Abstract
The common ragweed (Ambrosia artemisiifolia L.; Asteraceae) is a North American native that is invading Eurasia. Besides its economic impact on crop yield, it presents a major health problem because of its highly allergenic pollen. The plant was imported inadvertently to Europe in the eighteenth century and has become invasive in several countries. By analyzing French and North American populations, it was previously shown that French populations were best described as a mixture of native sources and that range expansion in France probably involved sequential bottlenecks. Here, our aim was to determine whether Eastern European populations of A. artemisiifolia originated from the previously established French populations or from independent trans-Atlantic colonization events. We used nuclear microsatellite markers to elucidate the relationships among populations from Eastern and Western Europe in relation to populations from a broad survey across the native North American range. We found that A. artemisiifolia from Eastern Europe did not originate from the earlier established French populations but rather represents multiple independent introductions from other sources, or introductions from a not yet identified highly diverse native population. Eastern European populations show comparable amounts of genetic variability as do previously characterized French and North American populations, but analyses of population structure clearly distinguish the two European groups. This suggests separate introductions in Eastern and Western Europe as well as divergent sources for these two invasions, possibly as a result of distinct rules for trade and exchange for Eastern Europe during most of the twentieth century.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Species introductions involve demographic and genetic bottlenecks when colonists are few and represent only a subset of the genetic variation available in populations in the native range (Husband and Barrett 1991). However populations of introduced and invasive plants do not always lose genetic variation during invasion (see Bossdorf et al. 2005; Dlugosch and Parker 2008; Puillandre et al. 2008 for reviews). Some invasive populations originate from multiple introductions, thereby amassing allelic variation from a broad range of different populations from the native range (Bossdorf et al. 2005; Ciosi et al. 2008; Facon et al. 2005; Genton et al. 2005a; Hufbauer and Sforza 2008; Lavergne and Molofsky 2007), though multiple introductions do not always lead to an increase in genetic variation (Durka et al. 2005). Altogether, successful introductions, i.e. those leading to naturalization and invasion of a new geographical area, often appear to involve relatively small bottlenecks, with the best predictor of establishment success being propagule pressure, both in terms of number of introductions and number of individuals released (Simberloff 2009). Therefore many species that successfully establish and invade are likely those that lost least variation in the process, though invasions of single genotypes are not unknown (Grimsby et al. 2007; Okada et al. 2009).
Invasive populations may also show altered genetic structure compared to those in the native range, either less, with homogenised mixes of representatives of several differentiated populations (Le Roux et al. 2008), or more (Ciosi et al. 2008; Marrs et al. 2008), with invasive populations being particular subsamples of well mixed populations from the native range. Studies that compare genetic structures of invasive populations and populations in the native range can help pinpoint the area of origin as well as elucidating patterns of colonisation and pathways of spread (Le Roux et al. 2006). Do invaders come from areas with similar climatic characteristics? Have invaders followed a stepping-stone from an initial unique introduction (Amsellem et al. 2000)? Do newly colonised populations at the front of the species range originate from already established ones in the invaded range (Genton et al. 2005a) or do they represent independent colonization events from the native range (Ciosi et al. 2008)? These questions are relevant for practical purposes of risk assessment and the efficacy of control measures but also will provide important insights into the evolutionary ecology of invasive and other species undergoing range expansion.
Here we investigate the genetic structure of populations of the invasive Ambrosia artemisiifolia L. (Asteraceae), a North American native that is invading Eurasia, found particularly in sunflower and corn fields, abandoned fields, disturbed areas and along roadsides (Bassett and Crompton 1975). This wind-pollinated monoecious annual plant is self-incompatible, thus showing an outcrossing mating system, even in colonizing populations (Friedman and Barrett 2008). During the past 15–20 years, the spread of common ragweed has become a major problem in some parts of France and a number of Eastern European countries, including Hungary, Croatia, Ukraine, Russia and Serbia (Kiss and Béres 2007).
