Abstract
Species distribution data provide critical baseline information for conservation planning and decision making. However, in many of the Earth’s most biodiverse regions, such data are lacking for many species. Here, we used ecological niche modeling and connectivity analyses to model distributions of endangered species and protected area connectivity across the Upper Guinea Forest (UGF) Global Conservation Hotspot of West Africa. We estimated the current distributions of African forest elephant Loxodonta cyclotis (Vulnerable), western chimpanzee Pan troglodytes verus (Critically Endangered), and pygmy hippopotamus Choeropsis liberiensis (Endangered) across the region and optimized connectivity in two main forest complexes in the region. We used occurrence data for the period 2010–2016 for the three species from two well-sampled national parks in Liberia (Sapo National Park and Gola National Park), and remotely sensed MODIS enhanced vegetation index data for the period 2010–2015. Our models predicted a total of 75,157 km2 of suitable habitat for chimpanzees in the region, 79,400 km2 for elephants, and 290,696 km2 for hippos. Of these areas, for chimpanzees, 30% of the area predicted falls within the boundaries of proposed or designated protected areas, and likewise 30% for elephants, and 19% for hippos. Liberia had the largest blocks of contiguous forest suitable for these species compared to other countries in the region but this forest was largely unprotected. This study identifies priority areas for biodiversity conservation and forest connectivity in the region, and reemphasizes the practicality of these tools to optimize conservation planning and implementation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biodiversity conservation is a global priority, yet garnering resources to meet global conservation priorities is challenging, in largest part owing to escalating threats to biodiversity, particularly in biodiversity-rich regions of the world (Squires 2014). These threats, including forest fragmentation, habitat degradation and destruction, overhunting, disease associated with human activities, and a myriad of negative impacts of climate change, are pushing endangered species to the brink of extinction (Thomas et al. 2004; Pimm et al. 2014).
To optimize biodiversity conservation and reduce threats in the wake of limited resources, conservation biologists have employed conservation triage (i.e., prioritize investments for maximum returns, given limited resources (Wilson et al. 2007; Bottrill et al. 2008, 2009), ecosystem/landscape level conservation, in situ species-level conservation, and ex situ captive breeding programs (Primack 2010). The debate over whether species-level strategies versus ecosystem/landscape-level approaches may lead to greater success continues unabated (Lindenmayer et al. 2007; Franklin 1993). Whereas Franklin (1993) argued that ecosystem/landscape level conservation is cost-efficient and holistic in species coverage (e.g., including microbes), Lindenmayer et al. (2007) argued that species-level strategies provide useful baseline information for policy and management, particularly for threatened species, keystone species, and invasive species, that have specific impacts on the environment. Carignan and Villard (2002), however, cautioned that, whatever the case may be, conservationists should select indicators carefully, to optimize biodiversity conservation.
Several habitat connectivity modeling approaches have been used in biodiversity science to optimize species and ecosystem conservation in a combined effort (Ball et al. 2009; McRae et al. 2008; Brás et al. 2013; Lehtomäki and Moilanen 2013; Cushman et al. 2018; Hearn et al. 2018; Khosravi et al. 2018; Zacarias and Loyola 2018). These approaches are generally based on least-cost analysis of resistance surfaces (e.g., habitat suitability, topography) to assess connectivity between distributional areas, but vary in input data requirements, scale of analysis, and conservation goals. Here, we explore the use of correlative ecological niche modeling to build resistance surfaces (niche models) for habitat connectivity analysis to identify spatial patterns of connectivity in the humid rainforest of West Africa. Correlative ecological niche modeling (ENM) uses known occurrences of a species and associated environmental data (e.g., temperature, precipitation) to identify areas of similar suitable environments for the species (Peterson et al. 2011). Ecological niche models are usually developed using machine-learning algorithms to characterize current and potential distributions of species in geographic and environmental spaces, and has been adopted widely in the scientific community, with notable applications in conservation, climate change, biogeography, and zoonotic disease research (Jennings and Veron 2015; Peterson et al. 2017; DeMatteo et al. 2017). A landscape connectivity approach, implemented in the Circuitscape program, which uses electronic circuit theory to predict patterns of animal movement between fragmented or heterogonous landscapes (e.g., connecting protected areas; McRae et al. 2008, 2016). Circuitscape uses resistance surfaces between fragmented range areas to map corridors with greater connectivity. Like ENM applications, several studies have applied Circuitscape to diverse challenges in large mammal conservation (Beier et al. 2011; Roever et al. 2013; Dickson et al. 2013).
