Abstract
Plant invasions can cause biotic homogenisation which can have cascading effects on the diversity of invaded ecosystems. These impacts on diversity are likely to be scale-dependent and thus affect different aspects of diversity (i.e. beta, gamma and alpha). For example, the widespread invasion of non-native pine trees causes a loss of plant gamma diversity; however, the effects of this invasion and co-invasion by ectomycorrhizal fungi on belowground fungal communities remain unknown. We established thirteen 400 m2 plots across a Pinus nigra density gradient in Canterbury, New Zealand. We sampled twenty-four soil samples from each plot and extracted and sequenced DNA for fungi from each sample independently, allowing determination of within-sample (α) and plot-scale (γ) diversity and turnover (β-diversity). Pine invasion was associated with a positive unimodal response in soil fungal β-diversity, reflected by an increase in saprotroph diversity at low pine density following a loss of this group of fungi at high pine densities. Pine invasion was also associated with an overall 47.7% loss of fungal α-diversity and a 50% loss of γ-diversity. Loss of diversity correlated to a shift from a saprotroph-dominated fungal community in low pine density plots to an ectomycorrhizal-dominated community in high pine density plots. However, despite the resulting dominance of ectomycorrhizal fungi, there was no increase in γ-diversity of ectomycorrhizal fungi as pine density increased. Our results support the concept that low-density invasions increase ecosystem heterogeneity and therefore beta-diversity, but that as aboveground plant communities become more homogenised there is a dramatic loss of fungal diversity across all scales that could inhibit recovery and restoration of invaded ecosystems.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pinaceae have been introduced globally for timber and pulp, amenity values, and erosion control (Richardson & Rejmánek, 2011). These introductions have led to wide-spread invasions, especially in the southern hemisphere, which have resulted in major ecological and economic problems. In the southern hemisphere, pines co-invade with ectomycorrhizal fungi from their native range (Dickie et al. 2010; Policelli et al. 2020), which through the global movement of fungal inoculum and soils are no longer a limiting factor for pine establishment within their invading range (Pringle et al. 2009). Co-invasion of pines and ectomycorrhizal fungi causes large shifts in biological diversity and ecosystem function (Nuñez & Dickie, 2014).
Biotic homogenisation is a major impact of the co-invasion of pines and ectomycorrhizal fungi (García et al. 2018) and has implications on the diversity of ecosystems (Socolar et al. 2016). For example, homogenisation is particularly discernible in aboveground communities when pines invade largely treeless systems because they can transform open grasslands and shrublands to monodominant closed canopy stands (Nuñez et al. 2017; Rundel et al. 2014). This is observed as a loss of plant diversity at high pine densities, despite pine increasing plant diversity at intermediate densities (Dickie et al. 2011; Sapsford et al. 2020). A decrease in aboveground plant diversity is a common trend in plant invasions at the local scale (Vila et al. 2011), but the impacts of these invasions on the diversity of soil fungal communities (Gomes et al. 2018), particularly saprotrophic communities, remain comparatively unknown.
The effect of pine invasions on fungal diversity may also depend on what scale is examined. Diversity at a plot-scale is a function of both the number of species at points within that plot, and the turnover in species composition between points (termed β-diversity) (Anderson et al. 2006). New Zealand grasslands are spatially heterogeneous in terms of plant community composition, comprising patches of different grass and shrub species, which may create diverse soil conditions supporting a high β-diversity of fungi. At the initial stages of pine invasion, where individual trees create isolated patches of novel soil conditions, environmental heterogeneity and β-diversity may be further increased. However, as pines continue to invade and form a closed, often mono-dominant, forest canopy, environmental heterogeneity and therefore fungal β-diversity may be greatly reduced, resulting in biotic homogenisation.
Since pines co-invade with ectomycorrhizal fungi, we may also see changes in both the composition and diversity of key functional guilds, especially when pines invade open grass- and shrublands. Open grass- and shrublands have distinct soil fungal communities from closed canopy forest stands (e.g. Chen et al. 2008; Wang et al. 2019). For example, as ectomycorrhizal fungi are largely absent from grasslands, we expect the effect of pine invasion on ectomycorrhizal fungal diversity to be positive. This positive effect may be small, as Hayward et al. (2015) suggested that pine invasion could be driven by a single species of ectomycorrhizal fungus, while Dickie et al. (2010) found that even well-established pine invasions were species poor (14 ectomycorrhizal operational taxonomic units (OTUs)). An increased ectomycorrhizal diversity is likely driven by the addition of late-stage fungal partners such as Amanita muscaria through succession (Vellinga et al. 2009; Vlk et al. 2020a), but ectomycorrhizal succession in southern hemisphere pine invasions has not been documented. Other functional groups, such as saprotrophs, also may respond to invasion. Saprotrophs are involved in the break-down of organic matter and dominate grassland fungal communities. However, in contrast to the arbuscular mycorrhizal fungi that dominate grasslands, ectomycorrhizal fungi have enzymatic capabilities to break-down organic matter and hence can inhibit saprotrophs (Fernandez & Kennedy, 2016; Lindahl & Tunlid, 2015; Orwin et al. 2011; Sterkenburg et al. 2018). We may expect that pine invasions may lead to the exclusion of saprotrophic fungi where ectomycorrhizal fungi become dominant (Clemmensen et al. 2015; Kyaschenko et al. 2017) and cause a decrease in saprotrophic fungal diversity. Certain species of ectomycorrhizal fungi (e.g. Cortinarius) may complement saprotrophs as decomposers to facilitate nutrient mobilisation from organic matter (Lindahl et al. 2021). However, the effects of Cortinarius spp. have been described for a boreal forest system, and the impacts of pine invasion on saprotroph diversity represent a major gap in our knowledge of the effects of pine invasion on fungal functional diversity.
In order to understand the impacts of pine invasions on fungal communities, we determined how invasion alters soil fungal community composition, specifically considering ectomycorrhizal fungi and saprotrophic fungal diversity at both point and plot scales. To accomplish this, we established 20 × 20 m plots across a pine invasion gradient and collected 24 soil samples within each plot. We sequenced DNA for fungi from each sample independently which allowed us to measure the spatial effects of pine invasion on fungal diversity (e.g. within-sample, plot-scale, and β-diversity).We hypothesized that an increased density of pine invasion would: (1) result in an initial increase (at low densities of pine) in β-diversity followed by a decrease (i.e. unimodal response as density of pine increased), (2) increase diversity of ectomycorrhizal fungi at the plot level through an accumulation of later successional species and (3) decrease diversity of saprotrophic fungi at the plot level.