The first Eurasian records of this species are from a herbarium specimen from Central France from 1863, and the species showed a gradual but continuous spread in this region, demonstrating continuous presence in the area of Lyon, France, which seems to be the focus of its current French distribution (Chauvel et al. 2006). The earliest recorded Eastern European records of this plant appear first 40 years later from Orsova, Romania, and about 20 years after that from the south-western part of Hungary (Kazinczi et al. 2008; Makra et al. 2005; Csontos et al. 2010). It is therefore possible that Eastern Europe was colonized from the already established French invasive populations, though an independent introduction from the native range of this species is also possible. Over the last 20 years additional populations have appeared and become established in the intervening areas of Switzerland, northern Italy and Austria, but these are thought to represent recent range expansion from one or the other long established centres of spread. The conquest of Eastern Europe, however, has been dramatically more rapid and complete than that of the West, with a larger occupied range and far denser populations, as revealed by pattern of airborne pollen density across Europe (Fig. 1; Makra et al. 2004; Makra et al. 2005). This then poses the question of whether the Eastern European invasion is caused by other more competitive genotypes or those better adapted to the conditions in Eastern Europe.
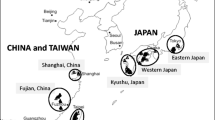
Annual counts of Ambrosia artemisiifolia pollen grains (a) averaged over 1995–2007 (reproduced with permission from Regula Gehrig, MeteoSwiss; source: EAN European Aeroallergen Network) as a proxy for the local distribution and abundance of this plant; sampling sites of A. artemisiifolia (b) in Western Europe (previously presented in Genton et al. 2005a) and Eastern Europe (new to this study). F France; HU Hungary; RO Romania; UA Ukraine; SCG Serbia and Montenegro
Here we extend our previous investigation of the population structure of invasive populations in France (Genton et al. 2005a). We have previously shown that the French invasive populations of A. artemisiifolia originated from a mixture of sources. French populations showed even higher within-population diversity than did native North American populations, they contained mixes of rare alleles found in distinct North American populations and assignment tests failed to identify a single area of origin in North America (Genton et al. 2005a). Chun et al. (2010) showed that historical French populations, reconstructed from herbarium specimens dating from the late nineteenth to early twentieth century, appeared to harbour lower allelic and genetic diversity than recent populations. Altogether, this suggests that ragweed seeds were introduced repeatedly, or as mixtures, from different parts of North America to France. However within France, ragweed populations at the front of the invasion, far from the original area of introduction near Lyon, France, are genetically less diverse, indicating that ragweed range expansion probably involves sequential bottlenecks from the primary introduction rather than subsequent new introductions (Genton et al. 2005a). Here we test whether Eastern European ragweed populations could have been founded from French populations involving sequential bottlenecks or whether Eastern European ragweed populations were introduced independently.
We analysed Eastern European populations of this invasive plant using microsatellites to address the following questions: (1) Were Eastern European populations founded from French populations? Are they characterised by a subset of the allelic diversity already found in the French populations? (2) Did they come independently from similar sources in North America, characterized by a similar amount of variation and similar allelic profiles, or from elsewhere? Here we compare our new data from Eastern European ragweed populations with data previously presented in Genton et al. (2005a).
Methods
Sampling and DNA extractions
Ragweed populations were sampled in six localities in Eastern Europe (Fig. 1; Table 1). In each sampling site leaves were collected from 30 individual plants at approximately 1.5 m spacing, according to the sampling method of Genton et al. (2005a), air-dried, and kept as herbarium materials. DNA was extracted from 10 to 15 mg dried leaf tissue using DNeasy Plant Mini Kits (QIAGEN) and then stored at −20°C.
Genetic data for a total of 12 North American and 10 French ragweed populations, obtained in a previously published study (Genton et al. 2005a), were also included in this work. The American populations sampled were located east of the Rocky Mountains, mostly from the East Coast and Great Lakes region of the USA and Canada, areas with a long history of commercial exchange with Western Europe. The French ragweed populations sampled were located in the Rhône-Alpes, Provence-Alpes-Côte-d’Azur and Bourgogne regions. The designations, locations and other data of the American and French ragweed populations included in this study are given in Genton et al. (2005a).
Microsatellite genotyping and analyses
We used the five nuclear microsatellite loci recently developed for A. artemisiifolia (Genton et al. 2005b). Polymerase chain reaction and allele resolution were carried out according to Genton et al. (2005a). Several samples from the study by Genton et al. (2005a), having the full range of alleles previously identified, were run on each gel to score allele sizes consistently between the previous and the present studies. New primers were designed for the locus Amb15 because of difficulties in amplification using the previous primers from Genton et al. (2005b). The new primer pair (Amb15-F2: aatccattccccacatcctt and Amb15-R2: gaggggttgggtcgagtaag) gave amplifications in a higher number of individuals.