Over the last 200 years, West Africa has been identified to rank among the world’s most heavily deforested regions, leaving its unique biota severely threatened (Allport 1991; Darwall et al. 2015). The remaining portions of the region’s well-known humid Upper Guinea lowland rainforest are highly fragmented; the exception is Liberia, currently possessing ~ 42% of its original forest in a relatively well-preserved, extensive, and continuous state (Liu et al. 2016). However, an increasing wave of investments from the agricultural and mining sectors raises concerns about threats to the remaining biodiversity (Primack 2010). Regional efforts have begun to address these threats, but little is known about the ecology and distributions of most West African species. If better documented and analyzed, distributional data could be leveraged to inform conservation programs and at least partially mitigate development-associated habitat fragmentation.
In this study, we integrated these two techniques to address questions about endangered species distributions, and protected area connectivity across the Upper Guinea Forest (UGF) of West Africa. Specifically, we aimed to (1) use correlative ENM to predict the current distributions of African forest elephant (Loxodonta cyclotis; Vulnerable), western chimpanzee (Pan troglodytes verus; Critically Endangered), and pygmy hippopotamus (Choeropsis liberiensis; Endangered) (hereafter referred to as chimpanzee, elephant, and hippo) across the humid lowland UGF of West Africa. Then, (2) we use the resulting models as surrogates to explore landscape connectivity and identify ideal movement corridors to connect protected areas across Liberia and neighboring countries (Guinea, Côte d’Ivoire, and Sierra Leone) using Circuitscape. We identified 14 suitable corridors to connect protected areas within two main forest complexes (Gola-Ziama and Sapo-Tai); these corridors likely represent practical candidate regions for applied conservation action.
Materials and methods
This study was conducted in the UGF, one of the two major humid tropical lowland rainforests blocks of western and central Africa (Fig. 1). We fitted ENM models in the portion of UGF from western Ghana to western Guinea to characterize suitable habitats for our target species, while forest connectivity analysis covered the portion of UGF within the Manor River Basin (Guinea, Sierra Leone, Liberia, and Côte d’Ivoire; Fig. 1). We used three severely threatened, but well-sampled/documented species (chimpanzee, elephant, hippo) to answer our questions. These species were selected because they are known to be flagship/umbrella species with large home ranges, have high ecological importance, well-sampled, and are of high global conservation priority.
Input data
Ecological niche modeling requires two types of input datasets: environmental data and georeferenced species occurrence data. Environmental data typically include layers summarizing aspects of temperature, precipitation, and/or vegetation, all of which may influence the survival of a species. Species occurrence data consist of unique, georeferenced locations where a species has been recorded (Peterson et al. 2011). Occurrence data were split into calibration and evaluation subsets (Peterson et al. 2011). The geographic space was divided into a calibration area (M), the area that has been accessible to the species over relevant time periods (we further reduced this area to the area from which we had available detailed sampling, Sapo National Park or Sapo NP), and a projection area (see below), a larger area of interest to which models are transferred in space or time (Peterson et al. 2011).
We used the NASA Moderate-resolution Imaging Spectroradiometer (MODIS) enhanced vegetation index (EVI) dataset at 250 m spatial and 16-day temporal resolutions to characterize the environmental landscape for the species (www.modis.gsfc.nasa.gov). MODIS vegetation indices have high spatial resolution and provide a realistic characterization of place and time specific environmental conditions (Bodbyl-Roels et al. 2011; Feilhauer et al. 2012; Peterson 2014). We downloaded 16-day composite MODIS Terra Satellite EVI data for the period 2010–2015- for West Africa. We chose EVI over Normalized Difference Vegetation Index (NDVI) because of EVI’s improved sensitivity over dense vegetation conditions, which is the case in our study region. Further processing of these data is described below.
