Abstract
MicroRNAs play important roles in carcinogenesis by negatively regulating the expression of target genes. Here we explore the biological function of miR-155 and the underlying mechanism in colorectal carcinoma. We validate, for the first time, that E2F2 is a direct target of miR-155 using western blot and a luciferase reporter assay and that miR-155 regulates the proliferation and cell cycle of colorectal carcinoma cells by targeting E2F2 using siRNA technology. We also found, for the first, time that E2F2 acts as a tumor suppressor in colorectal carcinoma. Overall, miR-155 plays an important role in colorectal carcinoma tumorigenesis by negative regulation of its targets including E2F2 and may be a potential therapeutic target for colorectal carcinoma treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal carcinoma is one of the most common malignancies with high incidence in western countries. Undesired cell proliferation usually becomes one of the main reasons of the genesis and development of colorectal carcinoma.
MicroRNAs (miRNAs) are a class of small, non-coding RNAs about 19–22 nt. Mature miRNAs bind to the “seed-matches” in the 3′-untranslated region (UTR) of target mRNAs through incomplete base-pairing, resulting in inhibition of translation or degradation of mRNA (Lagos-Quintana et al. 2001; Meltzer 2005; Foshay 2007). Bioinformatics studies have estimated that miRNAs regulate more than 30 % of human genes (Carrington 2003; Bartel 2004). miRNAs are involved in many biological processes including proliferation, differentiation and migration. The host gene of microRNA-155 (miR-155) is part of the B cell integration cluster (bic) which is involved in the progression of lymphoma, and the relationship between miR-155 and lymphoma has been well established (Calame 2007; Vigorito et al. 2007; O’Connell et al. 2010). Aberrant expression of miR-155 has been found in various cancers including colorectal carcinoma (Volinia et al. 2006; Faraoni et al. 2009). However, the biological function and downstream targets of miR-155 in colorectal carcinoma are still unclear (Zhang et al. 2013).
miRNAs usually function by repressing target genes. E2F transcription factor 2 (E2F2) is expressed at very low level in colorectal carcinoma (Xanthoulis and Tiniakos 2013), where it is negatively related with the high expression level of miR-155 (Volinia et al. 2006). E2F2 is a member of the E2F family which plays significant roles in proliferation, apoptosis and differentiation (DeGregori 2002). E2Fs usually bind to Rb protein, and the Rb/E2F complexes regulate the expression of genes involved in the progress of the cell cycle, DNA synthesis, checkpoint control and apoptosis (Dimova and Dyson 2005; McClellan et al. 2007). E2F2 belongs to the “activator” E2Fs which activate downstream targets and promotes cell growth. However, “activator” E2Fs can also act as a “suppressor” depending on the context (Johnson and DeGregori 2006). E2F2 represses cell cycle regulators to maintain quiescence (Infante et al. 2008) and represses Myc-induced lymphomagenesis (Opavsky et al. 2007; Pusapati et al. 2010) but little is known about its role in colorectal carcinoma.
Here, functions of miR-155 on colorectal carcinoma proliferation and cell cycle have been explored with new potential targets of miR-155 being identified. The roles of E2F2 in proliferation and cell cycle of colorectal carcinoma cells were also investigated. These findings provided evidence for hypothesis that miR-155 regulated the proliferation and cell cycle of colorectal carcinoma cells by targeting E2F2.
Materials and methods
Cell culture
Human colorectal carcinoma cell line, SW-620, and human embryonic kidney cells line, HEK-293T, were grown in Dulbecco’s modified Eagle’s medium (DMEM) with 10 % (v/v) fetal bovine serum (FBS) at 37 °C in a humidified atmosphere with 5 % (v/v) CO2. Human colorectal carcinoma cell line, HCT-116, was cultured in RPMI 1640 media with 10 % (v/v) FBS at 37 °C in a humidified atmosphere with 5 % (v/v) CO2.
Transfection
Cells were harvested and seeded in a six-well plate at 105/well. Four microgram DNA or 100 pmol RNA were transfected into the cells using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s protocol.
Cell viability assay
Cell viability assay was performed to test the effects of miR-155 and E2F2 on the proliferation of colorectal carcinoma cells. After treatment of transfection, 6,000 cells of each sample were plated in a 96-well plate in triplicate, cells was tested for proliferation every 24 h using the MTT and the absorbance at 490 nm was measured by microplate reader.
