaluated using receiver operating characteristic (ROC) curves. Among all quantified metabolites, spermine showed the best area under ROC curves (AUC), 0.766 [95% confidence interval (CI) 0.671–0.840] (Fig. 4a). To assess the predictive ability of combinations of multiple metabolites, an MLR model was developed. Stepwise feature selection selected spermine and ribulose-5-phosphate (Ru5P) from the metabolites showing Q-value < 0.05 (Table 2). The developed MLR model yielded an AUC of 0.790 (95% CI 0.699–0.859) (Fig. 4a). The spermine and MLR models were evaluated by CV with three division ratios (k-fold, k = 2, 5, 10), and the median AUC values after 200 CVs were almost constant, 0.752–754 and 0.774–0.775, respectively. The difference between the upper and lower 95% CI was small, e.g., 0.747–0.751 and 0.766–0.771 for the spermine and MLR models, respectively, in the case of k = 2. Small differences were also observed in k = 5 and 10 (Fig. 5).
Discrimination ability to differentiate IC from C. a ROC curves. AUC values are summarized in Table 4. b Predicted probability of IC using ADTree + Bagging. P-values calculated by Kruskal–Wallis test are shown. ***P-values < 0.001 and ****P-values < 0.0001 (Dunn’s post test)
We also developed an ADTree model and integrated multiple ADTree models generated by bagging methods (ADTree + Bagging). The boosting and bagging numbers were optimized at 7 and 9, respectively. The ADTree and multiple ADTree models yielded AUC values of 0.880 (95% CI 0.798–0.931) and 0.919 (95% CI 0.838–0.961), respectively. The former model is depicted in Fig. 6a. The concept of the ADTree + Bagging model is described in Fig. 6b. The ADTree + Bagging model included nine ADTree models, and the averaged value of each ADTree was used for prediction. The number of parameters used in this model is summarized in Fig. 6c. The generalization ability of the spermine model and the other three models were evaluated by resampling tests (Fig. 7). The median AUC values after 200 resamplings increased for the spermine (AUC = 0.772), MLR (AUC = 0.796), ADTree (AUC = 0.834), and ADTree + Bagging (AUC = 0.864) models. These AUC values showed significant differences in each other model (P < 0.01, Kruskal–Wallis test with Dunn’s post test). The differences between the ROC curves of the spermine model and the other combined models are summarized in Table 3. Figure 4b showed the predicted probabilities of IC calculated by ADtree + Bagging model.
Machine learning models. a Structure of an ADTree. b The concept of the ADTree with bagging model. c The number of variables used in the ADTree models with bagging methods. This algorithm consists of a root node and multiple simple decision trees in which an index is associated with each leaf node, and its final predictive value is the sum of the indices of the leaf nodes fulfilling the condition of the patients
AUC values to discriminate IC from C. AUC values were generated by resampling of the spermine model and three mathematical models. Horizontal bars represent the 95th, 75th, 50th (median), 25th, and 5th percentiles. Dots indicate outliers. The Kruskal–Wallis test with Dunn’s post test yielded adjusted P-values. ****P < 0.0001 and **P < 0.01
Metabolite comparisons in the analysis of each subtype (luminal A-like, luminal B-like, HER2-positive, and triple-negative) showed that five metabolite levels were significantly different between the luminal A-like and B-like subtypes, while N-acetylneuraminate was only significantly different between luminal A-like and triple-negative subtypes. No metabolites were significantly different among the other subtypes (Fig. 8).
Heatmap showing salivary metabolite concentration in four cancer subtypes. Luminal A-like (LA), luminal B-like (LB), HER2-positive (HER2), and triple-negative (TN). Metabolites showing P-values < 0.05 (Mann–Whitney test) in comparisons between C and IC were detected. The mean concentration of each metabolite was divided by the average of those in IC. Higher and lower concentrations are indicated in red and blue, respectively. White indicates the averaged concentrations in IC. Round black dots (fully filled) indicate metabolites with P-values < 0.05 (Kruskal–Wallis test)
Discussion
The aim of this study was to discriminate breast cancer patients from healthy controls using saliva metabolomics. Charged hydrophilic metabolites were comprehensively analyzed using CE-TOFMS, and polyamines were profiled with CE-TOFMS and their measurements optimized using LC-QQQMS to achieve more sensitize quantification. Patients with breast cancer showed higher concentrations of polyamines and amino acids (Figs. 1, 2) in saliva than controls. Figure 3 indicated that the elevation of these salivary metabolites was specific to IC. In general, concentrations of polyamines and their acetylated forms are elevated in cancer tissues. Although our reprocess could reduce the chance to identify some metabolites to specific IC subgroups, our data also showed high concentrations of polyamines and their acetylated forms after eliminating some metabolites according to our exclusion criteria. Therefore, we think our reprocess is appropriate. Elevated concentration of salivary amino acids was consistent with another report [29]. Lactate, an end product of glycolysis, was included in our oral cancer saliva data [14]. Our previous study found that carnitine and choline were elevated in saliva collected from patients with oral cancer [37].