We used primary occurrence data collected over the period 2010–2016, as part of ongoing systematic biomonitoring projects and other surveys across Sapo NP in southeastern Liberia and Gola National Park (Gola NP) in northwestern Liberia (e.g., Tweh et al. 2015; Fig. 1). All models were calibrated in Sapo NP and tested in Gola NP (Fig. 1). Sapo NP was chosen as our calibration area (M) because it is well-sampled and contains high-quality primary occurrence data for each of our species of interest; Gola NP was well-sampled and independent of Sapo NP. All data were collected as part of systematic surveys that used GPS for establishing geographic coordinates. Occurrence data for both Sapo NP and Gola NP were imported into ArcGIS and converted to point shapefiles. We used SDM Toolbox to subsample high-density cells and remove duplications in occurrence data (Brown 2014). At the end of the cleaning exercise, we calibrated our models with a total of 40 occurrence points for hippo, 120 for chimpanzee, and 59 for elephant.
Data processing and analysis
The original set of MODIS EVI data layers was mosaicked and cleaned in ArcGIS version 10.3. Cloudy images or artifacts were removed and excluded from further analysis, leaving images for 105 time periods that were included in model fitting. We applied principal components analysis (PCA) to the remaining 105 images to reduce dimensionality and correlation among data layers. PCA extracts the major axes of variation in a dataset and compresses them into fewer orthogonal variables or principal components, with the first PC explaining the most variation in the dataset (Abdi and Williams 2010). In our case, the first 46 PCs accounted for 95% of the variation in the data. These 46 PCs were converted to ASCII file format for ENM. We used the extract by mask (batch) tool in ArcGIS to extract Sapo NP (the model calibration area) from the 46 PCs which were used in calibrating models.
Ecological niche modeling
To estimate the current distribution of our focal species, we used Maxent version 3.3.3 k, a machine-learning software platform widely used to model ecological niches of species (Phillips et al. 2006; Phillips and Dudík 2008). Before fitting our models, we further reduced the 46 PCs for each species using the delete-one jackknife approach in Maxent with each species’ occurrence data and the 46 PCs (Wold et al. 1987; Abdi and Williams 2010; Shcheglovitova and Anderson 2013). To do this step, we set the random test percentage to 50%, output format to logistic, and left all other features at default. Once the variable set was reduced, we desired to assess many calibration conditions to be able to choose optimal models. As a result, we used six combinations of feature classes (Linear, Quadratic, Product, Threshold and Hinge) and five regularization multiplier values (0.1, 0.5, 1, 2, and 5) in Maxent (Phillips and Dudík 2008; Shcheglovitova and Anderson 2013; Muscarella et al. 2014). We further explored two environmental data sets for each species: one set using the jackknife-selected PCs, and another using the first nine PCs (1–9) that accounted for 85% of the variation in our dataset from the 46 PCs. We used these nine PCs separately because we observed that most of the PCs selected by the jackknife approach were not within the first ten PCs for all the species. We also used transects surveyed in Sapo NP as bias surfaces for each model to reflect survey effort (Fourcade et al. 2014), which constituted another variable in the model selection process.
To choose optimal models, we assessed model significance, performance, and complexity, in candidate models for each species. We used ENMtools (Warren et al. 2010) to generate scores for sample-corrected Akaike Information Criterion (AICc) for each model to assess complexity. We used Niche Toolbox for partial ROC significance test (http://shiny.conabio.gob.mx:3838/nichetoolb2/; Peterson et al. 2008) and we calculated an omission rate for each species (based on a 5% training presence threshold). Also, we calculated the proportion of areas predicted by each model and used this quantity in a subsequent binomial significance test. Best models for each species were selected based on (1) partial ROC and binomial significance tests, (2) minimum omission rates, and (3) model complexity (Table 1), in that order. Best models were projected across the region to predict suitable habitats for each species, both with and without clamping and extrapolation. We then took the median of the medians of the final models for each species and applied a 5% minimum training presence threshold to produce a binary distribution map. Finally, we used Mobility Oriented-Parity (MOP) analysis based on 10% sampling of the reference region to test for extrapolation in transfers of model predictions (Owens et al. 2013).