Materials and methods
Site and sampling method
We studied a non-native Pinus nigra invasion in a previously grazed grassland on a river terrace at Mount Barker, Canterbury, New Zealand (43°21′38″S, 171°37′10″E; 615 m elevation; Supporting Information Fig. S1), which is the same site previously measured by Dickie et al. (2011). The stands across this invasion were all approximately the same age, with varying densities resulting from the natural episodic pattern of seed dispersal by prevailing winds and turbulence patterns from a single source (Caplat et al. 2010). The largest trees are approximately 30 years old and 18 m tall. Increased pine density at this site has been previously shown to cause no change in fungal biomass as measured by phospholipid fatty acids and hyphal lengths in soil (Dickie et al. 2011). However, increased pine density does cause a marginally significant decline in the ratio of fungi to bacteria, a significant decline in arbuscular mycorrhizal fatty acid markers, and a substantial increase in bacterial-feeding relative to fungal-feeding nematodes (Dickie et al. 2011). Vegetation cover is described in Dickie et al. (2011). We sampled thirteen 20 × 20 m plots across a gradient of P. nigra total stand basal area (m2 ha−1) (hereafter “density”; Fig. S1) that had previously been established (see Dickie et al. (2011) for methods of plot establishment). Basal area was calculated from stem diameter within each plot.
Within each plot we collected 24 soil samples of c. 65 mm diameter of the A-horizon to c. 100 mm depth (Fig. 1), using 24 hand trowels that had been sterilised for 10 min in a 10% bleach solution and rinsed. The location of each sample was pre-determined on a regular grid (Fig. 1), with no deviation required from these pre-determined locations. Soils from a single point was placed into individual sterile bags and stored at 4 °C for up to 6 h before being frozen at − 20 °C for 2 to 3 days before DNA extraction.
Soil sampling design of each of thirteen 20 m × 20 m plots at Mount Barker in Canterbury, South Island. Twenty-four soil samples were collected (1 m to the left of each green circle) from each plot and DNA was independently extracted, sequenced and analysed for each sample. In our experimental set-up, plot-scale diversity was used as a proxy for γ-diversity as it was our largest study area (i.e. sum of all OTUs in the plot indicated by large purple circle), within-sample diversity was used as a proxy for α-diversity as it was our smallest study area (i.e. sum of OTUs as indicated by small purple circles around green circles) and β-diversity was the variation in diversity among the smallest study areas (i.e. γ/ α)
DNA extraction
DNA was extracted from all soil samples using a DNeasy PowerSoil Pro Kit (Qiagen, Hilden, Germany) as per the manufacturer's protocol. Briefly, soil samples were mixed in their bags and 0.25 g was removed for DNA extractions. The fungal amplicon library was prepared in a one-step PCR using fungal specific primers fITS7 (5’-GTG ART CAT CGA ATC TTT G -3’) and ITS4 (5’-TCC TCC GCT TAT TGA TAT GC-3’) which amplify the ITS2 region (universal genetic barcode for fungi; Ihrmark et al. 2012). PCRs were based on 20 µl reactions using 1 µl of PCR product, 0.4 µl of the fITS7 primer and 1.6 µl of the ITS4 primer. The parameters for this protocol are as follows: 5 min at 94 °C; 35 cycles of 30 s at 94 °C, 30 s at 57 °C and 30 s at 72 °C; 7 min at 72 °C before cooling to 4 °C. The ITS4 reverse primer included both Illumina adapted and index sequences (for identification of sequenced amplicons) and the fITS7 primer included Illumina adaptor sequences. PCR products were visualized on 1% agarose gels and concentrations normalized using SequelPrep (Thermofisher). Uniquely indexed libraries (n = 312) along with positive and negative controls were pooled for the sequencing run, which was performed on an Illumina MiSeq using 2 × 250 paired end V2 chemistry following the manufacturer’s recommendations.
Bioinformatic analysis
Paired-end reads (with adaptor sequence removed) were merged using USEARCH v11 (Edgar, 2010) with a minimum overlap length of 50 bp with no gaps allowed in the merged alignments. Sequence deconvolution such as quality control and clustering was carried out using VSEARCH (https://github.com/torognes/vsearch.wiki.git). Specifically, sequences less than 200 bp and low mean quality (< 20) were removed. Sequences that passed quality control were clustered into operational taxonomic units (OTUs). OTUs were identified by searching against the public database provided by UNITE (Nilsson et al. 2018; UNITE Community 2017).
Ecological information of each OTU was based primarily on FUNGuild (https://github.com/UMNFuN/FUNGuild) following the user’s manual where assignments were made on ranks of ‘probable’ and ‘highly probable’ (Nguyen et al. 2016). Where there was no match or an OTU was ranked as ‘possible’, literature searches were conducted and OTUs assigned to a guild based on published literature. As it is difficult to assign function to fungi found in soil, assignments were conservative. Thus, if more than one guild was applicable to an OTU or if no information was found, we classified these unknown OTUs in an "undefined fungus" guild.