DCIS has a very good prognosis compared to IC [38]. To solve one of the current issues about overdiagnosis and overtreatment of screen-detected DCIS [39, 40], several clinical trials are now in progress to evaluate the safety of active surveillance for low-risk DCIS [41, 42]. Therefore, discriminating IC from controls is more beneficial than discriminating DCIS from controls and we built predictive models to discriminate IC from controls without using the metabolites profile of DCIS group in our study.
Among the quantified metabolites, spermine showed the highest AUC values for discriminating IC from C. The combined MLR model consisting of spermine and Ru5P (Table 2) showed better AUC values (Table 4) than each component model alone. Features were selected using the threshold P = 0.05 to eliminate redundant elements, and only these two metabolites remained. This indicates the positive correlation between other metabolites and spermine and/or Ru5P, suggesting less additional predictive abilities. In fact, no significant difference was observed between the ROC curves of the spermine and MLR models (Table 3). Spermine alone showed high enough predictive ability, but other combination methods should be utilized to enhance the predictive ability of multiple metabolites.
The ADTree model showed better AUC values than spermine and MLR model (Table 4). Furthermore, ADTree + Bagging showed the best AUC values (Table 4). Only this model showed significant differences in ROC curves compared to those of all other models (Table 3). Compared to the MLR model, the features of the ADTree + Bagging model are difficult to evaluate due to their complexity. However, spermine and Ru5P were connected to the root node in the ADTree model (Fig. 6a), which indicated that the concentrations of these metabolites were always used in prediction. Thus, these metabolites contributed greatly to prediction in single ADTree models. Since the ADTree + Bagging model is complicated (Fig. 6b), we simply counted the usage of each metabolite in the model. Spermine and Ru5P were ranked first and second in the ADTree + Bagging model (Fig. 6c). Taken together, even in this machine learning method, spermine and Ru5P were important predictive factors in differentiating IC from C.
Activation of polyamine synthesis in tumor tissue and spread to the surrounding environment has been well described [43]. Ornithine is a precursor metabolite of polyamine in the urea cycle. Ornithine decarboxylase (ODC) (EC 4.1.1.17) converts ornithine to putrescine, and putrescine is converted to spermidine by spermidine synthase (SRM) and S-adenosylmethionine, which is provided by methionine pathways. Spermidine/spermine-N1-acetyltransferase (SSAT) (EC 2.3.1.57) acetylates these polyamines. Therefore, concentrations of polyamines and their acetylated forms are elevated in cancer tissues. Mutation of adenomatous polyposis coli (APC) function results in the upregulation of MYC, which induces ODC activation [44, 45]. MYC mutation is generally observed in various human cancers, and elevation of polyamines has been reported in such cancers. We previously confirmed the drastic changes in the metabolic profiles of colon cancer tissues caused by MYC mutation compared to those caused by several other oncogene mutations [46]. For example, the elevation of N1, N12-diacetylspermine has been repeatedly reported in blood and urine samples from patients with breast, colon, or lung cancers [47,48,49,50]. We previously found that various polyamines are elevated in the saliva of patients with pancreatic cancers [31]. Therefore, a combination of multiple markers is preferable to enhance specificity.
Recently, metabolomics has been employed to analyze saliva samples collected from patients with breast cancer. Scores combining quantified salivary polyamines have been positively correlated with breast cancer stage [25]. The scoring equation used in that study contained spermine and N1-acetylspermidine with positive coefficients, indicating a correlation between the elevation of these metabolites and tumor burden of breast cancer, which is consistent with our observations. However, N1-acetylspermine was used with a negative coefficient, inconsistently with our observations. One possible reason is the use of N1-acetylspermine as a confounding factor in the equation, as this metabolite was positively correlated with spermine. Recently, hydrophilic interaction chromatography-MS was utilized to profile metabolites in saliva collected from patients with breast cancers and revealed that various metabolites were elevated in phospholipid catabolism, such as lysophoshatidylcholine and phosphatidylcholine [28]. These metabolites were not observed using our methods.
