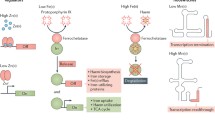
Abstract
Bacterial infections cause severe medical problems worldwide, resulting in considerable death and loss of capital. With the ever-increasing rise of antibiotic-resistant bacteria and the lack of development of new antibiotics, research on metal-based antimicrobial therapy has now gained pace. Metal ions are essential for survival, but can be highly toxic to organisms if their concentrations are not strictly controlled. Through evolution, bacteria have acquired complex metal-management systems that allow them to acquire metals that they need for survival in different challenging environments while evading metal toxicity. Metalloproteins that controls these elaborate systems in the cell, and linked to key virulence factors, are promising targets for the anti-bacterial drug development. Among several metal-sensory transcriptional regulators, the ArsR–SmtB family displays greatest diversity with several distinct metal-binding and nonmetal-binding motifs that have been characterized. These prokaryotic metolloregulatory transcriptional repressors represses the expression of operons linked to stress-inducing concentrations of metal ions by directly binding to the regulatory regions of DNA, while derepression results from direct binding of metal ions by these homodimeric proteins. Many bacteria, e.g., Mycobacterium tuberculosis, Bacillus anthracis, etc., have evolved to acquire multiple metal-sensory motifs which clearly demonstrate the importance of regulating concentrations of multiple metal ions. Here, we discussed the mechanisms of how ArsR–SmtB family regulates the intracellular bioavailability of metal ions both inside and outside of the host. Knowledge of the metal-challenges faced by bacterial pathogens and their survival strategies will enable us to develop the next generation drugs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
ArsR–SmtB family members possess a highly conserved DNA recognition helix-turn-helix (HTH) motif and bind as homodimers to their operator/promoter (O/P) region, repressing the expression of operons, in absence of metal ions, associated with the metal ion sequestration or efflux in both gram-negative and gram-positive bacteria, while derepresses the operons in presence of toxic concentrations of heavy metal ions, allowing these organisms to survive in challenging environments (Shi et al. 1994; Busenlehner et al. 2003; Osman and Cavet 2010) (Fig. 1). Some of the members also found to control virulence factors (Saha and Chakrabarti 2006; Zhao et al. 2010), sulfur oxidation (Mandal et al. 2007), hypoxia (Guimarães et al. 2011), prodigiosin biosynthesis (Gristwood et al. 2011), oxidative stress response (Ehira et al. 2010), bioluminescence (Gueuné et al. 2008), biofilm formation (Mac Aogáin et al. 2012), etc.
Seven major families of soluble metal-sensing transcriptional regulators have been identified in bacteria (Waldron and Robinson 2009), and are designated based upon their founding member(s): ArsR–SmtB (Huckle et al. 1993; Eicken et al. 2003), MerR (Brocklehurst et al. 1999; Outten et al. 2000), CsoR-RcnR (Liu et al. 2007a; Smaldone and Helmann 2007), CopY (Strausak and Solioz 1997), DtxR (Guedon and Helmann 2003), Fur (Gaballa and Helmann 1998; Ahn et al. 2006) and NikR (Dosanjh and Michel 2006; Wang et al. 2009a). The ArsR–SmtB family displays the greatest diversity among others, with thirteen distinct metal-sensing and two non metal-sensing motifs identified so far (Table 1). These have been designated α3 (Wu and Rosen 1991, 1993; Shi et al. 1994), α3N (Liu et al. 2005), α5 (Huckle et al. 1993; Kuroda et al. 1999; Singh et al. 1999), α3N–α5 (Thelwell et al. 1998; Sun et al. 2001; Busenlehner et al. 2002a), α5c (Campbell et al. 2007), α53 (Cavet et al. 2002), α4c (Cavet et al. 2003; Wang et al. 2005), α4c2 (Wang et al. 2010), α3N–2 (Ordóñez et al. 2008), α5–4 (Qin et al. 2007), α55 (Li et al. 2016a), α2–α52 (Slyemi et al. 2013; Moinier et al. 2014), α3–4 (Wang et al. 2006), α33 (non-metal binding motif) (Ehira et al. 2010) and α2–α5 (non-metal binding motif) (Saha and Chakrabarti 2006; Saha et al. 2006) based upon the location of the site where metal ions bind within the protein fold (Fig. 2; Table 1). The metal-sensing motif names originated from the typical structural fold (α1–α2–α3–α4–β1–β2–α5) observed in SmtB protein (Cook et al. 1998) (Fig. 3).
Sensory (metal/nonmetal-binding) sites in ArsR–SmtB family repressors. Alignment of representative sequences for the different sensory sites (highlighted in black) involving α5 (Synechococcus elongatus SmtB, SmtB_SE; Bacillus subtilis CzrA, CzrA_BS), α3N–α5 (Synechocystis sp. ZiaR, ZiaR_SS; Oscillatoria brevis BxmR, BxmR_OB), α3N (Staphylococcus aureus CadC, CadC_SA; Nostoc sp. AztR, AztR_NS), α5c (Mycobacterium tuberculosis NmtR, NmtR_MT), α53 (M. tuberculosis KmtR, KmtR_MT), α4c (M. tuberculosis CmtR, CmtR_MT), α4c2 (Streptomyces coelicolor CmtR, CmtR_SC), α5–4 (Acidithiobacillus ferrooxidans ArsR, ArsR_AF), α55 (Bacteroides vulgatus ArsR, ArsR_BV), α3N–2 (Corynebacterium glutamicum ArsR, ArsR_CG), α3 (Escherichia coli ArsR, ArsR_EC), α33 (C. glutamicum CyeR, CyeR_CG), α2–α5 (Vibrio cholerae HlyU, HlyU_VC), α2–α52 (Thiomonas arsenitoxydans AioF, AioF_TA) and α3–4 (Streptomyces sp. ArsR1, ArsR_SS) sites are shown. The α3N site in SmtB and the α5 site in CadC are not required for metal-responsiveness (highlighted in grey). The secondary structure determined for SmtB is shown on top
a Typical structural fold (α1–α2–α3–α4–β1–β2–α5) of an ArsR–SmtB family homodimeric repressor (structure of Synechococcus elongatus SmtB shown here). Two sub-units are shown in black and grey colors respectively. Secondary structural elements are indicated in numbers. b 90° rotated image of the above SmtB structure. The position of α3N and α5 sites are indicated by arrows. The figures were created with PyMOL (DeLano 2002)
ArsR–SmtB family members sense a wide variety of metal ions like As, Sb and Bi (ArsR, Escherichia coli), Zn (SmtB, Synechococcus sp.; ZiaR, Synechocystis sp.), Cd, Pb and Zn (CadC, Staphylococcus aureus; AztR, Anabaena sp.), Cd and Pb (CmtR, Mycobacterium tuberculosis), Zn and Co (CzrA, Bacillus subtilis), Ni and Co (NmtR, KmtR, M. tuberculosis) or Cu, Ag, Zn and Cd (BxmR, Oscillatoria brevis). Among 15 identified motifs in ArsR–SmtB family, only α2–α5 (HlyU, Vibrio cholerae; BigR, Xylella fastidiosa and Agrobacterium tumefaciens; PigS, Serratia sp.; SoxR, Pseudaminobacter salicylatoxidans; YgaV, E. coli) and α33 (CyeR, Corynebacterium glutamicum) motifs found not to sense any metal ions, but control transcription via novel redox switches (Saha and Chakrabarti 2006; Mandal et al. 2007; Gueuné et al. 2008; Ehira et al. 2010; Guimarães et al. 2011; Gristwood et al. 2011; Mukherjee et al. 2014, 2015). Some of the ArsR–SmtB family repressors (PagR, Bacillus anthracis; PyeR, Pseudomonas aeruginosa) do not have apparent metal-sensory sites yet controls transcription via unidentified novel methods (Zhao et al. 2010; Mac Aogáin et al. 2012). More than 82,000 ArsR–SmtB family members are found in the InterPro database (Finn et al. 2017), yet only a handful of the proteins was characterized in this group, indicates the possibility of discovering new and novel metal-sensory sites in this group. Several pathogenic bacteria (e.g., B. anthracis, M. tuberculosis, A. tumefaciens, B. cereus, etc.) and non-pathogenic soil bacteria (e.g., Microbacterium oxydans, Amycolatopsis keratiniphila, etc.) found to possess multiple ArsR–SmtB members in their genomes. This clearly indicates that these bacteria use not only several novel mechanisms to withstand toxic levels of various metal ions in the environment, but may also use them to their advantage to evade host-mediated immunity.
Metals are essential for survival of organisms yet slight changes in the concentration would make them toxic to the cells. Understanding the mechanisms of how bacteria use metals for their advantage would enable us to better prepare for the attacks of pathogenic bacteria, especially from antibiotic-resistant strains, by developing novel methods (e.g., antibacterial metal-nanoparticles, etc.) that use metals at our advantage. Therefore, it is essential to identify new novel metal-sensory sites in ArsR–SmtB family of transcriptional repressors and discover new mechanisms of transcriptional regulation. The knowledge gathered from these studies would help us to develop new-age drugs in response to the attacks of pathogenic microorganisms.
Characteristics of ArsR–SmtB family of transcriptional regulators
The ArsR–SmtB family of transcriptional metallorepressors represses the expression of genes/operons associated to stress-inducing concentrations of different heavy metal ions. Direct binding of metal ions by this group of homodimeric metal-sensors, remove them from their cognate O/P DNA, results in the derepression of the corresponding genes/operons (Busenlehner et al. 2003; Osman and Cavet 2010) (Fig. 1).
Among different families of metal-sensing transcriptional regulators that have so far been identified in archaea and bacteria (Wang et al. 2004; Waldron and Robinson 2009; Osman and Cavet 2010), the ArsR–SmtB family displays most diversity with as many as thirteen metal-sensing and two nonmetal-sensing motifs that have been identified till date (end of 2016). These metal-sensing motifs have been designated as α3, α3N, α3N–2, α33, α3–4, α5, α5c, α53, α5–4, α55, α3N–α5, α4c, α4c2, α2–α5 and α2–α52 (Table 1), based upon the locations of sensory amino acids within the known secondary structures of the proteins of the ArsR–SmtB family (Table 1; Fig. 3).
Several studies found that, in both metal-bound and metal-free states, ArsR–SmtB metallorepressors are weakly dissociable homodimers (Kar et al. 1997; Busenlehner et al. 2001, 2002a; Pennella et al. 2003) and each homodimer binds two metal ions either in the dimeric interface (inter) or within each monomer (intra) (Table 1).
α3 motif
One of the founding members of the ArsR–SmtB family is the plasmid-borne (Gladysheva et al. 1994; Bruhn et al. 1996) or chromosomally encoded (Diorio et al. 1995) ArsR that senses As(III), Sb(III) or Bi(III) (Wu and Rosen 1991; Gladysheva et al. 1994; Oden et al. 1994), with the sensory motif CxCx2C in α3 helix, and represses transcription of the ars operon in E. coli (Wu and Rosen 1991, 1993; Shi et al. 1994). The ars operon in E. coli plasmids R46 and R773 contain arsR, arsD, arsA, arsB and arsC genes, while the chromosomally encodes ars operon has all these except arsD and arsA genes (Rosen 1990; Busenlehner et al. 2003). The ArsC protein catalyzes reduction of arsenate As(V) to arsenite As(III) (Gladysheva et al. 1994) and the metallochaperone ArsD transports As(III) to ArsAB for extrusion (Lin et al. 2006). ArsAB encodes an arsenite-efflux system composed of secondary carrier protein ArsB and an anion-translocating ATPase ArsA (Rosen 1999). ArsA and ArsD proteins are always found together in bacterial and archaeal ars operons, which indicates the possibility that arsRDABC operon may have evolved from arsRBC operon by acquiring arsA and arsD genes together as a unit (Rosen 1999; Lin et al. 2006). E. coli plasmid R773 has three cysteines (Cys32, Cys34 and Cys37) in α3 helix that comprises the CxCx2C metal-sensory motif and form the trigonal metal-coordination complex (O’Halloran 1993). Only two cysteines (Cys32 and Cys34) are essential to produce the conformational changes, upon metal binding, that help to release the repressor from its cognate DNA and start transcription by RNA polymerase (Shi et al. 1996). Interestingly, the E. coli plasmid R773 ars operon found to show increased resistance to tellurite (Turner et al. 1992), which is not observed with the chromosomal ars operon (Cai et al. 1998), but whether this was due to the failure of tellurite to induce ars operon is not clear.
Like E. coli, several other bacteria and archaea found to encode ArsR protein (Table 2), e.g., ArsR proteins from S. aureus Plasmid pI258 (Ji and Silver 1992), Staphylococcus xylosus Plasmid pSX267 (Rosenstein et al. 1992), Yersinia enterocolitica Plasmid pYVe227 (Neyt et al. 1997), B. subtilis 168 (Sato and Kobayashi 1998), Pseudomonas aeruginosa (Cai et al. 1998), Acidiphilium multivorum AIU 301 Plasmid pKW301 (Suzuki et al. 1998), Synechocystis sp. PCC 6803 (López-Maury et al. 2003), Serratia marcescens Plasmid R478 (Ryan and Colleran 2002), Shigella flexneri 2457T (Vorontsov et al. 2007), Lactobacillus plantarum Plasmid pWCFS103 (van Kranenburg et al. 2005), Bacillus sp. CDB3 (Yu et al. 2015), Ferroplasma acidarmanus Fer1 (Gihring et al. 2003; Baker-Austin et al. 2007), Campylobacter coli (Noormohamed and Fakhr 2013), C. jejuni (Wang et al. 2009b), C. lari (Matsuda et al. 2016), ArsR1 and ArsR2 proteins from Pantoea sp. IMH (Wang et al. 2016), ArsR2 from Pseudomonas putida KT2440 (Fernández et al. 2016), ArsR1 and ArsR2 from Geobacillus kaustophilus HTA426 (Cuebas et al. 2011), ArsRC protein from Leptospirillum ferriphilum (Tuffin et al. 2006) and ArsR1 from Ochrobactrum tritici SCII24T (Branco et al. 2008). While most of these proteins have CxCx2C sensory motif at α3, some show variations (Table 2). Interestingly, the archaeal protein ArsR from F. acidarmanus found to have CxCx2C sensory motif in the C-terminal region instead of usual α3 helix (Gihring et al. 2003; Baker-Austin et al. 2007). L. ferriphilum arsRC genes are unusual in that they form one continuous ORF and encodes a 297 aa long fusion protein (Tuffin et al. 2006).
α5 motif
Another founding member of the ArsR–SmtB family, Synechococcus sp. PCC 7942 SmtB, functions as a transcriptional repressor that in the absence of Zn(II) represses transcription of smtA gene, which encodes a class II metallothionein protein SmtA involved in sequestering excess metal ions inside the cell (Huckle et al. 1993; Turner and Robinson 1995). Although Zn(II) is the preferred metal ion for SmtB, it also senses Cd(II), Cu(II), Co(II), Hg(II), Ni(II), Au(II) and Ag(I) with variable affinities (Turner and Robinson 1995). SmtB has two metal-sensory motifs (α3N and α5; Fig. 3) that binds metal ions, although α5 site is the regulatory one and α3N is the non-regulatory site (VanZile et al. 2002a, b). The α5 site binds Zn(II) ion tightly via Asp104 and His106 residues of one subunit and, His117 and Glu120 from other subunit in a tetrahedral symmetry with a consensus ‘DxHx10Hx2(E/H)’ sensory motif in α5 (VanZile et al. 2000). The non-regulatory α3N site binds metal ions by Cys61 and Asp64 residues from the α3 helix (Cx2D) of one subunit and, Cys14 and His18 residues (VanZile et al. 2002a) with the motif ‘Cx3H’ in N-terminal region (Table 1). Another well studied member of the α5 group (Table 2) is CzrA from S. aureus 912 (Kuroda et al. 1999) and B. subtilis (Harvie et al. 2006). Unlike SmtB, CzrA has only α5 sensory site with the typical ‘DxHx10Hx2H’ motif. S. aureus CzrA represses czrAB operon that codes for the repressor itself and CzrB protein which functions as a cation diffusion facilitator (CDF) antiporter efflux pump (Cherezov et al. 2008). The expression of czrAB operon is induced by metal ions with variable affinity, Zn(II)>Co(II)≫Ni(II) (Pennella et al. 2003). The B. subtilis CzrA represses its own transcription, cadA (a P-type ATPase) and czcD-trkA (czcD and trkA encodes a CDF and a cation exporter, respectively) operon with variable degrees (Harvie et al. 2006). Rv2358 from M. tuberculosis is another protein that belongs to the α5 group (Table 2) with the ‘DxHx10Hx2E’ motif, represses Rv2358-furB (zur) operon, which encodes the repressor itself and a zinc uptake regulator FurB (Zur) (Milano et al. 2004; Canneva et al. 2005; Maciag et al. 2007).
α3N motif
The cadmium resistant cad operon, originally identified in S. aureus plasmid pI258, has two genes cadC (transcriptional repressor) and cadA (P-type ATPase) (Novick and Roth 1968; Nucifora et al. 1989). Similar to SmtB, CadC has both α3N (N-terminal Cx3C and α3 CxC) and α5 (DxHx10Hx2E) sensory sites (Sun et al. 2001). Only α3N site is the regulatory site (Busenlehner et al. 2002a) in CadC, whereas in SmtB α5 is the regulatory one (VanZile et al. 2002a, b). The regulatory α3N site may adopt tetrahedral (Busenlehner et al. 2001) or trigonal (Busenlehner et al. 2002a) coordination complex and senses a wide range of metal ions like Cd(II), Bi(III), Co(II), Zn(II), Pb(II), and Hg(II) (Endo and Silver 1995; Busenlehner et al. 2002a, b), while non-regulatory α5-motif binds Zn(II) and Co(II) metal ions (Busenlehner et al. 2002a; Ye et al. 2005). Residues Cys7 and Cys11 in N-terminal of one subunit, and Cys58 and Cys60 in α3 from other subunit form the inter-subunit association via the metal ions (Wong et al. 2002). Out of four cysteine residues CadC have, only Cys7, Cys58 and Cys60 are required for its biological activity (Busenlehner et al. 2002a).