Statistical analysis
Fungal diversity
All analyses were performed using R version 3.5.0 (R Core Team 2020). To determine changes in within-sample (α), beta (β) and plot-scale (γ) diversity, we used the following equation:
In our experimental set-up (Fig. 1), plot-scale diversity was used as a proxy for γ-diversity as it was our largest study area, within-sample diversity was used as a proxy for α-diversity as it was our smallest study area and β-diversity was the variation in diversity among the smallest study areas (i.e. soil samples) (Crist et al. 2003). To determine soil fungal α-diversity of each soil sample, the OTUs were rarefied based on the minimum number of sequences observed per sample (1128) using the rrarefy function in the vegan package (Oksanen et al. 2015; Fig. S2; Fig. S3). To model α-diversity as a function of pine density of each plot, a linear mixed model was used with function lmer in the package lme4. To incorporate the dependency among plots, plot was used as a random intercept. γ-diversity was calculated taking the sum of OTUs present in each plot and modelled as a function of pine density of each plot using a linear model using function lm. β-diversity was calculated rearranging Eq. (1) to solve for β and modelled as a function of pine density of each plot using a linear model using function lm. Model assumptions were verified by visually inspecting residuals for assumptions of normality and homoscedasticity (Zuur and Ieno 2016). The above was completed for the whole fungal community and the two most dominant functional guilds: ectomycorrhizal fungi and saprotrophs. The proportion of sequences that were saprotrophs might be expected to decline due to dilution by ectomycorrhizal sequences, resulting in a decrease in sequencing depth for this group. In order to ensure that any observed changes in saprotroph diversity were not biased by a loss of sequence numbers, we also rarefied saprotrophs (minimum number of reads: 435) separately.
Community composition
To determine differences in the community composition of soil fungi across the pine density gradient, we used the Bray–Curtis dissimilarity index to calculate dissimilarly values from fungal OTU relative abundance data. Bray–Curtis dissimilarity values were calculated with the vegdist function in package vegan (Oksanen et al. 2015). Communities were visualised using non-metric multidimensional scaling (NMDS) using function metaMDS in package vegan. To test for community differences across the pine density gradient, permutational multivariate analysis of variance (PERMANOVA) with 9999 permutations using the function adonis was used in package vegan (Oksanen et al. 2015). The null hypothesis of a PERMANOVA is the centroids of the groups are equivalent for all groups. The above was repeated for communities of ectomycorrhizal fungi only and saprotroph fungi only. To determine the differences in the community composition of ectomycorrhizal fungi only, a threshold was placed on the OTU relative abundance data due to grassland plots (i.e. no pines present) having no ectomycorrhizal fungi. This threshold removed samples with ≤ 2 ectomycorrhizal OTUs. To determine the difference in dispersion (i.e. distance to centroid) among low density plots and high density plots we used function betadisper in package vegan. The function TukeyHSD in package stats was used to measure paired differences among plots.
Our most abundant OTU belonged to a Sistotrema sp. and has not previously been described in pine invasions. In addition, the genus Sistotrema is polyphyletic and ecologically complicated, including endophytes (Potvin et al. 2012), saprotrophs and ectomycorrhizal fungi (Policelli et al. 2020). In order to resolve the identity of Sistotrema in New Zealand pine invasions, and particularly whether an ectomycorrhizal status was supported, we conducted a phylogenetic analysis of pine-associated Cantherellales in New Zealand, along with representative sequences of native and non-native Sistotrema, Clavulina, Membranomyces, and Hydnum. Sequences were downloaded from GenBank, aligned using MAFFT version 7, and phylogenetic analysis carried out using MrBayes on XSEDE through the CIPRES Science Gateway version 3.3, with nucleotide substitution models determined using JModeltest 2.1.10.
Functional guilds
To determine changes in the proportion of reads (i.e. dominance) of functional guilds as a function of pine density of each plot we used a binomial generalized linear mixed model function glmer in the package lme4. The number of sequences belonging to each functional guild as a proportion of the total number of sequences in each sample was the response variable. The fixed covariate included P. nigra density and to incorporate the dependency among plots, plot was used as a random intercept. Model assumptions were verified by visually inspecting residuals for assumptions of normality and homoscedasticity (Zuur and Ieno 2016).
Results
After quality control, we had a library of soil fungi consisting of 2436 fungal OTUs (Supporting Information Table S1). Of the 2436 OTUs, 41% were assigned as saprotrophs, 5.6% as ectomycorrhizal fungi, 5.3% as plant pathogens, 4.3% were assigned to other mycorrhizal guilds (i.e. arbuscular mycorrhizal fungi, ericoid mycorrhizal fungi and orchid mycorrhizal fungi) and 3.5% as endophytes. Due to dual functions and many OTUs not assigned to genus, 34.9% of OTUs were undefined and were not assigned to a functional guild (70.8% of these OTUs were not assigned to genus and the remaining 29.2% had dual functions). The remaining 5.4% were assigned as lichenized fungi, lichen parasites, fungal parasites, biotrophs and animal pathogens.
Fungal diversity
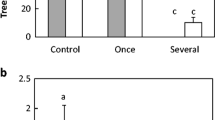
Beta-diversity had a unimodal response to P. nigra density (estimate = − 1.173, t = − 2.709, p = 0.022; Fig. 2a).Within-sample (α) diversity decreased by 47.7% as P. nigra density increased (estimate = − 0.459, t value = − 7.993, p < 0.001; Fig. 2b). Plot-level (γ) diversity decreased by 50.1% as P. nigra density increased (estimate = − 2.592, t value − 12.60, p < 0.001; Fig. 2c).
Diversity of soil fungal communities: a Beta (β) diversity (variation of diversity among soil samples within each plot) as a function of pine density (represented as basal area of Pinus nigra (m2/ha)). b Alpha (α) diversity (diversity of each soil sample within each plot) as a function of pine density. Open circles and colours demonstrate α-diversity of 24 samples within each plot and filled diamonds represent the mean α-diversity of each plot. c Gamma (γ) diversity (diversity of each plot) as a function of pine density
β-diversity of ectomycorrhizal fungi decreased (estimate = − 0.99, t = − 7.594, p < 0.0001) in a loglinear relationship to P. nigra density (Fig. 3a). Within-sample (α) diversity increased logarithmically (estimate = 0.796, t-value = 6.894, p < 0.001) as a function of P. nigra density (Fig. 3b) whereas plot-scale (γ) diversity did not change (p = 0.324; Fig. 3c). P. nigra density had an opposite effect on the diversity of saprotrophs which mimicked changes in overall fungal diversity patterns. β-diversity of saprotrophs had a unimodal response (estimate = − 0.987, t = − 2.542, p = 0.029) to P. nigra density (Fig. 3d). Results for saprotrophs were qualitatively similar if based on rarefaction of just saprotroph sequences, rather than the whole dataset, except that the unimodal response of β-diversity was no longer significant (p = 0.083; not shown). Both α- (estimate = − 0.139, t = − 5.942, p < 0.0001) and γ-diversity (estimate = − 0.963, t = − 7.936, p < 0.0001) of saprotrophs decreased as a function of P. nigra density (Fig. 3e, f).
