Abstract
Oncocytic carcinomas are composed of mitochondrion-rich cells. Though recognised by the WHO classification as a histological special type of breast cancer, their status as a discrete pathological entity remains a matter of contention. Given that oncocytic tumours of other anatomical sites display distinct clinico-pathological and molecular features, we sought to define the molecular genetic features of mitochondrion-rich breast tumours and to compare them with a series of histological grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type. Seventeen mitochondrion-rich breast carcinomas, including nine bona fide oncocytic carcinomas, were profiled with antibodies against oestrogen, progesterone and androgen receptors, HER2, Ki67, GCDFP-15, chromogranin, epithelial membrane antigen, cytokeratin 7, cytokeratin 14, CD68 and mitochondria antigen. These tumours were microdissected and DNA extracted from samples with >70% of tumour cells. Fourteen cases yielded DNA of sufficient quality/quantity and were subjected to high-resolution microarray comparative genomic hybridisation analysis. The genomic profiles were compared to those of 28 grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type. Oncocytic and other mitochondrion-rich tumours did not differ significantly between themselves. As a group, mitochondrion-rich carcinomas were immunophenotypically heterogenous. Recurrent copy number changes were similar to those described in unselected breast cancers. However, unsupervised and supervised analysis identified a subset of mitochondrion-rich cancers, which often displayed gains of 11q13.1-q13.2 and 19p13. Changes in the latter two chromosomal regions have been shown to be associated with oncocytic tumours of the kidney and thyroid, respectively, and host several nuclear genes with specific mitochondrial function. Our results indicate that in a way akin to oncocytic tumours of other anatomical sites, at least a subset of mitochondrion-rich breast carcinomas may be underpinned by a distinct pattern of chromosomal changes potentially relevant for mitochondria accumulation and constitute a discrete molecular entity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Accumulation of mitochondria resulting in mitochondrion-rich neoplastic cells and in oncocytic tumours has been reported in numerous anatomical sites, in particular thyroid, salivary glands and kidney. Abnormal accumulation of mitochondria confers to the neoplastic cells an abundant granular eosinophilic cytoplasm that is distinctive under the microscope. When the accumulation of mitochondria is widespread among the tumour cells, and generally accompanied by complete loss of their polarity due to the increased mitochondrial mass, a mitochondrion-rich tumour is called oncocytic (from the Greek word “to swell”) [1].
Oncocytic breast carcinomas have been reported [2–5] and the latest WHO classification of breast cancer recognises them as one of the histological special types of breast cancer, defining it as a breast carcinoma composed of >70% of oncocytes [6]. Early reports underscored the abundant cytoplasm rich in deeply eosinophilic granules in the tumour cells and their strong immunoreactivity to anti-mitochondrial antibodies [2, 3]. Electron microscopy of some samples confirmed the cytoplasmic accumulation of mitochondria [2, 3].
The scanty reports of oncocytic breast carcinomas [3, 4, 7] suggested that the frequency of mitochondria accumulation in breast carcinomas has been underestimated. In fact, we have recently shown that mitochondria accumulation is relatively common in breast carcinomas and that oncocytic breast carcinomas are far from rare [5]. The reasons for the failure to appreciate that mitochondria accumulation frequently occurs in breast carcinoma are varied. Among them is the common assumption that cells with abundant eosinophilic cytoplasm in breast carcinoma are the result of apocrine differentiation, while other breast tumours with eosinophilic granular cytoplasm fall into the neuroendocrine or acinic special histological types [5].
Mitochondrion-rich and oncocytic tumours of the thyroid, kidney, salivary and pituitary glands, liver, lung and rectum have been shown to harbour distinct clinico-pathological features, including sometimes different response to treatment (e.g. poor response to radioactive iodide treatment in thyroid oncocytic tumours), as well as different molecular profiles [8–25].
We hypothesised that mitochondrion-rich breast carcinomas would be characterised by a distinct constellation of molecular features. To address this question 17 mitochondrion-rich breast carcinomas, including seven cases that we have classified as bone fide oncocytic breast carcinomas in our recent report [5], were typed by immunohistochemistry and subjected to high-resolution microarray comparative genomic hybridisation. The aims of this study were (i) to define the chromosomal DNA alterations of mitochondrion-rich breast carcinomas (ii) to compare them with those of histological grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type.
Materials and methods
Case selection: oncocytic and mitochondrion-rich breast carcinomas
Seventeen cases, nine oncocytic and eight mitochondrion-rich breast carcinomas, were retrieved from the files of the Department of Haematology and Oncological Sciences at Bellaria Hospital, University of Bologna, Italy. All cases constitute part of a larger cohort described by our group in a separate publication with specific criteria for the definition of mitochondrion-rich and oncoytic breast carcinoma (see Ref. [5] and Supplementary Material and Methods). Histological grade was assessed on all cases according to the Nottingham grading system [26]. The study was approved by the local ethical committees.
Immunohistochemistry
Immunohistochemistry was performed using standardised automated procedures (Ventana-Benchmark, Tucson, Arizona, USA with Ultravision LP Detection System HRP Polymer—LabVision, Freemont, CA, USA) as previously described [5] with antibodies raised against oestrogen receptor, progesterone receptor, HER2, androgen receptor, Ki67, GCDFP-15, chromogranin, EMA, cytokeratin (CK) 7, CK14, CD68 and mitochondria (see Supplementary Table 1).
Positive and negative (omission of primary antibody and use of a IgG-matched immune serum) controls were included in each slide run. Immunohistochemical scoring was performed semiquantitatively according to previously published criteria (see Refs. [5, 27, 28] and Supplementary Material and Methods).
Microdissection and DNA extraction
All tumours, including cases and controls, were microdissected to ensure >70% of purity of neoplastic cells. Microdissection was performed as previously described [29]. DNA was extracted using the DNeasy Kit (Qiagen Ltd, Crawley, UK) according to the manufacturer’s recommendations. DNA concentration was measured with the PicoGreen® assay as per the manufacturer’s instructions (Invitrogen, Paisley, UK) [30].
Controls: invasive ductal carcinomas of no special type
From the series of 17 cases, only 14 (seven oncocytic and seven mitochondrion-rich carcinomas) cases provided DNA of enough quality/quantity to perform aCGH. As there are several lines of evidence that histological grade and oestrogen receptor (ER) status are significantly associated with the complexity of the genomes of breast cancers [29, 31–37], a series of 28 grade- and ER status-matched invasive ductal carcinomas of no special type (IDC-NST) was retrieved from the files of the Molecular Pathology of the Breakthrough Breast Cancer Research Centre as control tumours. All cases were reviewed by two of the authors (FCG and JSR-F) to ensure the absence of oncocytic features.
