Abstract
Founder effects and genetic drift can reduce the genetic diversity and alter the genetic composition of introduced species during the processes of population establishment and spread. Thus, founder effects are of particular concern for introduced commercial populations (usually founded from few individuals) and for the natural populations they interact with. Bullfrogs were initially introduced in China for aquaculture purposes and escapes from farms have established many feral populations. Most of the bullfrog farms currently operative have been founded from a limited number of descendents from the original introductions, providing an excellent framework to elucidate the importance of founder effects and genetic diversity in the establishment and persistence of invasive species introduced for commercial purposes. We sequenced a region of the mitochondrial cytochrome b gene in 510 samples collected from feral and farm individuals across China and compared them to populations in their native range. Only two haplotypes (H43 and H7) were identified, and H43 identified in this study for the first time was present at high frequency in both feral and commercial populations. We show a significant difference in the relative frequency of the two identified haplotypes in commercial and feral populations, and suggest that sequential founding events are responsible for the emerging widely distributed new haplotype and the observed differences in genetic structure between bullfrog populations. Our findings indicate that lack of genetic diversity does not necessarily impair the colonizing ability of invasive species and highlight the potential threat posed by introduced commercial populations, given their unique genetic makeup.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biological invasions pose one of the most serious threats to biodiversity in both natural and agricultural ecosystems (Pimentel et al. 2005; Sakai et al. 2001; Simberloff and Stiling 1996; Strayer et al. 2006). The occurrence of non-native invasion has accelerated to an extraordinary degree due to the globalization of economy and caused serious economic loss (Floerl et al. 2009; Vitousek et al. 1996; Westphal et al. 2008). In the United States approximately $120 billion are lost each year as a consequence of the 50,000 nonnative species (Pimentel et al. 2005). In many cases, the invasive process involves population bottlenecks and founding events, especially in species introduced for biological control or agriculture purposes with a limited number of initial founders (Hufbauer et al. 2004; Simberloff 2009). Furthermore, subsequent dispersals (for example as seedings or stock individuals for mass-production) can result in additional founder events. This stochastic sampling effect can lead to distinct genetic compositions between introduced and source populations and among the introduced populations themselves (Delaney et al. 2009; Hinomoto et al. 2006). In contrast, in cases where the founders come from multiple introductions, the genetic diversity can be higher than in the original native populations and hybridization among individuals from different sources can help the invasive process by facilitating the adaptation to new conditions (Kolbe et al. 2008).
The American bullfrog, Lithobates catesbeianaus (also known as Rana catesbeiana), is listed among the “100 of the World’s Worst Invasive Alien Species” in the Global Invasive Species Database (ISSG 2008). Native to eastern North America, bullfrogs have been widely introduced throughout the world, and established populations have been reported in many regions (Lever 2003). Bullfrogs can affect native amphibian populations through predation and competition for food and habitat (Kats and Ferrer 2003; Kiesecker and Blaustein 1998; Kiesecker et al. 2001; Lawler et al. 1999; Pearl et al. 2004; Li et al. 2011a). Furthermore, recent studies have also shown that bullfrogs are asymptomatic carriers of the pathogenic fungus Batrachochytrium dendrobatidis (Bd) that is deadly to other frog species (Daszak et al. 2004; Garner et al. 2006), the continuous trading of bullfrogs contributing to the spread of Bd. (Fisher and Garner 2007; Garner et al. 2006). In fact, the chytrid has been detected both in bullfrogs sampled in the field and in captivity in China (Bai et al. 2010).
Bullfrogs were first introduced into mainland China in the late 1950s from Cuba and Japan as a source of food (Li and Xie 2002), although the founders introduced from Japan failed to reproduce (Liu et al. 2009). As these were the only recorded introductions, bullfrogs in China are assumed to have source single original source (Cuba) (Liu et al. 2009). Within a period of 50 years after the first documented introduction, bullfrog aquaculture has gone through 3 waves of development (Bai 1989; Zeng 1998). The first wave was in the early 1960s following the first introduction of bullfrogs. The duration of this boom was short and interrupted for social reasons in 1966 when all bullfrog farms were shut down for about twenty years (Bai 1989; Zeng 1998). In spite of that, some bullfrogs escaped from farms established feral populations in Hanshou county of Hunan Province (Zeng 1998). The second wave of growth was in 1980s, when bullfrog aquaculture became a big business benefiting from improved husbandry techniques and economic reforms in China. In this case, the founders were not newly introduced but originated from descendants of the feral bullfrog populations established in 1960s in the Hunan Province (Bai 1989; Zeng 1998). During this period simple enclosures were employed by many households (Liu et al. 2009), favoring frequent escapes and the establishment of new feral bullfrog populations in different regions of China (Liu and Li 2009). With the development of aquaculture, these small scale breeding farms were eliminated gradually, and the bullfrog aquaculture ran into the third and mature period. Currently, the remaining bullfrog farms consist of elaborate enclosures, usually surrounded by brick and concrete wall, more effective in preventing escapes (Liu and Li 2009).