Our study has several limitations. Polyamine concentrations in biofluids are affected by dietary intake [51] and various diseases [52]. Other metabolites, such as amino acids, also fluctuate according to lifestyle and environmental factors [34]. Even when combining multiple markers, the effects of these factors should be minimized to realize accurate determination. The developed discrimination model should be compared with other cancer data to evaluate its specificity to breast cancer. This study tightly controlled the sampling conditions, especially considering fasting, which affects salivary metabolomics profiles [33]. A less stringent sampling protocol should be evaluated for the establishment of a screening model. We did not investigate family history or BRCA1/2 status in this study. These factors are important when considering the risk of breast cancer, so we need to take them into account in the future study.
In conclusion, we analyzed metabolites in saliva samples collected from patients with breast cancer and assessed their ability to discriminate among C, DCIS, and IC. Both CE-MS and LC–MS were used to identify and quantify a variety of hydrophilic metabolites in the samples. The metabolites showing higher fold changes between C and IC were not elevated in DCIS, indicating that they were elevated in IC alone. To enhance the discrimination ability of the concentration patterns of multiple metabolites, we utilized MLR and ADTree models. The MLR model showed higher accuracy than spermine model, despite there being no significant difference in their AUC values. The ADTree and ADTree + Bagging models showed even higher AUC values than spermine alone. Interestingly, both the MLR and machine learning-based models included spermine and Ru5P as predictive factors. Concentration patterns of salivary metabolites along with sophisticated computational classification technology can contribute to non-invasive breast cancer screening. Salivary metabolomics should be conducted before mammography. In other words, salivary metabolomics is considered to be useful for the selection of subject who should receive breast cancer screening with mammography and/or ultrasound. In the future, metabolomics could be used to recommend a biopsy to patients with suspicious mammography.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
International Agency for Research on Cancer. http://gco.iarc.fr/. Accessed 11 Jan 2019
Cancer Incidence in Five Continents, CI5plus. IARC Cancer Base No.9. Lyon, International Agency for Research on Cancer. http://ci5.iarc.fr. Accessed 11 Jan 2019
World Health Organization mortality database. http://www.who.int/healthinfo/mortality_data/en/. Accessed 11 Jan 2019
Matsuda A, Matsuda T, Shibata A, Katanoda K, Sobue T, Nishimoto H (2014) Cancer incidence and incidence rates in Japan in 2008: a study of 25 population-based cancer registries for the Monitoring of Cancer Incidence in Japan (MCIJ) project. Jpn J Clin Oncol 44(4):388–396
Habbema JD, van Oortmarssen GJ, van Putten DJ, Lubbe JT, van der Maas PJ (1986) Age-specific reduction in breast cancer mortality by screening: an analysis of the results of the Health Insurance Plan of Greater New York study. J Natl Cancer Inst 77(2):317–320
OECD Health Statistics https://doi.org/10.1787/health-data-en. Accessed 11 Jan 2019
Miller AB, Wall C, Baines CJ, Sun P, To T, Narod SA (2014) Twenty five year follow-up for breast cancer incidence and mortality of the Canadian National Breast Screening Study: randomised screening trial. BMJ 348:g366
Shah TA, Guraya SS (2017) Breast cancer screening programs: review of merits, demerits, and recent recommendations practiced across the world. J Microsc Ultrastruct 5(2):59–69
Johns LE, Coleman DA, Swerdlow AJ, Moss SM (2017) Effect of population breast screening on breast cancer mortality up to 2005 in England and Wales: an individual-level cohort study. Br J Cancer 116(2):246–252
Johns LE, Swerdlow AJ, Moss SM (2018) Effect of population breast screening on breast cancer mortality to 2005 in England and Wales: a nested case-control study within a cohort of one million women. J Med Screen 25(2):76–81
Kaczor-Urbanowicz KE, Martin Carreras-Presas C, Aro K, Tu M, Garcia-Godoy F, Wong DT (2017) Saliva diagnostics—Current views and directions. Exp Biol Med (Maywood) 242(5):459–472