Diversity of the two most dominant functional guilds. Ectomycorrhizal fungi: a Beta (β) diversity (variation of diversity among soil samples within each plot) as a function of basal area of Pinus nigra (m2/ha) (proxy for P. nigra density). b Alpha (α) diversity (diversity of each soil sample within each plot) as a function of P. nigra density. Open circles and colours demonstrate α-diversity of 24 samples within each plot and filled diamonds represent the mean alpha diversity of each plot. c Gamma (γ) diversity (total diversity of each plot) as a function of P. nigra density. Saprotrophic fungi: d β-diversity, e α-diversity, f γ-diversity
Community composition
The community composition of soil fungi differed significantly (F1,12 = 8.72, p = 0.0001) as a function of P. nigra density. This trend was similar for ectomycorrhizal fungi only (F1,12 = 4.06, p = 0.014; Fig. 4a) and saprotrophic fungi only (F1,12 = 8.94, p = 0.0001; Fig. 4b). Communities found at low density of pine did not overlap with communities found at high densities of P. nigra (Fig. 4a, b). Plots having intermediate pine density were intermediate between samples from high density and no pine plots, but some individual soil samples overlapped each of the two end points. However, this was only observed in the saprotroph community. Plots of intermediate pine density in the ectomycorrhizal fungal community overlapped substantially with plots of high pine density. The amount of dispersion (i.e. distance to centroid) differed significantly (F12,297 = 13.32, p < 0.0001) as a function of P. nigra density. Specifically, plots with no pine and low pine density had greater dispersion in comparison to plots with high density of pine (p < 0.0001).
Non-metric multidimensional scaling (NMDS) plot demonstrating the community composition of soil fungi across a pine density gradient where a is ectomycorrhizal fungi only and b is saprotrophic fungi only. Individual points represent fungal communities of individual samples within a plot. Plots are represented by colour-coded ellipses where light blue on the right of each figure are plots of no to low pine density and dark blue are plots of high pine density
Our most abundant OTU was a Sistotrema sp. and a strong match to the most abundant OTU identified on invasive P. contorta by Dickie et al. (2010), previously identified only as “Cantharellales” (GQ906424, GQ906425), to another “Cantharellales” sequence from P. contorta collected in Oregon (AY880947; Ashkannejhad & Horton, 2006), and to an ectomycorrhizal fungus collected from Arizona on Pinus ponderosa (MG761648; Bowman & Arnold, 2018). These sequences all match a recent collection of a Sistotrema sporocarp collected from pine needles in California, USA (GenBank KP81424, submitted in 2015 by L. Rosenthall and T. Bruns). The results support the polyphyletic nature of Sistotrema (Fig. S4), which includes endophytic and saprotrophic clades, but Sistotrema OTUs from this study all clustered with ectomycorrhizal isolates (Table S1; Fig. S4).
Functional guilds
Fungal functional guilds also shifted strongly as P. nigra density increased (Fig. 5a). Here we present results on functional guilds that represent greater than 1% of the total reads. The proportion of reads belonging to the ectomycorrhizal guild increased logarithmically (estimate = 0.887, z-value = 10.34, p < 0.0001) as a function of P. nigra density. The proportion of saprotrophs (p = 0.347) and ericoid mycorrhizal fungi (p = 0.995) did not change with P. nigra density whereas the proportion of endophytes logarithmically decreased (estimate = − 0.379, z-value = − 6.609, p < 0.0001) as a function of P. nigra density. Proportion of plant pathogens (estimate = − 0.008, z-value = − 2.889, p = 0.004) decreased linearly as a function of density of P. nigra.
Functional guild composition (proportion of all reads (out of 1) of guilds that represented greater than 1% of total reads) of soil fungi as a function of Pinus nigra density (represented as basal area (m2/ha)). a Lines indicate mean fitted values extracted from the best fit binomial model and ribbons represent standard errors about the means. b Proportion of reads belonging to ectomycorrhizal fungi represented by genera that contain at least 1% of ectomycorrhizal fungi reads as a function of Pinus nigra density. c Proportion of reads belonging to saprotrophic fungi represented by genera that contain at least 1% of saprotroph fungi reads as a function of Pinus nigra density. * indicates those species that significantly changed in abundance as a function of P. nigra density. Number in [] demonstrate number of OTUs in each genus
Specific genera of ectomycorrhizal fungi showed significant, but contrasting, changes in abundance as P. nigra density increased (Fig. 5b). Here we present results from the genera that represent greater than 1% of the ectomycorrhizal reads. The proportion of reads of Sistotrema spp. logarithmically increased (estimate = 0.128, t-value = 2.639, p = 0.023) as a function of P. nigra density. Amanita spp. (estimate = 0.012, t = value = 5.588, p = 0.0002), Tricholoma spp. (estimate = 0.01, t-value = 2.92, p = 0.014), a Tylospora sp. (estimate = 0.005, t-value = 3.862, p = 0.003) and Wilcoxina spp. (estimate = 0.003, t-value = 2.828, p = 0.016) significantly increased linearly as P. nigra density increased. The proportion of reads of Suillus spp. did not significantly change (p = 0.616) across the pine density gradient.
Specific genera of saprotroph fungi also showed significant, but contrasting, changes in abundance as P. nigra density increased (Fig. 5c). Here we present results from the genera that represent greater than 1% of the saprotroph reads. The proportion of reads of four genera decreased as a function of pine density. Specifically, the proportion of reads of Cladophialophora (estimate = − 0.000078, t-value = − 3.173, p = 0.002), Clavaria (estimate = − 0.000027, t-value = − 3.173, p = 0.009), Peziza (estimate = − 0.000034, t-value = − 2.255, p = 0.045) and Pleotrichocladium spp. (estimate = − 0.000175, t-value = − 3.418, p = 0.006) significantly decreased as a function of P. nigra density. Sagenomella (estimate = 0.00054, t = value = 4.786, p = 0.0006) and Saitozyma spp. (estimate = 0.00054, t = value = 6.497, p = 0.00004) significantly increased linearly as P. nigra density increased.