Other than S. aureus, CadC protein was found in several other organisms (Table 2), e.g., Listeria monocytogenes Plasmid pLm74 (Lebrun et al. 1994), Lactococcus lactis Plasmid pND302 (Liu et al. 1997), Stenotrophomonas maltophilia D457R (Alonso et al. 2000), Bacillus stearothermophilus LV (Nerey Md Mdel et al. 2002), Bacillus firmus (Ivey et al. 1992) and Streptococcus thermophilus 4134 (Schirawski et al. 2002). While CadC from S. aureus, B. firmus, B. stearothermophilus and S. maltophilia has both α3N and α5 sites, L. monocytogenes, L. lactis and S. thermophilus CadC contain only α3N site (Table 2).
Other bacterial proteins that belong to the α3N group (Table 2) are ArsR from Desulfovibrio desulfuricans G20 (Li and Krumholz 2007), ArsR1 from Pseudomonas putida KT2440 (Fernández et al. 2016), ArsR2 from Streptomyces sp. FR-008 Plasmid pHZ227 (Wang et al. 2006), ArsRC2 from Microbacterium sp. A33 (Achour-Rokbani et al. 2010), Rv2642 from M. tuberculosis (Li et al. 2016b), AztR from Nostoc sp. (Liu et al. 2005) and AseR from B. subtilis 168 (Harvie et al. 2006) senses not only Cd(II) but also As(III), Sb(III), Bi(III), Pb(II), Zn(II), etc. ArsR1 and ArsR2 protein from archaeon Halobacterium sp. Plasmid pNRC100 (Wang et al. 2004) also have signatures of α3N group and senses As(III) and Sb(III). Sometimes, it is difficult to ensure whether a protein belong to α3 or α3N motif as two cysteine residues in the α3 helix are enough for metalloregulation (Shi et al. 1996). For example, ArsR1 from P. putida (Fernández et al. 2016) and AseR from B. subtilis (Harvie et al. 2006) both have one N-terminal cysteine residue and CxC motif in α3 helix (Table 2). The position of N-terminal cysteine is not far from α3 helix and compared to SmtB sequence that cysteine residue would fall in α1 helix, unless both α1 and α2 helices are much shorter in length compared to SmtB (Fig. 2). Further experiments are required to correctly ascertain the function of N-terminal cysteine residue in these proteins and place them in the correct group.
The ars operon of Microbacterium sp. has an unusual arsRC2 fusion gene (Achour-Rokbani et al. 2010). This kind of fusion of the ArsR and ArsC proteins has been previously described in L. ferriphilum (Tuffin et al. 2006). The C-terminal region of 331 aa long ArsRC2 protein is related to putative arsenate reductases while the N-terminal portion has homology to transcriptional repressors of the ArsR–SmtB family. The N-terminal ArsR-domain contains a putative arsenite binding signature (ESCVCDL), almost identical to that of E. coli ArsR (ELCVCDL), and a contiguous DNA binding site with wHTH motif (Gladysheva et al. 1994; Achour-Rokbani et al. 2010). This kind of unusual fusion might reduce the problem of the diffusion of arsenic to the inducer attachment site and enhance the efficiency of transcription in response to arsenate.
α3N–α5 motif
Zn(II)-sensor ZiaR from Synechocystis PCC 6803, represses ziaA which encodes a heavy metal transporting P-type ATPase, has both α3N (N-terminal Cx5H and α3 CxC) and α5 (DxHx10Hx2E) metal sensory sites (Thelwell et al. 1998). Another member of α3N–α5 group, Oscillatoria brevis BxmR represses the expression of bxa1 (encoding a CPx-ATPase metal transporter), bxmR and bmtA (a heavy metal sequestering metallothionein) (Liu et al. 2004). BxmR binds to both monovalent, Ag(I) and Cu(I), and divalent, Zn(II) and Cd(II), metal ions and interestingly, also found to be induced by thiol oxidants diamide and H2O2 (Hirose et al. 2006). While both α3N and α5 sensory sites are essential for the inducer responsiveness of ZiaR (Thelwell et al. 1998), for BxmR either α3N (senses copper, cadmium, silver and zinc) or α5 (senses zinc) site is sufficient for zinc mediated regulation (Liu et al. 2008). Unlike other ArsR–SmtB sensors, BxmR can adopt an extended range of coordination chemistries (trigonal or tetrahedral) due to the presence of multiple metal-sensing residues in its α3N site (Hx7Cx3Hx3C in N-terminal and CxC in the α3 helix) that can sense a wide range of metals while α5 is primarily restricted to Zn(II) sensing (Liu et al. 2008).
α5c motif
NmtR, a Ni(II)/Co(II)-sensing repressor, was the first ArsR–SmtB family member that has been characterized in M. tuberculosis, represses nmt operon that contains nmtA gene which encodes a P-type ATPase metal efflux pump (Cavet et al. 2002). NmtR binds to Ni(II), Co(II) and Zn(II) with varying sensitivity, Ni(II)>Co(II)>Zn(II), and Zn(II) is not a potent allosteric regulator of DNA binding as compared to Ni(II) or Co(II) (Pennella et al. 2003). Interestingly, in cyanobacterium Synechococcus PCC 7942, NmtR-mediated repression was found to be only alleviated by Co(II) and not Ni(II), despite Ni(II) is known to be the most effective inducer in M. tuberculosis, which indicates that cytosolic metal concentrations in different hosts can influence the metal-responsiveness of these transcriptional repressors (Cavet et al. 2002). NmtR requires six residues (Asp91, His93, His104, His107, His109 and His116; Fig. 2) for Ni(II)- or Co(II)-responsiveness in vivo (Cavet et al. 2002). Out of which four (Asp91, His93, His104 and His107) residues are provided by the α5 helices of two monomers and the extra two residues are extended by the C-terminal extensions in NmtR homodimer forming α5c sites with DxHx10Hx2HxHx6H motif (Pennella et al. 2003). Interestingly, the N-terminal ‘Gly2-His3-Gly4’ residues, in M. tuberculosis NmtR, are found to form an alternate metal-sensory site with Asp91, His93, His104 and His107 residues, replacing the C-terminal His109 and His116 amino acids (Reyes-Caballero et al. 2011; Lee et al. 2012). The mutant of N-terminal His3 has been found to be significantly more sensitive to Zn(II)-mediated regulation than the Co(II)-mediated one which indicates that His3 has a direct role in this Ni(II)/Co(II)-mediated allosteric switch (Reyes-Caballero et al. 2011).
α53 motif
After NmtR, KmtR is the second novel Ni(II)/Co(II)-sensing ArsR–SmtB family member characterized from M. tuberculosis (Campbell et al. 2007). KmtR represses transcription of Rv2025c, encoding a CDF-family metal exporter and its own gene. KmtR-dependent repression was alleviated by binding to Ni(II) or Co(II). Although, both KmtR and NmtR binds Ni(II) and Co(II), KmtR binds these metal ions much tighter than that of NmtR suggesting importance of sensing variable concentrations of these metals by M. tuberculosis. In KmtR, His88, Glu101, His102, His110, and His111 form a new sensory site α53 (Table 1) with the motif ‘Hx12EHx7HH’ (Campbell et al. 2007).
α5–4 motif
The ars operon in Acidithiobacillus ferrooxidans is controlled by an As(III)-responsive transcriptional repressor, ArsR. Interestingly, As(III) binding site in A. ferrooxidans ArsR has no resemblance to the traditional α3 sensory motif found in plasmid R773 of E. coli and others (Table 1). Instead, it has three cysteine residues, Cys95, Cys96, and Cys102, constituting a unique As(III)-sensory site (CCx6C) at α5-helix designated α5–4 (Qin et al. 2007), where Cys95 and Cys96 residues in the α5 helix form a trigonal coordination metal-binding site with C-terminal Cys102 residue.
Several other bacteria found to possess α5–4 sensory site, e.g., ArsR2 and ArsR3 from O. tritici SCII24T (Branco et al. 2008), ArsR1, ArsR2, ArsR3 and ArsR4 from A. tumefaciens 5A (Kang et al. 2016), ArsR proteins from Pannonibacter indicus HT23 (Bandyopadhyay and Das 2016), Chromobacterium violaceum ATCC 12472 (Azevedo et al. 2008; Arruda et al. 2016), Acidithiobacillus caldus (Kotze et al. 2006), A. caldus TnAtcArs (Tuffin et al. 2005), L. ferriphilum TnLfArs (Tuffin et al. 2006) and Sinorhizobium meliloti Rm1021 (Yang et al. 2005). The consensus motif for α5–4 site is either CCx4-6C or CCx15C at and near α5 helix (Table 1).
α55 motif
ArsR from Bacteroides vulgatus ATCC 8482, obligate anaerobe and a common member of the human gut microbiota, is found to be very sensitive to the organoarsenicals methylarsenite MA(III), and arsenite As(III). This arsenic-inducible transcriptional repressor of the ars operon in B. vulgatus confers high resistance to MAs(III), followed by As(III), suggests that this organism maintains an ars operon as the result of dietary exposure to inorganic arsenic (Li et al. 2016a). In B. vulgatus ArsR, Cys99 residue from α5, and C-terminal Cys106 and Cys107 residues are predicted to form a new As(III)-sensory site (Cx6CC), designated α55 (Li et al. 2016a), contrary to α5–4 sensory site where two cysteine residues from α5 helix constitute trigonal metal-binding site with one C-terminal cysteine (Qin et al. 2007).
α3N–2 motif
C. glutamicum is one of the most arsenic-resistant microorganisms known and can grow in presence of elevated concentrations of arsenite or arsenate (Ordóñez et al. 2005). ArsR1 and ArsR2 proteins from C. glutamicum, binds As(III) or Sb(III) by a cysteine triad composed of Cys15, Cys16, and Cys55 residues comprising the CCx38C sensory motif (Table 1) (Ordóñez et al. 2005, 2008). This binding motif is distinctly different from other characterized ArsR–SmtB family regulators (Sun et al. 2001; Gladysheva et al. 1994; Turner and Robinson 1995).
α33 motif (nonmetal-binding)
C. glutamicum CyeR, is a unique redox-sensing transcriptional regulator that binds to the intergenic region between cyeR and cye1 (encodes an old yellow enzyme family protein), induced by oxidative stress (Ehira et al. 2010). CyeR does not bind any metal ions, but in the presence of oxidants such as diamide and H2O2, the DNA-binding activity of CyeR is found to be destabilized (Ehira et al. 2010). It has two cysteine residues (Cys36 and Cys43), with the sensory motif Cx6C in and close to α3 helix, but only Cys43 found to have a role in redox regulation (Ehira et al. 2010).
α3–4 motif
Plasmid pHZ227 in Streptomyces sp., encodes an As(III)-sensing ArsR1 protein, that represses arsRBOCT operon, has a unique metal sensory site designated α3–4, not observed in classical members of ArsR–SmtB family (Wang et al. 2006). ArsR1 is predicted to sense arsenite via Cx2H motif in the α3 helix and one cysteine residue located between β1 and β2 strands of wHTH DNA-binding region (Fig. 2).
α4c motif
The Cd(II)/Pb(II)-sensing CmtR in M. tuberculosis is structurally distinct from the other Cd(II)/Pb(II) sensor CadC of S. aureus plasmid pI258 in a way that CmtR binds Cd(II) or Pb(II) via coordination by α4 sensors (Cys57 and Cys61) and C-terminal Cys102 forming a distinct α4c site instead of α3N in CadC (Cavet et al. 2003). CmtR represses cmt operon encoding CmtA which is closely related to S. aureus CadA (Yoon et al. 1991) and E. coli ZntA (Rensing et al. 1997), and encodes Zn(II)/Cd(II)/Pb(II) P-type ATPase efflux pumps (Cavet et al. 2003). C-terminal residue Cys102 functions as a key allosteric metal-sensor in CmtR that influences disassembly of CmtR-cmt O/P oligomeric complexes in the presence of metal ions (Wang et al. 2005). Metal-dependent expression from CmtR-cmt and NmtR-nmt O/P revealed that CmtR is insensitive to Ni(II) and NmtR is insensitive to Cd(II) or Pb(II) (Cavet et al. 2003). MerR of Streptomyces lividans 1326 functions as a repressor and has α4c motif (Brünker et al. 1996). MerR binds in the intercistronic region between two operons and negatively regulate several genes, including a mercuric reductase merA and an organolyase merB (Rother et al. 1999). The repression is alleviated by binding of mercuric ions Hg(II) to the MerR (Brünker et al. 1996). Interestingly, in all the cases of mercury resistances which are mediated by Hg(II) reduction, the genes are usually regulated by activator proteins, except MerR of S. lividans that function as a repressor, a hallmark of ArsR–SmtB family regulators (Brünker et al. 1996).
α4c2 motif
S. coelicolor CmtR, in contrast to M. tuberculosis CmtR, binds Pb(II) or Cd(II) by forming two pairs of sulfur-rich coordination complexes per dimer (Wang et al. 2010), instead of one pair in M. tuberculosis (Cavet et al. 2003). While, metal-sensory site 1 resembles exactly to the α4c site of M. tuberculosis CmtR, the second metal-binding site is coordinated by the C-terminal Cys110 and Cys111 residues. Site 1 binds Cd(II) tightly than Pb(II) and mediates transcriptional derepression, in contrast, site 2 ligands Cys110 and Cys111 only show Cd(II)-responsiveness (Wang et al. 2010). The residue Cys24 from α2 helix is predicted to be the third thiolate ligand to complete the trigonal coordination structure at metal site 2 with C-terminal Cys110 and Cys111 residues, but Cys24 does not have any regulatory role as its absence has no influence on the Cd(II)-responsiveness at site 2 (Wang et al. 2010).
α2–α5 motif (nonmetal-binding)
HlyU protein from V. cholerae and V. vulnificus positively regulates the expression of hemolysin hlyA (Williams and Manning 1991; Williams et al. 1993) and RTX toxin rtxA1 (Liu et al. 2007b) genes, respectively, by binding directly to their cognate DNA upstream of the genes (Liu et al. 2007b; Mukherjee et al. 2015). In V. vulnificus, HlyU activates transcription of rtxA1 toxin gene by acting as a repressor of H-NS which negatively regulates the expression of the rtxA1 gene (Liu et al. 2009a). H-NS not only represses the transcription of the RTX toxin and its transport system, but also found to directly inhibit transcription of hemolysin gene hlyA. However, transcriptional silencing of the hlyA gene is found to be counteracted by the V. cholerae transcriptional activator HlyU (Wang et al. 2015). Therefore, HlyU acts as a repressor of another repressor H-NS (Liu et al. 2009a; Wang et al. 2015). HlyU found to be a member of ArsR–SmtB family, but it does not have any metal-sensory residues or motifs typical of the ArsR–SmtB family and constitutes a unique group, designated α2–α5 that does not sense metals (Saha and Chakrabarti 2006). Molecular dynamics (MD) simulation studies on V. cholerae HlyU reveal that the DNA binding residues tend to move away from the DNA bases when the distance between the Cys38 (in α2) and Cys104 (in α5) residues was small. In contrast, in the DNA bound form, the distance between Cys38 and Cys104 increases during simulation indicating the presence of a redox switch. The DNA-bound reduced form is responsible for activating hlyA gene and in the presence of an oxidizing agent repression is established (Mukherjee et al. 2015). Also, under oxygen-limiting conditions (e.g., host intestines, etc.) V. cholerae was predicted to use the redox switch for increased expression of virulence genes (Liu et al. 2011).
In Vibrio anguillarum, two gene clusters vah1 and rtxACHBDE are found to be responsible for the hemolytic and cytotoxic activities in fish, and are positively regulated by the HlyU protein like other bacterial HlyU proteins (Li et al. 2011). These two gene clusters are again silenced by the negative regulatory action of H-NS protein and V. anguillarum HlyU act to alleviate that repression by acting as a repressor of H-NS (Mou et al. 2013).
BigR in X. fastidiosa and A. tumefaciens is structurally similar to V. cholerae HlyU and found to undergo similar DNA-binding and release using a redox switch. In the reduced DNA-bound form of BigR, the two critical cysteine residues (Cys42 and Cys108 in α2 and α5 helices, respectively) found to be wide apart while the oxidized form indicates a reduction in distance between these residues due to the formation of disulfide bridge that results in dissociation of BigR from its cognate DNA (Guimarães et al. 2011). BigR binds to the ‘BigR-box’ in the Xylella and Agrobacterium promoters, and strongly represses transcription of an operon (encodes BigR, membrane proteins and beta-lactamase-like hydrolase BLH) responsible for biofilm formation (Barbosa and Benedetti 2007). BigR is found to be easily reduced, but difficult to oxidize as two unbound cysteine residues are not very accessible and a hydrogen sulfide-induced reactive oxygen is predicted to oxidize BigR (Guimarães et al. 2011) (Fig. 4).
Model of redox-responsive gene regulation. The reduced form of the protein binds to the promoter region and represses the expression of downstream genes in the operon. In the presence of an oxidizing agent, oxidized protein displaces from the promoter region. The RNA polymerase binds DNA and subsequently induces gene expression
In purple photosynthetic bacterium Rhodobacter capsulatus, transcriptional repressor SqrR functions as a master regulator of sulfide-dependent gene expressions. In absence of H2S, with two cysteins (Cys41 and Cys107) that are in reduced form, SqrR binds the promoter region and represses the expression of sulfide-responsive genes (SRGs). In presence of H2S, reactive sulfur species (RSS) promotes the formation of a sulfide bond between Cys41 and Cys107 residues, thereby inhibiting the ability of SqrR to bind to the promoter and derepression occurs (Shimizu et al. 2017) (Fig. 4).
PigS is another ArsR–SmtB family transcriptional regulator, belong to α2–α5 group, which represses expression of the red-pigmented prodigiosin antibiotic genes via the control of divergent operons in Serratia sp. (Gristwood et al. 2011). YgaV is an autoregulated, tributyltin (TBT)-inducible repressor found in E. coli that represses ygaVP operon. The ygaVP operon encodes YgaP protein, which is a membrane-associated protein with sulfur transferase (rhodanese) activity (Gueuné et al. 2008). The dimeric thiosulfate-inducible repressor SoxR binds cooperatively to the promoters regulating expression of the sulfur oxidation sox operon in P. salicylatoxidans (Mandal et al. 2007).
