Abstract
BRCA1 germline mutation carriers usually develop ER, PR and HER2 negative breast carcinoma. Somatic BRCA1 mutations are rare in sporadic breast cancers, but other mechanisms could impair BRCA1 functions in these tumors, particularly in triple-negative breast carcinomas (TNBCs). Id4, a helix-loop-helix DNA binding factor, blocks BRCA1 gene transcription in vitro and could downregulate BRCA1 in vivo. We compared Id4 immunoreactivity in 101 TNBCs versus 113 non-TNBCs, and correlated the results with tumor morphology and immunoreactivity for CK5/6, CK14, EGFR, and androgen receptor (AR). Id4 was present in 76 out of 101 (75 %) TNBCs: 40 (40 %) TNBCs displayed Id4 positivity in >50 % of neoplastic cells, 23 (23 %) in 5–50 %, and 13 (13 %) in <5 %. In contrast, only 6 (5 %) of 113 non-TNBCs showed focal Id4 positivity, limited to fewer than 5 % of the tumor (p < 0.0001). Id4 expression significantly associated with high histologic grade (p = 0.0002) and mitotic rate (p = 0.006). Id4 decorated all 12 TNBCs with large central acellular zone of necrosis in our series, with positive staining in 10–90 % of the cells. Id4 signal strongly correlated with cytokeratin CK14 reactivity (p < 0.0001), but not with CK5/6 and EGFR. All apocrine carcinomas in our series were positive for AR and most for EGFR, but they were negative for CK5/6, CK14, and Id4, with only two exceptions. Our results document substantial expression of Id4 in most TNBCs, which could result in functional downregulation of BRCA1 pathways in these tumors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Triple-negative breast carcinomas (TNBCs), defined by lack of expression for estrogen receptor (ER), progesterone receptor (PR) and human epidermal growth factor receptor 2 (HER2), represent about 10–15 % of all breast cancers [1]. They are frequent in BRCA1 germline mutation carriers [2], and common in women under 40 years of age and in African-American women [3]. TNBCs show aggressive clinical behavior characterized by early recurrence, frequent brain and lung involvement [3], and poor survival [4]. These tumors demonstrate a high rate of pathologic complete response post-neoadjuvant treatment [5, 6], but patients with residual disease have poor survival [6] and early relapse is common [7]. At present, TNBCs constitute the most challenging group of breast cancers due to the lack of effective targeted therapies. Morphologically, TNBCs are a very heterogeneous group, which consists predominantly of high grade invasive ductal carcinoma of no special type (NOS), but includes special subtypes such as medullary, metaplastic, and apocrine carcinomas. Gene expression studies have classified breast cancers into specific molecular subtypes, including basal-like breast carcinomas which show significant morphologic and immunophenotypic overlap with TNBCs [8]. The use of immunohistochemical stains for ER, HER2, basal cytokeratins CK 5/6 or CK 14 and EGFR in different combinations has been proposed as a surrogate method for the identification of basal-like breast carcinomas [9, 10].
Breast cancers arising in BRCA1 germline mutation carriers are usually triple-negative and display basal-like gene expression profiles [11]. Similarities between breast cancers occurring in BRCA1 germline mutation carriers and sporadic TNBCs suggest that BRCA1 inactivation also plays a role in the pathogenesis of some sporadic tumors [12–15]. Silencing of the BRCA1 gene by promoter methylation has been reported in some non-familial medullary carcinoma and metaplastic carcinomas [12, 16]. Furthermore, Turner et al. [12] showed a two-fold reduction of BRCA1 mRNA levels in basal-like breast cancers compared to matched non-basal-like carcinomas.
Id4 is a negative regulator of the basic helix-loop-helix group of transcription factors [17], and downregulates the BRCA1 promoter in vitro [18]. Recent studies have also demonstrated a regulatory interplay of Id4 with microRNAs (including miR 335 and miR 9) and p53 [19–22]. We have previously reported an inverse correlation between Id4 mRNA and ER immunoreactivity in normal breast epithelium and in ER-positive breast carcinomas [23]. Turner et al. [12] have demonstrated a 9.1-fold increase in Id4 mRNA levels in basal-like carcinoma compared to control tumors by PCR analysis. These data suggest that Id4 could play an important role in the functional downregulation of BRCA1 in TNBCs. In this study, we used immunohistochemistry to assess the expression of Id4 in TNBCs and correlated Id4 reactivity with tumor morphology, basal phenotype, and patient outcome.