Using FSTAT version 2.9.3.2 (Goudet 1995), we estimated (1) the means ± SD over all loci of the following genetic variation indices: allelic richness (R S ; El Mousadik and Petit 1996), observed heterozygosities and expected heterozygosities, respectively H O and H E (Nei 1987; the latter also referred to as Nei’s gene diversity); (2) F-statistics (Weir and Cockerham 1984). Deviations from Hardy-Weinberg proportions were assessed using FSTAT (Goudet 1995) with P-values being corrected for table-wide significance levels (α = 0.05) using the sequential Bonferroni method (Rice 1989). We also computed the means over all loci of the number of alleles (A) and the number of rare alleles (A R ), i.e. with allelic frequencies below 0.1. Finally, we tested for linkage disequilibria using Fisher exact tests in FSTAT (Goudet 1995) with P-values being corrected for table-wide significance levels (α = 0.05) using the sequential Bonferroni method (Rice 1989).
Genetic distances among populations were measured with the Populations program (http://bioinformatics.org/~tryphon/populations/), using the chord distance of Cavalli-Sforza and Edwards (1967). The distance matrix was submitted to a Principal Coordinate Analysis (PCoA), as implemented in Genalex (Peakall and Smouse 2006).
Comparison of genetic diversity and population differentiation
To compare the values of within-population genetic variation indices (A, A R , H E , F IS ) and population differentiation (F ST ) between the native range, France and Eastern Europe, we used FSTAT. For each index FSTAT computes the average over loci and populations for each group and then the squared difference between these two averages. Significance of this difference is then assessed using a permutation test: the whole sample is allocated at random to the three groups, keeping the number of populations constant in each group.
To compare the means of numbers of null genotypes among French, North American and Eastern European populations, pairwise mean comparisons were carried out using Student t-tests with the software JMP (SAS Institute Inc, SAS Campus Drive, Cary NC). The locus Amb15 was not included in these comparisons because different primer pairs had been used for genotyping Eastern European populations (see above), precisely because the number of null alleles was high using the first primer pair. Amb15 was however used in all other analyses as we could make the correspondence between the same alleles amplified using the two different primer pairs.
Analyses of molecular variance (AMOVAs) were performed using Arlequin version 3.0 (Excoffier et al. 2005), with variation being partitioned within- and among-populations, and distance among genotypes calculated as the number of different alleles. This method was preferred over the method that uses the squared number of repeat difference between genotypes to calculate distances (Slatkin 1995) to avoid biases due to departures from the stepwise model of microsatellite evolution.
Population subdivision and assignment tests
We used Structure version 2.3.1 (Pritchard et al. 2000) to identify the source populations of Eastern European invasive populations and to examine population subdivision. The model implemented allowed information on sampling location (LocPrior model; Hubisz et al. 2009), admixture, and correlation in allele frequencies (Falush et al. 2003). Burn-in length was set at 150,000 iterations followed by a run phase of 750,000 iterations. We employed a hierarchical approach to detect all layers of population structure (Coulon et al. 2008; Evanno et al. 2005; Rollins et al. 2009). Analyses were first conducted on the total dataset, using the region of origin of individuals as prior information to assist clustering. We then repeated analyses on each of the K groups inferred in the previous step, using the population of origin of individuals as prior information. We set the number of populations (K) from 1 to 8, but not higher, as the results for the first level of analysis showed that setting K values from 3 to 8 already led to clusters without geographical coherence and with admixed ancestry, which is typical of too high a cluster number. For all levels of analysis, we performed 30 independent runs for each value of K. Results were analysed with clumpp version 1.1.2 (Jakobsson and Rosenberg 2007) using the Greedy algorithm for 1 ≤ K≤4 and the Fast-Greedy algorithm for K > 4, with random input order and 10,000 permutations. Distinct modes among runs were identified by finding sets of runs with less than 85% similarity in the G′ pairwise similarity matrix (‘modes’ refer to distinct clustering solutions represented within the set of replicate cluster analyses). clumpp was used again to align outputs of the runs with the same clustering mode and to provide average cluster membership coefficients across aligned runs. The optimal K value was determined using the method of Evanno et al. (2005) based on the rate of change in the log probability of data between successive K values.