α2–α52 motif
Acidophilic and facultative chemoautotrophic bacterium Thiomonas arsenitoxydans encodes an ArsR–SmtB family metalloprotein AioF, stabilized by As(III) or As(V), but not Sb(III) or Mo(VI), binds to aioBA operon at two distinct places and induces its expression (Slyemi et al. 2013; Moinier et al. 2014). There are three cysteine residues, Cys53 in the α2 helix, and Cys111 and Cys115 in the α5 helix (Fig. 2), in AioF that binds As(III) or As(V) constitute the α2–α52 sensory motif (Slyemi et al. 2013) designated here. ArsR–SmtB family members known to repress the transcription of metal-resistant operons in the absence of the metal ions and derepresses them in their presence. Interestingly, contrary to ArsR–SmtB metalloregulators, AioF specifically activates transcription of the aioBA operon in presence of metal ions (Moinier et al. 2014).
None/unknown motifs
This group includes non-classical ArsR-family regulators that do not have obvious metal-sensory motifs and involved in processes which are independent of metal-sensing or resistance (Komeda et al. 1996; Ellermeier et al. 2006; Li et al. 2008; Keese et al. 2010; Gao et al. 2011, 2012; Mac Aogáin et al. 2012) (Table 2). Also, these regulators do not have any associated metal resistance genes such as metallothionins, metal reductases, or metal efflux pumps, which are usually linked with classical ArsR–SmtB family proteins (Busenlehner et al. 2003).
In spore-forming bacterium B. subtilis, SdpC induces the synthesis of an immunity protein SdpI that protects toxin-producing cells from being killed under starvation. SdpI is encoded by the sdpIR operon which is repressed by SdpR (Ellermeier et al. 2006). PyeR in P. aeruginosa negatively regulates pyeR-pyeM-xenB operon (pyeM encodes a major facilitator superfamily membrane transporter and xenB encodes an old yellow enzyme family reductase) and is the second ArsR regulator, after BigR (Guimarães et al. 2011), found to be involved in biofilm formation (Mac Aogáin et al. 2012). In R. meliloti (Kondorosi et al. 1991) and R. leguminosarum (Li et al. 2008) expression of the nodulation genes are repressed by a the NolR protein. In Rhodococcus rhodochrous, nhlBA operon encodes a nitrile hydratase (L-NHase) whose expression is repressed by the protein NhlD. In the presence of inducer amide, NhlC inhibits the repressor NhlD, leading to the expression of L-NHase, while in the absence of amide NhlC could not inhibit NhlD, leading to the repression of the L-NHase expression by NhlD (Komeda et al. 1996). In B. anthracis plasmid pXO1, PagR negatively controls expression of the pagAR operon that encodes a toxin gene pagA. PagR also represses transcription of atxA, a positive regulator of pagA (Hoffmaster and Koehler 1999). Rv2034 and Ms6762 proteins in M. tuberculosis and Mycobacterium smegmatis, respectively, positively regulates the expression of dosR, phoP and groEL2 genes and represses own genes (Gao et al. 2011, 2012).
Archaeon Methanococcus jannaschii protein MJ223 (Ray et al. 2003), hyperthermophilic archaeon Pyrococcus horikoshii proteins PH1061 (Okada et al. 2006) and PH1932 (Itou et al. 2008) have structural features common to ArsR–SmtB family. Phr protein from the hyperthermophilic archaeon Pyrococcus furiosus inhibits transcription of its own gene, a small heat shock protein Hsp20 and an AAA+ ATPase (Vierke et al. 2003; Keese et al. 2010).
Structural studies on ArsR–SmtB family of sensory proteins
ArsR–SmtB family members are included in InterPro database (Finn et al. 2017) with profile IPR001845 (HTH ArsR-type DNA-binding domain), in PROSITE database (Sigrist et al. 2013) with profile PS50987 (HTH_ARSR_2), in Pfam database (Finn et al. 2016) with profile PF01022 (HTH_5), in PRINTS database (Attwood et al. 2012) with profile PR00778 (HTHARSR) and in SMART database (Letunic et al. 2015) with profile SM00418 (HTH_ARSR). Several X-ray crystal and NMR structures of ArsR–SmtB family members have been solved over the years and these 3d structures help us to understand how conformational changes mediated by key sensory residues drive transcriptional regulation in this metallorepressor family (Table 3).
A highly conserved ‘ELCV(C/G)D’ motif named ‘metal-binding box’ was originally identified in the members of ArsR–SmtB family (Shi et al. 1994). This motif contains residues that directly involved in binding metal ions and several ArsR–SmtB repressors have been shown to use residues from this ‘metal-box’ including ArsR and CadC proteins (Endo and Silver 1995; Bruhn et al. 1996). However, with the discovery of several new and unique members the list of metal-sensory motifs found in this family expanded (Table 1). The X-ray structure of apo-SmtB found to contain the ‘ELCVGD’ motif in the α3 helix as part of the wHTH (α3-turn-α4) DNA-binding motif (Cook et al. 1998). The 2.2 Å resolution structure of SmtB from Synechococcus sp. strain PCC7942 is the first three-dimensional (3d) structure of ArsR–SmtB family that was solved by X-ray crystallography (Cook et al. 1998). This apo-SmtB structure showed that the protein is an elongated dimer with a twofold axis of symmetry consisting of 5 α-helices and 2 β-strands arranged into α1–α2–α3–α4–β1–β2–α5 motif (Fig. 5a). The dimeric interface is formed between the two N-terminal α1 and two C-terminal α5 helices. The DNA recognition helix-turn-helix (HTH) domain (α3-turn-α4), specially the α4 helix is highly conserved and is a distinguishing characteristic of the ArsR–SmtB family repressors. Helix 4 (α4) is also termed as the DNA recognition helix (αR) that binds to the DNA at the major groove, and β1/β2-strands form the wing similar to other winged-HTHs. This wHTH domain (α3-turn-αR) has strong structural similarity to other bacterial transcriptional regulators like catabolite activator protein CAP (Schultz et al. 1991), Fe(III)-regulated diphtheria toxin repressor DtxR (Pohl et al. 1999), LysR-family proteins (Mukherjee et al. 2009; Alanazi et al. 2013), MerR-family proteins (Shewchuk et al. 1989; Huang et al. 2016), etc. The SmtB crystal structure does not have any metal ion, however, experiments with mercuric acetate derivative suggested a total of four metal-binding sites in the dimeric repressor (Cook et al. 1998). The two metal-sensory motifs in SmtB, α3N (type 1) and α5 (type 2), binds four metal ions (Fig. 5a), however, Zn(II)-sensing α5 site is the regulatory site that controls derepression while α3N is non-regulatory in nature (VanZile et al. 2002a, b). The structure of the Zn-bound SmtB repressor shows that both α5 Zn(II)-binding sites of the homodimer must be filled in order to change the dynamics of the DNA-bound SmtB that drives the derepression (Eicken et al. 2003).
Representative 3d structures of ArsR–SmtB family of transcriptional repressors showing different sensory sites. a Structure of S. elongatus SmtB (PDB ID: 1R22) showing two α5 Zn(II) binding sites shared between two subunits (in black and grey). One α5 site is shown (inset) where histidine and glutamic acid residues (in grey) from one subunit and histidine and aspartic acid residues (in black) from another subunit binds a single Zn(II) metal ion. Non-regulatory α3N metal binding site is also indicated in the figure. Secondary structural elements of SmtB are indicated in numbers. b Structure of S. aureus CzrA (PDB ID: 2M30) showing two α5 sites which senses Zn(II). c Structure of M. tuberculosis NmtR (PDB ID: 2LKP) indicating two α5c sites which senses Ni(II). d Structure of M. tuberculosis CmtR (PDB ID: 2JSC) showing two α4c Cd(II) binding sites. The inset shows two cysteine residues from one subunit (in grey) and one cysteine from another subunit (in black) binds a single Cd(II) metal ion. e Structure of Pyrococcus horikoshii PH1061 (PDB ID: 1UB9) showing resemblance of ArsR–SmtB family fold. f Structure of Xylella fastidiosa BigR showing α2–α5 nonmetal-binding site. The inset shows reduced (PDB ID: 3PQJ) and oxidized (PDB ID: 3PQK) cysteine residues
S. aureus CzrA and Synechococcus SmtB share nearly 35% sequence identity, but CzrA lacks the N-terminal extension and residues that comprise the α3N metal-sensing site in SmtB (Fig. 5b) (VanZile et al. 2002a, b). In S. aureus, upon Zn(II)-binding to α5, CzrA functions to halt the internal dynamics of DNA–protein interaction, such that the Zn(II)-bound form became energetically unfavorable and is no longer binds to the O/P site (Eicken et al. 2003; Chakravorty et al. 2013). The solution structure of CzrA bound to czr operator sequence, reveals how two allosteric states of the protein work. Upon Zn(II)-binding, high-affinity DNA-bound closed-state switch to low-affinity open-state and results in derepression (Arunkumar et al. 2009). The allosteric pathway that controls derepression is regulated by several residues (His97, His67, Val66 and Leu68) in and around the sensory α5-helix in CzrA (Campanello et al. 2013).
The X-ray crystal structure of homodimeric S. aureus CadC shows that each monomer has six α-helices and three β-strands with α0–α1–α2–β0–α3–α4–β1–β2–α5 motif (Ye et al. 2005). CadC has an extra helix in N-terminal, designated α0, and one extra beta strand, designated β0, in contrast to SmtB (Cook et al. 1998). Although, CadC has both type 1 and type 2 metal-sensory sites like SmtB (Cook et al. 1998), but only type 1 site is required for metalloregulation (Ye et al. 2005) where in SmtB type 2 site is the regulatory one (VanZile et al. 2002a, 2002b). The α3N type 1 metal-binding site in CadC is composed of N-terminal Cys7 and Cys11 residues from one monomer, and Cys58 and Cys60 residues from another monomer. Even though Zn(II)/Cd(II)-sensing type 2 site is not regulatory in CadC, it is similar to type 2 site in SmtB (Turner et al. 1996). Mutagenesis results suggest that the Arg87 residue stabilizes type 2 site in SmtB by forming hydrogen bond and in CadC the residue corresponding to SmtB Arg87 is Gly84. Therefore, in CadC, Gly84 residue do not make any significant changes in the orientation of type 2 site, and hence, binding of Zn(II) to the type 2 site is non-regulatory (Kandegedara et al. 2009). CadC and SmtB might be evolutionary intermediates between ArsR and CzrA. ArsR uses only type 1 site for metalloregulation, while CzrA uses only type 2 site. CadC and SmtB both have type 1 and 2 sites, yet CadC uses only type 1 and SmtB uses only type 2 site for metalloregulation.
Although M. tuberculosis Ni(II)/Co(II)-sensor NmtR (Fig. 5c) suggested to bind metal ions with four α5 residues, Asp91, His93, His104, and His107, and two C-terminal residues, His109 and His116 (α5c motif) (Cavet et al. 2002), the recent apo-NmtR NMR structure along with molecular dynamics simulations proposed an alternative Ni(II)-coordination model that involves the N-terminal ‘Gly2-His3-Gly4’ motif and only four α5 ligands with no contribution from two C-terminal residues His109 and His116 previously suggested (Lee et al. 2012).
M. tuberculosis sensor CmtR responds in vivo to Cd(II) or Pb(II) by α4 residues Cys57 and Cys61, and C-terminal Cys102 (α4c motif). The NMR structure (Fig. 5d) shows that CmtR has a relatively weak affinity towards DNA and the unstructured C-terminal tail becomes less mobile in the metal-bound form than the apo-CmtR due to the recruitment of Cys102 as one of the metal–ligand (Banci et al. 2007).
Crystal structures of the transcriptional activator HlyU from V. vulnificus (Nishi et al. 2010) and V. cholerae (Mukherjee et al. 2014) have been elucidated which suggests the existence of a redox switch in transcriptional regulation instead of metallogeulation typical to ArsR–SmtB family of repressors. In V. cholerae HlyU, two cysteines Cys38 (in α2) and Cys104 (in α5) are found in the dimeric interface with a distance between the two being less than 5 Å in the oxidized form (Cys38 was found to be modified as sulfenic acid), while the reduced form shows a distance more than 5 Å. The presence of a redox switch is much more clearly observable in X. fastidiosa BigR X-ray crystal structures (Guimarães et al. 2011). In the reduced DNA-bound form of BigR, two cysteine residues (Cys42 and Cys108) are more than 9 Å apart, while in the oxidized form Cys42 and Cys108 are disulfide-linked (Fig. 5f). BigR cannot bind to the DNA in oxidized form due to altered orientation of two important residues Met18 and Tyr104, and more than 30% reduction in interface area compared to the reduced form. Formation of the disulfide bridge involving Cys42 and Cys108 induces conformational changes in the wHTH DNA-binding interface of BigR homodimer, results in loss of DNA binding (Guimarães et al. 2011).
The crystal structure of the B. anthracis PagR was solved at 1.8 Å resolution and the DNA–protein model suggests that the homodimer binds to DNA with a bend of approximately 40˚ (Zhao et al. 2010). The X-ray crystal structures of NolR in apo- and DNA-bound form indicates the importance of conformational switching of Gln56 residue in the DNA-recognition helix that senses target DNA sequence variations and influence nodulation and symbiosis in S. fredii. Although like B. anthracis PagR, S. fredii NolR does not show any substantial change in conformations between apo- and DNA-bound forms (Lee et al. 2014).
P. furiosus Phr protein is the first characterized heat shock transcription factor in archaea with ArsR–SmtB family signature. The X-ray crystal structure showed some surprising features. The N-terminal domain of Phr has similarity towards bacterial ArsR–SmtB family, while its C-terminal domain was found to resemble eukaryotic BAG domain (Liu et al. 2007c). X-ray crystal structures of few other proteins from archaea have been solved, e.g., M. jannaschii protein MJ223 (Ray et al. 2003), P. horikoshii PH1061 (Okada et al. 2006) and PH1932 (Itou et al. 2008), and they all show resemblance to ArsR–SmtB family of bacterial proteins (Fig. 5e).
Characteristics of DNA-binding region
ArsR–SmtB metallorepressors predominantly exists as homodimers in both metal-bound (weak-affinity towards cognate DNA) and metal-free (high-affinity towards cognate DNA) states in solution (Kar et al. 1997; Busenlehner et al. 2001, 2002a; Pennella et al. 2003). Most of these homodimers bind to at or near O/P sites of repressed operons which contain one or more imperfect inverted repeats with a distinct ‘12-2-12’ architecture (Table 4). The smt operon of Synechococcus sp. contains two imperfect inverted 12-2-12 repeats, termed ‘S1/S2’ and ‘S3/S4’, which overlaps the −10 and −35 regions of the RNA polymerase binding site (Huckle et al. 1993; Turner and Robinson 1995). The S1/S2 inverted repeat is required for the Zn(II)-mediated metalloregulation of smtA by α5-sensor SmtB, while the S3/S4 repeat is non-regulatory (Erbe et al. 1995; Turner et al. 1996). Each SmtB homodimer expect to bind to a single 12-2-12 inverted repeat, as both the homodimer and the inverted repeat are approximately two-fold symmetic, with two HTH motifs (α3-turn-α4) of the homodimer interacts with consecutive major grooves in the DNA-bound state, but this scenario would suggests significant bending of the DNA (~30˚) around the minor groove (Cook et al. 1998). Although the entire O/P region of smtA containing both S1/S2 and S3/S4 repeats was found to bind two SmtB homodimers, yet interestingly, both S1/S2 and S3/S4 repeats separately found to bind two homodimers each, which suggest the possibility of smt O/P forming a looped-structure stabilized by dimer–dimer interactions (Kar et al. 2001; VanZile et al. 2002b). Other α5 sensors, S. aureus and B. subtilis CzrA also bind to czr O/P with an imperfect 12-2-12 inverted repeat similar to smt O/P (Kuroda et al. 1999; Singh et al. 1999; Harvie et al. 2006).
Like the smtA and czr O/Ps, the M. tuberculosis nmt O/P contains a single 12-2-12 inverted repeat where α5c-sensor NmtR has been shown to bind tightly to repress transcription (Cavet et al. 2002; Pennella et al. 2003). Interestingly, another M. tuberculosis protein α53-sensor KmtR binds to a ‘13-4-13’ inverted repeat at the O/P region instead of 12-2-12 palindromic sequence (Campbell et al. 2007).
S. aureus α3N-sensor CadC protects the O/P site of the cad operon with a 12-2-12 imperfect repeat similar to that of the smt operon (Endo and Silver 1995; Busenlehner et al. 2003). A single CadC homodimer binds to the cad O/P (Busenlehner et al. 2001), however, at low salt concentrations two CadC dimers found to bind the DNA (Busenlehner et al. 2002a). Also, two distinct CadC-DNA complexes found to form at higher concentrations of the homodimer (Endo and Silver 1995). CadC proteins from other bacteria also bind to similar 12-2-12 repeats (Ivey et al. 1992; Lebrun et al. 1994; Liu et al. 1997; Schirawski et al. 2002). Other α3N-sensors like AztR from Nostoc sp. and Rv2642 from M. tuberculosis also bind to similar inverted repeats (Liu et al. 2005; Li et al. 2016b). Interestingly, M. tuberculosis Rv2642 represses several genes by binding to a 16-bp core palindromic region (TTTGATA-TA-TGTCAAA) which could be a part of extended 12-2-12 repeats (Table 4) (Li et al. 2016b). Another α3N-sensor B. subtilis AseR shows similar DNA-binding properties (Harvie et al. 2006). Microbacterium sp. ArsRC2 fusion-protein binds to larger inverted repeats (17-6-17 and 10-5-10) (Achour-Rokbani et al. 2010) and neither of these palindromes resemble the ArsR binding regions identified previously (Wu and Rosen 1993; Rosenstein et al. 1994; Xu et al. 1996).
Like α3N- and α5-sensors, α3N–α5 sensory proteins, e.g., Synechocystis sp. ZiaR, O. brevis BxmR, M. tuberculosis Rv2358 and M. smegmatis Ms2358, found to have similar DNA-binding characteristics (Table 4) (Thelwell et al. 1998; Liu et al. 2004; Canneva et al. 2005).