Total RNA extraction and quantitative real time PCR (qRT-PCR)
Total RNA was extracted from each sample after transfection using Trizol. The miR-155 expression level was detected by SYBR Green qRT-PCR method with a Hsa-miR-155 Detection Kit and EzOmics SYBR qPCR Kit (Biomics, Jiangsu, China) using a StepOne real-time PCR system (Applied Biosystems, Foster City, CA), miR-155 expression level was normalized to U6 snRNA and the miR-155 relative expression level of each group was calculated using the 2−∆∆Ct method.
Luciferase reporter assay
Luciferase reporter assay was used to test whether the miR-155 bound directly to the 3′-UTR of E2F2 mRNA. Fragments of E2F2 mRNA containing wild type (WT) or mutant (MUT) miR-155 binding site were cloned respectively into the pMIR-REPORT luciferase reporter vector (Ambion, San Diego, CA). For luciferase reporter assays, HEK-293T cells, that had high efficiency of co-transfection, were seeded in a 24-well plate (2 × 104cells per well) and 0.2 μg of either the constructed luciferase reporter plasmid WT or MUT, together with 0.2 μg β-galactosidase (β-gal) control plasmid (Ambion) plus either 20 pmol miR-155 mimics (155 M) or its corresponding negative control (MNC) were co-transfected into HEK-293T cells using Lipofectamine 2000 48 h after transfection. Luciferase activity and β-gal activity were measured using a luciferase assay system and β-gal enzyme assay system (Promega), respectively, according to the manufacturers’ protocols. The luciferase activity was normalized to β-gal.
Western blotting
Cells were harvested 48 h after transfection and lysed in RIPA lysis buffer on ice. Protein concentration was measured using BCA method. 40 μg protein of each sample was subjected to 12 % (w/v) SDS-PAGE. The separated proteins were transferred to a PVDF membrane (Millipore). Membranes were blocked and then incubated with primary antibodies against E2F2 (Epitomics, Burlingame, CA), survivin (Bioworld, Atlanta, GA, USA) or β-actin (Bioworld) followed by incubation with horseradish peroxidase-conjugated secondary antibodies. The protein expression level was normalized to β-actin.
Cell cycle analysis
Cells were harvested 48 h after transfection, resuspended in 1 ml ice-cold 70 % (v/v) ethanol and stored overnight at 4 °C. Then cells were stained by a cell cycle detection kit (Beyotime, Jiangsu, China) according to the manufacturer’s protocols. After incubation in dark at 37 °C for 30 min, cells were analyzed on a FACScalibur machine (Becton Dickinson) with CellQuest software. The percentage of cells in each phase of the cell cycle was analyzed by ModFit LT 4.0 software.
EdU incorporation assays
EdU incorporation assays was used to determine the cells in S phase with a Cell-light EdU Apollo 567 In Vitro Kit (Ribobio, Guangzhou, China) according to the manufacturer’s protocol. Forty eight hours after transfection, 50 mM EdU was added to each well, and the cells were cultured for additional 2 h at 37 °C. Cells were then fixed with 4 % (v/v) paraformaldehyde for 30 min and permeabilized with 0.5 % Triton X-100 for 10 min at room temperature. After washing with PBS, 100 μl Apollo staining solution was added to each well, and the cells were incubated for 30 min at room temperature. The cells were subsequently stained with 100 μl Hoechst 33,342 for 30 min and visualized under an laser scanning confocal microscope.
Statistics
All experiments were performed independently at least three times. The data was presented as mean ± SD. Student’s t test was performed for comparisons between two groups. P < 0.05 was considered to be significant.