Microarray comparative genomic hybridisation (aCGH)
The aCGH platform used for this study was constructed at the Breakthrough Breast Cancer Research Centre and comprises ~32,000 BAC clones tiled across the genome. This type of BAC array platform has been shown to be as robust as, and to have comparable resolution with high density oligonucleotide arrays [38–40]. Labelling, hybridisation, washes, image acquisition and aCGH data analysis were carried out as previously described (see Refs. [29, 30, 34, 37, 41–45] and Supplementary Material and Methods). Low level gain was defined as a cbs-smoothed Log2 ratio of between 0.12 and 0.45, corresponding to approximately 3–5 copies of the locus, whilst gene amplification was defined as having a log2 ratio >0.45, corresponding to more than 5 copies. Hierarchical cluster analysis was performed as previously described [37].
Results
Histopathological features of oncocytic and mitochondrion-rich breast carcinomas
The histopathological and clinical features of the 17 cases included in this study are summarised in Table 1 and illustrated in Fig. 1. Detailed morphological and survival analyses were performed and recently described in a separate publication [5]. Briefly, oncocytic and mitochondrion-rich breast carcinomas preferentially displayed pushing borders and were composed of oncocytic cells arranged in solid sheets and nests (Fig. 1a, b). The nuclei tended to be irregular and pleomorphic, often with discrete nucleoli (Fig. 1c). According to our recent report [5], we defined mitochondrion-rich breast carcinomas those tumours with strong positivity for the anti-mitochondrial antibody in 30% to 70% of the tumour cells or moderate positivity in more than 50% of them; mitochondrion-rich tumours showing strong positivity for the anti-mitochondrial antibody in at least 70% of the tumour cells were considered bona fide oncocytic breast carcinomas (Fig. 1d; Table 2). Oncocytic and non-oncocytic mitochondrion-rich breast carcinomas did not differ significantly between themselves apart from the stronger cytoplasmic eosinophilia and more widespread positivity for the anti-mitochondrial antibody observed in the oncocytic tumours [5]. Despite the original observations suggesting that oncocytic carcinomas would have low grade nuclear features [3], the majority of the tumours analysed here were of histological grade 3 (71%) and none was classified as of histological grade 1.
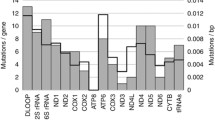
Morphological, immunohistochemical and genetic features of oncocytic and mitochondrion-rich breast carcinomas. Representative micrographs of histological sections (a–c) and immunohistochemistry with anti-mitochondrial antibody (d) of an oncocytic carcinoma. Frequency plots of chromosomal gains and losses (e) and amplifications (f) in 14 mitochondrion-rich breast carcinomas. In e, the proportion of tumours in which each clone is gained (green bars) or lost (red bars) is plotted (Y-axis) for each BAC clone according to genomic location (X-axis). In f, the proportion of tumours in which each clone is amplified (green bars) is plotted (Y-axis) for each BAC clone according to genomic location (X-axis). (Color figure online)
Immunophenotype of oncocytic and mitochondrion-rich breast carcinomas
Results of the immunohistochemistry are summarised in Table 2. No significant differences were observed between the oncocytic and the other mitochondrion-rich breast carcinomas in relation to their immunophenotype (data not shown). As a group, the seventeen cases were predominantly positive for hormone receptors: 76% were oestrogen receptor-positive and 47% progesterone receptor-positive. HER2 3+ immunohistochemical overexpression was found in 29%; all of these cases harboured HER2 gene amplification as defined by microarray comparative genomic hybridisation. According to the three major molecular subtypes of breast cancer described by Carey et al. [46], 65% of mitochondrion-rich breast carcinomas belonged to the luminal, 29% to the HER2 and 6% to the triple negative phenotype.
Genomic profiling of oncocytic and mitochondrion-rich breast carcinomas
Out of the 17 cases included in this study, aCGH was performed in seven oncocytic and seven mitochondrion-rich breast carcinomas. Three cases were excluded due to poor quality or insufficient DNA. Similar to what observed at the morphological and immunohistochemical levels [5], no significant differences were detected when comparing the genomic profiles of oncocytic and non-oncocytic mitochondrion-rich carcinomas (data not shown). Therefore, the genomic analysis of the bona fide oncocytic and of the other mitochondrion-rich carcinomas will be described together as a single entity under the name of mitochondrion-rich breast carcinomas.
Accordingly to the diversity observed at the immunophenotypical level, our aCGH analysis revealed that all mitochondrion-rich breast carcinomas are also heterogeneous at the genetic level. A varying level of genetic instability was identified, with an average of 18.61% of the genome showing either losses, gains or amplifications (range 9.65–34.32%). In agreement with a high prevalence of tumours of histological grade 3, the majority (78%) of mitochondrion-rich carcinomas displayed a complex architectural pattern of genomic aberrations as defined by Hicks et al. [47]. Regions of recurrent gains, losses and amplifications are shown in Fig. 1e, f, and summarised in Supplementary Table 2. The most frequent (i.e. ≥50%) copy number aberrations included gains of 1q, 5p, 8q, 11q, 16p and 17q and losses of 1p, 8p, 11q-qter, 16q, 17p and 22q and were in a way similar to those previously described in unselected breast carcinomas of high histological grade [33, 34]. The most frequent amplification was at 17q12-q21.2, encompassing HER2 amplicon and present in four out of 14 (29%) cases. Additional recurrent amplifications (i.e., in 2 or more cases) were found on 17q23, 17q25, 8p12-p11.22 and 8q22.3-q24.3.
Comparison with histological grade and oestrogen receptor status-matched invasive ductal carcinomas of no special type
The genomic profiles of 14 mitochondrion-rich breast carcinomas were compared with those of 28 grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type used as controls. Mitochondrion-rich carcinomas displayed a lower proportion of the genome altered by losses, gains or amplifications, however, this difference did not reach statistical significance (18.61% mitochondrion-rich carcinomas vs. 23.47% invasive ductal carcinomas of no special type, heteroscedastic t-test two-tailed P value = 0.164). Unsupervised hierarchical cluster analysis based on categorical aCGH data derived from 31,157 BACs revealed that a subset of the 14 mitochondrion-rich carcinomas consistently clustered together (Fig. 2). This mitochondrion-rich breast carcinomas cluster included eight mitochondrion-rich tumours and two invasive ductal carcinomas of no special type (IDC-NST5 and IDC-NST20 (Fig. 2) and was significantly enriched for carcinomas with a mitochondrion-rich phenotype (Fisher’s exact test, two-tailed P value < 0.001).