Distribution of private alleles
Because rare alleles can be powerful in identifying sources and routes of migrations, we also examined the distribution of private alleles, i.e. alleles unique to a single population or combination of populations. We used the program ADZE (Szpiech et al. 2008), which implements a rarefaction procedure for counting alleles private to populations while adjusting for differences in sample size across populations. We calculated the mean number of private alleles, averaging across loci, for each of three regional groupings of populations (North America, France, Eastern Europe) and each of three combined sets of two regional groupings. Calculations were performed using a standardized sample size of n = 252 (126 individuals times two chromosomes), corresponding to the smallest number of observations per regional grouping under consideration.
Results
Genetic diversity, F-statistics and Hardy Weinberg equilibrium
The five nuclear microsatellite loci had a total of 95 alleles in the six Eastern European populations analysed. Diversity indices, F-statistics estimates (Weir and Cockerham 1984) and results of exact Hardy-Weinberg tests (Guo and Thompson 1992) are presented in Table 1 for Eastern European populations and can be found in Genton et al. (2005a) for North American and French populations. Significant heterozygote deficiencies were detected in all populations, yielding significant positive F IS values. These values were greater in Eastern European populations than in North American ones and in France (P < 0.01, Fig. 2). Tests for linkage disequilibria were all non-significant. Genetic variation, measured as number of alleles, number of rare alleles or expected heterozygosity, was not significantly different among the native range, France and Eastern Europe (Fig. 3).
To assess whether the higher level of F IS in Eastern European populations could be due to the presence of more null alleles, we compared the proportion of null genotypes (i.e. giving no amplification in one or a few loci but giving bands in other loci; thus being assumed to be homozygotes for null alleles) among Eastern European, French and North American populations. Eastern European populations had a significant higher mean proportion of null genotypes (mean ± SE = 15.9 ± 2.0), than French (mean ± SE = 8.5 ± 1.5) or North American (mean ± SE = 10.2 ± 1.4) populations, which did not differ significantly from each other.
Comparison of variability distribution among populations between North America, Western Europe and Eastern Europe
We first performed an analysis of molecular variance (AMOVA) on Eastern European populations to divide the genetic variance into within- and among population components. Results indicated that most genetic variation in Eastern Europe was within, rather than among, populations (Table 2), as was the case in the native range and in France (Genton et al. 2005a). The percentage of variation attributed to among-population differentiation was 8.79% in Eastern Europe (Table 2), i.e. higher than in France (4.81%) and in North America (6.39%).
Distribution of private alleles
The number of private alleles was calculated for regional groupings of populations and their combinations (Fig. 4). At the scale of regions, estimates were higher in Eastern Europe (mean ± SE = 5.29 ± 2.50) than in France (mean ± SE = 1.84 ± 0.95) and North America (mean ± SE = 0.73 ± 0.51). In analyses on combinations of regions, private allele richness was higher in the North America/France combination (mean ± SE = 4.14 ± 2.03) than in the North America/Eastern Europe (mean ± SE = 1.45 ± 0.72) and France/Eastern Europe (mean ± SE = 1.28 ± 0.72) combinations. The fact that the proportion of private alleles is the highest in North America and France when these regions are combined while it is the lowest when regions are analysed individually suggests that North American and Western European populations are more similar to each other in terms of allelic profiles than they are to Eastern European populations.
Genetic relationships among populations
Chord genetic distances were calculated among populations, and the resulting distance matrix was subjected to PCoA (Fig. 5). The first principal coordinate (explaining 41.7% of total variation) partitioned populations into two groups with populations from France and North America in one group, and populations from Eastern Europe in the other. The second and third principal coordinates (explaining 15.1 and 12.9% of total variation, respectively) did not reveal any obvious further correspondence between genetic distances and the geographical origin of populations. The only remarkable feature that emerges from the decomposition of variance along these axes is that the third coordinate separated the Ukrainian population from other Eastern European populations.