Materials and methods
Patient selection and tissue microarrays (TMAs)
The study was approved by the Institutional Review Board. TNBCs were defined as having ER and PR nuclear staining in <1 % of the tumor cells, and either negative (0 or 1+) HER2 staining or equivocal (2+) HER2 staining and no HER2 gene amplification by FISH [24, 25]. A search of our pathology database identified patients with TNBC who had surgical excision of the primary carcinoma at our center between 2002 and 2004. Our study cohort consists of 101 patients with tissue blocks of the primary TNBC available for immunohistochemical analysis. Information about patient age, race and clinical follow-up was obtained from the electronic medical records. One of the study pathologists (YHW) reviewed all available slides for the patients in the study and selected representative tumor slides, which were reviewed together with the other study pathologist (EB). The morphologic characteristics of the TNBCs (tumor type, size, grade, growth pattern, presence or absence of associated in situ carcinoma, calcifications, necrosis, lymphocytic infiltrate, lymphovascular invasion, and lymph node metastasis) were noted. The lymphocytic infiltrate associated with invasive carcinoma was scored according to Kreike et al. [26] as absent = no notable lymphocytic infiltrate; minimal = scattered lymphocytes (<10 lymphocytes/400× power field); moderate = lymphocytes easily identified, but no large aggregates; extensive = large aggregates of lymphocytes in >50 % of the tumor. As reference cases, we used 113 non-TNBCs, which included 11 ER−/HER2+ carcinomas (HER2 3+ by immunohistochemistry or HER2 gene amplified by FISH) and 102 consecutive non-TNBCs (84 ER+/HER2-; 17 ER+/HER2+; 1 ER−/HER2+). The 102 tumors, available as triplicate 0.6 mm cores in TMAs, were all greater than 1 cm in size. The 1 cm cutoff had been arbitrarily chosen before preparation of the TMAs to ensure availability of residual carcinoma for possible future clinical use.
Immunohistochemistry
We performed immunoperoxidase stains for Id4, CK5/6, CK14, EGFR, AR, ER, PR, and HER2, with appropriate positive and negative controls for each staining. Normal breast epithelium expressing Id4 was used as positive control for Id4 staining [23]. Sources and dilutions of the primary antibodies are summarized in Table 1. ER, PR, and HER2 stains were repeated in all TNBCs and re-assessed according to the current American Society of Clinical Oncology/College of American Pathologist guideline recommendations [24, 25]. Immunoperoxidase studies for Id4 were performed on whole tissue sections (1–2 representative tumor blocks from each case) of all TNBCs and of 11 ER−/HER2+ breast carcinomas, and on TMAs of the remaining 102 non-TNBCs. The results of Id4 immunoreactivity on TMAs were validated by immunoperoxidase stain on whole tissue sections of 23/102 (22.5 %) reference tumors, including all non-TNBCs showing any Id4 positivity in the TMAs.
We recorded the percentage of Id4-positive tumor cells in each case and defined Id4 positivity as nuclear staining in at least 5 % of the tumor cells. EGFR was graded according to the scoring guidelines for HER2, and EGFR overexpression was defined as 2+ and 3+ reactivity [27, 28]. Carcinomas showing any cytoplasmic immunoreactivity for CK5/6 and CK14 were classified as positive for these markers.
Statistical analysis
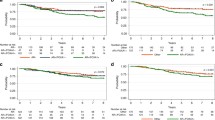
Clinicopathological characteristics of the TNBC and non-TNBC cohorts were compared using the t test or χ2 or if warranted, the Fisher’s exact test or Wilcoxon rank sum test. Similar tests were used to compare the clinicopathological characteristics in the TNBC cohort, by Id4 positivity, where Id4 positivity was defined at a 5 % cutoff. Kaplan–Meier methods were used to estimate distant metastasis-free survival and overall survival. Distant metastasis-free survival was defined as (a) the time from diagnosis to distant metastasis, or (b) the time to death or (c) time to last follow-up, according to patient status. Overall survival was defined as the time from initial diagnosis to death or to last follow-up. The log-rank test was used to assess survival differences between groups with low and high Id4 levels again using 5 % of tumor cells with positive staining as the cutoff; 5 % cutoff was chosen based on the results of exploratory analysis using different threshold values (data not shown). For all analyses, a p value <0.05 was considered as statistically significant. All statistical analysis was performed in SAS 9.2 and R 2.11.1.
Results
Study cohort
All patients in the study were women, with mean age of 54 years (range 29–95). Patients with TNBC and non-TNBC were similar with regard to age, tumor size, and length of follow-up (Table 2). Lymph node metastases were less common in patients with TNBCs than in patients with non-TNBCs, but visceral metastases were more frequent in TNBCs. Table 2 summarizes the clinicopathologic features of TNBCs and non-TNBCs. Information about patient ethnicity was available for all patients with TNBC: 86 (85 %) had been classified as Caucasian, including 21 (21 %) Jewish patients; 11 (11 %) were African-American and 4 (4 %) were Asian.
Morphology of TNBCs and non-TNBCs
Most TNBCs were invasive ductal carcinomas NOS (86/101; 85 %). Twelve of 101 (12 %) carcinomas had a large central acellular zone of necrosis occupying more than 30 % of the tumor mass (LCAZ) [29, 30]. Eleven tumors (11 %) were apocrine carcinomas by morphological assessment, and showed abundant eosinophilic cytoplasm and prominent nucleoli. [31] The remaining TNBCs consisted of two metaplastic carcinomas (2 %), and two carcinomas with medullary features (2 %).