The ars O/P sequence of E. coli and other bacteria also contains an imperfect 12-2-12 repeat, similar to the cad O/P, sensed by α3-sensor ArsR proteins (Gralla 1990; Ji and Silver 1992; Wu and Rosen 1993; Rosenstein et al. 1994; Xu et al. 1996; Sato and Kobayashi 1998; Suzuki et al. 1998; Wang et al. 2009b; Yu et al. 2015; Páez-Espino et al. 2015). Gel mobility shift experiments suggest that E. coli ArsR form only one DNA–protein complex (Wu and Rosen 1993; Xu et al. 1996). Acidophilic archaeon F. acidarmanus also predicted to have an imperfect 12-2-12 repeat adjacent to the TATA-box regions of arsRB operon (Baker-Austin et al. 2007).
Interestingly, several ArsR–SmtB family proteins do not conform to the ‘12-2-12’ rule as observed in α3, α3N, α5, α5c, α53 and α3N–α5-sensory repressors. The α3N–2 sensor ArsR1 from C. glutamicum, bind to two regions (S1 and S2) of 30 bp each (with 10 bp palindromic region present between S1 and S2) at O/P (from −7 to −37 bp and −47 to −77 bp) of the arsB gene (Table 4) (Ordóñez et al. 2008). S. lividans MerR (α4c) also binds to two sites spanning 28–31 bp regions with inverted repeats at the O/P site, but not similar to 12-2-12 repeat (Rother et al. 1999). Another α4c-sensor M. tuberculosis CmtR binds to an unusually long 90 bp protected region (from −80 to +10 bp) having 4 inverted repeats of 14–15 nt each with a ‘T(A/G)TAA-N4–5-T(T/G)ATA’ consensus at the O/P region (Chauhan et al. 2009). AioF from T. arsenitoxydans (α2–α52-sensor) binds to two long regions of 60–74 nt without any inverted repeats in the aioX-aioB intergenic region that overlaps O/P sites (Moinier et al. 2014). Similarly, α5–4-sensor A. ferrooxidans ArsR protects a 28 nt long region without any inverted repeat (Table 4). Interestingly, the protected region by A. ferrooxidans ArsR is found between −60 and −86 nt relative to the start site of the arsB gene (outside RNA polymerase binding sites) that it represses (Qin et al. 2007).
Non-metal sensor protein (α2–α5-sensor) like V. cholerae HlyU, binds to a region (31-nt long region with a 17-nt core palindrome) of about 150 bp away from the O/P of hlyA gene that it controls (Mukherjee et al. 2015). HlyU from V. vulnificus also recognizes a 42 nt long region, with imperfect palindrome, about 400 bp away from the rtxA1 transcription start site (Liu et al. 2009a). Interestingly, V. anguillarum HlyU binds to far upstream of the RNA polymerase binding site, but its 18–22 nt protected regions contain 5 bp direct repeats of ‘TAAAA’ instead of inverted repeats found in HlyU proteins from V. cholerae and V. vulnificus (Li et al. 2011). Similarly, other α2–α5-sensors like BigR, SoxR, PigS and SqrR binds to variable sized regions at the O/P sites with inverted or direct repeats (Table 4) (Mandal et al. 2007; Barbosa and Benedetti 2007; Gristwood et al. 2011; Mandal and Das Gupta 2012; Shimizu et al. 2017).
The non-classical group of ArsR–SmtB family proteins (NolR, SdpR, Phr, PyeR, Rv2034, etc.) that do not have any metal-sensory motif binds and protect variable regions at O/P sites with or without inverted repeats (Table 4) (Cren et al. 1995; Ellermeier et al. 2006; Li et al. 2008; Keese et al. 2010; Gao et al. 2011, 2012; Mac Aogáin et al. 2012).
Evolution of metal-sensory motifs
The most important organizing principle in biology is the universal tree of life that separates the living world into three domains—Archaea, Bacteria and Eucarya (Woese et al. 1990). Presently accepted theory of evolution suggests that the life on planet Earth might have evolved from a hot climatic condition (Wächtershäuser 2000, 2002; Schwartzman and Lineweaver 2004) and therefore, a hyperthermophile may have been the last common ancestor of life before the divergence of three primary domains (Schwartzman and Lineweaver 2004).
Though it has been observed that three domains are very dissimilar and the differences that separate them being of a more profound nature, but most of archaeal and bacterial lineages have an extensive history of horizontal or lateral gene transfer. Horizontal gene transfer (HGT), a widely-recognized adaptation mechanism in prokaryotes, can be defined as the sharing of genetic material from one individual to another that are not in a vertical or parent-offspring relationship (Soucy et al. 2015). Initially HGT was often associated with pathogenicity and antibiotic resistance in a microbial world, but the reach of HGT was far beyond this. It has been interesting to note that how the gene content in different domains of the universal tree of life has been shaped thoroughly by HGT among microbial world, between microbes and eucarya, and even among multicellular eukaryotes (Soucy et al. 2015). The domain archaea comprise of most of the hyperthermophiles while the bacterial kingdom also contains many. Several metabolic processes in archaea are found to be similar to bacterial systems (Laksanalamai et al. 2004) and also, many transcriptional regulators discovered in bacteria have homologs in the archaeal genome (Bell and Jackson 2001; Bell 2005; Geiduschek and Ouhammouch 2005), suggesting the possibility of HGT among domains. Also, archaeal members found to encode a large number of proteins with the HTH DNA-binding motifs whose sequences are highly similar to the bacterial HTH DNA-binding domains rather than to eukaryotic counterparts, and this relationship between archaeal and bacterial transcriptional regulators might have been occurred due to multiple HGT events (Aravind and Koonin 1999).
The classical view of the universal tree of life suggests that the Archaea and the Eukarya have a common ancestor, however, the origin of Eucarya remains controversial (Gribaldo et al. 2010). It can be stated that Eucarya are mainly the evolutionary chimeras of bacterial and archaeal cells that arose via endosymbiotic fusion (Soucy et al. 2015), but the mechanism by which eucarya interchange genes with prokaryotes are less clear (Gribaldo et al. 2010). Interestingly, the archaeal heat shock regulator Phr from P. furiosus is found to be a molecular chimera having N-terminal wHTH DNA-binding domain resembling bacterial wHTH motif and C-terminal domain that resembles eukaryotic BAG domain, suggesting HGT from hyperthermophiles to mesothermophiles (Liu et al. 2007c).
The ArsR–SmtB family of transcriptional regulators showed a great diversity with different types of sensory sites (Fig. 6) and in future we expect more unique members of this family will be discovered. The founding members of the ArsR–SmtB family repressor proteins (e.g., E. coli ArsR) had only type 1 site with α3 motif and binding of As(III) to the protein results in derepression. The As(III) binding site in E. coli ArsR is composed of only Cys32 and Cys34, while Cys37 is the non-regulatory residue (Shi et al. 1996), with CxCx2C motif in α3 helix (Table 1; Fig. 6d). While CxCx2C motif is the predominant one in α3-sensors, a wide range of variations is also observed in different bacteria (Table 2). A wide array of ArsR–SmtB family repressors sense cysteine residues for metal-mediated derepression, which indicates that the type 1 sites are the ancestral regulatory sites and type 2 sites might have evolved later (Kandegedara et al. 2009).
Cartoon representation of functionally characterized ArsR–SmtB family proteins depicting different sensory motifs. a α5, b α5c/α53, c α5–4/α55, d α3, e α3N, f α3N–α5, g α3N–2, h α4c, i α4c2, j α2–α52, k α2–α5, l α33 and m α3–4. Metal ions are denoted as spheres. Cysteine residues are marked as filled circles. Two subunits (I and II) of the protein are indicated in grey and black colors, respectively. N- and C-terminal ends of each subunit are indicated and for each subunit α-helices are numbered from 1 to 5. Dotted line represents N- or C-terminal extensions. The dotted triangle with ‘w’ letter represents the wing comprising β1–β2 strands in between α4 and α5 helices. n Model of the evolution of type 2 metal-binding site from type 1 site. Square box represents regulatory metal-binding site, star represents non-regulatory metal-binding site and dotted-line represents N-terminal extension of the protein
The α3N-sensory proteins (e.g., B. subtilis AseR, Nostoc sp. AztR, etc.) may have evolved from the α3-sensory ones by acquiring the N-terminal extension which provide one or two metal sensors apart from two cysteines that are contributed by the α3 helix similar to E. coli ArsR (Fig. 6e) (Liu et al. 2005; Harvie et al. 2006). The α3N-sensors have different combinations of residues (one or two cysteine residues or cysteine and histidine residues in N-terminal) contributes to the trigonal or tetragonal α3N site (Table 1; Fig. 2). Another α3N-sensor, S. aureus CadC not only has the regulatory α3N site, but also has a non-regulatory type 2 α5 site (Sun et al. 2001; Busenlehner et al. 2002a). Interestingly, CadC from L. monocytogenes has only α3N site and no α5 site (Lebrun et al. 1994), while CadC from L. lactis or S. thermophilus have partial α5 sites (Liu et al. 1997; Schirawski et al. 2002) (Table 2), which suggests a progression of type 2 α5 site in proteins with α3N motif (Fig. 6n). The α3N–α5 sensors (Fig. 6f), O. brevis BxmR or Synechocystis sp. ZiaR may be considered as intermediates between E. colil ArsR (type 1 site) and S. aureus CzrA (type 2 site) as they contain both true type 1 and 2 sites (Thelwell et al. 1998; Liu et al. 2004). Contrary to S. aureus CadC, although Synechococcus sp. SmtB has both α3N and α5 sites, α5 site is the regulatory site and α3N is the non-regulatory one (VanZile et al. 2002a, b). As all these proteins (ArsR, CadC, SmtB, etc.) have true type 1 site or remnants of type 1 site, they might have evolved from a common ancestor (Ye et al. 2005). Although, Synechocystis sp. ZiaR, O. brevis BxmR and Synechococcus sp. SmtB all have α3N and α5 sites, for ZiaR both sites are essential, for BxmR either one and for SmtB only α5 site is required for metal-mediated regulation (Thelwell et al. 1998; VanZile et al. 2002a, b; Liu et al. 2008), again indicating the progression of true type 2 site (e.g., S. aureus CzrA; Fig. 6a) by losing the type 1 site from a type 1–2 dual sensor (Fig. 6n) (Ye et al. 2005). The α5c or α53 sensors may have evolved from α5-sensory proteins (Fig. 6b) (Cavet et al. 2002; Campbell et al. 2007).
Interestingly, α3-sensor proteins have a length between 84 and 118 aa (e.g., true type 1 sensor E. coli ArsR has 117 aa), α3N-sensors have between 119 and 136 aa (e.g., type 1 and pseudo type 2 sensor S. aureus CadC has 122 aa), α3N–α5-sensors with 132-136 aa (e.g., dual type 1-2 sensor ZiaR has 132 aa) and α5-sensors have between 103 and 135 aa (e.g., pseudo type 1 and type 2 sensor SmtB has 122 aa; true type 2 sensor CzrA has 106 aa) (Table 2), indicating that the length of a protein has a direct relationship with number of sensory sites. The α3-sensors usually do not have long N-terminal extensions, therefore, length of α3-sensors are less than the α3N sensors with long N-terminal extensions providing metal ligands. Subsequently, α3N–α5 sensors are relatively longer to accommodate both type 1 and type 2 sites. Again, α5-sensors in the course of evolution lost type 1 site and became smaller in terms of protein length (Table 2).
The Arg87 residue in SmtB stabilizes the Zn-sensory type 2 site by forming hydrogen bond and in CadC the corresponding residue is Gly84 which do not make any significant changes in the conformation of type 2 site and subsequently turn into a non-regulatory site (Fig. 2) (Kandegedara et al. 2009). The α5 sensor CzrA also has arginine residue at the corresponding position, while α3/α3N-sensors ArsR, AseR, etc. have glycine residue. Interestingly, a single point mutation (guanine to cytosine) can substitute a glycine residue into an arginine residue suggesting that it is easy for the Nature to convert a non-regulatory type 2 site to a regulatory one (Kandegedara et al. 2009).
A phylogenetic tree of a subset of ArsR–SmtB family proteins show that they have evolved from a common evolutionary ancestor, with three distinct clusters—As(III)-sensor α3-group, Zn(II)-sensor α5-group and non-metal sensor α2–α5 group (Fig. 7). The α3- and α5-sensors, with type 1 and 2 sites respectively, belong to two distinct clusters. Also the redox-sensor α2–α5 proteins form a distinct group apart from the α3/α5-sensors which indicates the possibility of coexistence of primitive ArsR–SmtB family proteins with or without metal-sensory residues. Interestingly, all α2–α5-sensors belong to the phylum proteobacteria (Fig. 7; Table 2) and as proteobacteria is relatively younger compared to other groups (e.g., Firmicutes, Chloroflexi, Actinobacteria, Cyanobacteria, etc.), which suggests that this group may have evolved from ancestral metal-sensory groups by losing metal-binding residues in the course of evolution (Saha and Chakrabarti 2006; Hug et al. 2016). CadC proteins having regulatory type 1 and non-regulatory type 2 sites, clusters with α3/α3N-sensors with type 1 regulatory site, while SmtB which has regulatory type 2 site and vestigial type 1 sites clusters with α5-sensors with type 2 regulatory sites (Fig. 7).
A phylogenetic tree (Bootstrap consensus tree) of the 46 sequences (representative sequences taken from Table 2), created using the Neighbor-Joining method (Saitou and Nei 1987). Sequence alignment was done using MUSCLE (Edgar 2004). The percentage of replicating trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) are shown (if the value is 50 or more) next to the branches (Felsenstein 1985). The evolutionary distances were computed using the number of differences method (Nei and Kumar 2000) and are in the units of the number of amino acid differences per sequence. All positions with less than 95% site coverage were eliminated. Evolutionary analyses were conducted in MEGA7 (Kumar et al. 2016). For each entry sensory site, scientific name, protein name and class (for archaea, classes are shown in bracket) are indicated. The metal-binding motifs sense different metals, present in different clusters, are indicated on the right. ‘UK’ stands for the ‘unknown’ sensory site. ‘NM’ stands for ‘nonmetal-binding’ sensory site
The α3 motif (CxCx2C), instead of its usual place in α3-helix in bacteria, is found at the C-terminal region of acidophilic iron-oxidizing archaeon F. acidarmanus ArsR which represses arsRB operon and the derepression results from binding of As(III) metal ion to the protein (Gihring et al. 2003; Baker-Austin et al. 2007). Very little information is available about ArsR–SmtB homologs in Archaea and F. acidarmanus ArsR with As(III)-binding (CxCx2C) motif in the C-terminal region may indicate a possibility of the existence of this motif in a different region in the ancestral proteins and bacteria may have acquired it by HGT from archaeal counterparts. ArsR1 and ArsR2 from halophilic archaeon H. salinarum have α3N motifs and senses As(III) or Sb(III) to derepress ars operons (Table 2) (Wang et al. 2004). Some other archaeon like M. jannaschii (Ray et al. 2003), P. furiosus (Vierke et al. 2003) and P. horikoshii (Okada et al. 2006; Itou et al. 2008) have ArsR–SmtB family members in their genome, but without any identifiable metal-sensory or redox-sensory motifs (Table 2). With very little information about archeal ArsR–SmtB members and a few bioinformatic analysis with limited datasets on bacteria (Busenlehner et al. 2003; Campbell et al. 2007; Harvie et al. 2006), the knowledge on the evolution of ArsR–SmtB family is incomplete and requires further study.
Even though the overall fold in ArsR–SmtB family members is conserved, the location of metal-sensory sites varied on the surface of these proteins. Some of these metal-sensory sites (e.g., α3, α5, etc.) may have evolved in the natural course of evolution from an ancestral protein, but some sensory sites (e.g., α3N–2, α55, etc.) are unrelated to the binding sites of other characterized ArsR–SmtB family members and may have evolved by convergent evolution in response to niche environmental pressures (Ordóñez et al. 2008).
Identification of new metal sensors
The tree of life is comprised of an enormous number of branches and an approximation of the universal tree of life to full scale is a gigantic task and remains elusive (Woese et al. 1990; Koonin 2014; Hug et al. 2016). The 16s rRNA was used to construct the phylogenetic relationship between microorganisms, but it was soon realized that analyzing different molecular markers or genes may lead to either conflicting phylogenies or phylogenetic incongruence among microorganisms by grouping species that are split by other morphological, physiological or molecular markers. So, to avoid this conflict, it is better to use the whole genome instead of a gene sequence and ample new methods were employed to create genome sequences that illuminate the identity of organisms and place them correctly in the tree of life in the context of their proper ecosystem and community (Brown et al. 2015; Castelle et al. 2015). The genomes of a large number of Proteobacteria, Actinobacteria and Firmicutes, including the environmental strains like Burkholderia ubonensis (Price et al. 2013), Streptomyces antibioticus (Wang et al. 2017), etc., human pathogens like M. tuberculosis H37Rv (Cole et al. 1998), B. anthracis (Vilas-Bôas et al. 2007), etc., and plant pathogens like R. leguminosarum (Ryu 2015), A. tumefaciens (Mansfield et al. 2012), etc., encode a large number of ArsR–SmtB family of transcriptional regulators, mostly of unknown functions (Table 5). Several archaea predominantly from phylum Euryarchaeota, like Methanosarcina mazei (Deppenmeier et al. 2002), Haloarcula amylolytica (Yang et al. 2007), etc. also express a large number of ArsR–SmtB family proteins (Table 5). This signifies the importance of this family member to modulate varying metal types and concentrations in different environmental conditions for survival and proliferation.
In the last 30 years, only a few different metal-sensing and non-metal binding sites have been characterized in ArsR–SmtB family of transcriptional repressors (Table 2). With the increase in the number of newly characterized genome sequences more proteins now show signature motifs of ArsR–SmtB family in various sequence databases. At present, in InterPro database (Finn et al. 2017), HTH ArsR-type DNA-binding domain (motif number IPR001845) includes more than 82,000 proteins that have ArsR–SmtB signatures which implies the possibility of a large number of undiscovered ArsR–SmtB family proteins with unique sensory motifs. Several bacteria and archaea encodes a large number of putative ArsR–SmtB family regulators (Table 5), many of which do not exactly fit to the α3 or α5 metal binding motifs (Table 5). For example, the Hg(II)-sensor MerR from S. lividans (Rother et al. 1999) has been classified as ArsR–SmtB family member based on the basis of having α4c motif like another ArsR–SmtB repressor M. tuberculosis CmtR that senses Cd(II) or Pb(II) (Wang et al. 2005). Phylogenetic analysis suggests the possibility of extensive convergent evolution among different groups and strongly argues against the belief which states that proteins sharing overall sequence similarity would sense same metal ions (Ordóñez et al. 2008).