Principal coordinate analysis of chord distance among populations. The first, second and third principal coordinates account for 41.7, 15.1 and 12.9% of the variation, respectively. NA North America; F France; HU Hungary; RO Romania; UA Ukraine; SCG Serbia and Montenegro. Squares represent US, circles West European and diamonds East European populations
In Structure analyses, the modal value of the ∆K statistic was found at K = 2, with North American and French populations in one cluster, and Eastern European populations in another (Fig. 6). Only a few individuals contradicted this pattern of clustering and had mixed membership in the two clusters or were not assigned to the cluster containing individuals from the region from which they were sampled. Generally, levels above K = 2 produced no new clusters corresponding to geographical structuring among North American and French genotypes, but instead introduced some heterogeneity in membership coefficients. Within Eastern European populations, no particular geographical pattern of clustering was detected, except that all individuals from Ukraine had consistently high membership in the same cluster and ended up individualizing in a separate cluster when K reached 6 (not shown). In the next level of analyses, we searched for possible additional layers of structure by repeating analyses on both of the K = 2 groups inferred in the previous step, using ‘populations’—and not ‘regions’—as prior information to assist clustering. In analyses of Eastern European samples, the modal value of ∆K was found at K = 2, with a secondary peak at K = 5. At K = 2, one cluster corresponded to individuals from Ukraine, and the other clusters grouped individuals from all other populations (not shown). At K = 5, individuals from populations from Serbia-Montenegro, Romania, and Ukraine (i.e. SCG-Z, RO-E, UA-K) had high membership proportions in separate clusters, individuals from Hungarian populations HU-Bi and HU-Bu had high membership in the same cluster, and individuals from Hungarian population HU-K had roughly equal membership in multiple clusters (Fig. 6). In analyses of French and North American samples, the modal value of ∆K was observed for K = 6, but neither K = 6 nor any other value of K yielded an obvious pattern of geographical clustering (Fig. 6). In support of a lack of population structure, average log posterior probabilities of data for K = 1 and K = 2 were very close (on average over runs from the main mode: Ln(P) = −10,512.2 and Ln(P) = −10,512.9 for K = 1 and K = 2, respectively), suggesting that a model with multiple populations is not significantly better than a model with a single population to represent data from French and North American samples.
Population structure of Ambrosia artemisiifolia inferred using the Structure program. The number of predefined clusters was K = 2 for the analysis of the total dataset. Subsequent hierarchical analyses (indicated by arrows) assumed K = 6 clusters for French and North American samples, and K = 5 clusters for Eastern European samples. Each individual is represented by a thin vertical line that is partitioned into two components according to the inferred membership in the two genetic clusters. Black lines separate genotypes from distinct populations. Vertical axis represents the membership proportions to the K clusters assumed, obtained using the clumpp program by averaging memberships across all runs corresponding to the main clustering mode. NA North America; F France; HU Hungary; RO Romania; UA Ukraine; SCG Serbia and Montenegro
Discussion
Invasion history of European ragweed
Overall population genetic variability of A. artemisiifolia, measured as expected heterozygosity or allelic richness, was similar in North America, Eastern and Western Europe. We found no evidence for the loss of genetic variation via sequential bottlenecks observed for other invasions (e.g. Amsellem et al. 2000; Henry et al. 2009; Puillandre et al. 2008). Therefore, in contrast to what was found in France, where populations at the current invasion front in the Bourgogne and PACA regions originate from the original populations of introduction in the East of Lyon (Genton et al. 2005a), Eastern European populations appear not to have originated from colonists from these older established French populations. Nor did Eastern European populations originate from a single source among the native populations sampled. The high genetic variability observed in Eastern European populations suggests either multiple sources of introduction, or introduction from a highly diverse source that we failed to sample (e.g. Muirhead et al. 2008). Multiple introductions have been inferred for many plant introductions studied to date (Bossdorf et al. 2005; Dlugosch and Parker 2008; Hufbauer and Sforza 2008), including the introduction of this same species to Western Europe (Genton et al. 2005a).