Ninety-one of 113 (81 %) non-TNBCs were invasive ductal carcinomas NOS; the rest included 9 (8 %) invasive lobular carcinomas, 6 (5 %) invasive mammary carcinomas with mixed ductal and lobular features, 4 (4 %) invasive micropapillary carcinomas, 2 (2 %) mixed mucinous carcinomas, and 1 (1 %) invasive papillary carcinoma.
Id4 expression in TNBCs versus non-TNBCs
Id4 immunoreactivity was detected in 76 out of 101 (75 %) TNBCs, but the percentage of Id4-positive cells varied widely. Sixty-three (62.4 %) of the 101 TNBCs showed Id4 positivity in 5 % or more of the neoplastic cells, including 40 (39.6 %) cases with nuclear staining in at least 50 % of the tumor (Fig. 1) and 23 (22.8 %) in 5–50 %. Thirteen (12.7 %) TNBCs displayed focal weak nuclear reactivity in fewer than 5 % of the tumor cells. Twenty-five TNBCs (24.8 %) had no nuclear stain for Id4.
Only 6 out of 113 (5 %) non-TNBCs (4 ER+/HER2− and 2 ER−/HER2+ tumors) showed some immunoreactivity for Id4. The staining was focal and limited to fewer than 5 % of the tumor cells in all six cases when confirmed on the whole tissue sections.
The difference in Id4 expression between TNBCs and non-TNBCs was statistically significant (p < 0.0001) independent of the percentage of Id4-positive tumor cells used as cutoff.
Id4 expression and tumor morphology in TNBCs
Id4 positivity in breast carcinomas is significantly associated with high histologic grade, however, all but 4 TNBCs had modified Bloom–Richardson grade 3. Id4 is also significantly associated with high mitotic rate (Table 3). All 12 TNBCs with LCAZ showed strong positivity for Id4 (Fig. 2), ranging from 10 to 90 % of the tumor cells, with 6 cases showing strong nuclear staining in 50 % or more of the tumor (Table 4).
We observed a significant inverse correlation between Id4 positivity and apocrine morphology. Out of 11 apocrine TNBCs, only two cases showed very focal Id4 staining, which was limited to only 1 and 5 % of the tumor. The 9 remaining apocrine carcinomas were completely negative for Id4 (Table 5). All 11 apocrine TNBCs displayed strong and diffuse immunoreactivity for AR.
Patient age, tumor size, lymphovascular invasion, nodal and visceral metastases did not correlate with Id4 positivity in TNBCs.
Id4 expression and basal-like markers in TNBCs
Of the 101 TNBCs, 47 (47 %) were positive for CK14, 50 (50 %) for CK5/6, and 68 (67 %) for EGFR. Seventy-eight TNBCs (77 %) were positive for CK5/6 and/or EGFR, and 63 (62 %) for CK5/6 and/or CK14. Positive reactivity for Id4 significantly associated with positivity for CK14 (Table 6), but showed no correlation with CK5/6 and EGFR, either alone or in combination. We observed a significant relationship between Id4 staining and positivity for CK5/6 and/or CK14, but this finding is probably secondary to the strong positive correlation between Id4 and CK14. A case demonstrating positive reactivity for Id4, CK14, CK5/6 and EGFR is illustrated in Fig. 3. All 12 TNBCs with LCAZ were positive for Id4 and CK14, and showed variable positivity for CK5/6 and EGFR (Table 4).
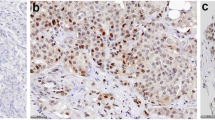
Expression of Id4 and of the basal markers CK5/6, CK14 and EGFR in a triple-negative breast carcinoma. a Tumor morphology (hematoxylin and eosin stain). b–e Id4, CK5/6, CK14, and EGFR immunoreactivity in the same tumor. Note that Id4 (b) CK5/6 (c) and CK14 (d) colocalizes in the same area, whereas EGFR positivity is present in a different area in f (×200)
All 11 apocrine TNBCs were negative for CK14 and 8/11 (73 %) were also negative for CK5/6. EGFR was positive in 10 out of 11 apocrine carcinomas, with 7 showing uniform and strong membranous immunoreactivity in >30 % of the tumor cells.
Id4 expression and patient ethnicity
The TNBCs of 17/21 (80 %) patients of Jewish ethnicity showed Id4 positivity in over 5 % of the tumor cells versus TNBCs in 46/80 (58 %) patients from other ethnic groups (p = 0.048). Aside from Jewish patients, the TNBC in patients of non-Jewish ethnicity, including African-American, did not show significant differences with regard to Id4 positivity.
Survival analysis
Follow-up data were available for 88/101 (87 %) patients with TNBC. The median follow-up time was 72.9 months (range 3.7–136.4 months). We found no statistically significant correlation between Id4 positivity and patient outcome. Overall survival and disease-free survival of patients with Id4 positive TNBC were not significantly different from those of patients with Id4 negative TNBC, independent of the percentage (5, 20 and 50 % or greater) of Id4 positivity used as cutoff for analysis.