With the discovery of more ArsR–SmtB family members with new sensory sites and based upon the presence or absence of one or more identified metal-sensing motifs, one can more accurately predict their ability to sense specific metals. In general, one metal-sensory motif (e.g., α3 or α5) correspond to specific metal (e.g., As or Zn), but an exception to this rule exists. The α3N proteins usually sense Cd(II) or Zn(II) (e.g., CadC, AztR, etc.), but M. tuberculosis Rv2642, D. desulfuricans ArsR, P. putida ArsR1, Streptomyces sp. ArsR2, B. subtilis AseR, etc. found to sense As(III) (Table 2). It is not apparent what factors in α3N proteins distinguish between a Cd(II)/Zn(II)-sensing site from a As(III)-sensing one, although it is possible that adjacent residues facilitate this facet of metal selectivity, such as the Arg87 residue that stabilizes the Zn(II)-sensory type 2 site in SmtB while in CadC the equivalent glycine residue convert the site into a non-regulatory one (Kandegedara et al. 2009). Similarly, M. tuberculosis CmtR with α4c motif senses Cd(II) or Pb(II), but another α4c group protein S. lividans MerR senses Hg(II) (Table 2).
ArsR–SmtB family repressors not only have different metal-sensory motifs, but also several members lack known metal-binding sites (Table 2). Further characterization of new and unique proteins in this family would enable us to understand the factors affecting metal-specificity in vivo. The characterization of ArsR–SmtB members which does not have known metal-sensory sites would help us to assign precise functions to the metal-sensory members in the ever-increasing number of homologues gathering in the sequence databases and metagenome datasets.
Conclusions
Metals (Na, Mg, K, Ca, Cr, Mn, Fe, Co, Ni, Cu, Zn, Se, Mo, etc.) are essential for the regular physiology and functions of all organisms. Approximately, half of all known proteins are predicted to require metal atoms for their structure and function (Gaballa and Helmann 1998; Andreini et al. 2004). Metals comprise relatively large portion of the periodic table and have a wide range of chemical properties that govern their sensitivity in the organism. A central metal ion binds to the atoms of donor ligands, such as oxygen, nitrogen and sulfur, through interactions that are often strong and selective (Haas and Franz 2009; Ma et al. 2009). The ability to sense metal ions in the environment is extremely important for the survival of pathogenic bacteria. Host organisms can both restrict access to essential metals from invading bacteria and use their innate toxicity of certain metals for effective bacterial killing (Weinberg 2009; White et al. 2009; Kehl-Fie and Skaar 2010; Shafeeq et al. 2011). In response, bacteria developed complex metal-regulatory systems to evade metal toxicity in hostile environments within the host or outside (Brocklehurst et al. 1999; Busenlehner et al. 2003).
The human body needs many metals like iron, copper, cobalt, zinc etc. for its survival. Most of them are required in a very low concentration and problem may arise if our body receives too much of them (Guengerich 2015). Concentrations of different metals in our body are essentially very low as compared to their occurrence in the environment. For example, the concentration of copper in the environment is about 50 mg/kg (Emsley 2003) but in the human body, it is maintained at a much lower concentration of about 1.7 mg/kg body weight (Velisek 2013). Similarly, the concentration of iron in the environment is nearly 700 fold more than what is found in the human body (Velisek 2013). This metal balance is maintained by homeostasis. A specific set of transporters present in the cell compartments is involved in maintaining the delicate balance of transport activities across the cell membrane. Hemochromatosis, Pica, Wilson’s and Menkes diseases are few examples which are associated with improper functioning of the homeostatic mechanism (Nelson 1999). Entry of metals in the body can also be regulated by the process of detoxication (the mechanism of preventing entry of damaging compounds in the body). Also, there are reports showing microbial sequestering of heavy metals by the intestinal microflora, which effectively reduce the metal absorption in the human body (Monachese et al. 2012; Breton et al. 2013).
The metallothioneins, a group of low-molecular-weight proteins, rich in sulfhydryl groups, serve as ligands for several essential and nonessential metals are also involved in regulation of metal concentration (Cherian and Goyer 1995). The expression of metallothionin genes are initiated by binding of metal transcription factor-1 (MTF-1) to the regulative region of metallothionin gene called metal responsive elements (MREs) (Grzywacz et al. 2015). In mammals, different types of metallothioneins are expressed and some are also tissue-specific (Sakulsak 2012). Ferritin is a storage protein for iron in reticuloendothelial cells of the liver, spleen, and bone. Transferrin is a glycoprotein that binds most of the ferric ion in plasma and plays a role in transporting iron (also aluminium and manganese) across cell membranes (Pillet et al. 2002). ArsR–SmtB family member SmtB from Synechococcus sp. regulates a class II metallothionein protein SmtA involved in sequestering excess metal ions inside the cell (Huckle et al. 1993). Another member O. brevis BxmR represses bmtA gene, which encodes a heavy metal sequestering metallothionein (Liu et al. 2004).
In recent times, the number of infections associated with conventional antibiotic-resistant microorganisms have increased multifold (e.g., multidrug resistant S. aureus) that fueled the search for new alternative anti-microbials in absence of new potent antibiotics in the market. Metal-nanoparticles, which use completely different mechanisms of antibacterial activity than the traditional antibiotics, provide a compelling alternative strategy to kill and restrict multidrug-resistant bacteria (Wright et al. 1998; Kaneko et al. 2007; Mikolay et al. 2010). Use of different types of metal-nanoparticles (made up of Zn, Ag, Cu, Fe, Al, Au, Mg, Ti, etc.), especially zinc and silver nanoparticles in particular (Kim et al. 2007; Yoon et al. 2007; Reddy et al. 2007; Padmavathy and Vijayaraghavan 2008; Simon-Deckers et al. 2009; Jiang et al. 2009; Tran et al. 2010), showed substantial reduction in both gram-positive (e.g., B. subtilis, S. aureus, Enterococcus faecium, B. megaterium, L. monocytogenes, etc.) and gram-negative (e.g., E. coli, klebsiella pneumoniae, Salmonella typhi, V. cholerae, Pseudomonas fluorescens, Salmonella enteritidis, S. typhimurium, etc.) bacterial viability (Feng et al. 2000; Koper et al. 2002; Gu et al. 2003; Panacek et al. 2006; Gil-Tomas et al. 2007; Jung et al. 2008; Nanda and Saravanan 2009; Perni 2009; Liu et al. 2009b; Jin et al. 2009; Jiang et al. 2009). Metal nanoparticles also showed substantial antiviral (Elechiguerra et al. 2005; Lu et al. 2008; Pinto et al. 2009; Di Gianvincenzo et al. 2010; Lara et al. 2010) and antifungal activities (Kim et al. 2009; Gajbhiye et al. 2009). The mechanisms of metal-toxicity mediated by metal-nanoparticles mainly relies on the loss of protein function (Calderón et al. 2009; Anjem and Imlay 2012; Xu and Imlay 2012), production of reactive oxygen species (Imlay et al. 1988; Touati et al. 1995; Nunoshiba et al. 1999; Banin et al. 2008; Warnes et al. 2012), impairment of membrane function (Yaganza et al. 2004; Zhang and Rock 2008; Hong et al. 2012), interfere with nutrient uptake (Fauchon et al. 2002; Pereira et al. 2008), or genotoxicity (Keyer and Imlay 1996; Linley et al. 2012).
On the other hand, the unsystematic widespread release of heavy metals into the soil and waters is a major health concern globally, as these cannot be broken down to non-toxic forms and therefore have long lasting effects on the ecosystem. Many of these metals are toxic even at very low concentrations. These are not only cytotoxic but also carcinogenic and mutagenic in nature (Giller et al. 1998; Mclaughlin et al. 1999; Yao et al. 2012). Toxic concentrations of metals, otherwise essential for life, disrupt various body functions and causes severe diseases like renal dysfunction, liver cirrhosis, bone weakness, heart failure, cerebral attack, memory loss, nephrosis, lung damage, chronic anemia, gastrointestinal irritations, vision loss, disability (Vinceti et al. 2001; Neustadt and Pieczenik 2007; Duda-Chodak and Baszczyk 2008; Ainza et al. 2010; Gulati et al. 2010), etc. Although, some heavy metals are essential for microorganisms, some microbes have, however adapted to tolerate high concentrations of metals and use them for their growth. The interactions between microorganisms and metal ions have significant environmental implications especially in bioremediation. That is why bioremediation of heavy metals by microorganisms has received a great deal of interest in recent times because of its beneficial and ecofriendly nature than any other conventional methods. Heavy metal biotransformation done by natural and genetically modified bacteria (e.g., E. coli, Methylococcus capsulatus, Pseudomonas sp., Ralstonia eutropha, Deinococcus radiodurans, Alcaligenes eutrophus, Bacillus sp., Enterobacter cloacae, Micrococcus sp., etc.) is found to be an effective alternative and provides a promising approach for the removal of a wide variety of ecotoxic heavy metals (Diels et al. 1995; Wang et al. 1997; Valls et al. 2000; Brim et al. 2000; Lopez et al. 2002; Ackerley et al. 2004; Zouboulis et al. 2004; Kostal et al. 2004; Iyer et al. 2005; Kiyono and Pan-Hou 2006; Congeevaram et al. 2007; Hasin et al. 2010).
Development of ecofriendly metal bioremediation technology, or metal-nanoparticle based antibacterial therapy are still in early stages of development and better understanding of how bacteria sense metals in various environments are important to develop further technology. Bacterial metal sensors, such as ArsR–SmtB repressors, detect surplus metal ions and modulate transcription of genes involved in metal uptake, efflux, sequestration, or detoxification (Tottey et al. 2005; Lucarelli et al. 2007). Several bacteria (e.g., M. tuberculosis, etc.) harbor not only one but multiple metal sensors in response to extracellular metal ions (Cole et al. 1998). Having several sensory sites for a range of diverse metals allows the pathogen to respond quickly to host mediated metal flux and help them to survive harsh environments. Understanding the mechanisms of how bacteria respond to various metals is of prime importance as this knowledge would benefit us in developing new and unconventional antibacterial treatments and also create a pollution free environment for our future generations.
Abbreviations
- O/P:
-
Operator/promoter
- CDF:
-
Cation diffusion facilitator
- MD:
-
Molecular dynamics
- wHTH:
-
Winged helix-turn-helix
- ORF:
-
Open reading frame
- aa:
-
Amino acids
- nt:
-
Nucleotides
- bp:
-
Base pairs
- HGT:
-
Horizontal gene transfer
References
Achour-Rokbani A, Cordi A, Poupin P, Bauda P, Billard P (2010) Characterization of the ars gene cluster from extremely arsenic-resistant Microbacterium sp. strain A33. Appl Environ Microbiol 76:948–955
Ackerley DF, Gonzalez CF, Keyhan M, Blake R, Matin A (2004) Mechanism of chromate reduction by the Escherichia coli protein, NfsA, and the role of different chromate reductases in minimizing oxidative stress during chromate reduction. Environ Microbiol 6:851–860
Ahn BE, Cha J, Lee EJ, Han AR, Thompson CJ, Roe JH (2006) Nur, a nickel-responsive regulator of the Fur family, regulates superoxide dismutases and nickel transport in Streptomyces coelicolor. Mol Microbiol 59:1848–1858
Ainza C, Trevors J, Saier M (2010) Environmental mercury rising. Water Air Soil Poll 205:47–48
Alanazi AM, Neidle EL, Momany C (2013) The DNA-binding domain of BenM reveals the structural basis for the recognition of a T-N11-A sequence motif by LysR-type transcriptional regulators. Acta Crystallogr D Biol Crystallogr 69:1995–2007
Alonso A, Sanchez P, Martínez JL (2000) Stenotrophomonas maltophilia D457R contains a cluster of genes from gram-positive bacteria involved in antibiotic and heavy metal resistance. Antimicrob Agents Chemother 44:1778–1782
Andreini C, Bertini I, Rosato A (2004) A hint to search for metalloproteins in gene banks. Bioinformatics 20:1373–1380
Anjem A, Imlay JA (2012) Mononuclear iron enzymes are primary targets of hydrogen peroxide stress. J Biol Chem 287:15544–15556
Aravind L, Koonin EV (1999) DNA-binding proteins and evolution of transcription regulation in the archaea. Nucleic Acids Res 27:4658–4670
Arruda LM, Monteiro LM, Silva-Rocha R (2016) The Chromobacterium violaceum ArsR arsenite repressor exerts tighter control on its cognate promoter than the Escherichia coli system. Front Microbiol 7:1851
Arunkumar AI, Campanello GC, Giedroc DP (2009) Solution structure of a paradigm ArsR family zinc sensor in the DNA-bound state. Proc Natl Acad Sci USA 106:18177–18182
Attwood TK, Coletta A, Muirhead G, Pavlopoulou A, Philippou PB, Popov I, Romá-Mateo C, Theodosiou A, Mitchell AL (2012) The PRINTS database: a fine-grained protein sequence annotation and analysis resource–its status in 2012. Database (Oxford) 2012:bas019
Azevedo JS, Silva-Rocha R, Silva A, Peixe Carepo MS, Cruz Schneider MP (2008) Gene expression of the arsenic resistance operon in Chromobacterium violaceum ATCC 12472. Can J Microbiol 54:137–142
Baker-Austin C, Dopson M, Wexler M, Sawers RG, Stemmler A, Rosen BP, Bond PL (2007) Extreme arsenic resistance by the acidophilic archaeon ‘Ferroplasma acidarmanus’ Fer1. Extremophiles 11:425–434
Banci L, Bertini I, Cantini F, Ciofi-Baffoni S, Cavet JS, Dennison C, Graham AI, Harvie DR, Robinson NJ (2007) NMR structural analysis of cadmium sensing by winged helix repressor CmtR. J Biol Chem 282:30181–30188
Bandyopadhyay S, Das SK (2016) Functional analysis of ars gene cluster of Pannonibacter indicus strain HT23(T) (DSM 23407(T)) and identification of a proline residue essential for arsenate reductase activity. Appl Microbiol Biotechnol 100:3235–3244
Banin E, Lozinski A, Brady KM, Berenshtein E, Butterfield PW, Moshe M, Chevion M, Greenberg EP, Banin E (2008) The potential of desferrioxamine-gallium as an anti-Pseudomonas therapeutic agent. Proc Natl Acad Sci USA 105:16761–16766
Barbosa RL, Benedetti CE (2007) BigR, a transcriptional repressor from plant-associated bacteria, regulates an operon implicated in biofilm growth. J Bacteriol 189:6185–6194
Bell SD (2005) Archaeal transcriptional regulation–variation on a bacterial theme? Trends Microbiol 13:262–265
Bell SD, Jackson SP (2001) Mechanism and regulation of transcription in archaea. Curr Opin Microbiol 4:208–213
Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW (2013) GenBank. Nucleic Acids Res 41:D36–D42
Branco R, Chung AP, Morais PV (2008) Sequencing and expression of two arsenic resistance operons with different functions in the highly arsenic-resistant strain Ochrobactrum tritici SCII24T. BMC Microbiol 8:95
Breton J, Daniel C, Dewulf J, Pothion S, Froux N, Sauty M, Thomas P, Pot B, Foligné B (2013) Gut microbiota limits heavy metals burden caused by chronic oral exposure. Toxicol Lett 222:132–138
Brim H, McFarlan SC, Fredrickson JK, Minton KW, Zhai M, Wackett LP, Daly MJ (2000) Engineering Deinococcus radiodurans for metal remediation in radioactive mixed waste environments. Nat Biotechnol 18:85–90
Brocklehurst KR, Hobman JL, Lawley B, Blank L, Marshall SJ, Brown NL, Morby AP (1999) ZntR is a Zn(II)-responsive MerR-like transcriptional regulator of zntA in Escherichia coli. Mol Microbiol 31:893–902
Brown CT, Hug LA, Thomas BC, Sharon I, Castelle CJ, Singh A, Wilkins MJ, Wrighton KC, Williams KH, Banfield JF (2015) Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523:208–211
Bruhn DF, Li J, Silver S, Roberto F, Rosen BP (1996) The arsenical resistance operon of IncN plasmid R46. FEMS Microbiol Lett 139:149–153