Previous studies in Europe and USA have shown that invasive bullfrogs are able to establish successful populations, even from a limited number of founders with low genetic variation (Ficetola et al. 2008; Funk et al. 2011). Unlike invasive bullfrogs in Europe and USA, the bullfrogs were introduced in China for aquaculture purposes only (Li and Xie 2002; Zeng 1998). However, limited information is available about the origin, frequency, numbers of individuals introduced and the subsequent spread of bullfrogs and across China. Furthermore, historical data cannot reveal the genetic changes occurring during the range expansion. Here, we sequenced a part of the mitochondrial cytochrome b gene of Chinese wild and farm bullfrogs, and compared them with populations from the bullfrog native range (Austin et al. 2004). We then used a combination of genetic data and historical records to (a) test the hypothesis that the genetic diversity of both feral and farm populations is limited and smaller than that in the natural range of the species as a consequence of the limited numbers of original founders (founder effect) and (b) to identify the potential source population of the bullfrogs in China.
Materials and methods
Sample collection
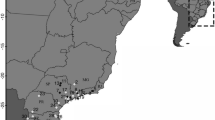
Surveys were conducted at 112 sites in six provinces of China during 2007–2010 (Fig. 1). Sampling sites were distributed at an altitudinal range of 12–2,700 m and a latitude range of 22.0°–30.3°N from the eastern to southwestern China (Fig. 1). All sites were in regions recorded as having bullfrog introductions for breeding purposes. To determine whether a site had been invaded by bullfrogs, we surveyed all accessible water bodies in each site for three consecutive nights with line transects (Li et al. 2011b). The third toe of the right hind-foot from each post-metamorphosed bullfrog was clipped if they were available (Bai et al. 2010). Tissue samples were then preserved separately in 70% EtOH and stored at −20°C in the laboratory. Since bullfrogs are known to be one of the main vectors of the deadly amphibian pathogen Batrachochytrium dendrobatidis (Bd) (Daszak et al. 2004; Fisher and Garner 2007; Garner et al. 2006), all the equipment, including boots and capture tools were disinfected before entering each site to prevent the spread of live Bd spores across sites.
Map showing feral and captive populations sampled in this study. Open circles denote sites where Lithobates catesbeianaus invaded but no individuals were captured, closed circles denote sites where L. catesbeianaus invaded and some individuals were captured; open triangles denote uninvaded sites; closed squares denote sampled food markets; closed stars denote sampled L. catesbeianaus farms. Some points are superimposed. Population assignment of feral individuals to the two haplotypes is represented by the shaded portion of the circle (H43: dark grey, H7: light grey; small circle: 1≤ sample size <10, maddile-sized circle: 10≤ sample size <20, big circle: 20≤ sample size)
Samples from captive bullfrogs were collected from six bullfrog farms and 23 food markets in 12 provinces and two municipalities during 2007–2011 (Fig. 1). At least five samples were collected per food market and twenty per bullfrog farm.