Follow-up data were available for all 113 patients with non-TNBC. The median follow-up time was 79 months (range 3–134 months). All 6 patients with non-TNBC showing focal Id4 positivity were alive with no evidence of disease at last follow-up (median 84 months, range 77–103 months). Due to the low number of Id4-positive non-TNBCs and lack of events in this group of patients, we did not perform statistical analysis to compare the outcome of Id4-positive versus Id4-negative non-TNBCs.
Discussion
TNBCs are a morphologically heterogeneous group of tumors. They consist mainly of high grade invasive ductal carcinoma of NOS, but include special subtypes such as medullary, metaplastic, and apocrine carcinomas. Morphologic features common to most TNBCs include nodular growth with a pushing border of invasion, poorly differentiated histology, high mitotic rate, prominent lymphocytic infiltrate, and extensive areas of necrosis such as geographic necrosis or LCAZ [29, 32].
Basal-like breast carcinomas are classified by gene expression profiling [33]. They overlap significantly with TNBCs, but the two groups of tumors are not identical [8]. Investigators have proposed the use of immunohistochemical markers as a surrogate method for the identification of basal-like breast carcinomas. Nielsen [34] and Livasy [32] identified basal-like breast cancer as ER and HER2 negative tumors that express CK5/6 and/or EGFR, whereas Rakha et al. [10] used positivity for a basal cytokeratin (CK5/6, CK17, CK14). At present, however, no immunohistochemical panel identifies all basal-like breast carcinomas with 100 % sensitivity and specificity. Homologous DNA repair is critically altered in most familial breast carcinomas associated with BRCA1 germline mutation, which constitute the prototype of basal-like carcinomas. Although somatic mutation of the BRCA1 gene is not commonly encountered in sporadic breast carcinomas, some of the morphologic features of BRCA1-deficient tumors [35] occur in TNBCs not associated with BRCA1 germline mutation. Functional inactivation of the BRCA1 gene could play an important role in the pathogenesis of these tumors.
Id4 is a member of the Id (inhibitor of DNA binding) family of proteins. Id proteins inhibit the functions of basic helix-loop-helix transcription factors by blocking the ability of these factors to bind DNA [17], and exert an important role in mammalian embryogenesis [36], angiogenesis [22, 37–39], and the maintenance of cancer stem cells [40]. Increased Id4 mRNA levels are found in small cell lung cancer [41], and positive Id4 nuclear immunoreactivity is present in glioblastoma [42] and malignant rhabdoid tumors [43]. Conversely, few studies have documented Id4 epigenetic inactivation by promoter hypermethylation in a wide range of human malignancies including mammary [44, 45], gastric [46], colorectal [47] and prostate adenocarcinoma [48], leukemia, [49] and lymphoma [50], suggesting that Id4 is a tumor-suppressor.
Id4 downregulates BRCA1 gene expression in vitro [18]. In turn, it appears that BRCA1 can downregulate the expression of Id4, as part of a regulatory loop balancing the expression of both genes [51].
microRNAs participate in the modulation of BRCA1 signal by Id4. microRNA-335 simultaneously upregulates the known BRCA1 activators ERα, IGF1R, SP1, and downregulates Id4, a BRCA1 repressor [19]. In one study, overexpression of microRNA-335 was associated with a significant increase of BRCA1 mRNA level and with marked reduction in Id4 mRNA, supporting a functional predominance of Id4 in BRCA1 gene regulation [19]. In the same study, microRNA-335 levels positively correlated with ER and BRCA1 expression in a group of 30 sporadic breast cancers [19]. Positive feedback regulation of microRNA-335 expression by estrogens was also documented in breast cancer cell line [19].
We have previously reported an inverse correlation between Id4 mRNA and ER positivity in the normal breast epithelium [23], consistent with a role of Id4 in the physiologic modulation of the mammary gland. We have also documented an inverse relationship between ER immunoreactivity and Id4 mRNA signal in a series of ER-positive breast carcinomas, and in ER-positive ductal carcinoma in situ and atypical ductal hyperplasia, non-obligate precursors of ER-positive breast cancer [23]. In this setting, Id4 appears to counterbalance the activities of BRCA1 and ER, opposing ER-driven tumorigenesis. Vice versa, Id4 expression could promote tumorigenesis via downregulation of BRCA1 in ER-negative carcinomas.
To evaluate this hypothesis, we assessed the expression of Id4 in TNBCs. Our data demonstrate that Id4 is highly expressed in TNBCs, in marked contrast to ER-positive cancers. We note, however, that Id4 levels in TNBCs vary widely with about 40 % of tumors showing positivity in more than half of the neoplastic cells and about one-fourth displaying positivity in 5–50 %. Our findings are in agreement with those of Turner et al. [12], who found that Id4 mRNA levels are 9.1-fold higher in basal-like breast cancer than in matched non-basal-like control tumors. Turner et al. identified basal-like carcinomas as positive for CK5/6, but our results show stronger correlation between Id4 and CK14 than between Id4 and CK5/6. We also observed that TNBCs with LCAZ, a subset of TNBCs with distinctive morphology and characterized by very poor prognosis [30, 52], show strong positivity for both Id4 and CK14.