Results
miR-155 promotes the proliferation and cell cycle progress of colorectal carcinoma cells
To investigate the effect of miR-155 on proliferation of colorectal carcinoma cells, we performed a cell viability assay. miR-155 mimic (155 M) was used to increase the expression level of miR-155 and miR-155 inhibitor (155I) was used to down-regulate miR-155 expression level. 155M or its corresponding negative control (MNC), 155I or its corresponding negative control (INC) was transfected into HCT-116 and SW-620, then the expression level of miR-155 was determined by qRT-PCR. After transfection with 155 M, miR-155 expression increased more than 150-fold and 350-fold, respectively, in HCT-116 and SW-620 compared with those transfected with MNC (Fig. 1a). After transfection with 155I, the miR-155 expression level decreased to 48 and 33 % respectively in HCT-116 and SW-620 compared with those transfected with INC (Fig. 1e). When transfected with 155 M, the proliferation of cells was significantly increased compared with cells transfected with MNC in both HCT-116 and SW-620 (Fig. 1b) whereas, after transfection with 155I, the proliferation of HCT-116 and SW-620 was inhibited significantly (Fig. 1f). These results suggested that miR-155 promoted the proliferation of colorectal carcinoma cells.
miR-155 promotes the proliferation and cell cycle of colorectal carcinoma cells. a, e qRT-PCR detected the expression level of miR-155 in HCT-116 and SW-620 cells transfected with miR-155 mimics (155 M) or miR-155 inhibitor (155I) compared with those transfected with its corresponding negative control (MNC or INC). The result are presented as fold changes of expression level. b Over-expression of miR-155 promoted the proliferation of HCT-116 and SW-620. c Over-expression of miR-155 induced cell cycle arrested at the S phase in both HCT-116 and SW-620. d Over-expression of miR-155 increased numbers of EdU-positive cells in HCT-116 and SW-620. f Down-regulation of miR-155 inhibits the proliferation of HCT-116 and SW-620. g Down-regulation of miR-155 induces cell cycle arrested at the G1 phase in both HCT-116 and SW-620. h Down-regulation of miR-155 decreases EdU-positive cells in HCT-116 and SW-620 (*P < 0.05, **P < 0.01)
To study whether miR-155 promoted the proliferation of colorectal carcinoma cells by affecting their cell cycle, cell cycle analysis and EdU incorporation assays were performed. Cell cycle analysis showed that after transfection with 155 M, cell cycle was arrested at the S phase compared with cells transfected with MNC in both HCT-116 and SW-620 (Fig. 1c). EdU incorporation assays showed that after transfection with 155 M, EdU-positive cells (red) increased to 1.75 and 1.77-fold, respectively, in HCT-116 and SW-620 compared with those transfected with MNC (Fig. 1d). When transfection was carried out with 155I, the cell cycle was arrested at G1 phase (Fig. 1g) and EdU-positive cells decreased to 49 and 54 % (Fig. 1h) compared with cells transfected with INC in both HCT-116 and SW-620. These results suggest that miR-155 promoted the progress of the cell cycle. Cell viability assay, cell cycle assay and EdU incorporation assays demonstrated that miR-155 promoted the proliferation and cell cycle progress of colorectal carcinoma cells.
E2F2 is a direct target of miR-155
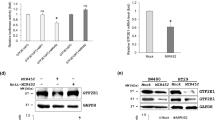
To explore the mechanism how miR-155 promoted proliferation and cell cycle process, we hypothesized that miR-155 regulated proliferation and cell cycle of colorectal carcinoma cells via its potential target E2F2 which was predicted by online software TargetScan (http://www.targetscan.org/) and miRDB (http://mirdb.org/miRDB/). Western blotting was performed to investigate the relationship between the expression level of miR-155 and E2F2. After transfection with 155 M, the E2F2 protein level decreased significantly to 49 and 18 % in HCT-116 and SW-620, respectively, compared with those transfected with MNC. After transfection with 155I, the E2F2 protein level, however, increased significantly to 2.3-fold and 3.1-fold in HCT-116 and SW-620, respectively, when compared with those transfected with INC (Fig. 2a–d). These results suggested that protein level of E2F2 can be negatively regulated by miR-155.
E2F2 is a direct target of miR-155. a–d Western blot analysis of HCT-116 and SW-620 for E2F2 protein expression after transfection, expression level of E2F2 was normalized to β-actin. e Sequence of wild type luciferase reporter plasmid wide type (WT) and its mutant type (MUT). f Luciferase reporter assay after transfection. Luciferase activities were normalized to β-gel (*P < 0.05, **P < 0.01)
To investigate whether E2F2 is a direct target of miR-155, a luciferase reporter assay was performed. According to the predicted miR-155 binding site on the 3′-UTR of E2F2 mRNA, a WT and MUT luciferase reporter plasmid were constructed (Fig. 2e). The WT or MUT luciferase reporter plasmid was co-transfected with either 155 M or MNC plus β-gal control plasmid. The relative luciferase activity of cells transfected with WT luciferase reporter plasmid plus 155 M decreased significantly to 54 % compared with that of cells transfected with WT luciferase reporter plasmid plus MNC (Fig. 2f). However, there was no significant difference between the relative luciferase activity of the cells cotransfected with MUT luciferase reporter plasmid plus either 155 M or MNC. These results suggest that miR-155 directly binds to 3′-UTR of E2F2 mRNA. Data from western blotting and the luciferase reporter assay demonstrated that E2F2 was a direct target of miR-155.