To identify suitable corridors for connectivity between suitable protected areas for our species of concern identified in our ENMs, we used 15 proposed and designated protected areas in the two major forest complexes in Liberia, Guinea, Sierra Leone, and Côte d’Ivoire (Fig. 3). These forest complexes together comprise > 60% of remaining forest in the UGF region. We used Circuitscape version 3.5 and its auxiliary software, Linkage Mapper (which uses least-cost corridor analysis to identify and map linkages between core areas) and Pinchpoint Mapper (which uses Circuitscape to identify pinch-points or bottlenecks in corridors produced by Linkage Mapper; McRae et al. 2016, 2008) to model connectivity between proposed and protected areas for each species within these forest complexes. Circuitscape uses electronic circuit theory concept to predict patterns of animal movement between fragmented or heterogeneous landscapes (McRae et al. 2008). To do this, it requires a resistance surface layer(s) and layers of the areas to be connected or core areas. We rescaled the continuous versions of our individual best models for each species from 250 m spatial resolution to 1 km resolution to reduce processing time, summed them using raster calculator tool in ArcGIS, and took the additive inverse (i.e., subtracting from unity) of each sum to recreate resistance surfaces. Our resistance values range between 1 and 100, representing low to high resistance. We used these resistance layers together with shapefiles of proposed and designated protected areas in the two forest complexes to build connectivity models (Fig. 3). To build networks and linkages between our core areas, we used pairwise analysis between core areas in each forest complex separately. We set the cut-off distance (cost-weighted corridor width) at 20 km.
Results
We calibrated a total of 120 models (5 values of regularization multiplier × 6 combinations feature classes x 2 sets of environmental variables x bias/no bias surface) for each species. Of the 120 models fitted for chimpanzees, three were selected as best. These models were significantly different from random and had relatively low omission rates < 28% (Table 1).
For elephants, three models were selected; however, none of the three models met our criteria for model selection. They had reasonable omission rates, but were not statistically significant, and were the most complex models of all species (Table 1). Eight models were selected for hippo, all of which were significantly different from random, less complex, and had low omission rates. All the best models for chimpanzees and elephants were models derived from the first PC variables, whereas the best models for hippos were those from jackknife-selected variables.
Our models predicted a total of 75,157 km2 of suitable habitats for chimpanzees in the region, 79,400 km2 for elephants, and 290,696 km2 for hippos (Fig. 2). Of these areas, for chimpanzees, 30% of the area predicted falls within the boundaries of proposed or designated protected areas, and likewise 30% for elephants, and 19% for hippos (Fig. 4). For both chimpanzees and elephants, Liberia held the largest area of suitable habitats (51 and 53% of total areas, respectively), followed by Côte d’Ivoire (33% for chimpanzees and 32% for elephants), and Ghana (12 and 11%; Fig. 4). Thirty-nine percent of suitable habitat for both species predicted in Liberia was within proposed or designated protected areas, whereas in Côte d’Ivoire, 48% was within the boundaries of protected areas; in Ghana, 14% fell within the boundaries of protected areas. Guinea and Sierra Leone had the least area of suitable habitats for both species, with Guinea hosting 1.3% for chimpanzees and 1.6% for elephants, and Sierra Leone hosting 2.7% for chimpanzees and 2.3% for elephants (Fig. 4). Côte d’Ivoire held the largest predicted area for hippos (35%), followed by Liberia 25%, and Ghana (19%; Fig. 4). Connectivity analysis showed that Liberia has the largest blocks of suitable contiguous forest compared to Côte d’Ivoire, Guinea, and Sierra Leone, which had much more fragmented forest landscapes (Fig. 3). The forest blocks are concentrated in southeastern Liberia, with Sapo NP connecting to Tai National Park in southwestern Côte d’Ivoire to the northwest, and in northwestern Liberia, with Gola NP connecting to Gola Rainforest National Park in Sierra Leone to the west, and Ziama Forest in Guinea to the north. Our connectivity analysis in the two forest complexes identified a total of 14 suitable corridors to connect proposed and designated protected areas in the two forest complexes, many of which overlapped among the three species (Fig. 3, Table 2).