Brünker P, Rother D, Sedlmeier R, Klein J, Mattes R, Altenbuchner J (1996) Regulation of the operon responsible for broad-spectrum mercury resistance in Streptomyces lividans 1326. Mol Gen Genet 251:307–315
Busenlehner LS, Cosper NJ, Scott RA, Rosen BP, Wong MD, Giedroc DP (2001) Spectroscopic properties of the metalloregulatory Cd(II) and Pb(II) sites of S. aureus pI258 CadC. Biochemistry 40:4426–4436
Busenlehner LS, Apuy JL, Giedroc DP (2002a) Characterization of a metalloregulatory bismuth(III) site in Staphylococcus aureus pI258 CadC repressor. J Biol Inorg Chem 7:551–559
Busenlehner LS, Weng TC, Penner-Hahn JE, Giedroc DP (2002b) Elucidation of the primary (α3N) and vestigial (α5) heavy metal binding sites in Staphylococcus aureus pI258 CadC: evolutionary implications for metal ion selectivity of ArsR/SmtB metal sensor proteins. J Mol Biol 319:685–701
Busenlehner LS, Pennella MA, Giedroc DP (2003) The SmtB/ArsR family of metalloregulatory transcriptional repressors: structural insights into prokaryotic metal resistance. FEMS Microbiol Rev 27:131–143
Cai J, Salmon K, DuBow MS (1998) A chromosomal ars operon homologue of Pseudomonas aeruginosa confers increased resistance to arsenic and antimony in Escherichia coli. Microbiology 144:2705–2713
Calderón IL, Elías AO, Fuentes EL, Pradenas GA, Castro ME, Arenas FA, Pérez JM, Vásquez CC (2009) Tellurite-mediated disabling of [4Fe-4S] clusters of Escherichia coli dehydratases. Microbiology 155:1840–1846
Campanello GC, Ma Z, Grossoehme NE, Guerra AJ, Ward BP, Dimarchi RD, Ye Y, Dann CE 3rd, Giedroc DP (2013) Allosteric inhibition of a zinc-sensing transcriptional repressor: insights into the arsenic repressor (ArsR) family. J Mol Biol 425:1143–1157
Campbell DR, Chapman KE, Waldron KJ, Tottey S, Kendall S, Cavallaro G, Andreini C, Hinds J, Stoker NG, Robinson NJ, Cavet JS (2007) Mycobacterial cells have dual nickel-cobalt sensors: sequence relationships and metal sites of metal-responsive repressors are not congruent. J Biol Chem 282:32298–32310
Canneva F, Branzoni M, Riccardi G, Provvedi R, Milano A (2005) Rv2358 and FurB: two transcriptional regulators from Mycobacterium tuberculosis which respond to zinc. J Bacteriol 187:5837–5840
Castelle CJ, Wrighton KC, Thomas BC, Hug LA, Brown CT, Wilkins MJ, Frischkorn KR, Tringe SG, Singh A, Markillie LM, Taylor RC, Williams KH, Banfield JF (2015) Genomic expansion of domain archaea highlights roles for organisms from new phyla in anaerobic carbon cycling. Curr Biol 25:690–701
Cavet JS, Meng W, Pennella MA, Appelhoff RJ, Giedroc DP, Robinson NJ (2002) A nickel-cobalt sensing ArsR-SmtB family repressor: contributions of cytosol and effector binding sites to metal selectivity. J Biol Chem 277:38441–38448
Cavet JS, Graham AI, Meng W, Robinson NJ (2003) A cadmium-lead sensing ArsR-SmtB repressor with novel sensory sites: complementary metal-discrimination by NmtR and CmtR in a common cytosol. J Biol Chem 278:44560–44566
Chakravorty DK, Wang B, Lee CW, Guerra AJ, Giedroc DP, Merz KM Jr (2013) Solution NMR refinement of a metal ion bound protein using metal ion inclusive restrained molecular dynamics methods. J Biomol NMR 56:125–137
Chauhan S, Kumar A, Singhal A, Tyagi JS, Krishna Prasad H (2009) CmtR, a cadmium-sensing ArsR-SmtB repressor, cooperatively interacts with multiple operator sites to autorepress its transcription in Mycobacterium tuberculosis. FEBS J 276:3428–3439
Cherezov V, Höfer N, Szebenyi DM, Kolaj O, Wall JG, Gillilan R, Srinivasan V, Jaroniec CP, Caffrey M (2008) Insights into the mode of action of a putative zinc transporter CzrB in Thermus thermophilus. Structure 16:1378–1388
Cherian MD, Goyer RA (1995) Part three, chapter 9, section A. In: Berthon G (ed) Handbook of metal-ligand interactions in biological fluids, vol 1. Marcel Dekker, Inc., New York, pp 648–654
Cole ST, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, Gordon SV, Eiglmeier K, Gas S, Barry CE 3rd, Tekaia F, Badcock K, Basham D, Brown D, Chillingworth T, Connor R, Davies R, Devlin K, Feltwell T, Gentles S, Hamlin N, Holroyd S, Hornsby T, Jagels K, Krogh A, McLean J, Moule S, Murphy L, Oliver K, Osborne J, Quail MA, Rajandream MA, Rogers J, Rutter S, Seeger K, Skelton J, Squares R, Squares S, Sulston JE, Taylor K, Whitehead S, Barrell BG (1998) Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393:537–544
Congeevaram S, Dhanarani S, Park J, Dexilin M, Thamaraiselvi K (2007) Biosorption of chromium and nickel by heavy metal resistant fungal and bacterial isolates. J Hazard Mater 146:270–277
Cook WJ, Kar SR, Taylor KB, Hall LM (1998) Crystal structure of the cyanobacterial metallothionein repressor SmtB: a model for metalloregulatory proteins. J Mol Biol 275:337–346
Cren M, Kondorosi A, Kondorosi E (1995) NolR controls expression of the Rhizobium meliloti nodulation genes involved in the core Nod factor synthesis. Mol Microbiol 15:733–747
Cuebas M, Villafane A, McBride M, Yee N, Bini E (2011) Arsenate reduction and expression of multiple chromosomal ars operons in Geobacillus kaustophilus A1. Microbiology 157:2004–2011
DeLano WL (2002) The PyMOL user manual. DeLano Scientific, San Carlos
Deppenmeier U, Johann A, Hartsch T, Merkl R, Schmitz RA, Martinez-Arias R, Henne A, Wiezer A, Bäumer S, Jacobi C, Brüggemann H, Lienard T, Christmann A, Bömeke M, Steckel S, Bhattacharyya A, Lykidis A, Overbeek R, Klenk HP, Gunsalus RP, Fritz HJ, Gottschalk G (2002) The genome of Methanosarcina mazei: evidence for lateral gene transfer between bacteria and archaea. J Mol Microbiol Biotechnol 4:453–461
Di Gianvincenzo P, Marradi M, Martínez-Avila OM, Bedoya LM, Alcamí J, Penadés S (2010) Gold nanoparticles capped with sulfate-ended ligands as anti-HIV agents. Bioorg Med Chem Lett 20:2718–2721
Diels L, Van Roy S, Somers K, Willems I, Doyen W, Mergeay M, Springael D, Leysen R (1995) The use of bacteria immobilized in tubular membrane reactors for heavy metal recovery and degradation of chlorinated aromatics. J Membr Sci 100:249–258
Diorio C, Cai J, Marmor J, Shinder R, DuBow MS (1995) An Escherichia coli chromosomal ars operon homolog is functional in arsenic detoxification and is conserved in gram-negative bacteria. J Bacteriol 177:2050–2056
Dosanjh NS, Michel SL (2006) Microbial nickel metalloregulation: NikRs for nickel ions. Curr Opin Chem Biol 10:123–130
Duda-Chodak A, Baszczyk U (2008) The impact of nickel on human health. J Elementol 13:685–696
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinform 5:113
Ehira S, Teramoto H, Inui M, Yukawa H (2010) A novel redox-sensing transcriptional regulator CyeR controls expression of an Old Yellow Enzyme family protein in Corynebacterium glutamicum. Microbiology 156:1335–1341
Eicken C, Pennella MA, Chen X, Koshlap KM, VanZile ML, Sacchettini JC, Giedroc DP (2003) A metal-ligand-mediated intersubunit allosteric switch in related SmtB/ArsR zinc sensor proteins. J Mol Biol 333:683–695
Elechiguerra JL, Bur JL, Morones JR, Camacho-Bragado A, Gao X, Lara HH, Yacaman MJ (2005) Interaction of silver nanoparticles with HIV-1. J Nanobiotechnol 3:6–10
Ellermeier CD, Hobbs EC, Gonzalez-Pastor JE, Losick R (2006) A three-protein signaling pathway governing immunity to a bacterial cannibalism toxin. Cell 124:549–559
Emsley J (2003) Nature’s building blocks: an A-Z guide to the elements. Oxford University Press, Oxford, pp 121–125
Endo G, Silver S (1995) CadC, the transcriptional regulatory protein of the cadmium resistance system of Staphylococcus aureus plasmid pI258. J Bacteriol 177:4437–4441
Erbe JL, Taylor KB, Hall LM (1995) Metalloregulation of the cyanobacterial smt locus: identification of SmtB binding sites and direct interaction with metals. Nucleic Acids Res 23:2472–2478
Fauchon M, Lagniel G, Aude JC, Lombardia L, Soularue P, Petat C, Marguerie G, Sentenac A, Werner M, Labarre J (2002) Sulfur sparing in the yeast proteome in response to sulfur demand. Mol Cell 9:713–723
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Feng QL, Wu J, Chen GQ, Cui FZ, Kim TN, Kim JO (2000) A mechanistic study of the antibacterial effect of silver ions on Escherichia coli and Staphylococcus aureus. J Biomed Mater Res 52:662–668
Fernández M, Morel B, Ramos JL, Krell T (2016) Paralogous regulators ArsR1 and ArsR2 of Pseudomonas putida KT2440 as a basis for arsenic biosensor development. Appl Environ Microbiol 82:4133–4144
Finn RD, Coggill P, Eberhardt RY, Eddy SR, Mistry J, Mitchell AL, Potter SC, Punta M, Qureshi M, Sangrador-Vegas A, Salazar GA, Tate J, Bateman A (2016) The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res 44:D279–D285
Finn RD, Attwood TK, Babbitt PC, Bateman A, Bork P, Bridge AJ, Chang HY, Dosztányi Z, El-Gebali S, Fraser M, Gough J, Haft D, Holliday GL, Huang H, Huang X, Letunic I, Lopez R, Lu S, Marchler-Bauer A, Mi H, Mistry J, Natale DA, Necci M, Nuka G, Orengo CA, Park Y, Pesseat S, Piovesan D, Potter SC, Rawlings ND, Redaschi N, Richardson L, Rivoire C, Sangrador-Vegas A, Sigrist C, Sillitoe I, Smithers B, Squizzato S, Sutton G, Thanki N, Thomas PD, Tosatto SC, Wu CH, Xenarios I, Yeh LS, Young SY, Mitchell AL (2017) InterPro in 2017-beyond protein family and domain annotations. Nucleic Acids Res 45:D190–D199
Gaballa A, Helmann JD (1998) Identification of a zinc-specific metalloregulatory protein, Zur, controlling zinc transport operons in Bacillus subtilis. J Bacteriol 180:5815–5821
Gajbhiye M, Kesharwani J, Ingle A, Gade A, Rai M (2009) Fungus-mediated synthesis of silver nanoparticles and their activity against pathogenic fungi in combination with fluconazole. Nanomedicine 5:382–386
Gao CH, Yang M, He ZG (2011) An ArsR-like transcriptional factor recognizes a conserved sequence motif and positively regulates the expression of phoP in mycobacteria. Biochem Biophys Res Commun 411:726–731
Gao CH, Yang M, He ZG (2012) Characterization of a novel ArsR-like regulator encoded by Rv2034 in Mycobacterium tuberculosis. PLoS One 7:e36255
Geiduschek EP, Ouhammouch M (2005) Archaeal transcription and its regulators. Mol Microbiol 56:1397–1407
Gihring TM, Bond PL, Peters SC, Banfield JF (2003) Arsenic resistance in the archaeon “Ferroplasma acidarmanus”: new insights into the structure and evolution of the ars genes. Extremophiles 7:123–130
Giller K, Witter E, McGrath SP (1998) Toxicity of heavy metals to microorganisms and microbial processes in agricultural soils: a review. Soil Biol Biochem 30:1389–1414
Gil-Tomas J, Tubby S, Parkin IP, Narband N, Dekker L, Nair SP, Wilson M, Streetc C (2007) Lethal photosensitisation of Staphylococcus aureus using a toluidine blue O–tiopronin–gold nanoparticle conjugate. J Mater Chem 17:3739–3746
Gladysheva TB, Oden KL, Rosen BP (1994) Properties of the arsenate reductase of plasmid R773. Biochemistry 33:7288–7293
Gralla JD (1990) Promoter recognition and mRNA initiation by Escherichia coli E sigma 70. Methods Enzymol 185:37–54
Gribaldo S, Poole AM, Daubin V, Forterre P, Brochier-Armanet C (2010) The origin of eukaryotes and their relationship with the Archaea: are we at a phylogenomic impasse? Nat Rev Microbiol 8:743–752
Gristwood T, McNeil MB, Clulow JS, Salmond GP, Fineran PC (2011) PigS and PigP regulate prodigiosin biosynthesis in Serratia via differential control of divergent operons, which include predicted transporters of sulfur-containing molecules. J Bacteriol 193:1076–1085
Grzywacz A, Gdula-Argasińska J, Muszyńska B, Tyszka-Czochara M, Librowski T, Opoka W (2015) Metal responsive transcription factor 1 (MTF-1) regulates zinc dependent cellular processes at the molecular level. Acta Biochim Pol 62:491–498
Gu H, Ho PL, Tong E, Wang L, Xu B (2003) Presenting vancomycin on nanoparticles to enhance antimicrobial activities. Nano Lett 3:1261–1263
Guedon E, Helmann JD (2003) Origins of metal ion selectivity in the DtxR/MntR family of metalloregulators. Mol Microbiol 48:495–506
Guengerich FP (2015) Introduction: metals in biology: metals at the host-pathogen interface. J Biol Chem 290:18943–18944
Gueuné H, Durand MJ, Thouand G, DuBow MS (2008) The ygaVP genes of Escherichia coli form a tributyltin-inducible operon. Appl Environ Microbiol 74:1954–1958
Guimarães BG, Barbosa RL, Soprano AS, Campos BM, de Souza TA, Tonoli CC, Leme AF, Murakami MT, Benedetti CE (2011) Plant pathogenic bacteria utilize biofilm growth-associated repressor (BigR), a novel winged-helix redox switch, to control hydrogen sulfide detoxification under hypoxia. J Biol Chem 286:26148–26157
Gulati K, Banerjee B, Bala Lall S, Ray A (2010) Effects of diesel exhaust, heavy metals and pesticides on various organ systems: possible mechanisms and strategies for prevention and treatment. Indian J Exp Biol 48:710–721
Haas KL, Franz KJ (2009) Application of metal coordination chemistry to explore and manipulate cell biology. Chem Rev 109:4921–4960
Harvie DR, Andreini C, Cavallaro G, Meng W, Connolly BA, Yoshida K, Fujita Y, Harwood CR, Radford DS, Tottey S, Cavet JS, Robinson NJ (2006) Predicting metals sensed by ArsR-SmtB repressors: allosteric interference by a non-effector metal. Mol Microbiol 59:1341–1356
Hasin AA, Gurman SJ, Murphy LM, Perry A, Smith TJ, Gardiner PE (2010) Remediation of chromium (VI) by a methane-oxidizing bacterium. Environ Sci Technol 44:400–405
Hirose K, Ezaki B, Liu T, Nakashima S (2006) Diamide stress induces a metallothionein BmtA through a repressor BxmR and is modulated by Zn-inducible BmtA in the cyanobacterium Oscillatoria brevis. Toxicol Lett 163:250–256
Hoffmaster AR, Koehler TM (1999) Autogenous regulation of the Bacillus anthracis pag operon. J Bacteriol 181:4485–4492
Hong R, Kang TY, Michels CA, Gadura N (2012) Membrane lipid peroxidation in copper alloy-mediated contact killing of Escherichia coli. Appl Environ Microbiol 78:1776–1784
Huang S, Liu X, Wang D, Chen W, Hu Q, Wei T, Zhou W, Gan J, Chen H (2016) Structural basis for the selective Pb(II) recognition of metalloregulatory protein PbrR691. Inorg Chem 55:12516–12519
Huckle JW, Morby AP, Turner JS, Robinson NJ (1993) Isolation of a prokaryotic metallothionein locus and analysis of transcriptional control by trace metal ions. Mol Microbiol 7:177–187
Hug LA, Baker BJ, Anantharaman K, Brown CT, Probst AJ, Castelle CJ, Butterfield CN, Hernsdorf AW, Amano Y, Ise K, Suzuki Y, Dudek N, Relman DA, Finstad KM, Amundson R, Thomas BC, Banfield JF (2016) A new view of the tree of life. Nat Microbiol 1:16048
Imlay JA, Chin SM, Linn S (1988) Toxic DNA damage by hydrogen peroxide through the Fenton reaction in vivo and in vitro. Science 240:640–642
Itou H, Yao M, Watanabe N, Tanaka I (2008) Crystal structure of the PH1932 protein, a unique archaeal ArsR type winged-HTH transcription factor from Pyrococcus horikoshii OT3. Proteins 70:1631–1634
Ivey DM, Guffanti AA, Shen Z, Kudyan N, Krulwich TA (1992) The cadC gene product of alkaliphilic Bacillus firmus OF4 partially restores Na+ resistance to an Escherichia coli strain lacking an Na+/H+ antiporter (NhaA). J Bacteriol 174:4878–4884
Iyer A, Mody K, Jha B (2005) Biosorption of heavy metals by a marine bacterium. Mar Pollut Bull 50:340–343
Ji G, Silver S (1992) Regulation and expression of the arsenic resistance operon from Staphylococcus aureus plasmid pI258. J Bacteriol 174:3684–3694
Jiang W, Mashayekhi H, Xing B (2009) Bacterial toxicity comparison between nano- and micro-scaled oxide particles. Environ Pollut 157:1619–1625
Jin T, Sun D, Su Y, Zhang H, Sue HJ (2009) Antimicrobial efficacy of zinc oxide quantum dots against Listeria monocytogenes, Salmonella enteritidis and Escherichia coli O157:h7. J Food Sci 74:46–52
Jung WK, Koo HC, Kim KW, Shin S, Kim SH, Park YH (2008) Antibacterial activity and mechanism of action of the silver ion in Staphylococcus aureus and Escherichia coli. Appl Environ Microbiol 74:2171–2178
Kandegedara A, Thiyagarajan S, Kondapalli KC, Stemmler TL, Rosen BP (2009) Role of bound Zn(II) in the CadC Cd(II)/Pb(II)/Zn(II)-responsive repressor. J Biol Chem 284:14958–14965