Analyses of the distribution of genetic variability brought additional insights to the history of the ragweed invasion. AMOVA, PCoA and Structure analyses revealed contrasted patterns of population structure in the introduced range: while populations from France appeared less differentiated than populations from the native range, a higher level of geographical structure was observed among Eastern European populations. These differences may be related to the population structure in the native range and the genetic makeup of founding propagules. The shallow population structure of introduced French populations suggests that they were all founded by genetically similar sources, either by a single introduction of mixed propagules followed by dissemination, or by multiple introductions coming from similar mixtures of sources (Genton et al. 2005a). Such a transformation of among population genetic variation into within population genetic variation due to several introductions from similar multiple sources has been reported in several other cases of biological invasions with multiple introductions (e.g. Kolbe et al. 2004; Lavergne and Molofsky 2007). By contrast, the higher population structure observed in Eastern European populations of A. artemisiifolia may correspond to the introduction of genetically differentiated propagules resulting from independent samplings either from similar highly diverse populations or from separate differentiated populations. Such a pattern of higher population structure in the invasive range has been reported for several organisms (e.g. Ciosi et al. 2008; Marrs et al. 2008), though it seems less frequent than the opposite pattern (Bossdorf et al. 2005).
Different origins and structure for Eastern and Western European invasive ragweed populations
Several lines of evidence indicate that the Eastern and Western European invasive ragweed populations originate, at least in part, from separate mixes of different native populations. Analyses of population structure revealed that Western and Eastern European populations were differentiated, and French populations of A. artemisiifolia appeared genetically much more similar to the sampled North American populations than did Eastern European populations, as indicated by the number of alleles private to the combination of French and North American populations, patterns of genetic distance among populations and assignment tests. The higher F IS values in Eastern Europe further support their distinctness. The plant is self incompatible and outcrossing in its native area (Friedman and Barrett 2008) and we have no indication of a breakdown of self incompatibility and shift in reproductive mode in Eastern Europe. Furthermore we found a higher proportion of null genotypes in these populations, so the significant F IS values point strongly to the existence of null alleles caused by mutations in the flanking regions of the microsatellites that are expected to evolve more slowly than repeat number, implying a longer history of independent evolution. Because the microsatellite markers were cloned from French populations (Genton et al. 2005b), genetically distant populations should harbour more null alleles. The fact that more null alleles seem to be present in Eastern Europe than in North America and France thus further indicate that Eastern European populations are genetically distinct from both the sampled North American and French ones. We also note that a scenario in which Eastern European populations originate, at least in part, from different native populations than those analysed by Genton et al. (2005a) is likely given the geopolitics of the latter half of the twentieth century that facilitated neither commercial nor human exchanges between Eastern Europe and North America or Western Europe.
Conclusion
Introductions from multiple sources and separate introductions to different sites have previously been reported in the literature (see introduction), but the pattern found for A. artemisiifolia is rarer, with two separate introductions on the same continent, different levels of population structure in different parts of the invasive range, and introductions from different but multiple sources each (see another example with the invasive grass Bromus tectorum; Novak and Mack 2001). This results in a similar level of variability in introduced and native populations, possibly yielding a considerable potential for rapid evolution in invasive populations. Nonetheless, we have little evidence for post-introduction adaptive evolution in this species. Genton et al. (2005c) investigated changes of introduced populations compared to native ones, but found no evidence for any evolutionary loss of defence against natural enemies despite strong enemy release in Europe, though there may have been an evolutionary change in the phenology of introduced populations, reflecting adaptation to higher latitudes in the introduced range.
Another remarkable aspect of the invasion of A. artemisiifolia in Europe is the possible role of the political context in the genetic structure and diversity on the invaded range. The wars and their consequences may indeed have set the stage for the introduction of genetically different sources in Western and Eastern Europe. The locations of the populations that gave rise to the Eastern populations remain to be identified.