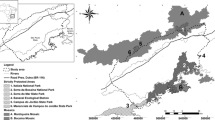
Suitable corridors for connectivity of proposed and protected areas predicted within the Sapo-Tai and Gola-Ziama forest complexes in West Africa based on least cost path. 1. Tai National Park 2. Krahn-Grebo National Park 3. Sapo National Park 4. Grand Kru Proposed Protected Area (PPA) 5. Senkwen PPA 6. Gbi PPA 7. Zwedru PPA 8. Gouin Park 9. Scio Park 10. Kpo PPA 11. Gola National Park 12. Foya PPA 13. Wologizi PPA 14. Ziama Forest Park 15. Mt. Yonon
Discussion
Model predictions
ENM provides a framework for estimating geographic distributions of species; however, ENMs are cast on geographic extents, and as such do not and cannot take into account finer-scale phenomena such as land use and land cover distributions. In ENM, the best models are those that meet set criteria of statistical significance, model complexity, and performance. However, in this study, we found no agreement between AICc and the other two criteria. Even though the AICc metric has been widely adopted in the field of ENM as a criterion (Shcheglovitova and Anderson 2013; Warren et al. 2013; Muscarella et al. 2014), we caution that it should be used with care and along with other criteria as used in this study. In particular, the best models selected for elephants did not meet our criteria, perhaps for two reasons. First, the species as sampled here within Sapo NP only, is a classic example of a “Wallace’s Dream species,” as described in Saupe et al. (2012). In this situation, a species’ distributional area is constrained by dispersal capabilities, and models for such species will not be significantly different from random because all habitats in the species’ calibration area are suitable. The second related reason could be attributed to our selection of Sapo NP as a calibration area, which was influenced by the availability of high-quality data. We recognize that this feature was a limitation of this study that should be carefully considered in future studies (Barve et al. 2011).
However, in spite of the statistical uncertainty in the elephant models, their biological explanation is relevant. Areas of suitable habitats predicted by the elephant models are similar to those predicted by chimpanzees, as we expected given the broader distribution of these species in the region. Unlike the related African elephant Loxodonta africana, African forest elephants inhabit the dense humid tropical rainforest of west and central Africa and have overlapping ranges with western chimpanzees. In Sapo NP, for example, these species inhabit the same sites, as evident by camera trapping records (Vogt 2012). In addition, we tested our chimpanzee models using independent data from several chimpanzee projects across the region. We found that our predictions strongly correlated with areas currently occupied by chimpanzees, including areas in Sierra Leone and Guinea, which are known important sanctuaries for chimpanzees and elephants in the region. A notable example is the Western Area Peninsula National Park in Sierra Leone, a designated World Heritage Site that hosts 80–90% of Sierra Leone’s terrestrial biodiversity, including chimpanzee (UNESCO World Heritage 2018), which was identified as a disjunct, but highly suitable area for chimpanzees.
In contrast, and perhaps of greatest concern in this study, hippo models met all of our criteria for statistical significance, model complexity, and performance, and yet the models seemed to be underfit and overly general. Of the three species, hippos use the most restricted habitats (aquatic forest habitats); as such, the widespread distributional potential of the species across the region compared to elephants and chimpanzees was unexpected. This could be attributed to the spatial bias in our dataset as articulated above and explained by Warren et al. (2013). Thus, we caution that the hippo models should be interpreted with care.
Overall, an important consideration in interpretation of ENM results is the movement capability (mobility) of a species. That is, uninhabited suitable areas predicted in these models can only be occupied if dispersal barriers do not prevent these species from colonizing them (Peterson et al. 2011). Therefore, providing suitable corridors that connect suitable habitats will enhance these dispersal abilities of these species to colonize uninhabited habitats.
Species distributions and forest connectivity
Using ENM to inform conservation planning is a relatively novel approach in Africa, particularly for West Africa (Iloh and Ogundipe 2016). This study predicted areas of suitable habitat for each of the three focal species across the western sector of the Upper Guinea Forest block, highlighting priority areas inside and outside of protected areas. One notable result was the congruence between predicted suitable habitats and the distribution of the remaining forest in the region, as reported in other studies (Allport 1991; Liu et al. 2016). Our models clearly showed that Liberia retains the largest contiguous portions of suitable habitats for these species, followed by Côte d’Ivoire. However, most Liberian forest areas remain unprotected, far short of its neighbors, which have all major forest patches designated as protected areas. Since 2006, the Liberian Government set a goal of protecting 30% of the country’s forest in a network of protected areas (NFRL 2006); this goal is yet to be realized, with just ~ 10% of the country’s forest presently designated as protected areas.