When subject to unsupervised gene array analysis, apocrine carcinomas consistently cluster together as a distinct subtype [26, 53]. In our study, most (8/11; 73 %) apocrine carcinomas were negative for the basal cytokeratin CK5/6, consistent with prior data [26]. All apocrine carcinomas in our study were also CK14 negative, but showed positive EGFR staining, consistent with a recent report by Vranic et al. [54]. We found that Id4 expression is extremely rare in apocrine TNBCs. Apocrine carcinomas are unique among TNBCs because they express AR, which documents activity of a hormonal pathway. Based on our prior observation that in breast carcinomas ER and Id4 are inversely correlated [23], we speculate that a similar inverse relationship also exists between Id4 and AR, but our hypothesis needs further testing.
No information regarding BRCA1 mutation status was available for the patients in our study. Even though in our series Id4 positivity was statistically higher in patients with TNBC and Jewish ethnicity, the latter group represents only a fifth of all our cases, whereas about 75 % of all TNBC were Id4 positive. BRCA1 (and BRCA2) mutations are identified in less than 10 % of all patients with breast cancer [55–57], and BRCA germline mutations carriers account for only about 11–12 % of young patients with TNBC [58, 59]. Based on these data, it is reasonable to assume that most of the patients in our series were not BRCA1 (or BRCA2) germline mutation carriers. Although it would be interesting to know the distribution of Id4 in TNBCs occurring in BRCA1 germline mutation carriers, the validity of our data documenting overexpression of Id4 in TNBCs remains unaltered.
Id4 positive TNBCs did not have worse clinical outcome than Id4 negative tumors. These data are consistent with the findings in two recent series, which have reported that the overall prognosis of TNBCs does not significantly differ in patients with or without BRCA-germline mutation [60, 61]. A recent study has reported high Id4 expression in the cancer stem cells of a mouse mammary cancer cell line. In the same study, knockdown of Id4 expression suppressed the stem cell properties [40]. The expression of stem cell markers is reported to be high in basal-like breast carcinoma compared to other breast cancer subtypes [62, 63]. Our finding of high Id4 expression in basal-like breast carcinoma links these observations. However, assuming that Id4 positivity in TNBC reflects a stem cell enriched population, the stem cell phenotype does not appear to predict clinical outcome, as we found no significant difference in overall survival and disease-free survival between Id4 positive and Id4 negative TNBCs.
In summary, our results document high expression of intranuclear Id4 protein in TNBCs, in contrast to most non-triple-negative tumors. These findings suggest that Id4 overexpression plays a role in the downregulation of BRCA1 in sporadic TNBCs of patients without BRCA1 germline mutation, and provide new insight into the biology of these tumors.
References
Dawson SJ, Provenzano E, Caldas C (2009) Triple negative breast cancers: clinical and prognostic implications. Eur J Cancer 45(Suppl 1):27–40
Lakhani SR, Van De Vijver MJ, Jacquemier J, Anderson TJ, Osin PP, McGuffog L, Easton DF (2002) The pathology of familial breast cancer: predictive value of immunohistochemical markers estrogen receptor, progesterone receptor, HER-2, and p53 in patients with mutations in BRCA1 and BRCA2. J Clin Oncol 20(9):2310–2318
Carey LA, Perou CM, Livasy CA, Dressler LG, Cowan D, Conway K, Karaca G, Troester MA, Tse CK, Edmiston S et al (2006) Race, breast cancer subtypes, and survival in the Carolina Breast Cancer Study. JAMA 295(21):2492–2502
Tischkowitz M, Brunet JS, Begin LR, Huntsman DG, Cheang MC, Akslen LA, Nielsen TO, Foulkes WD (2007) Use of immunohistochemical markers can refine prognosis in triple negative breast cancer. BMC Cancer 7:134
Rouzier R, Perou CM, Symmans WF, Ibrahim N, Cristofanilli M, Anderson K, Hess KR, Stec J, Ayers M, Wagner P et al (2005) Breast cancer molecular subtypes respond differently to preoperative chemotherapy. Clin Cancer Res 11(16):5678–5685