Laboratory analysis
DNA was extracted following the procedure described by Goka et al. (2009) with some modifications. In brief, approximately 3 mg (wet weight) of each toe-clip or mouthparts were placed in a 2 mL disposable centrifuge tube, 100 μL of lysis buffer containing 1 mg/mL proteinase K, 0.01 M NaCl, 0.1 M EDTA, 0.01 M Tris–HCl (pH 8.0) and 0.5% Nonidet P-40 was added to the tube. The microtube was shaken for 1 min using vortex mixer at room temperature (about 20°C), after a brief centrifugation to recover all material from the bottom of the tube, the tube was incubated at 50°C for 120 min and at 95°C for 20 min. Finally, the microtube was centrifuged at 12,000 rpm for 3 min in cold temperatures, and the extract was diluted one-tenth of its original concentration for polymerase chain reaction (PCR) amplification (Goka et al. 2009). A 591-bp segment of the cytochrome b mitochondrial gene was amplified as described in Ficetola et al. (2008). The PCR products were then separated on agarose gel electrophoresis (2% agarose gels). Finally, PCR products were directly sequenced using the same primers used for the amplification (Beijing Genomics Institute, Beijing, China). The sequenced fragment includes the 408-bp amplified by Austin et al. (2004) in the native range bullfrogs.
Data analysis
To make our results comparable with previous phylogeography analyses of bullfrogs in their native range (Austin et al. 2004), we used only 408-bp of the sequenced 591-bp for the following analysis. Sequences were aligned using Clustal X (Thompson et al. 1997), and unique haplotypes were identified using DnaSP 5.10 (Rozas et al. 2003). Number of haplotypes, haplotype diversity (Hd) and nucleotide diversity (π) within each introduced populations were estimated by ARLEQUIN ver3.5 (Excoffier and Lischer 2010). φST values were calculated to measure the genetic differentiation among feral bullfrog populations. Field sampling sites from each island in Zhoushan Archipelago were merged into a single biological meaningful population for their geographical proximity (Yang 2003).
Two basic methods were used to infer the most likely population(s) from which the invasive bullfrogs in China originated (Muirhead et al. 2008; Wares et al. 2005). Firstly, the putative native source was identified from the relationships among haplotypes between native and invasive populations based on the minimum spanning network. Networks were constructed based on statistical parsimony implemented in TCS 1.21 (Clement et al. 2000). In order to make a full comparison between our data and previously published work we also included haplotype information from the invaded Europe and Oregon range in the analysis Secondly, the similarity of haplotype frequencies among four native groups and invasive populations in China was assessed through Analysis of Molecular Variance (AMOVA) (Excoffier et al. 1992), and the native group with the highest level of similarity with invasive bullfrogs was considered the most likely population of origin. For AMOVA, the native range of bullfrogs was divided into four biologically meaningful groups based on the geographical distribution of cyt b clade groups (Austin et al. 2004; Ficetola et al. 2008). These four areas were the western group (clade 3–1 and subclades 2–1 and 2–2; native populations 19, 20, 21, 31, 32, 34 in Austin et al. 2004), the eastern group (clade 3–2 and subclades 2–3 and 2–5; populations 5, 7, 9, 25, 26, 28, 35, 36, 37, 38, 39, 40, 41, 42), the northeastern group (presence of the subclade 2–5 only within the clade 3–2; populations 1, 2, 3, 4, 6, 15, 16, 17, 24), and the area of overlap between western and eastern haplotypes (contemporary presence of haplotypes from both clades 3–1 and 3–2, and several different subclades; populations 8, 10, 11, 12, 13, 14, 18, 22, 23, 27, 29, 30, 33). The geographical distribution of the cyt b clades and geographic distribution of the populations is showed in Fig 3 and Fig 8 respectively in Austin et al. (2004).
Results
Bullfrogs were detected in 64 out of 112 surveyed sites. In total 288 bullfrog samples were collected from 42 invaded sites covering three provinces in China: 152 samples originated from 32 sites on 7 islands in the Zhoushan Archipelago of Zhejiang Province in eastern China and 136 samples came from 10 sites of Sichuan Province and Yunnan Province in southwestern China (Fig. 1, Table 1, Appendix 1—Electronic Supplementary Material). In addition, 222 samples were collected from captive bullfrogs: 133 individuals from 23 food markets and 89 from six bullfrog farms (Fig. 1, Table 2).
Only two haplotypes were found in the 510 samples (288 from feral populations and 222 from captive individuals) collected across China, while 41 haplotypes were identified from 215 samples collected from the native ranges and at least two haplotypes were detected in almost all native populations where more than one individual was sampled (Austin et al. 2004). Nucleotide diversity in feral bullfrogs from China was significantly lower than in those from the native ranges (Mann–Whitney U 15, 40 = 154, P = 0.006; populations where only a single individual was sampled were excluded from analysis).