Discussion
Fungal diversity
Co-invasion of Pinus nigra and associated ectomycorrhizal fungi caused a positive unimodal response in fungal species turnover or β-diversity. In addition, we observed a significant loss of both soil fungal α- and γ-diversity, and decreased community dispersion. Approximately half of the fungal diversity was lost at both within-sample and plot-scale as pines invaded grasslands. This is contradictory to other invasions; for example, the invasion of Alliaria petiolate (garlic mustard) in the northeastern USA resulted in an increase in α-diversity in invaded sites compared to uninvaded sites (Anthony et al. 2017; Gomes et al. 2018). Invaded sites also had lower β-diversity in comparison to uninvaded sites (Anthony et al. 2017). The loss in diversity at both within-sample and plot-scale, in our study, could be due to pines having highly specific interactions with their co-introduced ectomycorrhizal fungi. Interestingly, the unimodal response of β-diversity of the fungal community may mirror changes in aboveground plant diversity (Davis et al. 2019; Sapsford et al. 2020). Plots with intermediate densities of pine would have patches of pines and patches of native vegetation and thus greater variation in diversity among the individual samples (Powell et al. 2013).
Despite contradictory effects of invasion on fungal diversity, plant invasions may lead to biotic homogenisation of soil fungal communities (Anthony et al. 2017; García et al. 2018). It has been demonstrated that homogenisation belowground occurs after certain types of disturbances such as land-use intensification (Dopheide et al. 2020; Gossner et al. 2016); however, biotic homogenisation is fairly unexplored in plant invasions. We observed less variation within the fungal community in invaded plots of high pine density suggesting community convergence as environmental conditions become more homogeneous (Anthony et al. 2017; Caruso et al. 2012). It is also possible, that just as pines homogenize the aboveground community through competition for light, their associated ectomycorrhizal fungi may homogenize the belowground community through competition with and inhibition of other fungi.
Changes in diversity across the pine invasion gradient differed across functional guilds. Although we observed a small increase in ectomycorrhizal fungal α-diversity with invasion, this effect was counterbalanced by a much larger loss of saprotrophic fungal α-diversity resulting in an overall loss of soil fungal α-diversity as pine density increased. It has not previously been demonstrated that pine invasions, which are increasingly widespread in the southern hemisphere, cause major losses of saprotrophic fungal diversity. Pine invasion at this site has been shown to cause no change in total fungal biomass (Dickie et al. 2011), hence the loss of saprotrophic diversity is not likely to represent a dilution effect in sequencing depth, which we also confirmed by testing diversity responses after rarefaction to a standard sequencing depth within saprotrophs. Rather, the results support the concept of saprotrophic exclusion as ectomycorrhizal proportional abundance increases. Competition between ectomycorrhizal and saprotrophic fungi is well established at a local scale (Bödeker et al. 2016). In the context of invasions, it has also been shown that suppression of ectomycorrhizal fungi by the non-mycorrhizal plant Alliaria petiolata can increase saprotrophic diversity (Anthony et al. 2017), supporting our suggestion that the loss of saprotroph diversity is due to ectomycorrhizal suppression rather than a more generic effect of plant invasion. Ectomycorrhizal fungi converged in composition, represented by a decrease in β-diversity of ectomycorrhizal fungi. Interestingly, this combined with the increase in α-diversity resulted in no net change in γ-diversity of ectomycorrhizal fungi. The decrease in β-diversity could be due to the dominance of key ectomycorrhizal fungi in high density plots, particularly Amanita spp., Sistotrema spp. and Tricholoma spp.
Community composition and functional guilds
Our results show that the co-invasion of Pinus nigra and associated ectomycorrhizal fungi causes fundamental shifts in ectomycorrhizal fungal communities, with a gradual accumulation from primarily Suillus at the lowest densities of pine to a diverse but entirely non-native community of ectomycorrhizal fungi at high pine densities. This integrates previously conflicting studies, supporting Hayward et al.’s suggestion of invasion driven by a single species (Hayward et al. 2015), but also the development of more complex communities over time (Dickie et al. 2010).
Some shifts in fungal communities occurred at very low densities of invasion into grassland (i.e. pine basal area of 6 m2/ha); for example, ectomycorrhizal fungi dominated early invaded communities with over 50% of the total sequence reads. We observed Suillus spp., Sistotrema spp. and a Tylospora sp.in grassland areas where P. nigra had yet to invade, suggesting ectomycorrhizal fungal inoculum was not a limiting factor in the success of P. nigra establishment into this grassland ecosystem (Pringle et al. 2009). Suillus is an important ectomycorrhizal species for the early establishment of pines in both plantations and its natural range (Chapela et al. 2001; Clemmensen et al. 2015) and is also considered the only species necessary to enable pine invasion (Hayward et al. 2015; Policelli et al. 2019).
Sistotrema was the most abundant OTU in our data, and dominant at low densities of invasion.The results support an ectomycorrhizal and non-native status, with a clustering of the New Zealand Pinus associated sequences, along with Pinus associated ectomycorrhizal sequences from Oregon and Arizona, USA and the sporocarp collection from California, and a possible relationship to Sistotrema alboleteum.
With increasing density of invasion, we also observed the introduction and increase in dominance of Amanita spp., Tricholoma spp. and Wilcoxina spp. The change in fungal communities with tree density is comparable to the ectomycorrhizal fungi associated with pines in their native ranges during later stages of pine succession (Vellinga et al. 2009; Vlk et al. 2020b). However, we observed only a subset of ectomycorrhizal fungal species that typically associate with pines in their native range (Hynson et al. 2013), reflecting species of ectomycorrhizal fungi associated with pine that may be more successful co-invaders in novel environments (Vlk et al. 2020b).
