Abstract
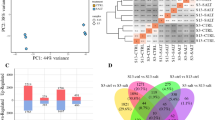
Plants respond differently to salinity stress due to their unique gene architectures. Among genes, transcription factors (TFs) regulate many physiological and biochemical processes by modulating the rate of transcription initiation of target genes. Modulation of TFs has been correlated to the salt adaptation of any given genotype. In order to identify the expression of eight TFs (belong to bHLH, CBF, MYB, WRKY, and Zpt2 families) in three annual Medicago genotypes (M. polymorpha cv. Ieze, M. laciniata cv. Shushtar, and M. laciniata cv. Gheshm) under salinity stress, the RT-qPCR analyses were performed. Attempts were also made to establish relationships between gene expression profiles and morpho-physiological traits in these genotypes. In response to salinity, cv. Ieze had minimal changes in biomass, the electrolyte leakage, H2O2 content, and the higher ratio of reduced to oxidized glutathione than the other genotypes. Furthermore, Ieze had lower accumulation of Na+ and less decrease in K+ content. Altogether, it is concluded that Ieze could be regarded as a salt tolerant genotype. Transcriptome profile showed considerable variation across Medicago genotypes and among plant tissues. Among five TFs, Zpt2-2 and CBF4 had higher expression in salt-tolerant genotypes suggesting these genes as good candidates in genetic improvement programs to produce stress-tolerant plants.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- EL:
-
electrolyte leakage
- GSH:
-
reduced glutathione
- GSSG:
-
oxidized glutathione
- RT-qPCR:
-
reverse transcription quantitative polymerase chain reaction
- TFs:
-
transcription factors
- TGSH:
-
total glutathione
References
Alexieva, V., Sergiev, S., Karanov, E.: The effect of drought and ultraviolet radiation on growth and stress markers in pea and wheat. — Plant Cell Environ. 24: 1337–1344, 2001.
Baloglu, M.C., Inal, B., Kavas, M., Unver, T.: Diverse expression pattern of wheat transcription factors against abiotic stresses in wheat species. — Gene 550: 117–122, 2014.
Becher, M., Talke, I.N., Krall, L., Kramer, U.: Cross-species microarray transcript profiling reveals high constitutive expression of metal homeostasis genes in shoots of the zinc hyperaccumulator Arabidopsis halleri. — Plant J. 37: 251–268, 2004.
Besseau, S., Li, J., Palva, E.T.: WRKY54 and WRKY70 cooperate as negative regulators of leaf senescence in Arabidopsis thaliana. — J. exp. Bot. 63: 2667–2679, 2012.
Cook, D.T., Burden, R.S.: Lipids modulation of plasma membrane-bound ATPase. — Physiol. Plant. 78: 153–159, 1990.
Dash, S., Van Hemert, J., Hong, L., Wise, R.P., Dickerson, J.A.: Plexdb: gene expression resources for plants and plant pathogens. — Nucl. Acid Res. 40: 1194–1201, 2012.
De Haan, R.L., Sheaffer, C.C., Samac, D.A., Moynihan, J.M., Barnes, D.K.: Evaluation of annual Medicago for upper midwest agro-ecosystems. — J. Agron. Crop Sci. 188: 417–425, 2002.
De Lorenzo, L., Merchan, F., Blanchet, S., Megías, M., Frugier, F., Crespi, M., Sousa, C.: Differential expression of the TFIIIA regulatory pathway in response to salt stress between Medicago truncatula genotypes. — Plant Physiol. 145: 1521–1532, 2007.
Dubos, C., Stracke, R., Grotewold, E., Weisshaar, B., Martin, C., Lepiniec, L.: MYB transcription factors in Arabidopsis. — Trends Plant Sci. 15: 573–581, 2010.
Guilfoyle, T.J., Hagen, G.: Auxin response factors. - Curr. Opin. Plant. Biol. 10: 453–460. 2007.
Hoagland, D.R., Arnon, D.I.: The water-culture method for growing plants without soil. — Circular Calif. Agr. Exp. Sta. 347: 32, 1950.
Karami, M., Rafiei, F., Shiran, B., Khodambashi, M.: Comparative response of annual Medicago spp. to salinity. — Russ. J. Plant Physiol. 62: 617–624, 2015.
Kodiara. K.S., Qin, F., Tran, L.F., Maruyama, K., Kidokoro, S., Fujita, Y., Yamaguchi-Shinozaki, K.: Arabidopsis Cys2/His2 zinc-finger proteins AZF1 and AZF2 negatively regulate abscisic acid-repressive and auxin-inducible genes under abiotic stress conditions — Plant Physiol. 157: 742–756. 2011.