Two closely related haplotypes (H43 and H7) were identified in both feral and captive bullfrogs, with only one silent nucleotide substitution different between them. Haplotype H7 was one of the most frequent in both the native and the other two invasive (Europe and Oregon) populations, while H43 (GenBank Accession no. JQ241268) had not been previously identified (Austin et al. 2004; Ficetola et al. 2008; Funk et al. 2011). Furthermore, H43 was present in high frequencies in samples from bullfrog farms, food markets and feral populations, amounting 98.9, 76.7 and 53.1% respectively. Among the samples collected in the farms, in five out of six farms a single haplotype (H43) was found and the only farm with variability had a low frequency of the haplotype H7 (1/18) (Table 2). The proportion of H43 in bullfrogs from the three different sources (bullfrog farms, food markets and feral populations) differed significantly (Chi-squared test, χ2 = 71.90, df = 2, P < 0.001). The proportion of H43 also differed significantly between bullfrogs from farms and food markets with frequencies of 98.9 and 76.7% respectively, (Chi-squared test, χ2 = 20.03, df = 1, P < 0.001). The proportion of H43 in food markets was also higher than that in wild populations (Chi-squared test, χ2 = 20.16, df = 1, P < 0.001).
Pairwise estimates of φST were not significant among six feral populations in Zoushan Archipelago after Bonferroni correction (italicised area in Table 3), suggesting they might derive from the same source of rearing stock, and thus the commercial populations that once existed in early 1990s on these island. In contrast, significant population differentiation was found among some sites of southwestern China, implying that multiple rearing stocks with different genetic compositions had been introduced into this region. However, we couldn’t rule out the possibility that the genetic differentiation among these populations were simply due to genetic drift.
Statistical parsimony network showed that haplotypes identified in the east and west native region were divided into two main groups (Fig. 2). The H43 and H7 haplotypes found in China clustered in the group of western haplotypes, suggesting that the invasive bullfrogs most likely originated from the western native region. Consistent with these results, AMOVA analysis also showed that the percentage of among group variation was smallest for western native populations (39.43%) when compared the four native groups with wild China (Table 4). The percentages of among group variations between Northeast, East and Overlap of native rage and wild China were larger, amounting 92.38, 86.62 and 64.29% respectively (Table 4). Overall, these results indicated that western native population has the highest probability to be the potential origin of invasive bullfrogs in China and that the genetic makeup of introduced bullfrogs has changed relative to their source populations during the process of introduction and spread.
Discussion
Only two haplotypes were identified both from wild (n = 288) and captive (n = 222) samples in China, while 41 haplotypes were present in 215 samples from native populations (Austin et al. 2004). This indicated that the molecular genetic diversity of invasive bullfrogs in China has decreased drastically compared to that of conspecifics in the native populations. Serious reductions of genetic diversity in invading areas have been commonly hypothesized as a result of low propagule pressures, especially the lack of multiple introductions from different native regions (Dlugosch and Parker 2008; Simberloff 2009). Our results match the historical records of bullfrog introductions in China, according to which bullfrogs were introduced only once during the late 1950s (Liu et al. 2009). Correspondingly, five haplotypes were recently identified in 174 samples from invasive feral populations in Europe (Ficetola et al. 2008), where at least 25 different introductions occurred between 1930s and late 1990s (Ficetola et al. 2007). The differences in genetic diversity of invasive bullfrogs between Europe and China is consistent with the hypothesis that multiple introductions can mitigate the loss of genetic diversity loss in invaders (Dlugosch and Parker 2008; Kolbe et al. 2004). Additionally, our results confirmed the ability of invasive bullfrogs to create new populations even with very low levels genetic diversity (Ficetola et al. 2008; Funk et al. 2011).