Interestingly, the proportion of DNA reads that were saprotrophs did not change across the invasion gradient, despite the large loss of diversity. However, two OTUs, matching Sagenomella (a filamentous ascomycete sometimes associated with pine soils (Fritze and Bååth 1993)) and Saitozyma sp. (a dominant basidiomycete soil yeast (Mašínová et al. 2017)), were found in higher proportions at high pine densities than in grassland plots, while a greater number of other OTUs declined. To our knowledge, our study is the first to identify saprotroph species associated with pine invasions into grasslands. Saprotrophs and ectomycorrhizal fungi interact to decompose organic matter (Fernandez and Kennedy 2016). With the observed decline in diversity of saprotrophs across the invasion gradient, it is possible the introduced ectomycorrhizal fungi are competitively excluding the majority of saprotrophic fungi (Anthony et al. 2017; Bödeker et al. 2016; Fernandez and Kennedy 2016). In the northern hemisphere where pines are native, saprotrophs and ectomycorrhizal fungi can occupy different vertical positions of the soil profile (Bödeker et al. 2016; Fernandez and Kennedy 2016). However, neither the uninvaded grassland nor pine invaded sites had much soil profile development, with a thin grass or pine needle litter layer directly over a well mixed A-horizon, which may have limited vertical niche partitioning as a mechanism for coexistence.
Conclusion
Co-invasion of pine and their ectomycorrhizal fungi caused an unexpected and massive loss in soil fungal diversity at point and plot scales, with an initial increase in heterogeneity followed by biotic homogenisation. The loss in diversity also corresponded to strong shifts in belowground fungal community composition and functional guilds, even at low densities of invasion. Such drastic changes will have ramifications for soil function and future plant establishment and could potentially inhibit recovery and restoration of co-invaded ecosystems. Legacies of pines (i.e. effects of pines that remain after their removal) are not easily reversed and thus, neither are the impacts of co-invaded ectomycorrhizal fungi (Sapsford et al. 2020). A common occurrence after removal of pines is the re-invasion of pines (as ectomycorrhizal fungal inoculums remains in the soil) in addition to invasion by other non-native species, specifically grasses (Dickie et al. 2014; Paul and Ledgard 2009), highlighting difficulties for native restoration. Native trees in this system are slower to establish than pines and other exotic plants, and the role of soil biota is not fully understood (Dickie et al. 2012; Wood et al. 2015). Invasive ectomycorrhizal fungi have been found in native beech forests in New Zealand (Orlovich and Cairney 2004). Therefore, the change in belowground communities we observed as a result of pine invasions could pose a problem for native restoration now and in the future. Furthermore, our results demonstrate the importance of considering both above- and belowground impacts of invasions.
Availability of data and material
The data that supports the findings of this study are available in the supporting information of this article.
References
Anderson MJ, Ellingsen KE, McArdle BH (2006) Multivariate dispersion as a measure of beta diversity. Ecol Lett 9(6):683–693. https://doi.org/10.1111/j.1461-0248.2006.00926.x
Anthony MA, Frey SD, Stinson KA (2017) Fungal community homogenization, shift in dominant trophic guild, and appearance of novel taxa with biotic invasion. Ecosphere 8(9):e01951. https://doi.org/10.1002/ecs2.1951
Ashkannejhad S, Horton TR (2006) Ectomycorrhizal ecology under primary succession on coastal sand dunes: interactions involving Pinus contorta, suilloid fungi and deer. New Phytol 169(2):345–354. https://doi.org/10.1111/j.1469-8137.2005.01593.x
Bödeker ITM, Lindahl BD, Olson Å, Clemmensen KE (2016) Mycorrhizal and saprotrophic fungal guilds compete for the same organic substrates but affect decomposition differently. Funct Ecol 30:1967–1978. https://doi.org/10.1111/1365-2435.12677
Bowman EA, Arnold AE (2018) Distributions of ectomycorrhizal and foliar endophytic fungal communities associated with Pinus ponderosa along a spatially constrained elevation gradient. Am J Bot 105(4):687–699. https://doi.org/10.1002/ajb2.1072
Caplat P, Coutts SR, Buckley Y (2010) Gone with the wind: high-resolution analysis of pine dispersal in New Zealand Mountains. In: Proceedings of the 17th Australasian weeds conference, pp 190–193
Caruso T, Hempel S, Powell JR, Barto EK, Rillig MC (2012) Compositional divergence and convergence in arbuscular mycorrhizal fungal communities. Ecology 93(5):1115–1124. https://doi.org/10.1890/11-1030.1
Chapela IH, Osher LJ, Horton TR, Henn MR (2001) Ectomycorrhizal fungi introduced with exotic pine plantations induce soil carbon depletion. Soil Biol Biochem 33(12):1733–1740. https://doi.org/10.1016/S0038-0717(01)00098-0
Chen CR, Condron LM, Xu ZH (2008) Impacts of grassland afforestation with coniferous trees on soil phosphorus dynamics and associated microbial processes: a review. For Ecol Manage 255(3):396–409. https://doi.org/10.1016/j.foreco.2007.10.040
Clemmensen KE, Finlay RD, Dahlberg A, Stenlid J, Wardle DA, Lindahl BD (2015) Carbon sequestration is related to mycorrhizal fungal community shifts during long-term succession in boreal forests. New Phytol 205(4):1525–1536. https://doi.org/10.1111/nph.13208
Crist TO, Veech JA, Gering JC, Summerville KS (2003) Partitioning species diversity across landscapes and regions: a hierarchical analysis of alpha, beta, and gamma diversity. Am Nat 162(6):734–743. https://doi.org/10.1086/378901