References
Amsellem L, Noyer JL, Le Bourgeois T, Hossaert-McKey M (2000) Comparison of genetic diversity of the invasive weed Rubus alceifolius Poir. (Rosaceae) in its native range and in areas of introduction, using amplified fragment length polymorphism (AFLP) markers. Mol Ecol 9:443–455
Bassett IJ, Crompton CW (1975) The biology of Canadian weeds. 11. Ambrosia artemisiifolia L. and A. psilostachya DC. Can J Plant Sci 55:463–476
Bossdorf O, Auge H, Lafuma L, Rogers W, Siemann E, Prati D (2005) Phenotypic and genetic differentiation between native and introduced plant populations. Oecologia 144:1–11
Cavalli-Sforza L, Edwards AWF (1967) Phylogenetic analysis: models and estimation procedures. Evolution 32:550–570
Chauvel B, Dessaint F, Cardinal-Legrand C, Bretagnolle F (2006) The historical spread of Ambrosia artemisiifolia L. in France from herbarium records. J Biogeogr 33:665–673
Chun YJ, Fumanal B, Laitung B, Bretagnolle F (2010) Gene flow and population admixture as the primary post-invasion processes in common ragweed (Ambrosia artemisiifolia) populations in France. New Phytol 185:1100–1107
Ciosi M, Miller NJ, Kim KS, Giordano R, Estoup A, Guillemaud T (2008) Invasion of Europe by the western corn rootworm, Diabrotica virgifera virgifera: multiple transatlantic introductions with various reductions of genetic diversity. Mol Ecol 17:3614–3627
Coulon A, Fitzpatrick JW, Bowman R, Stith BM, Makarewich CA, Stenzler LM, Lovette IJ (2008) Congruent population structure inferred from dispersal behaviour and intensive genetic surveys of the threatened Florida scrub-jay (Aphelocoma coerulescens). Mol Ecol 17:1685–1701
Csontos P, Vitalos M, Barina Z, Kiss L (2010) Early distribution and spread of Ambrosia artemisiifolia in Central and Eastern Europe. Bot Helv 120:75–78
Dlugosch KM, Parker IM (2008) Founding events in species invasion: genetic variation, adaptive evolution, and the role of multiple introductions. Mol Ecol 17:431–449
Durka W, Bossdorf O, Prati D, Auge H (2005) Molecular evidence for multiple introductions of garlic mustard (Alliaria petiolata, Brassicaceae) to North America. Mol Ecol 14:1697–1706
El Mousadik A, Petit R (1996) High level of genetic differentiation for allelic richness among populations of the argan tree (Argania spinosa (L.) Skeels) endemic to Morocco. Theor Appl Genet 92:832–839
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Facon B, Jarne P, Pointier J-P, David P (2005) Hybridization and invasiveness in the freshwater snail Melanoides tuberculata: hybrid vigour is more important than increase in genetic variance. J Evol Biol 18:524–535
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 164:1567–1587
Friedman J, Barrett SCH (2008) High outcrossing in the annual colonizing species Ambrosia artemisiifolia (Asteraceae). Ann Bot 101:1303–1309
Genton BJ, Shykoff JA, Giraud T (2005a) High genetic diversity in French invasive populations of common ragweed Ambrosia artemisiifolia as a consequence of multiple sources of introduction. Mol Ecol 14:4275–4285
Genton BJ, Jonot O, Thévenet D, Fournier E, Blatrix R, Vautrin D, Solignac M, Giraud T (2005b) Isolation of five polymorphic microsatellite loci, using an enrichment protocol, in the invasive weed Ambrosia artemisiifolia (Asteraceae). Mol Ecol Notes 5:381–383
Genton BJ, Kotanen PM, Cheptou P-O, Adolphe C, Shykoff JA (2005c) Enemy release but no evolutionary loss of defence in a plant invasion: an intercontinental reciprocal experiment. Oecologia 146:404–414
Goudet J (1995) Fstat (version 1.2): a computer program to calculate F-statistics. J Hered 86:485–486
Grimsby JL, Tsirelson D, Gammon MA, Kesseli R (2007) Genetic diversity and clonal vs. sexual reproduction in Fallopia spp. (Polygonaceae). Am J Bot 94:957–964
Guo SW, Thompson EA (1992) Performing the exact test of Hardy-Weinberg proportions for multiple alleles. Biometrics 48:361–372
Henry P, Le Lay G, Goudet J, Guisan A, Jahodová S, Besnard G (2009) Reduced genetic diversity, increased isolation and multiple introductions of invasive giant hogweed in the western Swiss Alps. Mol Ecol 18:2819–2831
Hubisz MJ, Falush D, Stephens M, Pritchard JK (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Resour 9:1322–1332