Three factors could have contributed greatly to the successful establishment of bullfrogs in China despite extreme low genetic diversity. Firstly, the high fertility of bullfrogs may allow them to survive demographic bottlenecks, as demonstrated by invasive bullfrogs in Europe where the most invasive populations derived from less than six females (Ficetola et al. 2008). Secondly, the invasive species characterized by initial small population size can experience an increase in the frequency of recessive homozygotes, which could increase the efficiency of selection against deleterious mutants, and thus the potential to purge inbreeding depression (Ficetola et al. 2011). Lastly, high propagule pressure due to repeated escapes from commercial populations. Liu and Li. (2009) found the establishment of bullfrog populations is positively correlated with the presence of simple aquaculture enclosures and the numbers of bullfrogs raised. The rare occurrences of feral bullfrogs in central China (Hubei and Hunan Provinces), where bullfrog aquaculture industry have flourished, is probably related to high hunting pressure (Liu and Li 2009), which may reduce bullfrog survival and the breeding chance of females because of higher hunting pressure on males that is easy to find by call identification in the field (Li et al. 2006). A recent study on invasive rainbow trout in Chile by Consuegra et al. (2011) found that propagule pressure—measured as the cumulative weighted distance to every salmonid farm within a 100-km radius—was a good predictor of the incidence of farm escapees, and that escapes could promote both the establishment and spread of feral population. These results not only underscore the ecological threats posed by aquaculture industry to wild populations, but also provide an alternative explanation of the well-known genetic paradox of why invaders become establishing successfully despite low genetic variation. The other identified potential solutions to the dilemma including high reproductive rates, purging deleterious alleles that cause inbreeding depression as mentioned above and multiple introduction events (Frankham 2005).
The different proportion of H43 in samples from captivity (85.6%) and wild (53.1%) indicated that extreme founding events had occurred during the mass-production process in captivity after 1990s, given that the majority of feral bullfrogs were descents of those escaped from farms with simple enclosures in 1980s–1990s (Liu and Li 2009; Zeng 1998). Subsequent founding events and population bottlenecks have probably occurred in commercial populations since 1990s as a consequence of the fluctuations of the bullfrog aquaculture industry. Thus, while only one or a few bullfrog farms were under operation when the bullfrog aquaculture was in depression, their descents were dispersed to many farms around the country at the flourishing period of the frog farming industry (Bai 1989). Although our results suggest that most farm populations are dominated by the H43 haplotype, the presence of H7 in many market samples such as food markets in Yuxi and Dali in Yunnan Province and Chengdu in Sichuan Province suggest that there must be some remaining genetic diversity in some of the unsampled farms, particularly in southwestern China. Thus, the results implied that the genetic composition of commercial bullfrog populations varied among farms, as for other introduced commercial populations such as the honey bee (Hymenoptera: Apidae) in the United States (Delaney et al. 2009). We cannot discard that the extremely high frequency of H43 found in farm populations could be an artifact of insufficient sampling, since only six farms were sampled and four of them were managed by the same aquaculture company in Huzhou, Zhejiang Province. However, 23 food markets distributed across 21 cities of 14 provinces were sampled (Table 2), and bullfrog farms are the only source of bullfrogs in food markets. Thus, samples from food markets could provide a better representation of commercial populations of bullfrog.
Both AMOVA and phylogeographic analyses indicate that western native populations are the most likely origin of invasive bullfrogs in China. However, a significant genetic difference still existed between bullfrog populations in western native and those in caught in the field in China (P = 0.0001) (Table 4). We suggest that the widely distributed new haplotype (H43) should be responsible for this differences, because AMOVA is more sensitive to the sequence and frequency changes of haplotypes (Wares et al. 2005). A recent study on the invasive Anolis sagrei in St Vincent also found the absence or lower frequency in the native range of particular haplotypes that were nearly fixed in the invasive range due to founder effects during the introduction and secondary establishment process, making it hard to identify the original population (Thorpe and Eales 2010). Both the intensity of sampling and the resolution of the molecular markers can strongly influence the accuracy of source population assignment (Dlugosch and Parker 2008; Muirhead et al. 2008). For this study, samples have been collected across large areas of both the native range and the invasive range. However, only a single locus in the mitochondrial cytochrome b gene was analysed, and the differentiation among some populations might not be fully resolved. The application of multiple variable markers would help in the finer discrimination among populations (Muirhead et al. 2008). Furthermore, since Cuba seems to be the initial origin of bullfrogs in China, the comparison of the genetic structure of samples from Cuba could provide a better understanding of the source and introduction history of bullfrogs in China.