Li, J., Brader, G., Kariola, T., Palva, E.T.: WRKY70 modulates the selection of signaling pathways in plant defense. — Plant J. 46: 477–491, 2006.
Li, D., Su, Z., Dong, J., Wang, T.: An expression database for roots of the model legume Medicago truncatula under salt stress. — BMC Genomic 11: 517–525, 2009.
Li, D., Zhang, Y., Hu, X., Shen, X., Ma, L., Su, Z., Dong, J.: Transcriptional profiling of Medicago truncatula under salt stress identified a novel CBF transcription factor MtCBF4 that plays an important role in abiotic stress responses. — BMC Plant Biol. 11: 109–128, 2011.
Li, J., Besseau, S., Petri, Toronen, P., Sipari, N., Kollist, H., Holm, L., Tapio Palva, E.: Defense-related transcription factors WRKY70 and WRKY54 modulate osmotic stress tolerance by regulating stomatal aperture in Arabidopsis. — New Phytol. 200: 457–472, 2013.
Lindemose, S., O’Shea, C., Jensen, M. K., Skriver, K.: Structure, function and networks of transcription factors involved in abiotic stress responses. — Int. J. mol. Sci. 14: 5842–5878, 2013.
López-Pérez, L., Martínez-Ballesta, M.D.C., Maurel, C., Carvajal, M.: Changes in plasma membrane lipids, aquaporins and proton pump of broccoli roots, as an adaptation mechanism to salinity. — Phytochemistry 70: 492–500, 2009.
Mansour, M.M.F.: Plasma membrane permeability as an indicator of salt tolerance in plants. — Biol. Plant. 57: 1–10, 2013.
Mansour, M.M.F., Salama, K.H.A.: Cellular basis of salinity tolerance in plants. — Environ. exp. Bot. 52: 113–122, 2004.
May, M.J., Leaver, C.J.: Oxidative stimulation of glutathione synthesis in Arabidopsis thaliana suspension culture. — Plant Physiol. 103: 157–166, 1993.
Merchan, F., De Lorenzo, L., Rizzo, S.G., Niebel, A., Manyani, H., Frugier, F., Crespi, M.: Identification of regulatory pathways involved in the reacquisition of root growth after salt stress in Medicago truncatula. — Plant Cell 51: 1–17, 2007.
Mittler, R., Kim, Y., Song, L., Coutu, J., Ciftci-Yilmaz, S., Lee, H., Stevenson, B., Zhu, J.K.: Gain-and loss-of-function mutations in Zat10 enhance the tolerance of plants to abiotic stress. — FEBS Lett. 580: 6537–6542, 2006.
Mittova, V., Tal, M., Volokita, M., Guy, M.: Up-regulation of the leaf mitochondrial and peroxisomal antioxidative systems in response to salt-induced oxidative stress in the wild salt-tolerant tomato species Lycopersicon pennellii. — Plant Cell Environ. 26: 845–856, 2003.
Munns, R.: Plant salt tolerance. - In: Sunkar, R. (ed.): Methods in Molecular Biology. Pp. 25–38. Springer, Berlin 2010.
Polle, A., Rennenberg, H.: Field studies on Norway spruce trees at high altitudes. II. Defence systems against oxidative stress in needles. — New Phytol. 121: 635–642, 1992.
Rafiei, F., Torabi, Z., Ebrahimie, E.: Type A response regulators are involved in the plant-microbe interaction. — Plant Omics J. 8: 178–182, 2015.
Rao, A., Ahmad, S.D., Sabir, S.M., Awan, S.I.: Potential antioxidant activities improve salt tolerance in ten varieties of wheat (Triticum aestivum L.). — Amer. J. Plant Sci. 4: 69–76, 2013.
Rushton, P.J., Somssich, I.E., Ringler, P., Shen, Q.J.: WRKY transcription factors. — Trends Plant Sci. 15: 247–258, 2010.
Saibo, N.J.M., Lourenco, T., Oliveira, M.M.: Transcription factors and regulation of photosynthetic and related metabolism under environmental stresses. — Ann. Bot. 103: 609–623, 2009.
Salentijn, E.M.J., Pereira, A., Angenent, G.C., Van der Linden, G.C., Krens, F., F., Smulders, M.J.M., Vosman, B.: Plant translational genomics: from model species to crops. — Mol. Breed. 20: 1–13, 2007.