Unsupervised hierarchical cluster analysis of mitochondrion-rich breast carcinomas and grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type. Dendogram and heatmap of hierarchical clustering analysis using aCGH categorical data derived from 31,157 BACs including 14 mitochondrion-rich breast carcinomas and 28 grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type. Rows: cbs-log2 ratios categorised as gains (red), losses (blue) and no change (white) for each BAC clone in genomic order. Note the presence of a mitochondrion-rich breast carcinoma-enriched cluster composed of eight mitochondrion-rich breast carcinomas and two invasive ductal carcinomas of no special type. (Color figure online)
We next retrieved a list of 694 genes from the most complete database of nuclear genes codifying for proteins with mitochondrial function (http://www.mitop.de:8080/mitop2/startHuman.do;jsessionid=3F1FAB85FAD7C67F2125FF55A0F11422; Supplementary Table 3) and the categorical aCGH data for each one of these genes in each tumour. These categorical data was subjected to hierarchical clustering analysis using the same parameters and clustering methods as above. A cluster significantly enriched for mitochondrion-rich breast carcinomas was also identified and composed of six out of the eight mitochondrion-rich carcinomas and of a case of invasive ductal carcinomas of no special type (IDC-NST20) which previously clustered together (Fisher’s exact test, two-tailed P value = 0.003; Supplementary Figure 1).
It is important to note that tumours pertaining to the mitochondrion-rich cluster—either using all probes (n = 10; eight mitochondrion-rich, four of which oncocytic, and two invasive ductal carcinomas of no special type, Fig. 2) or probes mapping to the mitochondria function-related genes (n = 7; six mitochondrion-rich, three of which oncocytic, and one invasive ductal carcinomas of no special type, Supplementary Figure 1)—did not otherwise differ from all remaining tumours (n = 32 and n = 35, respectively) in terms of their clinico-pathological parameters, including those associated with genomic aberration patterns in breast cancer, i.e. histological grade, ER and HER2 status [31–35] (Fisher’s exact test, two-tailed P value > 0.05; Supplementary Table 4). Thus, these results provide strong circumstantial evidence that a subset of mitochondrion-rich breast carcinomas displays a distinct pattern of genetic aberrations.
Multi-Fisher’s exact text adjusted for false discovery revealed the regions differentially lost, gained or amplified between 14 mitochondrion-rich breast carcinomas and 28 grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type (Fig. 3 and Supplementary Table 5a and 5b). Among the regions significantly more frequently gained in mitochondrion-rich carcinomas were 5p13.33, 11q13.1-13.2, 17q25.3, 16p13.3 and 19p13 (multi-Fisher’s exact test, adjusted P value < 0.05; Fig. 3a). Although 11q13.1-13.2 was preferentially gained in mitochondrion-rich carcinomas, amplification of a downstream region on 11q (i.e. 11q13.3 encompassing CCND1) was significantly more prevalent in grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type (Fig. 3b). Surprisingly no mitochondrion-rich breast carcinoma displayed amplification of CCND1, an oncogene frequently amplified in oestrogen receptor-positive tumours [34, 48, 49]. Therefore, although the majority (76%) of mitochondrion-rich breast carcinomas express oestrogen receptor, they are not characterised by some of the changes typically found in this subgroup of breast tumours.
Comparative analysis of the prevalence of chromosomal gains, losses and amplifications in mitochondrion-rich breast carcinomas and grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type. a Gains and losses: top frequency plot of gains and losses in 14 mitochondrion-rich breast carcinomas, middle frequency plot of gains and losses in 28 grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type, bottom multi-Fisher’s comparisons between 14 mitochondrion-rich breast carcinomas and 28 grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type. b Amplifications: top frequency plot of amplifications in 14 mitochondrion-rich breast carcinomas, middle frequency plot of amplifications in 28 grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type, bottom multi-Fisher’s comparisons between 14 mitochondrion-rich breast carcinomas and 28 grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type. In frequency plots, the proportion of tumours in which each clone is gained (green bars), lost (red bars) or amplified (green bars) is plotted (Y-axis) for each BAC clone according to genomic location (X-axis). Multi-Fisher’s exact tests were performed with cbs log2 ratios for each clone, and those with an adjusted P-value of less than 0.05 are plotted (inverse log10, y-axis) according to genomic location (x-axis). (Color figure online)
Mitochondrion-rich cluster tumours harbour distinct genetic aberrations
Given that unsupervised cluster analysis revealed that a subset of invasive carcinomas enriched for tumours with mitochondrion-rich phenotype consistently clustered together independently of histological type or of other clinico-pathological features, we aimed to identify the constellation of copy number changes underpinning this group of tumours. For clarity we will hereafter refer to the eight mitochondrion-rich tumours that clustered together shown in Fig. 2 as the “mitochondrion-rich cluster”. We first compared the genomic profiles of this mitochondrion-rich cluster with the profiles of all remaining tumours included in this study (n = 34, six mitochondrion-rich carcinomas not belonging to the mitochondrion-rich cluster and the 28 invasive ductal carcinomas controls). The mitochondrion-rich cluster displayed a significantly lower proportion of the genome altered by copy number changes (12.00% mitochondrion-rich cluster vs. 24.16% remaining tumours, heteroscedastic t-test two-tailed P value = 0.002). Interestingly, the 11q13.1-13.2 region was gained in all but one samples of the mitochondrion-rich cluster. Multi-Fisher’s exact text adjusted for false discovery detected similar differences to the ones described above, such as gains of 11q13.1-13.2, 17q25.3, 16p13.3 and 19p13.3 as significantly associated with the mitochondrion-rich cluster (Multi-Fisher’s exact test, adjusted P value < 0.05; Fig. 4a, b and Supplementary Table 6a and 6b).