Hufbauer RA, Sforza R (2008) Multiple introductions of two invasive Centaurea taxa inferred from cpDNA haplotypes. Divers Distrib 14:252–261
Husband BC, Barrett SCH (1991) Colonization history and population genetic structure of Eichhornia paniculata in Jamaica. Heredity 66:287–296
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Kazinczi G, Béres I, Novák R, Bíró K, Pathy Z (2008) Common ragweed (Ambrosia artemisiifolia): A review with special regards to the results in Hungary. I. Taxonomy, origin and distribution, morphology, life cycle and reproduction strategy. Herbologia 9:55–91
Kiss L, Béres I (2007) Anthropogenic factors behind the recent population expansion of common ragweed (Ambrosia artemisiifolia L.) in Eastern Europe: is there a correlation with political transitions? J Biogeogr 33:2156–2157
Kolbe JJ, Glor RE, Schettino LRG, Lara AC, Larson A, Losos JB (2004) Genetic variation increases during biological invasion by a Cuban lizard. Nature 431:177–181
Lavergne S, Molofsky J (2007) Increased genetic variation and evolutionary potential drive the success of an invasive grass. Proc Natl Acad Sci USA 104:3883–3888
Le Roux JJ, Wieczorek AM, Ramadan MM, Tran CT (2006) Resolving the native provenance of invasive fireweed (Senecio madagascariensis Poir.) in the Hawaiian Islands as inferred from phylogenetic analysis. Divers Distrib 12:694–702
Le Roux JJ, Wieczorek AM, Meyer J-Y (2008) Genetic diversity and structure of the invasive tree Miconia calvescens in Pacific Islands. Divers Distrib 14:935–948
Makra L, Juhász M, Borsos E, Béczi R (2004) Meteorological variables connected with airborne ragweed pollen in Southern Hungary. Int J Biometeorol 49:37–47
Makra L, Juhász M, Béczi R, Borsos E (2005) The history and impacts of airborne Ambrosia (Asteraceae) pollen in Hungary. Grana 44:57–64
Marrs RA, Sforza R, Hufbauer RA (2008) When invasion increases population genetic structure: a study with Centaurea diffusa. Biol Invasions 10:561–572
Muirhead JR, Gray DK, Kelly DW, Ellis SM, Heath DD, Macisaac HJ (2008) Identifying the source of species invasions: sampling intensity vs. genetic diversity. Mol Ecol 17:1020–1035
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Novak SJ, Mack RN (2001) Tracing plant introduction and spread: genetic evidence from Bromus tectorum (Cheatgrass). Bioscience 51:114–122
Okada M, Lyle M, Jasieniuk M (2009) Inferring the introduction history of the invasive apomictic grass Cortaderia jubata using microsatellite markers. Divers Distrib 15:148–157
Peakall ROD, Smouse PE (2006) Genalex 6: genetic analysis in excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Puillandre N, Dupas S, Dangles O, Zeddam J-L, Capdevielle-Dulac C, Barbin K, Torres-Leguizamon M, Silvain J-F (2008) Genetic bottleneck in invasive species: the potato tuber moth adds to the list. Biol Invasions 10:319–333
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Rollins LA, Woolnough AP, Wilton AN, Sinclair R, Sherwin WB (2009) Invasive species can’t cover their tracks: using microsatellites to assist management of starling (Sturnus vulgaris) populations in Western Australia. Mol Ecol 18:1560–1573
Simberloff D (2009) The role of propagule pressure in biological invasions. Annu Rev Ecol Evol Syst 40:81–102
Slatkin M (1995) A measure of population subdivision based on microsatellite allele frequency. Genetics 139:457–462
Szpiech ZA, Jakobsson M, Rosenberg NA (2008) ADZE: a rarefaction approach for counting alleles private to combinations of populations. Bioinformatics 24:2498–2504
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Acknowledgments
We are grateful to Tünde Jankovics and Vera Hayova for their help in collecting ragweed samples, to Bernard Clot from MeteoSwiss, and we thank anonymous reviewers for their suggestions. TG acknowledges a grant ANR 07-BDIV-003 and LK an Invited Professorship from University Paris-Sud. A part of this work was supported by grants of the Hungarian Scientific Research Fund (OTKA T046841 and IN67377).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gladieux, P., Giraud, T., Kiss, L. et al. Distinct invasion sources of common ragweed (Ambrosia artemisiifolia) in Eastern and Western Europe. Biol Invasions 13, 933–944 (2011). https://doi.org/10.1007/s10530-010-9880-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-010-9880-y