Kaneko Y, Theondel M, Olakanmi O, Britigan BE, Singh PK (2007) The transition metal gallium disrupts Pseudomonas aeruginosa iron metabolism and has antimicrobial and antibiofilm activity. J Clin Investig 117:877–888
Kang YS, Brame K, Jetter J, Bothner BB, Wang G, Thiyagarajan S, McDermott TR (2016) Regulatory activities of four ArsR proteins in Agrobacterium tumefaciens 5A. Appl Environ Microbiol 82:3471–3480
Kar SR, Adams AC, Lebowitz J, Taylor KB, Hall LM (1997) The cyanobacterial repressor SmtB is predominantly a dimer and binds two Zn2+ ions per subunit. Biochemistry 36:15343–15348
Kar SR, Lebowitz J, Blume S, Taylor KB, Hall LM (2001) SmtB-DNA and protein-protein interactions in the formation of the cyanobacterial metallothionein repression complex: Zn2+ does not dissociate the protein-DNA complex in vitro. Biochemistry 40:13378–13389
Keese AM, Schut GJ, Ouhammouch M, Adams MW, Thomm M (2010) Genome-wide identification of targets for the archaeal heat shock regulator phr by cell-free transcription of genomic DNA. J Bacteriol 192:1292–1298
Kehl-Fie TE, Skaar EP (2010) Nutritional immunity beyond iron: a role for manganese and zinc. Curr Opin Chem Biol 14:218–224
Keyer K, Imlay JA (1996) Superoxide accelerates DNA damage by elevating free-iron levels. Proc Natl Acad Sci USA 93:13635–13640
Kim JS, Kuk E, Yu KN, Kim JH, Park SJ, Lee HJ, Kim SH, Park YK, Park YH, Hwang CY, Kim YK, Lee YS, Jeong DH, Cho MH (2007) Antimicrobial effects of silver nanoparticles. Nanomedicine 3:95–101
Kim KJ, Sung WS, Suh BK, Moon SK, Choi JS, Kim JG (2009) Antifungal activity and mode of action of silver nano-particles on Candida albicans. Biometals 22:235–242
Kiyono M, Pan-Hou H (2006) Genetic engineering of bacteria for environmental remediation of mercury. J Health Sci 52:199–204
Komeda H, Kobayashi M, Shimizu S (1996) A novel gene cluster including the Rhodococcus rhodochrous J1 nhlBA genes encoding a low molecular mass nitrile hydratase (L-NHase) induced by its reaction product. J Biol Chem 271:15796–15802
Kondorosi E, Pierre M, Cren M, Haumann U, Buiré M, Hoffmann B, Schell J, Kondorosi A (1991) Identification of NolR, a negative transacting factor controlling the nod regulon in Rhizobium meliloti. J Mol Biol 222:885–896
Koonin EV (2014) Carl Woese’s vision of cellular evolution and the domains of life. RNA Biol 11:197–204
Koper O, Klabunde J, Marchin G, Klabunde KJ, Stoimenov P, Bohra L (2002) Nanoscale powders and formulations with biocidal activity toward spores and vegetative cells of Bacillus Species, viruses, and toxins. Curr Microbiol 44:49–55
Kostal JRY, Wu CH, Mulchandani A, Chen W (2004) Enhanced arsenic accumulation in engineered bacterial cells expressing ArsR. Appl Environ Microbiol 70:4582–4587
Kotze AA, Tuffin IM, Deane SM, Rawlings DE (2006) Cloning and characterization of the chromosomal arsenic resistance genes from Acidithiobacillus caldus and enhanced arsenic resistance on conjugal transfer of ars genes located on transposon TnAtcArs. Microbiology 152:3551–3560
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Kuroda M, Hayashi H, Ohta T (1999) Chromosome-determined zinc-responsible operon czr in Staphylococcus aureus strain 912. Microbiol Immunol 43:115–125
Laksanalamai P, Whitehead TA, Robb FT (2004) Minimal protein-folding systems in hyperthermophilic archaea. Nat Rev Microbiol 2:315–324
Lara HH, Ayala-Nunez NV, Turrent LI, Padilla CR (2010) Mode of antiviral action of silver nanoparticles against HIV-1. J Nanobiotechnol 8:1–9
Lebrun M, Audurier A, Cossart P (1994) Plasmid-borne cadmium resistance genes in Listeria monocytogenes are similar to cadA and cadC of Staphylococcus aureus and are induced by cadmium. J Bacteriol 176:3040–3048
Lee CW, Chakravorty DK, Chang FM, Reyes-Caballero H, Ye Y, Merz KM Jr, Giedroc DP (2012) Solution structure of Mycobacterium tuberculosis NmtR in the apo state: insights into Ni(II)-mediated allostery. Biochemistry 51:2619–2629
Lee SG, Krishnan HB, Jez JM (2014) Structural basis for regulation of rhizobial nodulation and symbiosis gene expression by the regulatory protein NolR. Proc Natl Acad Sci USA 111:6509–6514
Letunic I, Doerks T, Bork P (2015) SMART: recent updates, new developments and status in 2015. Nucleic Acids Res 43:D257–D260
Li X, Krumholz LR (2007) Regulation of arsenate resistance in Desulfovibrio desulfuricans G20 by an arsRBCC operon and an arsC gene. J Bacteriol 189:3705–3711
Li F, Hou B, Hong G (2008) Symbiotic plasmid is required for NolR to fully repress nodulation genes in Rhizobium leguminosarum A34. Acta Biochim Biophys Sin 40:901–907
Li L, Mou X, Nelson DR (2011) HlyU is a positive regulator of hemolysin expression in Vibrio anguillarum. J Bacteriol 193:4779–4789
Li J, Mandal G, Rosen BP (2016a) Expression of arsenic resistance genes in the obligate anaerobe Bacteroides vulgatus ATCC 8482, a gut microbiome bacterium. Anaerobe 39:117–123
Li Q, Li C, Xie L, Zhang C, Feng Y, Xie J (2016b) Characterization of a putative ArsR transcriptional regulator encoded by Rv2642 from Mycobacterium tuberculosis. J Biomol Struct Dyn 5:1–9
Lin YF, Walmsley AR, Rosen BP (2006) An arsenic metallochaperone for an arsenic detoxification pump. Proc Natl Acad Sci USA 103:15617–15622
Linley E, Denyer SP, McDonnell G, Simons C, Maillard JY (2012) Use of hydrogen peroxide as a biocide: new consideration of its mechanisms of biocidal action. J Antimicrob Chemother 67:1589–1596
Liu CQ, Khunajakr N, Chia LG, Deng YM, Charoenchai P, Dunn NW (1997) Genetic analysis of regions involved in replication and cadmium resistance of the plasmid pND302 from Lactococcus lactis. Plasmid 38:79–90
Liu T, Nakashima S, Hirose K, Shibasaka M, Katsuhara M, Ezaki B, Giedroc DP, Kasamo K (2004) A novel cyanobacterial SmtB/ArsR family repressor regulates the expression of a CPx-ATPase and a metallothionein in response to both Cu(I)/Ag(I) and Zn(II)/Cd(II). J Biol Chem 279:17810–17818
Liu T, Golden JW, Giedroc DP (2005) A zinc(II)/lead(II)/cadmium(II)-inducible operon from the cyanobacterium anabaena is regulated by AztR, an alpha3N ArsR/SmtB metalloregulator. Biochemistry 44:8673–8683
Liu M, Alice AF, Naka H, Crosa JH (2007a) The HlyU protein is a positive regulator of rtxA1, a gene responsible for cytotoxicity and virulence in the human pathogen Vibrio vulnificus. Infect Immun 75:3282–3289
Liu T, Ramesh A, Ma Z, Ward SK, Zhang L, George GN, Talaat AM, Sacchettini JC, Giedroc DP (2007b) CsoR is a novel Mycobacterium tuberculosis copper-sensing transcriptional regulator. Nat Chem Biol 3:60–68
Liu W, Vierke G, Wenke AK, Thomm M, Ladenstein R (2007c) Crystal structure of the archaeal heat shock regulator from Pyrococcus furiosus: a molecular chimera representing eukaryal and bacterial features. J Mol Biol 369:474–488
Liu T, Chen X, Ma Z, Shokes J, Hemmingsen L, Scott RA, Giedroc DP (2008) A Cu(I)-sensing ArsR family metal sensor protein with a relaxed metal selectivity profile. Biochemistry 47:10564–10575
Liu M, Naka H, Crosa JH (2009a) HlyU acts as an H-NS antirepressor in the regulation of the RTX toxin gene essential for the virulence of the human pathogen Vibrio vulnificus CMCP6. Mol Microbiol 72:491–505
Liu Y, He L, Mustapha A, Li H, Hu ZQ, Lin M (2009b) Antibacterial activities of zinc oxide nanoparticles against Escherichia coli O157:H7. J Appl Microbiol 107:1193–1201
Liu Z, Yang M, Peterfreund GL, Tsou AM, Selamoglu N, Daldal F, Zhong Z, Kan B, Zhu J (2011) Vibrio cholerae anaerobic induction of virulence gene expression is controlled by thiol-based switches of virulence regulator AphB. Proc Natl Acad Sci USA 108:810–815
Lopez A, Lazaro N, Morales S, Margues AM (2002) Nickel biosorption by free and immobilized cells of Pseudomonas fluorescens 4F39: a comparative study. Water Air Soil Pollut 135:157–172
López-Maury L, Florencio FJ, Reyes JC (2003) Arsenic sensing and resistance system in the cyanobacterium Synechocystis sp. strain PCC 6803. J Bacteriol 185:5363–5371
Lu L, Sun RW, Chen R, Hui CK, Ho CM, Luk JM, Lau GK, Che CM (2008) Silver nanoparticles inhibit hepatitis B virus replication. Antivir Ther 13:253–262
Lucarelli D, Russo S, Garman E, Milano A, Meyer-Klaucke W, Pohl E (2007) Crystal structure and function of the zinc uptake regulator FurB from Mycobacterium tuberculosis. J Biol Chem 282:9914–9922
Ma Z, Jacobsen FE, Giedroc DP (2009) Coordination chemistry of bacterial metal transport and sensing. Chem Rev 109:4644–4681
Mac Aogáin M, Mooij MJ, McCarthy RR, Plower E, Wang YP, Tian ZX, Dobson A, Morrissey J, Adams C, O’Gara F (2012) The non-classical ArsR-family repressor PyeR (PA4354) modulates biofilm formation in Pseudomonas aeruginosa. Microbiology 158:2598–2609
Maciag A, Dainese E, Rodriguez GM, Milano A, Provvedi R, Pasca MR, Smith I, Palù G, Riccardi G, Manganelli R (2007) Global analysis of the Mycobacterium tuberculosis Zur (FurB) regulon. J Bacteriol 189:730–740
Mandal S, Das Gupta SK (2012) Interactions of sulfur oxidation repressor with its promoters involve different binding geometries. Arch Microbiol 194:737–747
Mandal S, Chatterjee S, Dam B, Roy P, Das Gupta SK (2007) The dimeric repressor SoxR binds cooperatively to the promoter(s) regulating expression of the sulfur oxidation (sox) operon of Pseudaminobacter salicylatoxidans KCT001. Microbiology 153:80–91
Mansfield J, Genin S, Magori S, Citovsky V, Sriariyanum M, Ronald P, Dow M, Verdier V, Beer SV, Machado MA, Toth I, Salmond G, Foster GD (2012) Top 10 plant pathogenic bacteria in molecular plant pathology. Mol Plant Pathol 13:614–629
Matsuda M, Kuribayashi T, Yamamoto S, Millar BC, Moore JE (2016) Transformation and characterization of an arsenic gene operon from urease-positive thermophilic Campylobacter (UPTC) in Escherichia coli. Folia Microbiol 61:57–62
Mclaughlin MJ, Parker DR, Clarke JM (1999) Metals and micronutrients-food safety issues. Field Crop Res 60:143–163
Mikolay A, Huggett S, Tikana L, Grass G, Braun J, Nies DH (2010) Survival of bacteria on metallic copper surfaces in a hospital trial. Appl Microbiol Biotechnol 87:1875–1879
Milano A, Branzoni M, Canneva F, Profumo A, Riccardi G (2004) The Mycobacterium tuberculosis Rv2358-furB operon is induced by zinc. Res Microbiol 155:192–200
Moinier D, Slyemi D, Byrne D, Lignon S, Lebrun R, Talla E, Bonnefoy V (2014) An ArsR/SmtB family member is involved in the regulation by arsenic of the arsenite oxidase operon in Thiomonas arsenitoxydans. Appl Environ Microbiol 80:6413–6426
Monachese M, Burton JP, Reid G (2012) Bioremediation and tolerance of humans to heavy metals through microbial processes: a potential role for probiotics? Appl Environ Microbiol 78:6397–6404
Mou X, Spinard EJ, Driscoll MV, Zhao W, Nelson DR (2013) H-NS is a negative regulator of the two hemolysin/cytotoxin gene clusters in Vibrio anguillarum. Infect Immun 81:3566–3576
Mukherjee D, Saha RP, Chakrabarti P (2009) Structural and unfolding features of HlyT, a tetrameric LysR type transcription regulator of Vibrio cholerae. Biochim Biophys Acta 1794:1134–1141
Mukherjee D, Datta AB, Chakrabarti P (2014) Crystal structure of HlyU, the hemolysin gene transcription activator, from Vibrio cholerae N16961 and functional implications. Biochim Biophys Acta 1844:2346–2354
Mukherjee D, Pal A, Chakravarty D, Chakrabarti P (2015) Identification of the target DNA sequence and characterization of DNA binding features of HlyU, and suggestion of a redox switch for hlyA expression in the human pathogen Vibrio cholerae from in silico studies. Nucleic Acids Res 43:1407–1417
Nanda A, Saravanan M (2009) Biosynthesis of silver nanoparticles from Staphylococcus aureus and its antimicrobial activity against MRSA and MRSE. Nanomedicine 5:452–456
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Nelson N (1999) Metal ion transporters and homeostasis. EMBO J 18:4361–4371
Nerey MDC, Pichuantes C, Mdel Nerey Md, Saavedra CP, Araya MA, Tantaleán JC, Vásquez CC (2002) Expression of Bacillus stearothermophilus LV cadmium resistance genes in Escherichia coli causes hypersensitivity to cadmium chloride. Curr Microbiol 45:187–190
Neustadt J, Pieczenik S (2007) Toxic-metal contamination: mercury. Integr Med 6:36–37
Neyt C, Iriarte M, Thi VH, Cornelis GR (1997) Virulence and arsenic resistance in Yersiniae. J Bacteriol 179:612–619
Nishi K, Lee HJ, Park SY, Bae SJ, Lee SE, Adams PD, Rhee JH, Kim JS (2010) Crystal structure of the transcriptional activator HlyU from Vibrio vulnificus CMCP6. FEBS Lett 584:1097–1102
Noormohamed A, Fakhr MK (2013) Arsenic resistance and prevalence of arsenic resistance genes in Campylobacter jejuni and Campylobacter coli isolated from retail meats. Int J Environ Res Public Health 10:3453–3464
Novick RP, Roth C (1968) Plasmid-linked resistance to inorganic salts in Staphylococcus aureus. J Bacteriol 95:1335–1342
Nucifora G, Chu L, Misra TK, Silver S (1989) Cadmium resistance from Staphylococcus aureus plasmid pI258 cadA gene results from a cadmium-efflux ATPase. Proc Natl Acad Sci USA 86:3544–3548
Nunoshiba T, Obata F, Boss AC, Oikawa S, Mori T, Kawanishi S, Yamamoto K (1999) Role of iron and superoxide for generation of hydroxyl radical, oxidative DNA lesions, and mutagenesis in Escherichia coli. J Biol Chem 274:34832–34837
Oden KL, Gladysheva TB, Rosen BP (1994) Arsenate reduction mediated by the plasmid-encoded ArsC protein is coupled to glutathione. Mol Microbiol 12:301–306
O’Halloran TV (1993) Transition metals in control of gene expression. Science 261:715–725
Okada U, Sakai N, Yao M, Watanabe N, Tanaka I (2006) Structural analysis of the transcriptional regulator homolog protein from Pyrococcus horikoshii OT3. Proteins 63:1084–1086
Ordóñez E, Letek M, Valbuena N, Gil JA, Mateos LM (2005) Analysis of genes involved in arsenic resistance in Corynebacterium glutamicum ATCC 13032. Appl Environ Microbiol 71:6206–6215
Ordóñez E, Thiyagarajan S, Cook JD, Stemmler TL, Gil JA, Mateos LM, Rosen BP (2008) Evolution of metal(loid) binding sites in transcriptional regulators. J Biol Chem 283:25706–25714
Osman D, Cavet JS (2010) Bacterial metal-sensing proteins exemplified by ArsR-SmtB family repressors. Nat Prod Rep 27:668–680
Outten FW, Outten CE, Hale J, O’Halloran TV (2000) Transcriptional activation of an Escherichia coli copper efflux regulon by the chromosomal MerR homologue, cueR. J Biol Chem 275:31024–31029
Padmavathy N, Vijayaraghavan R (2008) Enhanced bioactivity of ZnO nanoparticles – an antimicrobial study. Sci Technol Adv Mat 9:35004–35010
Páez-Espino AD, Durante-Rodríguez G, de Lorenzo V (2015) Functional coexistence of twin arsenic resistance systems in Pseudomonas putida KT2440. Environ Microbiol 17:229–238
Panacek A, Kvitek L, Prucek R, Kolar M, Vecerova R, Pizurova N, Sharma VK, Nevecna T, Zboril R (2006) Silver colloid nanoparticles: synthesis, characterization, and their antibacterial activity. J Phys Chem 110:16248–16253
Pennella MA, Shokes JE, Cosper NJ, Scott RA, Giedroc DP (2003) Structural elements of metal selectivity in metal sensor proteins. Proc Natl Acad Sci USA 100:3713–3718
Pereira Y, Lagniel G, Godat E, Baudouin-Cornu P, Junot C, Labarre J (2008) Chromate causes sulfur starvation in yeast. Toxicol Sci 106:400–412
Perni S (2009) The antimicrobial properties of light-activated polymers containing methylene blue and gold nanoparticles. Biomaterials 30:89–93
Pillet S, Fournier M, Measures LN, Bouquegneau JM, Cyr DG (2002) Presence and regulation of metallothioneins in peripheral blood leukocytes of grey seals. Toxicol Appl Pharmacol 185:207–217
Pinto DB, Shukla S, Perkas N, Gedanken A, Sarid R (2009) Inhibition of herpes simplex virus type 1 infection by silver nanoparticles capped with mercaptoethane sulfonate. Bioconjugate Chem 20:1497–1502
Pohl E, Holmes RK, Hol WG (1999) Crystal structure of a cobalt-activated diphtheria toxin repressor-DNA complex reveals a metal-binding SH3-like domain. J Mol Biol 292:653–667