Wang X, Kaczor-Urbanowicz KE, Wong DT (2017) Salivary biomarkers in cancer detection. Med Oncol 34(1):7
Zhang A, Sun H, Wang X (2012) Saliva metabolomics opens door to biomarker discovery, disease diagnosis, and treatment. Appl Biochem Biotechnol 168(6):1718–1727
Ishikawa S, Sugimoto M, Kitabatake K, Sugano A, Nakamura M, Kaneko M et al (2016) Identification of salivary metabolomic biomarkers for oral cancer screening. Sci Rep 6:31520
Rapado-Gonzalez O, Majem B, Muinelo-Romay L, Lopez-Lopez R, Suarez-Cunqueiro MM (2016) Cancer salivary biomarkers for tumours distant to the oral cavity. Int J Mol Sci 17(9):1531
de Abreu PD, Areias VR, Franco MF, Benitez MC, do Nascimento CM, de Azevedo CM et al (2013) Measurement of HER2 in saliva of women in risk of breast cancer. Pathol Oncol Res 19(3):509–513
Streckfus C, Bigler L, Tucci M, Thigpen JT (2000) A preliminary study of CA15-3, c-erbB-2, epidermal growth factor receptor, cathepsin-D, and p53 in saliva among women with breast carcinoma. Cancer Invest 18(2):101–109
Navarro MA, Mesia R, Diez-Gibert O, Rueda A, Ojeda B, Alonso MC (1997) Epidermal growth factor in plasma and saliva of patients with active breast cancer and breast cancer patients in follow-up compared with healthy women. Breast Cancer Res Treat 42(1):83–86
Brooks MN, Wang J, Li Y, Zhang R, Elashoff D, Wong DT (2008) Salivary protein factors are elevated in breast cancer patients. Mol Med Rep 1(3):375–378
Cavaco C, Pereira JAM, Taunk K, Taware R, Rapole S, Nagarajaram H et al (2018) Screening of salivary volatiles for putative breast cancer discrimination: an exploratory study involving geographically distant populations. Anal Bioanal Chem 410(18):4459–4468
Al-Muhtaseb SI (2014) Serum and saliva protein levels in females with breast cancer. Oncol Lett 8(6):2752–2756
Liu X, Yu H, Qiao Y, Yang J, Shu J, Zhang J et al (2018) Salivary glycopatterns as potential biomarkers for screening of early-stage breast cancer. EBioMedicine 28:70–79
Zhang L, Xiao H, Karlan S, Zhou H, Gross J, Elashoff D et al (2010) Discovery and preclinical validation of salivary transcriptomic and proteomic biomarkers for the non-invasive detection of breast cancer. PLoS ONE 5(12):e15573
Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M (2010) Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics 6(1):78–95
Takayama T, Tsutsui H, Shimizu I, Toyama T, Yoshimoto N, Endo Y et al (2016) Diagnostic approach to breast cancer patients based on target metabolomics in saliva by liquid chromatography with tandem mass spectrometry. Clin Chim Acta 452:18–26
Tsutsui H, Mochizuki T, Inoue K, Toyama T, Yoshimoto N, Endo Y et al (2013) High-throughput LC-MS/MS based simultaneous determination of polyamines including N-acetylated forms in human saliva and the diagnostic approach to breast cancer patients. Anal Chem 85(24):11835–11842
Wang X, Zhao X, Chou J, Yu J, Yang T, Liu L et al (2018) Taurine, glutamic acid and ethylmalonic acid as important metabolites for detecting human breast cancer based on the targeted metabolomics. Cancer Biomark 23(2):255–268
Zhong L, Cheng F, Lu X, Duan Y, Wang X (2016) Untargeted saliva metabonomics study of breast cancer based on ultra performance liquid chromatography coupled to mass spectrometry with HILIC and RPLC separations. Talanta 158:351–360
Cheng F, Wang Z, Huang Y, Duan Y, Wang X (2015) Investigation of salivary free amino acid profile for early diagnosis of breast cancer with ultra performance liquid chromatography-mass spectrometry. Clin Chim Acta 447:23–31
Asai Y, Itoi T, Sugimoto M, Sofuni A, Tsuchiya T, Tanaka R et al (2018) Elevated polyamines in saliva of pancreatic cancer. Cancers 10(2):43
Asai Y, Itoi T, Sugimoto M, Sofuni A, Tsuchiya T, Tanaka R et al (2018) Elevated polyamines in saliva of pancreatic cancer. Cancers (Basel) 10(2):43
Tomita A, Mori M, Hiwatari K, Yamaguchi E, Itoi T, Sunamura M et al (2018) Effect of storage conditions on salivary polyamines quantified via liquid chromatography-mass spectrometry. Sci Rep 8(1):12075
Ishikawa S, Sugimoto M, Kitabatake K, Tu M, Sugano A, Yamamori I et al (2017) Effect of timing of collection of salivary metabolomic biomarkers on oral cancer detection. Amino Acids 49(4):761–770