Comparative analysis of the prevalence of chromosomal gains, losses and amplifications in the mitochondrion-rich cluster tumours and all remaining tumours, and grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type. a Gains and losses in the mitochondrion-rich cluster (n = 8 mitochondrion-rich breast carcinomas) and all remaining tumours (n = 6 mitochondrion-rich breast carcinomas and 28 grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type): top frequency plot of gains and losses in the mitochondrion-rich cluster, middle frequency plot of gains and losses in all remaining tumours, bottom multi-Fisher’s comparisons between the mitochondrion-rich cluster and all remaining tumours. b Amplifications in the mitochondrion-rich cluster and all remaining tumours: top frequency plot of amplifications in the mitochondrion-rich cluster, middle frequency plot of amplifications in all remaining tumours, bottom multi-Fisher’s comparisons between the mitochondrion-rich cluster and all remaining tumours. c Gains and losses in the mitochondrion-rich cluster and 16 grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type: top frequency plot of gains and losses in the mitochondrion-rich cluster, middle frequency plot of gains and losses in 16 grade- and oestrogen receptor status-matched invasive ductal carcinomas of no special type, bottom multi-Fisher’s comparisons between the mitochondrion-rich cluster and 16 grade and oestrogen receptor status-matched invasive ductal carcinomas of no special type. d Amplifications the mitochondrion-rich cluster and 16 grade and oestrogen receptor status-matched invasive ductal carcinomas of no special type: top frequency plot of amplifications in the mitochondrion-rich cluster, middle frequency plot of amplifications in 16 grade and oestrogen receptor status-matched invasive ductal carcinomas of no special type, bottom multi-Fisher’s comparisons between mitochondrion-rich cluster and 16 grade and oestrogen receptor status-matched invasive ductal carcinomas of no special type. In frequency plots, the proportion of tumours in which each clone is gained (green bars), lost (red bars) or amplified (green bars) is plotted (Y-axis) for each BAC clone according to genomic location (X-axis). Multi-Fisher’s exact tests were performed with cbs log2 ratios for each clone, and those with an adjusted P value of less than 0.05 are plotted (inverse log10, y-axis) according to genomic location (x-axis). (Color figure online)
We next tested whether the genomic profiles of the mitochondrion-rich cluster would differ from those of 16 invasive ductal carcinomas of no special type selected among the 28 controls to match the grade and the oestrogen receptor status of the eight mitochondrion-rich cluster tumours. In agreement with the previous analysis, aCGH revealed a significantly lower proportion of the genome altered by copy number changes in the mitochondrion-rich cluster than in grade- and oestrogen status-matched invasive ductal carcinomas of no special type (12.00% mitochondrion-rich cluster vs. 27.42% invasive ductal carcinomas of no special type, heteroscedastic t-test two-tailed P value = 0.017). Akin to the results of the comparison between mitochondrion-rich cluster tumours and all remaining cancers, comparison between mitochondrion-rich cluster tumours and grade- and oestrogen status-matched invasive ductal carcinomas of no special type revealed gains of 11q13.1-13.2, 17q25.3, 16p13.3 and 19p13.3 to be significantly more prevalent in the mit-rich cluster (multi-Fisher’s exact test, adjusted P value < 0.05; Fig. 4c, d and Supplementary Table 7a and 7b).
To further determine the characteristics of the tumours from the mitochondrion-rich cluster we compared the clinico-pathological, immunophenotypic and genetic features of the mitochondrion-rich cluster (8 tumours) with the mitochondrion-rich carcinomas that did not cluster with them (n = 6). No significant difference, including the level of reactivity to anti-mitochondria antibody, was found between the two groups of tumours (Fisher’s exact test, two-tailed P value > 0.05; Supplementary Table 8). Nevertheless, and in spite of the limited sample size, the mitochondrion-rich cluster still displayed a significantly lower proportion of the genome harbouring changes (12.00% mitochondrion-rich cluster vs. 27.42% remaining mitochondrion-rich breast carcinomas, heteroscedastic t-test two-tailed P value < 0.001) and significant regions of differential copy number changes were identified, which in part overlapped with the differences described above (Multi-Fisher’s exact test, adjusted P value < 0.05; Supplementary Figure 2 and Supplementary Table 9a and 9b). For instance, the mitochondrion-rich cluster harboured more often gains of 11q13.1-q13.2 and less frequently displayed losses of 16q12-q24.
Nuclear genes with mitochondrial function localised in the differentially altered chromosomal DNA regions of mitochondrion-rich carcinomas and of the mitochondrion-rich cluster subgroup
Taken together, the above results demonstrate that at least a subgroup of mitochondrion-rich breast carcinomas appears to be distinct from grade and oestrogen receptor status-matched invasive ductal carcinomas of no special type in terms of their patterns of copy number aberrations. We identified several nuclear genes with mitochondrial function according to current databases (http://www.mitop.de:8080/mitop2/startHuman.do;jsessionid=3F1FAB85FAD7C67F2125FF55A0F11422; Supplementary Table 3, see above) in the chromosomal DNA regions differentially altered in mitochondrion-rich carcinomas and in the mitochondrion-rich cluster subgroup. The proteins coded by these genes include components of the respiratory chain (NDUFS8, SDHA, ATP5I, NDUFV1, ATP5D, NDUFS7, UQCR, COX4NB, COX4I1), mitochondrial ribosomal proteins (MRPL11, MRPL12, MRPL14, MRPL49, MRPL54), enzymes (PRDX5, BDH1, PC, PYCR1, ACP6), proteins with a role in regulation of the mitochondrial genome (TK2, DDX28, NUDT1, SIRT3, NUDT8) or of mitochondrial structure (FIBP, SMCP, GPX4), transporter and carrier proteins (CCS, TRAP1, SLC25A10, PDZK1, SLC25A22) as well as proteins involved in apoptosis (DNAJA3, GLRX2, MRPL41, BAD, TERT). There was no clear prevalence of gene groups with specific functional roles (Supplementary Table 10).
Discussion
We present the first molecular immunophenotypic and high-resolution genetic analysis of mitochondrion-rich breast carcinomas, including seven oncocytic tumours. Our data indicate that they display immunophenotypic chromosomal DNA patterns similar to those described in unselected breast carcinomas.
However, differentially altered chromosomal DNA regions compared with matched controls were identified for both the fourteen mitochondrion-rich carcinomas and for a subgroup of eight of them that clustered together after both unsupervised and supervised analyses.
A region commonly gained in mitochondrion-rich breast carcinoma but not previously linked to a mitochondrion-rich/oncoytic phenotype is 5p13.33 where Telomerase Reverse Transcriptase (TERT, the catalytic core of telomerase) has been mapped. This finding is particularly intriguing, in light of the recently established positive link between telomere maintenance, TERT and mitochondrial biogenesis and function. In fact TERT can localise to the mitochondrion where it modulates its activity [50].
The differentially altered chromosomal regions host a number nuclear genes with established mitochondrial function, such as SDHA, also localised to 5p13.33. SDHA codes for subunit A of complex II of the respiratory chain (succinate dehydrogenase). Although SDHA is commonly gained in our mitochondria-rich carcinomas, SDH is felt to act as a tumour suppressor because its loss induces pseudohypoxia. In fact, inactivating mutations of SDHB, SDHC, SDHD (subunits B, C, D of succinate dehydrogenase), and rarely SDHA mutations, cause hereditary paragangliomas and pheochromocytomas [51].
Remarkably, a region selectively lost in mitochondrion-rich breast carcinoma is 16q21 where the mitochondrial isoform of Timidine Kinase (TK2) has been mapped. Defects in TK2 gene have been associated to a myopathic mitochondrial DNA depletion syndrome, characterised by a reduction in copy number of mitochondrial DNA [52].
The mitochondrion-rich breast carcinoma cluster tumours differed at the genetic level from the remaining mitochondrion-rich breast carcinomas and from matched non-mitochondrion-rich control cases. This subset displayed lower levels of genetic instability and more frequently harboured focal gains of 11q13.1-13.2 and 19p13, which were present in 88 and 63% of the cases, respectively.