Price EP, Sarovich DS, Webb JR, Ginther JL, Mayo M, Cook JM, Seymour ML, Kaestli M, Theobald V, Hall CM, Busch JD, Foster JT, Keim P, Wagner DM, Tuanyok A, Pearson T, Currie BJ (2013) Accurate and rapid identification of the Burkholderia pseudomallei near-neighbour, Burkholderia ubonensis, using real-time PCR. PLoS One 8:e71647
Qin J, Fu HL, Ye J, Bencze KZ, Stemmler TL, Rawlings DE, Rosen BP (2007) Convergent evolution of a new arsenic binding site in the ArsR/SmtB family of metalloregulators. J Biol Chem 282:34346–34355
Ray SS, Bonanno JB, Chen H, de Lencastre H, Wu S, Tomasz A, Burley SK (2003) X-ray structure of an M. jannaschii DNA-binding protein: implications for antibiotic resistance in S. aureus. Proteins 50:170–173
Reddy KM, Feris K, Bell J, Wingett DG, Hanley C, Punnoose A (2007) Selective toxicity of zinc oxide nanoparticles to prokaryotic and eukaryotic systems. Appl Phys Lett 90:2139021–2139023
Rensing C, Mitra B, Rosen BP (1997) The zntA gene of Escherichia coli encodes a Zn(II)-translocating P-type ATPase. Proc Natl Acad Sci USA 94:14326–14331
Reyes-Caballero H, Lee CW, Giedroc DP (2011) Mycobacterium tuberculosis NmtR harbors a nickel sensing site with parallels to Escherichia coli RcnR. Biochemistry 50:7941–7952
Rosen BP (1990) The plasmid-encoded arsenical resistance pump: an anion-translocating ATPase. Res Microbiol 141:336–341
Rosen BP (1999) Families of arsenic transporters. Trends Microbiol 7:207–212
Rosenstein R, Peschel A, Wieland B, Götz F (1992) Expression and regulation of the antimonite, arsenite, and arsenate resistance operon of Staphylococcus xylosus plasmid pSX267. J Bacteriol 174:3676–3683
Rosenstein R, Nikoleit K, Götz F (1994) Binding of ArsR, the repressor of the Staphylococcus xylosus (pSX267) arsenic resistance operon to a sequence with dyad symmetry within the ars promoter. Mol Gen Genet 242:566–572
Rother D, Mattes R, Altenbuchner J (1999) Purification and characterization of MerR, the regulator of the broad-spectrum mercury resistance genes in Streptomyces lividans 1326. Mol Gen Genet 262:154–162
Ryan D, Colleran E (2002) Arsenical resistance in the IncHI2 plasmids. Plasmid 47:234–240
Ryu CM (2015) Against friend and foe: type 6 effectors in plant-associated bacteria. J Microbiol 53:201–208
Saha RP, Chakrabarti P (2006) Molecular modeling and characterization of Vibrio cholerae transcription regulator HlyU. BMC Struct Biol 6:24
Saha RP, Basu G, Chakrabarti P (2006) Cloning, expression, purification, and characterization of Vibrio cholerae transcriptional activator, HlyU. Protein Expr Purif 48:118–125
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sakulsak N (2012) Metallothionein: an overview on its metal homeostatic regulation in mammals. Int J Morphol 30:1007–1012
Sato T, Kobayashi Y (1998) The ars operon in the skin element of Bacillus subtilis confers resistance to arsenate and arsenite. J Bacteriol 180:1655–1661
Schirawski J, Hagens W, Fitzgerald GF, van Sinderen D (2002) Molecular characterization of cadmium resistance in Streptococcus thermophilus strain 4134: an example of lateral gene transfer. Appl Environ Microbiol 68:5508–5516
Schultz SC, Shields GC, Steitz TA (1991) Crystal structure of a CAP-DNA complex: the DNA is bent by 90 degrees. Science 253:1001–1007
Schwartzman DW, Lineweaver CH (2004) The hyperthermophilic origin of life revisited. Biochem Soc Trans 32:168–171
Shafeeq S, Yesilkaya H, Kloosterman TG, Narayanan G, Andrew PW, Kuipers OP, Morrissey JA (2011) The cop operon is required for copper homeostasis and contributes to virulence in Streptococcus pneumoniae. Mol Microbiol 81:1255–1270
Shewchuk LM, Verdine GL, Walsh CT (1989) Transcriptional switching by the metalloregulatory MerR protein: initial characterization of DNA and mercury (II) binding activities. Biochemistry 28:2331–2339
Shi W, Wu J, Rosen BP (1994) Identification of a putative metal binding site in a new family of metalloregulatory proteins. J Biol Chem 269:19826–19829
Shi W, Dong J, Scott RA, Ksenzenko MY, Rosen BP (1996) The role of arsenic-thiol interactions in metalloregulation of the ars operon. J Biol Chem 271:9291–9297
Shimizu T, Shen J, Fang M, Zhang Y, Hori K, Trinidad JC, Bauer CE, Giedroc DP, Masuda S (2017) Sulfide-responsive transcriptional repressor SqrR functions as a master regulator of sulfide-dependent photosynthesis. Proc Natl Acad Sci USA 114:2355–2360
Sigrist CJ, de Castro E, Cerutti L, Cuche BA, Hulo N, Bridge A, Bougueleret L, Xenarios I (2013) New and continuing developments at PROSITE. Nucleic Acids Res 41:D344–D347
Simon-Deckers A, Loo S, Mayne-L’hermite M, Herlin-Boime N, Menguy N, Reynaud C, Gouget B, Carrière M (2009) Size-, composition- and shape-dependent toxicological impact of metal oxide nanoparticles and carbon nanotubes toward bacteria. Environ Sci Technol 43:8423–8429
Singh VK, Xiong A, Usgaard TR, Chakrabarti S, Deora R, Misra TK, Jayawal RK (1999) ZntR is an autoregulatory protein and negatively regulates the chromosomal zinc resistance operon znt of Staphylococcus aureus. Mol Microbiol 33:200–207
Slyemi D, Moinier D, Talla E, Bonnefoy V (2013) Organization and regulation of the arsenite oxidase operon of the moderately acidophilic and facultative chemoautotrophic Thiomonas arsenitoxydans. Extremophiles 17:911–920
Smaldone GT, Helmann JD (2007) CsoR regulates the copper efflux operon copZA in Bacillus subtilis. Microbiology 153:4123–4128
Soucy SM, Huang J, Gogarten JP (2015) Horizontal gene transfer: building the web of life. Nat Rev Genet 16:472–482
Strausak D, Solioz M (1997) CopY is a copper-inducible repressor of the Enterococcus hirae copper ATPases. J Biol Chem 272:8932–8936
Sun Y, Wong MD, Rosen BP (2001) Role of cysteinyl residues in sensing Pb(II), Cd(II), and Zn(II) by the plasmid pI258 CadC repressor. J Biol Chem 276:14955–14960
Suzuki K, Wakao N, Kimura T, Sakka K, Ohmiya K (1998) Expression and regulation of the arsenic resistance operon of Acidiphilium multivorum AIU 301 plasmid pKW301 in Escherichia coli. Appl Environ Microbiol 64:411–418
Thelwell C, Robinson NJ, Turner-Cavet JS (1998) An SmtB-like repressor from Synechocystis PCC 6803 regulates a zinc exporter. Proc Natl Acad Sci USA 95:10728–10733
Tottey S, Harvie DR, Robinson NJ (2005) Understanding how cells allocate metals using metal sensors and metallochaperones. Acc Chem Res 38:775–783
Touati D, Jacques M, Tardat B, Bouchard L, Despied S (1995) Lethal oxidative damage and mutagenesis are generated by iron in delta fur mutants of Escherichia coli: protective role of superoxide dismutase. J Bacteriol 177:2305–2314
Tran N, Mir A, Mallik D, Sinha A, Nayar A, Webster TJ (2010) Bactericidal effect of iron oxide nanoparticles on Staphylococcus aureus. Int J Nanomedicine 5:277–283
Tuffin IM, de Groot P, Deane SM, Rawlings DE (2005) An unusual Tn21-like transposon containing an ars operon is present in highly arsenic-resistant strains of the biomining bacterium Acidithiobacillus caldus. Microbiology 151:3027–3039
Tuffin IM, Hector SB, Deane SM, Rawlings DE (2006) Resistance determinants of a highly arsenic-resistant strain of Leptospirillum ferriphilum isolated from a commercial biooxidation tank. Appl Environ Microbiol 72:2247–2253
Turner JS, Robinson NJ (1995) Cyanobacterial metallothioneins: biochemistry and molecular genetics. J Ind Microbiol 14:119–125
Turner RJ, Hou Y, Weiner JH, Taylor DE (1992) The arsenical ATPase efflux pump mediates tellurite resistance. J Bacteriol 174:3092–3094
Turner JS, Glands PD, Samson AC, Robinson NJ (1996) Zn2+-sensing by the cyanobacterial metallothionein repressor SmtB: different motifs mediate metal-induced protein-DNA dissociation. Nucleic Acids Res 24:3714–3721
UniProt Consortium (2015) UniProt: a hub for protein information. Nucleic Acids Res 43:D204–D212
Valls M, Atrian S, de Lorenzo V, La F (2000) Engineering a mouse metallothionein on the cell surface of Ralstonia eutropha CH34 for immobilization of heavy metals in soil. Nat Biotechnol 18:661–665
van Kranenburg R, Golic N, Bongers R, Leer RJ, de Vos WM, Siezen RJ, Kleerebezem M (2005) Functional analysis of three plasmids from Lactobacillus plantarum. Appl Environ Microbiol 71:1223–1230
VanZile ML, Cosper NJ, Scott RA, Giedroc DP (2000) The zinc metalloregulatory protein Synechococcus PCC7942 SmtB binds a single zinc ion per monomer with high affinity in a tetrahedral coordination geometry. Biochemistry 39:11818–11829
VanZile ML, Chen X, Giedroc DP (2002a) Structural characterization of distinct alpha3N and alpha5 metal sites in the cyanobacterial zinc sensor SmtB. Biochemistry 41:9765–9775
VanZile ML, Chen X, Giedroc DP (2002b) Allosteric negative regulation of smt O/P binding of the zinc sensor, SmtB, by metal ions: a coupled equilibrium analysis. Biochemistry 41:9776–9786
Velisek J (2013) The chemistry of food. John Wiley & Sons, Hoboken
Vierke G, Engelmann A, Hebbeln C, Thomm M (2003) A novel archaeal transcriptional regulator of heat shock response. J Biol Chem 278:18–26
Vilas-Bôas GT, Peruca AP, Arantes OM (2007) Biology and taxonomy of Bacillus cereus, Bacillus anthracis, and Bacillus thuringiensis. Can J Microbiol 53:673–687
Vinceti M, Wei ET, Malagoli C, Bergomi M, Vivoli G (2001) Adverse health effects of selenium in humans. Rev Environ Health 16:233–251
Vorontsov II, Minasov G, Brunzelle JS, Shuvalova L, Kiryukhina O, Collart FR, Anderson WF (2007) Crystal structure of an apo form of Shigella flexneri ArsH protein with an NADPH-dependent FMN reductase activity. Protein Sci 16:2483–2490
Wächtershäuser G (2000) Origin of life. Life as we don’t know it. Science 289:1307–1308
Wächtershäuser G (2002) Discussing the origin of life. Science 298:747–749
Waldron KJ, Robinson NJ (2009) How do bacterial cells ensure that metalloproteins get the correct metal? Nat Rev Microbiol 7:25–35
Wang CL, Michels PC, Dawson SC, Kitisakkul S, Baross JA, Keasling JD, Clark DS (1997) Cadmium removal by a new strain of Pseudomonas aeruginosa in aerobic culture. Appl Environ Microbiol 63:4075–4078
Wang G, Kennedy SP, Fasiludeen S, Rensing C, DasSarma S (2004) Arsenic resistance in Halobacterium sp. strain NRC-1 examined by using an improved gene knockout system. J Bacteriol 186:3187–3194
Wang Y, Hemmingsen L, Giedroc DP (2005) Structural and functional characterisation of Mycobacterium tuberculosis CmtR, a PbII/CdII-sensing SmtB/ArsR metalloregulatory repressor. Biochemistry 44:8976–8988
Wang L, Chen S, Xiao X, Huang X, You D, Zhou X, Deng Z (2006) arsRBOCT arsenic resistance system encoded by linear plasmid pHZ227 in Streptomyces sp. strain FR-008. Appl Environ Microbiol 72:3738–3742
Wang L, Jeon B, Sahin O, Zhang Q (2009a) Identification of an arsenic resistance and arsenic-sensing system in Campylobacter jejuni. Appl Environ Microbiol 75:5064–5073
Wang SC, Dias AV, Zamble DB (2009b) The “metallo-specific” response of proteins: a perspective based on the Escherichia coli transcriptional regulator NikR. Dalton Trans 14:2459–2466
Wang Y, Kendall J, Cavet JS, Giedroc DP (2010) Elucidation of the functional metal binding profile of a Cd(II)/Pb(II) sensor CmtR(Sc) from Streptomyces coelicolor. Biochemistry 49:6617–6626
Wang H, Ayala JC, Benitez JA, Silva AJ (2015) RNA-seq analysis identifies new genes regulated by the histone-like nucleoid structuring protein (H-NS) affecting Vibrio cholerae virulence, stress response and chemotaxis. PLoS One 10:e0118295
Wang L, Zhuang X, Zhuang G, Jing C (2016) Arsenic resistance strategy in Pantoea sp. IMH: organization, function and evolution of ars genes. Sci Rep 6:39195
Wang F, Fu SN, Bao YX, Yang Y, Shen HF, Lin BR, Zhou GX (2017) Kitamycin C, a new antimycin-type antibiotic from Streptomyces antibioticus strain 200-09. Nat Prod Res 28:1–6
Warnes SL, Caves V, Keevil CW (2012) Mechanism of copper surface toxicity in Escherichia coli O157:H7 and Salmonella involves immediate membrane depolarization followed by slower rate of DNA destruction which differs from that observed for gram-positive bacteria. Environ Microbiol 14:1730–1743
Weinberg ED (2009) Iron availability and infection. Biochim Biophys Acta 1790:600–605
White C, Lee J, Kambe T, Fritsche K, Petris MJ (2009) A role for the ATP7A copper-transporting ATPase in macrophage bactericidal activity. J Biol Chem 284:33949–33956
Williams SG, Manning PA (1991) Transcription of the Vibrio cholerae haemolysin gene, hlyA, and cloning of a positive regulatory locus, hlyU. Mol Microbiol 5:2031–2038
Williams SG, Attridge SR, Manning PA (1993) The transcriptional activator HlyU of Vibrio cholerae: nucleotide sequence and role in virulence gene expression. Mol Microbiol 9:751–760
Woese CR, Kandler O, Wheelis ML (1990) Towards a natural system of organisms: proposal for the domains archaea, bacteria, and eucarya. Proc Natl Acad Sci USA 87:4576–4579
Wong MD, Lin YF, Rosen BP (2002) The soft metal ion binding sites in the Staphylococcus aureus pI258 CadC Cd(II)/Pb(II)/Zn(II)-responsive repressor are formed between subunits of the homodimer. J Biol Chem 277:40930–40936
Wright JB, Lam K, Burell RE (1998) Wound management in an era of increasing bacterial antibiotic resistance: a role for topical silver treatment. Am J Infect Control 26:572–577
Wu J, Rosen BP (1991) The ArsR protein is a trans-acting regulatory protein. Mol Microbiol 5:1331–1336
Wu J, Rosen BP (1993) Metalloregulated expression of the ars operon. J Biol Chem 268:52–58
Xu FF, Imlay JA (2012) Silver(i), mercury(ii), cadmium(ii), and zinc(ii) target exposed enzymic iron-sulfur clusters when they toxify Escherichia coli. Appl Environ Microbiol 78:3614–3621
Xu C, Shi W, Rosen BP (1996) The chromosomal arsR gene of Escherichia coli encodes a trans-acting metalloregulatory protein. J Biol Chem 271:2427–2432
Yaganza ES, Rioux D, Simard M, Arul J, Tweddell RJ (2004) Ultrastructural alterations of Erwinia carotovora subsp. atroseptica caused by treatment with aluminum chloride and sodium metabisulfite. Appl Environ Microbiol 70:6800–6808
Yang HC, Cheng J, Finan TM, Rosen BP, Bhattacharjee H (2005) Novel pathway for arsenic detoxification in the legume symbiont Sinorhizobium meliloti. J Bacteriol 187:6991–6997
Yang Y, Cui HL, Zhou PJ, Liu SJ (2007) Haloarcula amylolytica sp. nov., an extremely halophilic archaeon isolated from Aibi salt lake in Xin-Jiang, China. Int J Syst Evol Microbiol 57:103–106
Yao ZT, Li JH, Xie HH, Yu CH (2012) Review on remediation technologies of soil contaminated by heavy metals. Proc Environ Sci 16:722–729
Ye J, Kandegedara A, Martin P, Rosen BP (2005) Crystal structure of the Staphylococcus aureus pI258 CadC Cd(II)/Pb(II)/Zn(II)-responsive repressor. J Bacteriol 187:4214–4221
Yoon KP, Misra TK, Silver S (1991) Regulation of the cadA cadmium resistance determinant of Staphylococcus aureus plasmid pI258. J Bacteriol 173:7643–7649
Yoon KY, Byeon JH, Park JH, Hwang J (2007) Susceptibility constants of Escherichia coli and Bacillus subtilis to silver and copper nanoparticles. Sci Total Environ 373:572–575
Yu X, Zheng W, Bhat S, Aquilina JA, Zhang R (2015) Transcriptional and posttranscriptional regulation of Bacillus sp. CDB3 arsenic-resistance operon ars1. PeerJ 3:e1230
Zhang YM, Rock CO (2008) Membrane lipid homeostasis in bacteria. Nature Rev Microbiol 6:222–233
Zhao H, Volkov A, Veldore VH, Hoch JA, Varughese KI (2010) Crystal structure of the transcriptional repressor PagR of Bacillus anthracis. Microbiology 156:385–391
Zouboulis AI, Loukidou MX, Matis KA (2004) Biosorption of toxic metals from aqueous solutions by bacteria strains isolated from metal-polluted soils. Process Biochem 39:909–916
Acknowledgements
This work was supported by an Early Career Research grant (ECR/2016/001598) from DST-SERB, India to Dr. Rudra P. Saha.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Saha, R.P., Samanta, S., Patra, S. et al. Metal homeostasis in bacteria: the role of ArsR–SmtB family of transcriptional repressors in combating varying metal concentrations in the environment. Biometals 30, 459–503 (2017). https://doi.org/10.1007/s10534-017-0020-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10534-017-0020-3