Although H43 has been firstly identified in China, there are three potential sources of H43. Firstly and most importantly, H43 could have been present as a rare haplotype in the United States and introduced to Cuba in 1916–1917 (Zeng 1998). The reasons why it was not detected by Austin et al. in 2004 could be either that it disappeared due to genetic drift or that it was only present at low frequencies. Secondly, H43 could have originated in Cuba as a result of mutation and have been introduced to China in late 1950s. Given that bullfrogs had been widely cultured across Cuba since early 1940s for food supply and international trade (Zeng 1998), founding events are also likely in Cuban commercial populations. Lastly, H43 could have derived from a mutation in H7 after bullfrogs were introduced to China and then increased in frequency due to several founder effects. Although the origin of H43 is still under debate, there is no doubt that sequential founding events should be responsible for the high frequency of H43 both in wild and captive populations in China.
Sequential founding events related to introduced commercial populations have important implications for the understanding and study of biological invasions. Firstly, rare alleles/haplotypes will be lost or overrepresented in these populations compared to the initial founders, making it difficult the inferences of the origin of the source populations through the analysis of genetic variation (Muirhead et al. 2008; Wares et al. 2005). Thus the interpretation of our results should be taken cautiously. Secondly, commercial populations can be critical as a source of continuing propagule pressure (Consuegra et al. 2011; Naylor et al. 2005). Delaney et al. (2009) found that the genetic makeup of commercial honey bees changed over a 10-year span due to artificial selection, and the commercial populations of Orius strigicollis in Japan were always genetically different from nearby wild populations (Hinomoto et al. 2006). These examples would lead us to expect that the commercial populations may act as genetic pools to the invasive colorizations, just as multiple introductions can do to promote the establishment and spread of invasive species (Allendorf and Lundquist 2003; Forsyth and Duncan 2001; Kolar and Lodge 2001; Lavergne and Molofsky 2007; Lockwood et al. 2005; Roman and Darling 2007; Simberloff 2009; Von Holle and Simberloff 2005). Our results reaffirm the difficulty of inferring the original source of invasive populations based on genetic analysis alone as a result of sequential founding events, and the potential threats posed by introduced commercial bullfrog populations, given their unique genetic makeup compared to feral populations.
References
Allendorf FW, Lundquist LL (2003) Introduction: population biology, evolution, and control of invasive species. Conserv Biol 17:24–30
Austin JD, Lougheed SC, Boag PT (2004) Discordant temporal and geographic patterns in maternal lineages of eastern north American frogs, Rana catesbeiana (Ranidae) and Pseudacris crucifer (Hylidae). Mol Phylogenet Evol 32:799–816
Bai JY (1989) The current issue of bullfrog breeding. China Fish 11:10–11
Bai CM, Garner TWJ, Li YM (2010) First evidence of Batrachochytrium dendrobatidis in China: discovery of chytridiomycosis in introduced American bullfrogs and native amphibians in the Yunnan Province, China. Ecohealth 7:127–134
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659
Consuegra S, Phillips N, Gajardo G, de Leaniz CG (2011) Winning the invasion roulette: escapes from fish farms increase admixture and facilitate establishment of non-native rainbow trout. Evol Appl 4:660–671
Daszak P, Strieby A, Cunningham AA, Longcore JE, Brown CC, Porter D (2004) Experimental evidence that the bullfrog (Rana catesbeiana) is a potential carrier of chytridiomycosis, an emerging fungal disease of amphibians. Herpetol J 14:201–207
Delaney DA, Meixner MD, Schiff NM, Sheppard WS (2009) Genetic characterization of commercial honey bee (Hymenoptera: Apidae) populations in the United States by using mitochondrial and microsatellite markers. Ann Entomol Soc Am 102:666–673