These two genomic regions (i.e. 11q13.1-13.2 and 19p13) harbour genes that encode proteins with a direct mitochondrial function (10 and 5 genes, respectively). For instance, overexpression of peroxiredoxin-5 (PRDX5), which maps to 11q13.1-13.2 and encodes a member of the peroxiredoxin family of anti-oxidant enzymes, has been associated with worse prognosis in breast cancers [53] and resistance to oxidative stress [54]. Interestingly, overexpression of PRDX1, another member of the same family, has been demonstrated in oncocytic tumours of the salivary glands, suggesting that detoxifying mechanisms may be up-regulated in these neoplasms likely carrying mitochondrial dysfunctions and electrons leakage [55]. It is plausible to think that the overexpression of anti-oxidant enzymes, observed to be differentially regulated according to mitochondrial damage [56], may exert a protective effect against reactive oxygen species and subsequent chromosomal DNA damage, resulting in the reduced genetic instability observed in our mitochondrion-rich breast carcinomas.
Gains of the 11q13.1-13.2 and 19p13 regions may in part explain the mitochondria accumulation in a subset of mitochondrion-rich breast carcinomas. Several lines of evidence indicate that these two regions have a specific etiological role for oncocytic tumours of other anatomical sites. Translocations between 11q13 and other chromosomal regions (e.g. t(5;11)(q35;q13); t(6;11)(p21;q13); t(7;11)(q11.2;q13)) appear to define a subset of renal oncocytomas [11, 12, 20, 24] and suggest that activation of genes mapping to 11q13 is potentially responsible for the oncogenic properties of these translocations. Among the genes mapping to 11q13 is the Cyclin D1 gene (CCND1) that could be the target of these rearrangements. However, not all renal oncocytomas harbouring 11q13 rearrangements display Cyclin D1 overexpression and the breakpoints most likely lie outside the CCND1 major translocation cluster breakpoint regions [11]. Interestingly, CCND1 was not amplified in our mitochondrion-rich tumours, in spite of its frequent amplification in breast carcinomas [34, 48, 49] and in our control cases. Although CCND1 was not amplified in our mitochondrion-rich carcinomas, CCDN1 expression may be altered in breast carcinoma by other mechanisms such as post-transcriptional chimeric CCDN1 mRNA [57]. Mitochondria are well known to play a key role in apoptosis. Among the chromosomal DNA regions significantly altered in our mitochondrion-rich carcinoma cluster was BAD (11q13.1), one of the main genes that control apoptosis. The 11q13.1 region was gained and since BAD antagonizes Bcl2 this finding suggests that the apoptosis pathway may be deregulated in our cases.
Thyroid oncocytic tumours display gains of 19p in up to 46% of the malignant cases [21] and have been associated with somatic missense mutations of GRIM19 [18] which maps to 19p13.11, a region close to the TCO (thyroid tumour with cell oxyphilia) locus that has been linked to familial thyroid tumours with oncocytic change [15]. Our findings on mitochondria-rich breast carcinoma underscore the relevance of 19p13 for the oncoytic phenotype.
Molecular explanations to account for the aberrant accumulation of mitochondria and resulting in the oncocytic appearance of tumour cells in other anatomical sites have been proposed by several studies. These include genetic changes of the nuclear [12, 18, 20, 21, 24] and mitochondrial [8, 14, 16–19] DNA. Particularly relevant are genetic alterations that cause disassembly of Complex I of the mitochondrial respiratory chain [8, 13, 14, 56, 58].
Given that a subgroup of mitochondrion-rich breast carcinomas appeared to harbour a distinctive pattern of chromosomal DNA aberrations we hypothesised that, in a way akin to oncocytic tumours of other anatomical sites, the mitochondrion-rich/oncocytic phenotype may depend in this tumour subgroup on genetic events affecting nuclear DNA. In the remaining mitochondrion-rich breast carcinomas the accumulation of mitochondria may be primarily due to the mitochondrial DNA alterations mentioned above.
We therefore attempted to test mitochondrial DNA alterations in all our mitochondrion-rich breast carcinomas but only in six cases we could amplify more than 10% of the mitochondrial DNA genome. In one case (case 11) there was a Thr to Ile substitution in the ND3 gene predicted to impair oxidative phosphorylation and—as in the case of many oncocytic tumours of other sites—belonging to Complex I. In the other cases sequencing data was too incomplete to allow any conclusion (Supplementary Table 11).
To the best of our knowledge this is the first report to show that high-resolution microarray comparative genomic hybridisation analysis associates specific regions of nuclear DNA unbalance with aberrant mitochondria accumulation in tumour cells. Given the limited sample size, our study should be perceived as exploratory and hypothesis generating. Further studies with larger cohorts are needed to confirm our findings and determine the potential clinical and biological implications of the identification of this special type of breast cancer. The mitochondrion-rich subset of carcinomas clustering together displayed higher nuclear DNA stability. If a lower genetic instability is considered as a parameter for a slower cell division and subsequent damage accumulation, one would actually expect a less aggressive behaviour for these cases. In fact, statistical analysis by our group has not shown a worse survival for mitochondrion-rich breast carcinomas, despite the common occurrence of unfavourable prognostic features (e.g. HER2 positivity, grade 3 tumours) [5]. It should be noted that some oncocytic tumours of other anatomical sites may be resistant to radiotherapy [22] and this may be considered when making treatment decisions.
As for the biological implications, our data underscore the relevance of nuclear DNA alterations for the mitochondrial biogenesis of tumour cells and the development of a mitochondria-rich/oncoytic phenotype. The relationship between the molecular changes leading to accumulation of mitochondria and neoplastic transformation is still largely obscure. It has nevertheless become recently clear that a common genetic feature of oncocytic tumours, regardless of their anatomical site, is the accumulation of high (homoplasmic or near homoplasmic) levels of biochemically damaging mutations within the mitochondrial DNA leading to a compensatory increase of the mitochondrial mass [8, 13, 14, 56, 58]. Particularly important are the alterations that affect Complex I of the mitochondrial respiratory chain [8, 13, 14, 56, 58]. Mitochondrial DNA mutations, particularly those involving Complex I, are well known to occur in many human tumours. For unclear reasons these mutations are sometimes selected for and accumulate in the mitochondrial genome of neoplastic cells of tumours that usually arise in organs with high metabolic activity, like endocrine glands (thyroid parathyroid, pituitary), kidney or salivary gland. The mutations result in a profound defect in oxidative phosphorylation but the defective neoplastic cells are unable to overcome the resulting pseudohypoxic condition by stabilizing HIF1a. They respond with the mitochondrial mass increase that defines the oncoytic phenotype [8, 13, 14, 56, 58].