Our connectivity analysis identified 14 corridors and three potential sites for establishment of new protected areas in both Gola-Ziama and Sapo-Tai forest complexes, several of which coincide and at least partially agree with results of previous analyses (Junker et al. 2015), reinforcing the importance of these areas as putative habitat corridors. The southeastern block of forest (in Liberia) connecting to southwestern Côte d’Ivoire (Sapo-Tai forest complex) should be considered a particularly significant regional priority. This forest block holds > 50% of the contiguous forest suitable for our focal species in the region. Interestingly, the Liberian side of this forest is largely unprotected, and is under increasing threats from oil palm (Elias guinesis) plantation expansion, artisanal mining, and logging (Freeman et al. 2018). We emphasize here that the potential to preserve this forest block should focus on these areas with urgency. We acknowledge current regional initiatives aimed at this forest block, one of which led to the recent (2017) designation of the Krahn-Grebo National Park. However, even though we applaud these efforts, they should be expanded to reflect the proportional significance of this area. Additionally, the Gola-Ziama forest complex, in northwestern Liberia (bordering Sierra Leone and Guinea), is notable for its potential conservation urgency, given that it is also largely unprotected and equally threatened. We recognize efforts of the last decade resulting in designation of Gola National Park in Liberia and Gola Rainforest National Park in Sierra Leone; the northern portion of this forest block, extending north to Ziama in Guinea, is characterized by highly heterogeneous natural landscapes (e.g., submontane forest, semi-deciduous forest), and many unique, high-value species for conservation. Additionally, it should be noted here that to optimize biodiversity conservation, conservation efforts should not only be focused on forest protection but should also address other major drivers of biodiversity loss, particularly hunting, a major threat to West African biodiversity to avoid the empty forest syndrome (Wilkie et al. 2011).
We show that the integration of ENM and connectivity analysis studies provides powerful practical tools to optimize conservation planning and identify priority areas objectively for implementation (conservation action), with the goal of optimizing connectivity among distributional areas of species. These tools, when adopted and applied appropriately, will accelerate knowledge of species distribution and conservation efforts in incompletely-known but highly biodiverse regions of the world. We emphasize that connecting suitable habitats for these species will enhance their dispersal abilities and survivability in the region.
References
Abdi H, Williams LJ (2010) Principal component analysis. Wiley Interdiscip Rev Comput Stat 2:433–459
Allport G (1991) The status and conservation of threatened birds in the Upper Guinea forest. Bird Conserv Int 1:53–74
Ball IR, Possingham HP, Watts M (2009) Marxan and relatives: software for spatial conservation prioritisation. In: Spatial conservation prioritisation: quantitative methods and computational tools, pp 185–195
Barve N, Barve V, Jiménez-Valverde A et al (2011) The crucial role of the accessible area in ecological niche modeling and species distribution modeling. Ecol Model 222:1810–1819
Beier P, Spencer W, Baldwin RF, McRAE BH (2011) Toward best practices for developing regional connectivity maps: regional connectivity maps. Conserv Biol 25:879–892
Bodbyl-Roels S, Peterson AT, Xiao X (2011) Comparative analysis of remotely-sensed data products via ecological niche modeling of avian influenza case occurrences in Middle Eastern poultry. Int J Health Geogr 10:1–21
Bottrill MC, Joseph LN, Carwardine J et al (2008) Is conservation triage just smart decision making? Trends Ecol Evol 23:649–654
Bottrill MC, Joseph LN, Carwardine J et al (2009) Finite conservation funds mean triage is unavoidable. Trends Ecol Evol 24:183–184
Brás R, Cerdeira JO, Alagador D, Araújo MB (2013) Linking habitats for multiple species. Environ Model Softw 40:336–339
Brown JL (2014) SDMtoolbox: a python-based GIS toolkit for landscape genetic, biogeographic and species distribution model analyses. Methods Ecol Evol 5:694–700