Dlugosch KM, Parker IM (2008) Founding events in species invasions: genetic variation, adaptive evolution, and the role of multiple introductions. Mol Ecol 17:431–449
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Ficetola GF, Coic C, Detaint M, Berroneau M, Lorvelec O, Miaud C (2007) Pattern of distribution of the American bullfrog Rana catesbeiana in Europe. Biol Invasions 9:767–772
Ficetola GF, Bonin A, Miaud C (2008) Population genetics reveals origin and number of founders in a biological invasion. Mol Ecol 17:773–782
Ficetola GF, Garner TWJ, Wang JL et al (2011) Rapid selection against inbreeding in a wild population of a rare frog. Evol Appl 4:30–38
Fisher MC, Garner TWJ (2007) The relationship between the emergence of Batrachochytrium dendrobatidis, the international trade in amphibians and introduced amphibian species. Fungal Biol Rev 21:2–9
Floerl O, Inglis GJ, Dey K, Smith A (2009) The importance of transport hubs in stepping-stone invasions. J Appl Ecol 46:37–45
Forsyth DM, Duncan RP (2001) Propagule size and the relative success of exotic ungulate and bird introductions to New Zealand. Am Nat 157:583–595
Frankham R (2005) Invasion biology—resolving the genetic paradox in invasive species. Heredity 94:385–385
Funk WC, Garcia TS, Cortina GA, Hill RH (2011) Population genetics of introduced bullfrogs, Rana (Lithobates) catesbeianus, in the Willamette Valley, Oregon, USA. Biol Invasions 13:651–658
Garner TWJ, Perkins MW, Govindarajulu P, Seglie D, Walker S, Cunningham AA, Fisher MC (2006) The emerging amphibian pathogen Batrachochytrium dendrobatidis globally infects introduced populations of the North American bullfrog, Rana catesbeiana. Biol Lett 2:455–459
Goka K, Yokoyama J, Une Y, Kuroki T, Suzuki K, Nakahara M, Kobayashi A, Inaba S, Mizutani T, Hyatt AD (2009) Amphibian chytridiomycosis in Japan: distribution, haplotypes and possible route of entry into Japan. Mol Ecol 18:4757–4774
Hinomoto N, Higaki T, Noda T (2006) Genetic diversity in field and commercial populations of Orius strigicollis (Poppius) (Heteroptera: Anthocoridae) measured by microsatellite markers. Appl Entomol Zool 41:499–506
Hufbauer RA, Bogdanowicz SM, Harrison RG (2004) The population genetics of a biological control introduction: mitochondrial DNA and microsatellite variation in native and introduced populations of Aphidus ervi, a parisitoid wasp. Mol Ecol 13:337–348
ISSG (2008) Invasive Species Specialist Group. http://www.issg.org/database/welcome/. Accessed 6 Jun 2011
Kats LB, Ferrer RP (2003) Alien predators and amphibian declines: review of two decades of science and the transition to conservation. Divers Distrib 9:99–110
Kiesecker JM, Blaustein AR (1998) Effects of introduced bullfrogs and smallmouth bass on microhabitat use, growth, and survival of native red-legged frogs (Rana aurora). Conserv Biol 12:776–787
Kiesecker JM, Blaustein AR, Miller CL (2001) Potential mechanisms underlying the displacement of native red-legged frogs by introduced bullfrogs. Ecology 82:1964–1970
Kolar CS, Lodge DM (2001) Progress in invasion biology: predicting invaders. Trends Ecol Evol 16:199–204
Kolbe JJ, Glor RE, Schettino LRG, Lara AC, Larson A, Losos JB (2004) Genetic variation increases during biological invasion by a Cuban lizard. Nature 431:177–181
Kolbe JJ, Larson A, Losos JB, de Queiroz K (2008) Admixture determines genetic diversity and population differentiation in the biological invasion of a lizard species. Biol Lett 4:434–437
Lavergne S, Molofsky J (2007) Increased genetic variation and evolutionary potential drive the success of an invasive grass. Proc Natl Acad Sci USA 104:3883–3888
Lawler SP, Dritz D, Strange T, Holyoak M (1999) Effects of introduced mosquitofish and bullfrogs on the threatened California red-legged frog. Conserv Biol 13:613–622
Lever C (2003) Naturalized reptiles and amphibians of the world. Oxford University Press, Oxford
Li Z, Xie Y (2002) Invasive alien species in China. China Forestry Publishing House, Beijing
Li YM, Wu ZJ, Duncan RP (2006) Why islands are easier to invade: human influences on bullfrog invasion in the Zhoushan archipelago and neighboring mainland China. Oecologia 148:129–136
Li YM, Ke ZW, Wang YH, Blackburn TM (2011a) Frog community responses to recent American bullfrog invasions. Curr Zool 57:83–92