Carey LA, Dees EC, Sawyer L, Gatti L, Moore DT, Collichio F, Ollila DW, Sartor CI, Graham ML, Perou CM (2007) The triple negative paradox: primary tumor chemosensitivity of breast cancer subtypes. Clin Cancer Res 13(8):2329–2334
Rakha EA, El-Sayed ME, Green AR, Lee AH, Robertson JF, Ellis IO (2007) Prognostic markers in triple-negative breast cancer. Cancer 109(1):25–32
Rakha EA, Elsheikh SE, Aleskandarany MA, Habashi HO, Green AR, Powe DG, El-Sayed ME, Benhasouna A, Brunet JS, Akslen LA et al (2009) Triple-negative breast cancer: distinguishing between basal and nonbasal subtypes. Clin Cancer Res 15(7):2302–2310
Nielsen TO, Hsu FD, Jensen K, Cheang M, Karaca G, Hu Z, Hernandez-Boussard T, Livasy C, Cowan D, Dressler L et al (2004) Immunohistochemical and clinical characterization of the basal-like subtype of invasive breast carcinoma. Clin Cancer Res 10(16):5367–5374
Rakha EA, El-Sayed ME, Green AR, Paish EC, Lee AH, Ellis IO (2007) Breast carcinoma with basal differentiation: a proposal for pathology definition based on basal cytokeratin expression. Histopathology 50(4):434–438
Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A, Deng S, Johnsen H, Pesich R, Geisler S et al (2003) Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA 100(14):8418–8423
Turner NC, Reis-Filho JS, Russell AM, Springall RJ, Ryder K, Steele D, Savage K, Gillett CE, Schmitt FC, Ashworth A et al (2007) BRCA1 dysfunction in sporadic basal-like breast cancer. Oncogene 26(14):2126–2132
Rhiem K, Todt U, Wappenschmidt B, Klein A, Wardelmann E, Schmutzler RK (2010) Sporadic breast carcinomas with somatic BRCA1 gene deletions share genotype/phenotype features with familial breast carcinomas. Anticancer Res 30(9):3445–3449
Garcia AI, Buisson M, Bertrand P, Rimokh R, Rouleau E, Lopez BS, Lidereau R, Mikaelian I, Mazoyer S (2011) Down-regulation of BRCA1 expression by miR-146a and miR-146b-5p in triple negative sporadic breast cancers. EMBO Mol Med 3(5):279–290
Turner N, Tutt A, Ashworth A (2004) Hallmarks of ‘BRCAness’ in sporadic cancers. Nat Rev Cancer 4(10):814–819
Esteller M, Silva JM, Dominguez G, Bonilla F, Matias-Guiu X, Lerma E, Bussaglia E, Prat J, Harkes IC, Repasky EA et al (2000) Promoter hypermethylation and BRCA1 inactivation in sporadic breast and ovarian tumors. J Natl Cancer Inst 92(7):564–569
de Candia P, Benezra R, Solit DB (2004) A role for Id proteins in mammary gland physiology and tumorigenesis. Adv Cancer Res 92:81–94
Beger C, Pierce LN, Kruger M, Marcusson EG, Robbins JM, Welcsh P, Welch PJ, Welte K, King MC, Barber JR et al (2001) Identification of Id4 as a regulator of BRCA1 expression by using a ribozyme-library-based inverse genomics approach. Proc Natl Acad Sci USA 98(1):130–135
Heyn H, Engelmann M, Schreek S, Ahrens P, Lehmann U, Kreipe H, Schlegelberger B, Beger C (2011) MicroRNA miR-335 is crucial for the BRCA1 regulatory cascade in breast cancer development. Int J Cancer 129(12):2797–2806
Dell’Orso S, Ganci F, Strano S, Blandino G, Fontemaggi G (2010) ID4: a new player in the cancer arena. Oncotarget 1(1):48–58
Jeon HM, Sohn YW, Oh SY, Kim SH, Beck S, Kim S, Kim H (2011) ID4 imparts chemoresistance and cancer stemness to glioma cells by derepressing miR-9*-mediated suppression of SOX2. Cancer Res 71(9):3410–3421
Fontemaggi G, Dell’Orso S, Trisciuoglio D, Shay T, Melucci E, Fazi F, Terrenato I, Mottolese M, Muti P, Domany E et al (2009) The execution of the transcriptional axis mutant p53, E2F1 and ID4 promotes tumor neo-angiogenesis. Nat Struct Mol Biol 16(10):1086–1093
de Candia P, Akram M, Benezra R, Brogi E (2006) Id4 messenger RNA and estrogen receptor expression: inverse correlation in human normal breast epithelium and carcinoma. Hum Pathol 37(8):1032–1041
Hammond ME, Hayes DF, Dowsett M, Allred DC, Hagerty KL, Badve S, Fitzgibbons PL, Francis G, Goldstein NS, Hayes M et al (2010) American Society of Clinical Oncology/College Of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol 28(16):2784–2795
Wolff AC, Hammond ME, Schwartz JN, Hagerty KL, Allred DC, Cote RJ, Dowsett M, Fitzgibbons PL, Hanna WM, Langer A et al (2007) American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. J Clin Oncol 25(1):118–145
Kreike B, van Kouwenhove M, Horlings H, Weigelt B, Peterse H, Bartelink H, van de Vijver MJ (2007) Gene expression profiling and histopathological characterization of triple-negative/basal-like breast carcinomas. Breast Cancer Res 9(5):R65