Carignan V, Villard M-A (2002) Selecting indicator species to monitor ecological integrity: a review. Environ Monit Assess 78:45–61
Cushman SA, Elliot NB, Bauer D et al (2018) Prioritizing core areas, corridors and conflict hotspots for lion conservation in southern Africa. PLoS ONE 13:e0196213
Darwall W, Polidoro B, Smith K, Somda J (2015) Ecosystem profile Guinean Forests of West Africa biodiversity hotspot. Critical Ecosystem Partnership Fund Report
DeMatteo KE, Rinas MA, Zurano JP et al (2017) Using niche-modelling and species-specific cost analyses to determine a multispecies corridor in a fragmented landscape. PLoS ONE 12:e0183648
Dickson BG, Roemer GW, McRae BH, Rundall JM (2013) Models of regional habitat quality and connectivity for pumas (Puma concolor) in the southwestern United States. PLoS ONE 8:e81898
Feilhauer H, He KS, Rocchini D (2012) Modeling species distribution using niche-based proxies derived from composite bioclimatic variables and MODIS NDVI. Remote Sens 4:2057–2075
Fourcade Y, Engler JO, Rödder D, Secondi J (2014) Mapping species distributions with MAXENT using a geographically biased sample of presence data: a performance assessment of methods for correcting samplingbias. PLoS ONE 9:e97122
Franklin JF (1993) Preserving biodiversity: species, ecosystems, or landscapes? Ecol Appl 3:202–205
Freeman B, Danjuma DF, Molokwu-Odozi M (2018) Status of globally threatened birds of Sapo National Park. Ostrich-J Afr Ornithol, Liberia. https://doi.org/10.2989/00306525.2018.1502694
Hearn AJ, Cushman SA, Goossens B et al (2018) Evaluating scenarios of landscape change for Sunda clouded leopard connectivity in a human dominated landscape. Biol Conserv 222:232–240
Iloh AC, Ogundipe O (2016) Using ecological niche modelsto plan conservation in a changing environment: a case for the plant Chasmanthera dependens Hochst (Menispermaceae) in West Africa. J Ecol Nat Environ 8:1–8
Jennings AP, Veron G (2015) Predicted distributions, niche comparisons, and conservation status of the spotted linsang and banded linsang. Mammal Res 60:107–116
Junker J, Boesch C, Freeman T et al (2015) Integrating wildlife conservation with conflicting economic land-use goals in a West African biodiversity hotspot. Basic Appl Ecol 16:690–702
Khosravi R, Hemami M-R, Cushman SA (2018) Multispecies assessment of core areas and connectivity of desert carnivores in central Iran. Divers Distrib 24:193–207
Lehtomäki J, Moilanen A (2013) Methods and workflow for spatial conservation prioritization using Zonation. Environ Model Softw 47:128–137
Lindenmayer DB, Fischer J, Felton A et al (2007) The complementarity of single-species and ecosystem-oriented research in conservation research. Oikos 116:1220–1226
Liu Z, Wimberly MC, Dwomoh FK (2016) Vegetation dynamics in the Upper Guinean Forest Region of West Africa from 2001 to 2015. Remote Sens 9:1–5
McRae BH, Dickson BG, Keitt TH, Shah VB (2008) Using circuit theory to model connectivity in ecology, evolution, and conservation. Ecology 89:2712–2724
McRae B, Shah V, Edelman A (2016) Circuitscape: modeling landscape connectivity to promote conservation and human health. Nat Conserv
Muscarella R, Galante PJ, Soley-Guardia M et al (2014) ENMeval: an R package for conducting spatially independent evaluations and estimating optimal model complexity for Maxent ecological niche models. Methods Ecol Evol 5:1198–1205
NFRL (2006) National Forestry Reform Law of 2006. http://www.fao.org/forestry/16151-05fd47b845599b5d3a594a9b0240dacff.pdf. Accessed 1 May 2018
Owens HL, Campbell LP, Dornak LL et al (2013) Constraints on interpretation of ecological niche models by limited environmental ranges on calibration areas. Ecol Model 263:10–18
Peterson AT (2014) Mapping disease transmission risk: enriching models using biogeography and ecology. Johns Hopkins University Press, Baltimore
Peterson AT, Papeş M, Soberón J (2008) Rethinking receiver operating characteristic analysis applications in ecological niche modeling. Ecol Model 213:63–72
Peterson AT, Soberón J, Pearson RG et al (2011) Ecological niches and geographic distributions. Princeton University Press, Princeton
Peterson AT, Campbell LP, Moo-Llanes DA et al (2017) Influences of climate change on the potential distribution of Lutzomyia longipalpis sensu lato (Psychodidae: Phlebotominae). Int J Parasitol 10–11:667–674