Li Y, Xu F, Guo Z, Liu X, Jin C, Wang Y, Wang S (2011b) Reduced predator species richness drives the body gigantism of a frog species on the Zhoushan Archipelago in China. J Anim Ecol 80:171–182
Liu X, Li Y (2009) Aquaculture enclosures relate to the establishment of feral populations of introduced species. PLoS ONE 4(7):e6199
Liu X, Li Y, McGarrity ME (2009) Geographical variation in body size and sexual size dimorphism of introduced American bullfrogs in southwestern China. Biol Invasions 12:2037–2047
Lockwood JL, Cassey P, Blackburn T (2005) The role of propagule pressure in explaining species invasions. Trends Ecol Evol 20:223–228
Muirhead JR, Gray DK, Kelly DW, Ellis SM, Heath DD, Macisaac HJ (2008) Identifying the source of species invasions: sampling intensity vs. genetic diversity. Mol Ecol 17:1020–1035
Naylor R, Hindar K, Fleming IA, Goldburg R, Williams S, Volpe J, Whoriskey F, Eagle J, Kelso D, Mangel M (2005) Fugitive salmon: assessing the risks of escaped fish from net-pen aquaculture. Bioscience 55:427–437
Pearl CA, Adams MJ, Bury RB, McCreary B (2004) Asymmetrical effects of introduced bullfrogs (Rana catesbeiana) on native Ranid frogs in Oregon. Copeia 2004:11–20
Pimentel D, Zuniga R, Morrison D (2005) Update on the environmental and economic costs associated with alien-invasive species in the United States. Ecol Econ 52:273–288
Roman J, Darling JA (2007) Paradox lost: genetic diversity and the success of aquatic invasions. Trends Ecol Evol 22:454–464
Rozas J, Sanchez-DelBarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Sakai AK, Allendorf FW, Holt JS, Lodge DM, Molofsky J, With KA, Baughman S, Cabin RJ, Cohen JE, Ellstrand NC, McCauley DE, O’Neil P, Parker IM, Thompson JN, Weller SG (2001) The population biology of invasive species. Ann Rev Ecol Syst 32:305–332
Simberloff D (2009) The role of propagule pressure in biological invasions. Annu Rev Ecol Evol Syst 40:81–102
Simberloff D, Stiling P (1996) Risks of species introduced for biological control. Biol Conserv 78:185–192
Strayer DL, Eviner VT, Jeschke JM, Pace ML (2006) Understanding the long-term effects of species invasions. Trends Ecol Evol 21:645–651
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Thorpe RS, Eales J (2010) Revealing the geographic origin of an invasive lizard: the problem of native population genetic diversity. Biol Invasions 12:77–86
Vitousek PM, DAntonio CM, Loope LL, Westbrooks R (1996) Biological invasions as global environmental change. Am Sci 84:468–478
Von Holle B, Simberloff D (2005) Ecological resistance to biological invasion overwhelmed by propagule pressure. Ecology 86:3212–3218
Wares J, Hughes A, Grosberg R (2005) Mechanisms that drive evolutionary change: insights from species introductions and invasions. In: Sax DF, Stachowicz JJ, Gaines SD (eds) Species invasions: insights into ecology, evolution, and biogeography. Sinauer Associates, Sunderland, pp 229–257
Westphal MI, Browne M, MacKinnon K, Noble I (2008) The link between international trade and the global distribution of invasive alien species. Biol Invasions 10:391–398
Yang Y-H (2003) Phylogeographic patterns and population history of the black-spotted pond frog Rana nigromaculata revealed by the mitochonrial cytochrome b gene, Dissertation, Chinese Academy of Science
Zeng XJ (1998) Prospect and strategy of bullfrog farming. Inland Fish 10:4–5
Acknowledgments
We thank Feng Xu, Lin Zhang, Supen Wang, Wei Zhu and Yu Luo for collecting samples. We are also grateful to James Fordyce and two anonymous reviewers for helpful comments on the draft of the manuscript. All of the collection and handling of amphibians was conducted under the approval of the Animal Care and Ethics Committee, Institute of Zoology, Chinese Academy of Sciences. This research was supported by grants from the “973” program (code: 2007CB411600) and the National Science Foundation (code: 30870312).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bai, C., Ke, Z., Consuegra, S. et al. The role of founder effects on the genetic structure of the invasive bullfrog (Lithobates catesbeianaus) in China. Biol Invasions 14, 1785–1796 (2012). https://doi.org/10.1007/s10530-012-0189-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-012-0189-x