Reis-Filho JS, Milanezi F, Steele D, Savage K, Simpson PT, Nesland JM, Pereira EM, Lakhani SR, Schmitt FC (2006) Metaplastic breast carcinomas are basal-like tumours. Histopathology 49(1):10–21
Dabbs DJ, Chivukula M, Carter G, Bhargava R (2006) Basal phenotype of ductal carcinoma in situ: recognition and immunohistologic profile. Mod Pathol 19(11):1506–1511
Tsuda H, Takarabe T, Hasegawa F, Fukutomi T, Hirohashi S (2000) Large, central acellular zones indicating myoepithelial tumor differentiation in high-grade invasive ductal carcinomas as markers of predisposition to lung and brain metastases. Am J Surg Pathol 24(2):197–202
Maiorano E, Regan MM, Viale G, Mastropasqua MG, Colleoni M, Castiglione-Gertsch M, Price KN, Gelber RD, Goldhirsch A, Coates AS (2010) Prognostic and predictive impact of central necrosis and fibrosis in early breast cancer: results from two International Breast Cancer Study Group randomized trials of chemoendocrine adjuvant therapy. Breast Cancer Res Treat 121(1):211–218
O’Malley FP, Bane AL (2004) The spectrum of apocrine lesions of the breast. Adv Anat Pathol 11(1):1–9
Livasy CA, Karaca G, Nanda R, Tretiakova MS, Olopade OI, Moore DT, Perou CM (2006) Phenotypic evaluation of the basal-like subtype of invasive breast carcinoma. Mod Pathol 19(2):264–271
Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS et al (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98(19):10869–10874
Nielsen MF, Caumo A, Chandramouli V, Schumann WC, Cobelli C, Landau BR, Vilstrup H, Rizza RA, Schmitz O (2004) Impaired basal glucose effectiveness but unaltered fasting glucose release and gluconeogenesis during short-term hypercortisolemia in healthy subjects. Am J Physiol Endocrinol Metab 286(1):E102–E110
Lakhani SR, Jacquemier J, Sloane JP, Gusterson BA, Anderson TJ, van de Vijver MJ, Farid LM, Venter D, Antoniou A, Storfer-Isser A et al (1998) Multifactorial analysis of differences between sporadic breast cancers and cancers involving BRCA1 and BRCA2 mutations. J Natl Cancer Inst 90(15):1138–1145
Jen Y, Manova K, Benezra R (1996) Expression patterns of Id1, Id2, and Id3 are highly related but distinct from that of Id4 during mouse embryogenesis. Dev Dyn 207(3):235–252
Lyden D, Young AZ, Zagzag D, Yan W, Gerald W, O’Reilly R, Bader BL, Hynes RO, Zhuang Y, Manova K et al (1999) Id1 and Id3 are required for neurogenesis, angiogenesis and vascularization of tumour xenografts. Nature 401(6754):670–677
Benezra R (2001) Role of Id proteins in embryonic and tumor angiogenesis. Trends Cardiovasc Med 11(6):237–241
Kuzontkoski PM, Mulligan-Kehoe MJ, Harris BT, Israel MA (2010) Inhibitor of DNA binding-4 promotes angiogenesis and growth of glioblastoma multiforme by elevating matrix GLA levels. Oncogene 29(26):3793–3802
Park SJ, Kim RJ, Nam JS (2011) Inhibitor of DNA-binding 4 contributes to the maintenance and expansion of cancer stem cells in 4T1 mouse mammary cancer cell line. Lab Anim Res 27(4):333–338
Kamalian L, Gosney JR, Forootan SS, Foster CS, Bao ZZ, Beesley C, Ke Y (2008) Increased expression of Id family proteins in small cell lung cancer and its prognostic significance. Clin Cancer Res 14(8):2318–2325
Zeng W, Rushing EJ, Hartmann DP, Azumi N (2010) Increased inhibitor of differentiation 4 (id4) expression in glioblastoma: a tissue microarray study. J Cancer 1:1–5
Venneti S, Le P, Martinez D, Xie SX, Sullivan LM, Rorke-Adams LB, Pawel B, Judkins AR (2011) Malignant rhabdoid tumors express stem cell factors, which relate to the expression of EZH2 and Id proteins. Am J Surg Pathol 35(10):1463–1472
Umetani N, Mori T, Koyanagi K, Shinozaki M, Kim J, Giuliano AE, Hoon DS (2005) Aberrant hypermethylation of ID4 gene promoter region increases risk of lymph node metastasis in T1 breast cancer. Oncogene 24(29):4721–4727
Noetzel E, Veeck J, Niederacher D, Galm O, Horn F, Hartmann A, Knuchel R, Dahl E (2008) Promoter methylation-associated loss of ID4 expression is a marker of tumour recurrence in human breast cancer. BMC Cancer 8:154
Chan AS, Tsui WY, Chen X, Chu KM, Chan TL, Li R, So S, Yuen ST, Leung SY (2003) Downregulation of ID4 by promoter hypermethylation in gastric adenocarcinoma. Oncogene 22(44):6946–6953