Our data supports the concept that mitochondria may pathologically accumulate as a result of nuclear DNA alterations, as well as of specific mitochondrial DNA changes, underlining the complex relationship between energy production, pseudohypoxia, mitochondrial function and tumour development [13, 18, 23, 56, 58]. It is also possible that nuclear DNA changes may influence or modulate the accumulation of mitochondria in tumours that have already acquired high level (e.g. homoplasmic) mitochondrial DNA mutations, while the nucleus may hold the answer in those cases in which the occurrence of the oncocytic phenotype can not be linked to mitochondrial DNA mutations.
In conclusion, mitochondrion-rich breast carcinomas are a group of tumours heterogeneous at the immunophenotypic level. At the genetic level, a subgroup of them has a distinctive pattern of chromosomal DNA aberrations detected by high-resolution array CGH and seems to constitute a discrete molecular entity. Importantly, this subgroup displays chromosomal changes in regions previously associated with oncocytic tumours of other anatomical sites that encode for nuclear genes relevant to mitochondrial function. These alterations may be important for the accumulation of mitochondria and determine the biologic features of the tumour. Further studies are warranted to determine the specific molecular events involved in the development of mitochondrion-rich breast cancers, a previously under-recognised group of breast carcinoma.
References
Tallini G (1998) Oncocytic tumours. Virchows Arch 433(1):5–12
Costa MJ, Silverberg SG (1989) Oncocytic carcinoma of the male breast. Arch Pathol Lab Med 113(12):1396–1399
Damiani S, Eusebi V, Losi L, D’Adda T, Rosai J (1998) Oncocytic carcinoma (malignant oncocytoma) of the breast. Am J Surg Pathol 22(2):221–230
Hamperl H (1972) Oncocyten und hyaline einschlusse in der menschlischen mamma. Virchows Arch 10:88–92
Ragazzi M, de Biase D, Betts CM, Farnedi A, Ramadan SS, Tallini G, Reis-Filho JS, Eusebi V (2010) Oncocytic carcinoma of the breast: frequency, morphology and follow up. Human Pathol 42(2):166–175
Tavassoli FA, Devilee P (eds) (2003) Tumours of the breast. Pathology and genetics. Tumours of the breast and female genital organs. International Agency for Research of Cancer (IARC), Lyon
Damiani S, Dina R, Eusebi V (1999) Eosinophilic and granular cell tumors of the breast. Semin Diagn Pathol 16(2):117–125
Gasparre G, Porcelli AM, Bonora E, Pennisi LF, Toller M, Iommarini L, Ghelli A, Moretti M, Betts CM, Martinelli GN, Ceroni AR, Curcio F, Carelli V, Rugolo M, Tallini G, Romeo G (2007) Disruptive mitochondrial DNA mutations in complex I subunits are markers of oncocytic phenotype in thyroid tumors. Proc Natl Acad Sci USA 104(21):9001–9006
Solis LM, Raso MG, Kalhor N, Behrens C, Wistuba II, Moran CA (2010) Primary oncocytic adenocarcinomas of the lung: a clinicopathologic, immunohistochemical, and molecular biologic analysis of 16 cases. Am J Clin Pathol 133(1):133–140
Sheu SY, Handke S, Brocker-Preuss M, Gorges R, Frey UH, Ensinger C, Ofner D, Farid NR, Siffert W, Schmid KW (2007) The C allele of the GNB3 C825T polymorphism of the G protein beta3-subunit is associated with an increased risk for the development of oncocytic thyroid tumours. J Pathol 211(1):60–66
Jhang JS, Narayan G, Murty VV, Mansukhani MM (2004) Renal oncocytomas with 11q13 rearrangements: cytogenetic, molecular, and immunohistochemical analysis of cyclin D1. Cancer Genet Cytogenet 149(2):114–119
Sukov WR, Ketterling RP, Lager DJ, Carlson AW, Sinnwell JP, Chow GK, Jenkins RB, Cheville JC (2009) CCND1 rearrangements and cyclin D1 overexpression in renal oncocytomas: frequency, clinicopathologic features, and utility in differentiation from chromophobe renal cell carcinoma. Hum Pathol 40(9):1296–1303
Gasparre G, Bonora E, Tallini G, Romeo G (2010) Molecular features of thyroid oncocytic tumors. Mol Cell Endocrinol 321(1):67–76
Bonora E, Porcelli AM, Gasparre G, Biondi A, Ghelli A, Carelli V, Baracca A, Tallini G, Martinuzzi A, Lenaz G, Rugolo M, Romeo G (2006) Defective oxidative phosphorylation in thyroid oncocytic carcinoma is associated with pathogenic mitochondrial DNA mutations affecting complexes I and III. Cancer Res 66(12):6087–6096
Canzian F, Amati P, Harach HR, Kraimps JL, Lesueur F, Barbier J, Levillain P, Romeo G, Bonneau D (1998) A gene predisposing to familial thyroid tumors with cell oxyphilia maps to chromosome 19p13.2. Am J Hum Genet 63(6):1743–1748
Lewis PD, Baxter P, Paul Griffiths A, Parry JM, Skibinski DO (2000) Detection of damage to the mitochondrial genome in the oncocytic cells of Warthin’s tumour. J Pathol 191(3):274–281
Lewis PD, Baxter PW, Griffiths AP, Parry JM, Skibinski DO (2000) Authors’ reply. Mitochondrial DNA damage and oncocytic neoplasia. J Pathol 192(4):562–563
Maximo V, Botelho T, Capela J, Soares P, Lima J, Taveira A, Amaro T, Barbosa AP, Preto A, Harach HR, Williams D, Sobrinho-Simoes M (2005) Somatic and germline mutation in GRIM-19, a dual function gene involved in mitochondrial metabolism and cell death, is linked to mitochondrion-rich (Hurthle cell) tumours of the thyroid. Br J Cancer 92(10):1892–1898
Maximo V, Sobrinho-Simoes M (2000) Mitochondrial DNA ‘common’ deletion in Hurthle cell lesions of the thyroid. J Pathol 192(4):561–562
Neuhaus C, Dijkhuizen T, van den Berg E, Storkel S, Stockle M, Mensch B, Huber C, Decker HJ (1997) Involvement of the chromosomal region 11q13 in renal oncocytoma: case report and literature review. Cancer Genet Cytogenet 94(2):95–98
Wada N, Duh QY, Miura D, Brunaud L, Wong MG, Clark OH (2002) Chromosomal aberrations by comparative genomic hybridization in hurthle cell thyroid carcinomas are associated with tumor recurrence. J Clin Endocrinol Metab 87(10):4595–4601