Sugimoto M, Saruta J, Matsuki C, To M, Onuma H, Kaneko M et al (2013) Physiological and environmental parameters associated with mass spectrometry-based salivary metabolomic profiles. Metabolomics 9(2):454–463
Sugimoto M, Kawakami M, Robert M, Soga T, Tomita M (2012) Bioinformatics tools for mass spectroscopy-based metabolomic data processing and analysis. Curr Bioinform 7(1):96–108
Freund Y, Mason L (1999) The alternating decision tree learning algorithm. Icml 1999:124–133
Wang Q, Gao P, Wang X, Duan Y (2014) Investigation and identification of potential biomarkers in human saliva for the early diagnosis of oral squamous cell carcinoma. Clin Chim Acta 427:79–85
Irene LW, James JD, Bernard F, Eleftherios PM, Stewart JA, Thomas BJ et al (2011) Long-term outcome of invasive ipsilateral breast tumor recurrences after lumpectomy in NSABP B-17 and B-24 randomized clinical trials for DCIS. J Natl Cancer Inst 103(6):478–488
Welch HG, Black WC (2010) Overdiagnosis in cancer. J Natl Cancer Inst 102(9):605–613
Welch HG (2009) Overdiagnosis and mammography screening. BMJ 339:b1425
Elshof LE, Tryfonidis K, Slaets L, van Leeuwen-Stok AE, Skinner VP, Dif N et al (2015) Feasibility of a prospective, randomised, open-label, international muticentre, phase III, non-inferiority trial to assess the saftey of active surveillance for low risk ductal carcinoma in site The LORD study. Eur J Cancer 51(12):1497–1510
Francis A, Thomas J, Fallowfield L, Wallis M, Bartlett JM, Brookes C et al (2015) Addressing overtreatment of screen detected DCIS; the LORIS trial. Eur J Cancer 51(16):2296–2303
Soda K (2011) The mechanisms by which polyamines accelerate tumor spread. J Exp Clin Cancer Res 30:95
Dejure FR, Eilers M (2017) MYC and tumor metabolism: chicken and egg. EMBO J 36(23):3409–3420
Gerner EW, Meyskens FL Jr (2004) Polyamines and cancer: old molecules, new understanding. Nat Rev Cancer 4(10):781–792
Satoh K, Yachida S, Sugimoto M, Oshima M, Nakagawa T, Akamoto S et al (2017) Global metabolic reprogramming of colorectal cancer occurs at adenoma stage and is induced by MYC. Proc Natl Acad Sci USA 114(37):E7697–E7706
Hiramatsu K, Takahashi K, Yamaguchi T, Matsumoto H, Miyamoto H, Tanaka S et al (2005) N(1), N(12)-Diacetylspermine as a sensitive and specific novel marker for early- and late-stage colorectal and breast cancers. Clin Cancer Res 11(8):2986–2990
Takahashi Y, Sakaguchi K, Horio H, Hiramatsu K, Moriya S, Takahashi K et al (2015) Urinary N1, N12-diacetylspermine is a non-invasive marker for the diagnosis and prognosis of non-small-cell lung cancer. Br J Cancer 113(10):1493–1501
Nakajima T, Katsumata K, Kuwabara H, Soya R, Enomoto M, Ishizaki T et al (2018) Urinary polyamine biomarker panels with machine-learning differentiated colorectal cancers, benign disease, and healthy controls. Int J Mol Sci 19(3):756
Wikoff WR, Hanash S, DeFelice B, Miyamoto S, Barnett M, Zhao Y et al (2015) Diacetylspermine is a novel prediagnostic serum biomarker for non-small-cell lung cancer and has additive performance with pro-surfactant protein B. J Clin Oncol 33(33):3880–3886
Vargas AJ, Ashbeck EL, Thomson CA, Gerner EW, Thompson PA (2014) Dietary polyamine intake and polyamines measured in urine. Nutr Cancer 66(7):1144–1153
Park MH, Igarashi K (2013) Polyamines and their metabolites as diagnostic markers of human diseases. Biomol Ther (Seoul) 21(1):1–9
Acknowledgements
We thank Editage (www.editage.jp) for English language editing.
Funding
This study was funded by JSPS KAKENHI Grant Numbers 16H05408 and 25461996, and research Grants from the Yamagata Prefectural Government and the City of Tsuruoka.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interests. M. Sunamura and M. Sugimoto hold unpaid advisory positions in a commercial organization. No other author declares non-financial competing interests.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the ethics committees of Keio University (No.20120143), Teikyo University (No.15-047-2), and Kitasato University Kitasato Institutional Hospital (No.17006) and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Murata, T., Yanagisawa, T., Kurihara, T. et al. Salivary metabolomics with alternative decision tree-based machine learning methods for breast cancer discrimination. Breast Cancer Res Treat 177, 591–601 (2019). https://doi.org/10.1007/s10549-019-05330-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-019-05330-9