Umetani N, Takeuchi H, Fujimoto A, Shinozaki M, Bilchik AJ, Hoon DS (2004) Epigenetic inactivation of ID4 in colorectal carcinomas correlates with poor differentiation and unfavorable prognosis. Clin Cancer Res 10(22):7475–7483
Vinarskaja A, Goering W, Ingenwerth M, Schulz WA (2011) ID4 is frequently downregulated and partially hypermethylated in prostate cancer. World J Urol. doi:10.1007/s00345-011-0750-8
Yu L, Liu C, Vandeusen J, Becknell B, Dai Z, Wu YZ, Raval A, Liu TH, Ding W, Mao C et al (2005) Global assessment of promoter methylation in a mouse model of cancer identifies ID4 as a putative tumor-suppressor gene in human leukemia. Nat Genet 37(3):265–274
Hagiwara K, Nagai H, Li Y, Ohashi H, Hotta T, Saito H (2007) Frequent DNA methylation but not mutation of the ID4 gene in malignant lymphoma. J Clin Exp Hematop 47(1):15–18
Welcsh PL, Lee MK, Gonzalez-Hernandez RM, Black DJ, Mahadevappa M, Swisher EM, Warrington JA, King MC (2002) BRCA1 transcriptionally regulates genes involved in breast tumorigenesis. Proc Natl Acad Sci USA 99(11):7560–7565
Jimenez RE, Wallis T, Visscher DW (2001) Centrally necrotizing carcinomas of the breast: a distinct histologic subtype with aggressive clinical behavior. Am J Surg Pathol 25(3):331–337
Farmer P, Bonnefoi H, Becette V, Tubiana-Hulin M, Fumoleau P, Larsimont D, Macgrogan G, Bergh J, Cameron D, Goldstein D et al (2005) Identification of molecular apocrine breast tumours by microarray analysis. Oncogene 24(29):4660–4671
Vranic S, Tawfik O, Palazzo J, Bilalovic N, Eyzaguirre E, Lee LM, Adegboyega P, Hagenkord J, Gatalica Z (2010) EGFR and HER-2/neu expression in invasive apocrine carcinoma of the breast. Mod Pathol 23(5):644–653
Peto J, Collins N, Barfoot R, Seal S, Warren W, Rahman N, Easton DF, Evans C, Deacon J, Stratton MR (1999) Prevalence of BRCA1 and BRCA2 gene mutations in patients with early-onset breast cancer. J Natl Cancer Inst 91(11):943–949
Ford D, Easton DF, Peto J (1995) Estimates of the gene frequency of BRCA1 and its contribution to breast and ovarian cancer incidence. Am J Hum Genet 57(6):1457–1462
Robertson L, Hanson H, Seal S, Warren-Perry M, Hughes D, Howell I, Turnbull C, Houlston R, Shanley S, Butler S et al (2012) BRCA1 testing should be offered to individuals with triple-negative breast cancer diagnosed below 50 years. Br J Cancer 106(6):1234–1238
Young SR, Pilarski RT, Donenberg T, Shapiro C, Hammond LS, Miller J, Brooks KA, Cohen S, Tenenholz B, Desai D et al (2009) The prevalence of BRCA1 mutations among young women with triple-negative breast cancer. BMC Cancer 9:86
Evans DG, Howell A, Ward D, Lalloo F, Jones JL, Eccles DM (2011) Prevalence of BRCA1 and BRCA2 mutations in triple negative breast cancer. J Med Genet 48(8):520–522
Bayraktar S, Gutierrez-Barrera AM, Liu D, Tasbas T, Akar U, Litton JK, Lin E, Albarracin CT, Meric-Bernstam F, Gonzalez-Angulo AM et al (2011) Outcome of triple-negative breast cancer in patients with or without deleterious BRCA mutations. Breast Cancer Res Treat 130(1):145–153
Lee LJ, Alexander B, Schnitt SJ, Comander A, Gallagher B, Garber JE, Tung N (2011) Clinical outcome of triple negative breast cancer in BRCA1 mutation carriers and noncarriers. Cancer 117(14):3093–3100
Honeth G, Bendahl PO, Ringner M, Saal LH, Gruvberger-Saal SK, Lovgren K, Grabau D, Ferno M, Borg A, Hegardt C (2008) The CD44+/CD24− phenotype is enriched in basal-like breast tumors. Breast Cancer Res 10(3):R53
Park SY, Lee HE, Li H, Shipitsin M, Gelman R, Polyak K (2010) Heterogeneity for stem cell-related markers according to tumor subtype and histologic stage in breast cancer. Clin Cancer Res 16(3):876–887
Disclosure
The authors declare that neither pharmaceutical nor industry support was provided for this work. No funding for this project was received from any of the following organizations: National Institutes of Health (NIH); Wellcome Trust; Howard Hughes Medical Institute (HHMI); or other(s).
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wen, Y.H., Ho, A., Patil, S. et al. Id4 protein is highly expressed in triple-negative breast carcinomas: possible implications for BRCA1 downregulation. Breast Cancer Res Treat 135, 93–102 (2012). https://doi.org/10.1007/s10549-012-2070-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-012-2070-0