Ambrosini-Spaltro A, Salvi F, Betts CM, Frezza GP, Piemontese A, Del Prete P, Baldoni C, Foschini MP, Viale G (2006) Oncocytic modifications in rectal adenocarcinomas after radio and chemotherapy. Virchows Arch 448(4):442–448
Sobrinho-Simoes M, Maximo V, Castro IV, Fonseca E, Soares P, Garcia-Rostan G, Oliveira MC (2005) Hurthle (oncocytic) cell tumors of thyroid: etiopathogenesis, diagnosis and clinical significance. Int J Surg Pathol 13(1):29–35
Sinke RJ, Dijkhuizen T, Janssen B, Olde Weghuis D, Merkx G, van den Berg E, Schuuring E, Meloni AM, de Jong B, Geurts van Kessel A (1997) Fine mapping of the human renal oncocytoma-associated translocation (5;11)(q35;q13) breakpoint. Cancer Genet Cytogenet 96(2):95–101
Maximo V, Soares P, Lima J, Cameselle-Teijeiro J, Sobrinho-Simoes M (2002) Mitochondrial DNA somatic mutations (point mutations and large deletions) and mitochondrial DNA variants in human thyroid pathology: a study with emphasis on Hurthle cell tumors. Am J Pathol 160(5):1857–1865. doi:10.1016/S0002-9440(10)61132-7
Elston CW, Ellis IO (1991) Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology 19(5):403–410
Goldhirsch A, Glick JH, Gelber RD, Coates AS, Thurlimann B, Senn HJ (2005) Meeting highlights: international expert consensus on the primary therapy of early breast cancer 2005. Ann Oncol 16(10):1569–1583
Wolff AC, Hammond ME, Schwartz JN, Hagerty KL, Allred DC, Cote RJ, Dowsett M, Fitzgibbons PL, Hanna WM, Langer A, McShane LM, Paik S, Pegram MD, Perez EA, Press MF, Rhodes A, Sturgeon C, Taube SE, Tubbs R, Vance GH, van de Vijver M, Wheeler TM, Hayes DF (2007) American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. J Clin Oncol 25(1):118–145
Marchio C, Iravani M, Natrajan R, Lambros MB, Savage K, Tamber N, Fenwick K, Mackay A, Senetta R, Di Palma S, Schmitt FC, Bussolati G, Ellis LO, Ashworth A, Sapino A, Reis-Filho JS (2008) Genomic and immunophenotypical characterization of pure micropapillary carcinomas of the breast. J Pathol 215(4):398–410
Geyer FC, Weigelt B, Natrajan R, Lambros MB, de Biase D, Vatcheva R, Savage K, Mackay A, Ashworth A, Reis-Filho JS (2010) Molecular analysis reveals a genetic basis for the phenotypic diversity of metaplastic breast carcinomas. J Pathol 220(5):562–573
Loo LW, Grove DI, Williams EM, Neal CL, Cousens LA, Schubert EL, Holcomb IN, Massa HF, Glogovac J, Li CI, Malone KE, Daling JR, Delrow JJ, Trask BJ, Hsu L, Porter PL (2004) Array comparative genomic hybridization analysis of genomic alterations in breast cancer subtypes. Cancer Res 64(23):8541–8549
Melchor L, Honrado E, Huang J, Alvarez S, Naylor TL, Garcia MJ, Osorio A, Blesa D, Stratton MR, Weber BL, Cigudosa JC, Rahman N, Nathanson KL, Benitez J (2007) Estrogen receptor status could modulate the genomic pattern in familial and sporadic breast cancer. Clin Cancer Res 13(24):7305–7313
Natrajan R, Lambros MB, Geyer FC, Marchio C, Tan DS, Vatcheva R, Shiu KK, Hungermann D, Rodriguez-Pinilla SM, Palacios J, Ashworth A, Buerger H, Reis-Filho JS (2009) Loss of 16q in high grade breast cancer is associated with estrogen receptor status: evidence for progression in tumors with a luminal phenotype? Genes Chromosomes Cancer 48(4):351–365
Natrajan R, Lambros MB, Rodriguez-Pinilla SM, Moreno-Bueno G, Tan DS, Marchio C, Vatcheva R, Rayter S, Mahler-Araujo B, Fulford LG, Hungermann D, Mackay A, Grigoriadis A, Fenwick K, Tamber N, Hardisson D, Tutt A, Palacios J, Lord CJ, Buerger H, Ashworth A, Reis-Filho JS (2009) Tiling path genomic profiling of grade 3 invasive ductal breast cancers. Clin Cancer Res 15(8):2711–2722
Geyer FC, Lopez-Garcia MA, Lambros MB, Reis-Filho JS (2009) Genetic characterization of breast cancer and implications for clinical management. J Cell Mol Med 13(10):4090–4103
Weigelt B, Baehner FL, Reis-Filho JS (2010) The contribution of gene expression profiling to breast cancer classification, prognostication and prediction: a retrospective of the last decade. J Pathol 220(2):263–280
Marchio C, Iravani M, Natrajan R, Lambros MB, Geyer FC, Savage K, Parry S, Tamber N, Fenwick K, Mackay A, Schmitt FC, Bussolati G, Ellis I, Ashworth A, Sapino A, Reis-Filho JS (2009) Mixed micropapillary-ductal carcinomas of the breast: a genomic and immunohistochemical analysis of morphologically distinct components. J Pathol 218(3):301–315
Coe BP, Ylstra B, Carvalho B, Meijer GA, Macaulay C, Lam WL (2007) Resolving the resolution of array CGH. Genomics 89(5):647–653
Gunnarsson R, Staaf J, Jansson M, Ottesen AM, Goransson H, Liljedahl U, Ralfkiaer U, Mansouri M, Buhl AM, Smedby KE, Hjalgrim H, Syvanen AC, Borg A, Isaksson A, Jurlander J, Juliusson G, Rosenquist R (2008) Screening for copy-number alterations and loss of heterozygosity in chronic lymphocytic leukemia—a comparative study of four differently designed, high resolution microarray platforms. Genes Chromosomes Cancer 47(8):697–711
Tan DS, Lambros MB, Natrajan R, Reis-Filho JS (2007) Getting it right: designing microarray (and not ‘microawry’) comparative genomic hybridization studies for cancer research. Lab Invest 87(8):737–754
Mackay A, Tamber N, Fenwick K, Iravani M, Grigoriadis A, Dexter T, Lord CJ, Reis-Filho JS, Ashworth A (2009) A high-resolution integrated analysis of genetic and expression profiles of breast cancer cell lines. Breast Cancer Res Treat 118(3):481–498
Natrajan R, Weigelt B, Mackay A, Geyer FC, Grigoriadis A, Tan DS, Jones C, Lord CJ, Vatcheva R, Rodriguez-Pinilla SM, Palacios J, Ashworth A, Reis-Filho JS (2010) An integrative genomic and transcriptomic analysis reveals molecular pathways and networks regulated by copy number aberrations in basal-like, HER2 and luminal cancers. Breast Cancer Res Treat 121(3):575–589