Davis KT, Callaway RM, Fajardo A, Pauchard A, Nuñez MA, Brooker RW, Maxwell BD, Dimarco RD, Peltzer DA, Mason B, Ruotsalainen S, McIntosh ACS, Pakeman RJ, Smith AL, Gundale MJ (2019) Severity of impacts of an introduced species corresponds with regional eco-evolutionary experience. Ecography 42(1):12–22. https://doi.org/10.1111/ecog.04014
Dickie IA, Bolstridge N, Cooper JA, Peltzer DA (2010) Co-invasion by Pinus and its mycorrhizal fungi. New Phytol 187(2):475–484. https://doi.org/10.1111/j.1469-8137.2010.03277.x
Dickie IA, Yeates GW, St. John MG, Stevenson BA, Scott JT, Rillig MC, Peltzer DA, Orwin KH, Kirschbaum MUF, Hunt JE, Burrows LE, Barbour MM, Aislabie J (2011) Ecosystem service and biodiversity trade-offs in two woody successions. J Appl Ecol 48(4):926–934. https://doi.org/10.1111/j.1365-2664.2011.01980.x
Dickie IA, Davis M, Carswell FE (2012) Quantification of mycorrhizal limitation in beech spread. N Z J Ecol 36:210–215
Dickie IA, St. John MG, Yeates GW, Morse CW, Bonner KI, Orwin K, Peltzer DA (2014) Belowground legacies of Pinus contorta invasion and removal result in multiple mechanisms of invasional meltdown. AoB Plants 6:plu056. https://doi.org/10.1093/aobpla/plu056
Dopheide A, Makiola A, Orwin KH, Holdaway RJ, Wood JR, Dickie IA (2020) Rarity is a more reliable indicator of land-use impacts on soil invertebrate communities than other diversity metrics. Elife 9:e52787. https://doi.org/10.7554/eLife.52787
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19):2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Fernandez CW, Kennedy PG (2016) Revisiting the ‘Gadgil effect’: do interguild fungal interactions control carbon cycling in forest soils? New Phytol 209:1382–1394. https://doi.org/10.1111/nph.13648
Fritze H, Bååth E (1993) Microfungal species composition and fungal biomass in a coniferous forest soil polluted by alkaline deposition. Microb Ecol 25(1):83–92. https://doi.org/10.1007/BF00182131
García RA, Franzese J, Policelli N, Sasal Y, Zenni RD, Nuñez MA, Taylor K, Pauchard A (2018) Non-native pines are homogenizing the ecosystems of South America. In: Rozzi R, May RH Jr, Chapin FS III, Massardo F, Gavin MC, Klaver IJ, Pauchard A, Nuñez MA, Simberloff D (eds) From biocultural homogenization to biocultural conservation. Springer International Publishing, Cham, pp 245–263. https://doi.org/10.1007/978-3-319-99513-7_15
Gomes SIF, Merckx VSFT, Hynson NA (2018) Biological invasions increase the richness of arbuscular mycorrhizal fungi from a Hawaiian subtropical ecosystem. Biol Invasions 20(9):2421–2437. https://doi.org/10.1007/s10530-018-1710-7
Gossner MM, Lewinsohn TM, Kahl T, Grassein F, Boch S, Prati D, Birkhofer K, Renner SC, Sikorski J, Wubet T, Arndt H, Baumgartner V, Blaser S, Blüthgen N, Börschig C, Buscot F, Diekötter T, Jorge LR, Jung K et al (2016) Land-use intensification causes multitrophic homogenization of grassland communities. Nature 540(7632):266–269. https://doi.org/10.1038/nature20575
Hayward J, Horton TR, Pauchard A, Nuñez MA (2015) A single ectomycorrhizal fungal species can enable a Pinus invasion. Ecology 96(5):1438–1444. https://doi.org/10.1890/14-1100.1
Hynson NA, Merckx VSFT, Perry BA, Treseder KK (2013) Identities and distributions of the co-invading ectomycorrhizal fungal symbionts of exotic pines in the Hawaiian Islands. Biol Invasions 15(11):2373–2385. https://doi.org/10.1007/s10530-013-0458-3
Ihrmark K, Bödeker ITM, Cruz-Martinez K, Friberg H, Kubartova A, Schenck J, Strid Y, Stenlid J, Brandström-Durling M, Clemmensen KE, Lindahl BD (2012) New primers to amplify the fungal ITS2 region—evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol Ecol 82(3):666–677. https://doi.org/10.1111/j.1574-6941.2012.01437.x
Kyaschenko J, Clemmensen KE, Hagenbo A, Karltun E, Lindahl BD (2017) Shift in fungal communities and associated enzyme activities along an age gradient of managed Pinus sylvestris stands. ISME J 11(4):863–874. https://doi.org/10.1038/ismej.2016.184
Lindahl BD, Tunlid A (2015) Ectomycorrhizal fungi – potential organic matter decomposers, yet not saprotrophs. New Phytol 205(4):1443–1447. https://doi.org/10.1111/nph.13201
Lindahl BD, Kyaschenko J, Varenius K, Clemmensen KE, Dahlberg A, Karltun E, Stendahl J (2021) A group of ectomycorrhizal fungi restricts organic matter accumulation in boreal forest. Ecol Lett 24(7):1341–1351. https://doi.org/10.1111/ele.13746
Mašínová T, Bahnmann BD, Větrovský T, Tomšovský M, Merunková K, Baldrian P (2017) Drivers of yeast community composition in the litter and soil of a temperate forest. FEMS Microbiol Ecol 93:223. https://doi.org/10.1093/femsec/fiw223
Nguyen NH, Song Z, Bates ST, Branco S, Tedersoo L, Menke J, Schilling JS, Kennedy PG (2016) FUNGuild: an open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol 20:241–248. https://doi.org/10.1016/j.funeco.2015.06.006
Nilsson RH, Larsson K-H, Taylor AFS, Bengtsson-Palme J, Jeppesen TS, Schigel D, Kennedy P, Picard K, Glöckner FO, Tedersoo L, Saar I, Kõljalg U, Abarenkov K (2018) The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res 47(D1):D259–D264. https://doi.org/10.1093/nar/gky1022
Nuñez MA, Dickie IA (2014) Invasive belowground mutualists of woody plants. Biol Invasions 16(3):645–661. https://doi.org/10.1007/s10530-013-0612-y
Nuñez MA, Chiuffo MC, Torres A, Paul T, Dimarco RD, Raal P, Policelli N, Moyano J, García RA, van Wilgen BW, Pauchard A, Richardson DM (2017) Ecology and management of invasive Pinaceae around the world: progress and challenges. Biol Invasions 19(11):3099–3120. https://doi.org/10.1007/s10530-017-1483-4