Phillips SJ, Dudík M (2008) Modeling of species distributions with Maxent: new extensions and a comprehensive evaluation. Ecography 31:161–175
Phillips SJ, Dudík M, Schapire RE (2006) A maximum entropy approach to species distribution modeling. In: Proceedings of the twenty-first international conference on Machine learning. ACM, p 83
Pimm SL, Jenkins CN, Abell R et al (2014) The biodiversity of species and their rates of extinction, distribution, and protection. Science 344:1246752
Primack RB (2010) Essentials of conservation biology. Sinauer Associates, Sunderland
Roever CL, van Aarde RJ, Leggett K (2013) Functional connectivity within conservation networks: delineating corridors for African elephants. Biol Conserv 157:128–135
Saupe EE, Barve V, Myers CE et al (2012) Variation in niche and distribution model performance: the need for a priori assessment of key causal factors. Ecol Model 237–238:11–22
Shcheglovitova M, Anderson RP (2013) Estimating optimal complexity for ecological niche models: a jackknife approach for species with small sample sizes. Ecol Model 269:9–17
Squires D (2014) Biodiversity conservation in Asia. Asia Pac Policy Stud 1:144–159
Thomas CD, Cameron A, Green RE et al (2004) Extinction risk from climate change. Nature 427:145–148
Tweh CG, Lormie MM, Kouakou CY et al (2015) Conservation status of chimpanzees Pan troglodytes verus and other large mammals in Liberia: a nationwide survey. Oryx 49:710–718
UNESCO World Heritage (2018) Western Area Peninsula National Park. In: UNESCO World Herit. Cent. https://whc.unesco.org/en/tentativelists/5741/. Accessed 18 Apr 2018
Vogt M (2012) Biomonitoring & research programme, Sapo National Park: updated plan for research opportunities and priorities in and around Sapo National Park. Fauna Flora Int Liberia
Warren DL, Glor RE, Turelli M (2010) ENMTools: a toolbox for comparative studies of environmental niche models. Ecography 33:607–611
Warren DL, Wright AN, Seifert SN et al (2013) Incorporating model complexity and spatial sampling bias into ecological niche models of climate change risks faced by 90 California vertebrate species of concern. Divers Distrib 20:334–343
Wilkie DS, Bennett EL, Peres CA, Cunningham AA (2011) The empty forest revisited. Ann N Y Acad Sci 1223(1):120–128
Wilson KA, Underwood EC, Morrison SA et al (2007) Conserving biodiversity efficiently: what to do, where, and when. PLoS Biol 5:e223
Wold S, Esbensen K, Geladi P (1987) Principal component analysis. Chemom Intell Lab Syst 2:37–52
Zacarias D, Loyola R (2018) Distribution modelling and multi-scale landscape connectivity highlight important areas for the conservation of savannah elephants. Biol Conserv 224:1–8. https://doi.org/10.1016/j.biocon.2018.05.014
Acknowledgements
This study was made possible by the contribution of data from Fauna & Flora International, Liberia (FFI) for Sapo NP; the Society for Conservation of Nature in Liberia (SCNL), Conservation Society of Sierra Leone (CSSL), and the Royal Society for the Protection of Birds (RSPB) for Gola forest in Liberia and Sierra Leone; and Wild Chimpanzee Foundation and the Pan African Programme at the Max-Planck Institute for other parts of the region. We are also very grateful to developers of the APES database (http://apesportal.eva.mpg.de/database) for providing access to these datasets. Finally, we are thankful to the participants of KU-EEB’s spring semester (2018) scientific writing working group class for critical reviews of previous drafts of this manuscript, Daniel Jiménez-Garcia and Lindsay Campbell for their advice on processing remote sensing data, and Luis Osorio for writing the R script used in the MOP analysis.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Samuel Cushman.
This article belongs to the Topical Collection: Forest and plantation biodiversity.
Rights and permissions
About this article
Cite this article
Freeman, B., Roehrdanz, P.R. & Peterson, A.T. Modeling endangered mammal species distributions and forest connectivity across the humid Upper Guinea lowland rainforest of West Africa. Biodivers Conserv 28, 671–685 (2019). https://doi.org/10.1007/s10531-018-01684-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10531-018-01684-6