Arriola E, Marchio C, Tan DS, Drury SC, Lambros MB, Natrajan R, Rodriguez-Pinilla SM, Mackay A, Tamber N, Fenwick K, Jones C, Dowsett M, Ashworth A, Reis-Filho JS (2008) Genomic analysis of the HER2/TOP2A amplicon in breast cancer and breast cancer cell lines. Lab Invest 88(5):491–503
Simpson PT, Reis-Filho JS, Lambros MB, Jones C, Steele D, Mackay A, Iravani M, Fenwick K, Dexter T, Jones A, Reid L, Da Silva L, Shin SJ, Hardisson D, Ashworth A, Schmitt FC, Palacios J, Lakhani SR (2008) Molecular profiling pleomorphic lobular carcinomas of the breast: evidence for a common molecular genetic pathway with classic lobular carcinomas. J Pathol 215(3):231–244
Reis-Filho JS, Drury S, Lambros MB, Marchio C, Johnson N, Natrajan R, Salter J, Levey P, Fletcher O, Peto J, Ashworth A, Dowsett M (2008) ESR1 gene amplification in breast cancer: a common phenomenon? Nat Genet 40(7):809–810; author reply 810–802
Carey LA, Dees EC, Sawyer L, Gatti L, Moore DT, Collichio F, Ollila DW, Sartor CI, Graham ML, Perou CM (2007) The triple negative paradox: primary tumor chemosensitivity of breast cancer subtypes. Clin Cancer Res 13(8):2329–2334
Hicks J, Krasnitz A, Lakshmi B, Navin NE, Riggs M, Leibu E, Esposito D, Alexander J, Troge J, Grubor V, Yoon S, Wigler M, Ye K, Borresen-Dale AL, Naume B, Schlicting E, Norton L, Hagerstrom T, Skoog L, Auer G, Maner S, Lundin P, Zetterberg A (2006) Novel patterns of genome rearrangement and their association with survival in breast cancer. Genome Res 16(12):1465–1479
Reis-Filho JS, Savage K, Lambros MB, James M, Steele D, Jones RL, Dowsett M (2006) Cyclin D1 protein overexpression and CCND1 amplification in breast carcinomas: an immunohistochemical and chromogenic in situ hybridisation analysis. Mod Pathol 19(7):999–1009
Elsheikh S, Green AR, Aleskandarany MA, Grainge M, Paish CE, Lambros MB, Reis-Filho JS, Ellis IO (2008) CCND1 amplification and cyclin D1 expression in breast cancer and their relation with proteomic subgroups and patient outcome. Breast Cancer Res Treat 109(2):325–335
Sahin E, Colla S, Liesa M, Moslehi J, Muller FL, Guo M, Cooper M, Kotton D, Fabian AJ, Walkey C, Maser RS, Tonon G, Foerster F, Xiong R, Wang YA, Shukla SA, Jaskelioff M, Martin ES, Heffernan TP, Protopopov A, Ivanova E, Mahoney JE, Kost-Alimova M, Perry SR, Bronson R, Liao R, Mulligan R, Shirihai OS, Chin L, DePinho RA (2011) Telomere dysfunction induces metabolic and mitochondrial compromise. Nature 470(7334):359–365. doi:10.1038/nature09787
Burnichon N, Briere JJ, Libe R, Vescovo L, Riviere J, Tissier F, Jouanno E, Jeunemaitre X, Benit P, Tzagoloff A, Rustin P, Bertherat J, Favier J, Gimenez-Roqueplo AP (2010) SDHA is a tumor suppressor gene causing paraganglioma. Hum Mol Genet 19(15):3011–3020. doi:10.1093/hmg/ddq206
Saada A, Shaag A, Mandel H, Nevo Y, Eriksson S, Elpeleg O (2001) Mutant mitochondrial thymidine kinase in mitochondrial DNA depletion myopathy. Nat Genet 29(3):342–344. doi:10.1038/ng751
Karihtala P, Mantyniemi A, Kang SW, Kinnula VL, Soini Y (2003) Peroxiredoxins in breast carcinoma. Clin Cancer Res 9(9):3418–3424
Shiota M, Izumi H, Miyamoto N, Onitsuka T, Kashiwagi E, Kidani A, Hirano G, Takahashi M, Ono M, Kuwano M, Naito S, Sasaguri Y, Kohno K (2008) Ets regulates peroxiredoxin1 and 5 expressions through their interaction with the high-mobility group protein B1. Cancer Sci 99(10):1950–1959
Demasi AP, Furuse C, Altemani A, Junqueira JL, Oliveira PR, Araujo VC (2009) Peroxiredoxin I is overexpressed in oncocytic lesions of salivary glands. J Oral Pathol Med 38(6):514–517
Porcelli AM, Ghelli A, Ceccarelli C, Lang M, Cenacchi G, Capristo M, Pennisi LF, Morra I, Ciccarelli E, Melcarne A, Bartoletti-Stella A, Salfi N, Tallini G, Martinuzzi A, Carelli V, Attimonelli M, Rugolo M, Romeo G, Gasparre G (2010) The genetic and metabolic signature of oncocytic transformation implicates HIF1alpha destabilization. Hum Mol Genet 19(6):1019–1032
Guerra E, Trerotola M, Dell’ Arciprete R, Bonasera V, Palombo B, El-Sewedy T, Ciccimarra T, Crescenzi C, Lorenzini F, Rossi C, Vacca G, Lattanzio R, Piantelli M, Alberti S (2008) A bicistronic CYCLIN D1-TROP2 mRNA chimera demonstrates a novel oncogenic mechanism in human cancer. Cancer Res 68(19):8113–8121. doi:10.1158/0008-5472.CAN-07-6135
Gasparre G, Romeo G, Rugolo M, Porcelli AM (2010) Learning from oncocytic tumors: why choose inefficient mitochondria? Biochim Biophys Acta. doi:10.1016/j.bbabio.2010.08.006
Acknowledgments
This study was funded by the Breakthrough Breast Cancer Research Centre and by the University of Bologna School of Medicine-Dipartimento di Ematologia e Scienze Oncologiche “L. e A. Seràgnoli”; the study was also supported in part by a research AIRC regional grant (code 1145) to GT and by Italian Government research grants to GT (Ricerca Fondamentale Orientata, number 20074zw8la) and GG (FIRB-’Futuro in Ricerca’ J31J10000040001); DdB was the recipient of a fellowship from the Centro Interdipartimentale di Ricerca sul Cancro “G. Prodi” (CIRC), University of Bologna.
Conflict of interest
The authors have no conflicts of interest to declare.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Felipe C. Geyer and Dario de Biase contributed equally to this work. Jorge S. Reis-Filho and Giovanni Tallini contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Geyer, F.C., de Biase, D., Lambros, M.B.K. et al. Genomic profiling of mitochondrion-rich breast carcinoma: chromosomal changes may be relevant for mitochondria accumulation and tumour biology. Breast Cancer Res Treat 132, 15–28 (2012). https://doi.org/10.1007/s10549-011-1504-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-011-1504-4