Sanchez, D.H., Pieckenstain, F.L., Szymanski, J., Erban, A., Bromke, M., Hannah, M., Udvardi, M.K.: Comparative functional genomics of salt stress in related model and cultivated plants identifies and overcomes limitations to translational genomics. — PloS ONE 6: e17094, 2011.
Schmittgen, T.D., Livak, K.J.: Analyzing real-time PCR data by the comparative CT method. — Nat. Protoc. 3: 1101–1108, 2008.
Sheaffer, C.C., Simmons, S.R., Schmitt, M.A.: Annual medic and berseem clover dry matter and nitrogen production in rotation with corn. — Agron. J. 93: 1080–1086, 2001.
Tausz, M., Sircelj, H., Grill, D.: The glutathione system as a stress marker in plant ecophysiology: is a stress-response concept valid? — J. exp. Bot. 55: 1955–1962, 2004.
Taji, T., Seki, M., Satou, M., Sakurai, T., Kobayashi, M., Ishiyama, K., Narusaka, Y., Narusaka, M., Zhu, J.K., Shinozaki, K.: Comparative genomics in salt tolerance between Arabidopsis and Arabidopsis-related halophyte salt cress using Arabidopsis microarray. — Plant Physiol. 135: 1697–1709, 2004.
Tester, M., Davenport, R.: Na+ tolerance and Na+ transport in higher plants. — Ann. Bot. 91: 503–527, 2003.
Tivoli, B., Baranger, A., Sivasithamparam, K., Barbetti, M.J.: Annual Medicago: from a model crop challenged by a spectrum of necrotrophic pathogens to a model plant to explore the nature of disease resistance. — Ann. Bot. 98: 1117–1128, 2006.
Vilanova, S., Cañizares, J., Pascual, L., Blanca, J.M., Díez, M.J., Prohens, J., Picó, B.: Application of genomic tools in plant breeding. — Curr. Genomics 13: 179–195, 2012.
Walsh, M.J., Delaney, R.H., Groose, R.W., Krall, J.M.: Performance of annual medic species (Medicago spp.) in Southeastern Wyoming. — Agron. J. 93: 1249–1256, 2001.
Wang, T.Z., Tian, Q.Y., Wang, B.L., Zhao, M.G., Zhang, W.H.: Genome variations account for different response to three mineral elements between Medicago truncatula ecotypes Jemalong A17 and R108. — BMC Plant Biol. 14: 122–130, 2014.
Weber, M., Harada, E., Vess, C., Roepenack-Lahaye, E.V., Clemens, S.: Comparative microarray analysis of Arabidopsis thaliana and Arabidopsis halleri roots identifies nicotianamine synthase, a ZIP transporter and other genes as potential metal hyperaccumulation factors. — Plant J. 37: 269–281, 2004.
Wilkings, O., Waldron, L., Nahal, H., Provart, N.J., Campbell, M.M.: Genotype and time of the day shape the Populus drought response. — Plant J. 60: 703–715, 2009.
Zahaf, O., Blanchet, S., De Zélicourt, A., Alunni, B., Plet, J., Laffont, C., de' Lorenzo, L., Imbeaud, S., Ichanté, J.L., Diet, A., Badri, M., Zabalza, A., Gonzalez, E.M., Delacroix, H., Gruber, V., Frugier, F., Crespi, M.: Comparative transcriptomic analysis of salt adaptation in roots of contrasting Medicago truncatula genotypes. — Mol. Plants 5: 1068–1081, 2012.
Zamani Babgohari, M., Niazi, A., Moghadam, A.A., Deihimi, T., Ebrahimie, E.: Genome-wide analysis of key salinitytolerance transporter (HKT1;5) in wheat and wild wheat relatives (A and D genomes). — In Vitro cell. dev. Biol. Plant. 49: 97–106, 2012.
Zang, X., Komatsu, S.: A proteomics approach for identifying osmotic-stress-related proteins in rice. — Phytochemistry 68: 426–437, 2007.
Author information
Authors and Affiliations
Corresponding author
Additional information
Acknowledgements: This study was supported by Iran National Science Foundation (grant No. INSF 90006335) and in part by Shahrekord University. Authors gratefully thank Dr. Fariborz Khajali from Shahrekord University for help with English editing.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Mokhtari, F., Rafiei, F., Shabani, L. et al. Differential expression pattern of transcription factors across annual Medicago genotypes in response to salinity stress. Biol Plant 61, 227–234 (2017). https://doi.org/10.1007/s10535-016-0666-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10535-016-0666-7