Oksanen J, Blanchet FG, Kindt FG, Legendre P, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Wagner H (2015) Vegan: community ecology package
Orlovich DA, Cairney JG (2004) Ectomycorrhizal fungi in New Zealand: current perspectives and future directions. N Z J Bot 42(5):721–738. https://doi.org/10.1080/0028825X.2004.9512926
Orwin KH, Kirschbaum MUF, John MGS, Dickie IA (2011) Organic nutrient uptake by mycorrhizal fungi enhances ecosystem carbon storage: a model-based assessment. Ecol Lett 14(5):493–502. https://doi.org/10.1111/j.1461-0248.2011.01611.x
Paul TSH, Ledgard NJ (2009) Vegetation succession associated with wilding conifer removal. N Z Plant Prot 62:374–379. https://doi.org/10.30843/nzpp.2009.62.4878
Policelli N, Bruns TD, Vilgalys R, Nuñez MA (2019) Suilloid fungi as global drivers of pine invasions. New Phytol 222(2):714–725
Policelli N, Horton TR, García RA, Naour M, Pauchard A, Nuñez MA (2020) Native and non-native trees can find compatible mycorrhizal partners in each other’s dominated areas. Plant Soil. https://doi.org/10.1007/s11104-020-04609-x
Potvin LR, Richter DL, Jurgensen MF, Dumroese RK (2012) Association of Pinus banksiana Lamb. and Populus tremuloides Michx. Seedling fine roots with Sistotrema brinkmannii (Bres.) J. Erikss. (Basidiomycotina). Mycorrhiza 22(8):631–638. https://doi.org/10.1007/s00572-012-0440-4
Powell KI, Chase JM, Knight TM (2013) Invasive plants have scale-dependent effects on diversity by altering species-area relationships. Science (new York, n.y.) 339(6117):316–318. https://doi.org/10.1126/science.1226817
Pringle A, Bever JD, Gardes M, Parrent JL, Rillig MC, Klironomos JN (2009) Mycorrhizal symbioses and plant invasions. Annu Rev Ecol Evol Syst 40(1):699–715. https://doi.org/10.1146/annurev.ecolsys.39.110707.173454
R Core Team (2020) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Richardson DM, Rejmánek M (2011) Trees and shrubs as invasive alien species—a global review. Divers Distrib 17(5):788–809. https://doi.org/10.1111/j.1472-4642.2011.00782.x
Rundel PW, Dickie IA, Richardson DM (2014) Tree invasions into treeless areas: mechanisms and ecosystem processes. Biol Invasions 16(3):663–675. https://doi.org/10.1007/s10530-013-0614-9
Sapsford SJ, Brandt AJ, Davis KT, Peralta G, Dickie IA, Gibson RD II, Green JL, Hulme PE, Nuñez MA, Orwin KH, Pauchard A, Wardle DA, Peltzer DA (2020) Towards a framework for understanding the context dependence of impacts of non-native tree species. Funct Ecol 34(5):944–955. https://doi.org/10.1111/1365-2435.13544
Socolar JB, Gilroy JJ, Kunin WE, Edwards DP (2016) How should beta-diversity inform biodiversity conservation? Trends Ecol Evol 31(1):67–80. https://doi.org/10.1016/j.tree.2015.11.005
Sterkenburg E, Clemmensen KE, Ekblad A, Finlay RD, Lindahl BD (2018) Contrasting effects of ectomycorrhizal fungi on early and late stage decomposition in a boreal forest. ISME J 12(9):2187–2197. https://doi.org/10.1038/s41396-018-0181-2
UNITE Community (2017) Full UNITE+INSD dataset. Version 01.12.2017. UNITE Community. https://doi.org/10.15156/BIO/587474
Vellinga EC, Wolfe BE, Pringle A (2009) Global patterns of ectomycorrhizal introductions. New Phytol 181(4):960–973. https://doi.org/10.1111/j.1469-8137.2008.02728.x
Vila M, Espinar JL, Hejda M, Hulme PE, Jarosik V, Maron JL, Pergl J, Schaffner U, Sun Y, Pysek P (2011) Ecological impacts of invasive alien plants: a meta-analysis of their effects on species, communities and ecosystems. Ecol Lett 14(7):702–708. https://doi.org/10.1111/j.1461-0248.2011.01628.x
Vlk L, Tedersoo L, Antl T, Větrovský T, Abarenkov K, Pergl J, Albrechtová J, Vosátka M, Baldrian P, Pyšek P, Kohout P (2020a) Early successional ectomycorrhizal fungi are more likely to naturalize outside their native range than other ectomycorrhizal fungi. New Phytol 227(5):1289–1293. https://doi.org/10.1111/nph.16557
Vlk L, Tedersoo L, Antl T, Větrovský T, Abarenkov K, Pergl J, Albrechtová J, Vosátka M, Baldrian P, Pyšek P, Kohout P (2020b) Alien ectomycorrhizal plants differ in their ability to interact with co-introduced and native ectomycorrhizal fungi in novel sites. ISME J 14(9):2336–2346. https://doi.org/10.1038/s41396-020-0692-5
Wang K, Zhang Y, Tang Z, Shangguan Z, Chang F, Jia F, Chen Y, He X, Shi W, Deng L (2019) Effects of grassland afforestation on structure and function of soil bacterial and fungal communities. Sci Total Environ 676:396–406. https://doi.org/10.1016/j.scitotenv.2019.04.259
Wood JR, Dickie IA, Moeller HV, Peltzer DA, Bonner KI, Rattray G, Wilmshurst JM (2015) Novel interactions between non-native mammals and fungi facilitate establishment of invasive pines. J Ecol 103(1):121–129. https://doi.org/10.1111/1365-2745.12345
Zuur AF, Ieno EN (2016) A protocol for conducting and presenting results of regression-type analyses. Methods Ecol Evol 7(6):636–645. https://doi.org/10.1111/2041-210X.12577
Funding
We thank the New Zealand Ministry of Business, Innovation and Employment for financial support through the Winning Against Wildings research programme.
Author information
Authors and Affiliations
Contributions
IAD and DAP developed the project concept, IAD and AW collected soil samples, AW extracted DNA, SJS led analyses and wrote the first draft of the manuscript, and all authors contributed substantially to revisions.
Corresponding author
Ethics declarations
Conflict of interest
We declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sapsford, S.J., Wakelin, A., Peltzer, D.A. et al. Pine invasion drives loss of soil fungal diversity. Biol Invasions 24, 401–414 (2022). https://doi.org/10.1007/s10530-021-02649-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-021-02649-7