E2F2 acts as a tumor suppressor in colorectal carcinoma
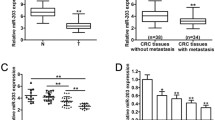
E2F2 plays an important role in the progress of the cell cycle. To investigate its role in colorectal carcinoma, we used a specific siRNA to knockdown E2F2. Western blotting was used to detect the protein expression level of E2F2 after transfection with the E2F2-siRNA. Protein expression level of E2F2 decreased to 23 and 28 %, respectively, in HCT-116 and SW-620 when transfected with E2F2-siRNA compared with those transfected with its corresponding negative control (SINC) (Fig. 3a). After transfection with E2F2-siRNA, cell proliferation of HCT-116 and SW-620 had increased markedly (Fig. 3b), the cell cycle was arrested at S phase (Fig. 3c) and EdU-positive cells increased to 1.35-fold and 1.63-fold (Fig. 3d), respectively, in HCT-116 and SW-620 compared with those transfected with SINC. These results demonstrate that down-regulation of E2F2 promotes the proliferation and cell cycle of colorectal carcinoma cells and that E2F2 acted as a tumor suppressor in colorectal carcinoma.
E2F2 acts as a tumor suppressor in colorectal carcinoma. a Western blot analysis of HCT-116 and SW-620 for E2F2 protein expression after transfection with E2F2 specific siRNA, expression level of E2F2 was normalized to β-actin. b Cell proliferation assay was carried out after transfection. Knockdown of E2F2 promoted the proliferation of HCT-116 and SW-620. c Knockdown of E2F2 in HCT-116 and SW-620 induced cell cycle arrested at S phase. d Knockdown of E2F2 in HCT-116 and SW-620 increased EdU-positive cells. e Western blot analysis of HCT-116 and SW-620 for survivin protein expression after transfection, expression level of survivin was normalized to β-actin (*P < 0.05, **P < 0.01)
To validate that E2F2 acts as a tumor suppressor in colorectal carcinoma cells, we detected the protein expression level of survivin, which is important for tumor survival. After transfection with E2F2-siRNA, western blotting was used to detect protein expression level of survivin. When E2F2 was knock down with specific E2F2-siRNA, the expression level of survivin increased to 2.9-fold and 3.5-fold, respectively, in HCT-116 and SW-620 comparing to those transfected with SINC (Fig. 3e). These results demonstrate that E2F2 represses the expression of survivin in colorectal carcinoma.
miR-155 regulates the proliferation and cell cycle of colorectal carcinoma cells by targeting E2F2
To investigate whether miR-155 affects the proliferation and cell cycle of colorectal carcinoma cells by repressing E2F2, a E2F2 specific siRNA was used to knockdown E2F2 when cells were transfected with 155 M or MNC. After transfection, the proliferation and cell cycle of HCT-116 and SW-620 were tested. 155 M plus SINC increased proliferation (Fig. 4a–d), induce cell cycle arrest at S phase (Fig. 4e, f) and increased EdU-positive cells (Fig. 4g, h) in both HCT-116 and SW-620 compared to cells transfected with MNC plus SINC. But 155 M plus siRNA failed to promote proliferation and cell cycle process and failed to increase EdU-positive cells in both HCT-116 and SW-620 compared to cells transfected with MNC plus SINC. This means that siRNA reversed the effect of 155 M on proliferation and the cell cycle. These results demonstrate that miR-155 regulates the proliferation and cell cycle of colorectal carcinoma cells by targeting E2F2.
miR-155 regulates the proliferation and cell cycle of colorectal carcinoma by targeting E2F2. a, c Knockdown of E2F2 in HCT-116 and SW-620 cells partially reverse the effect of miR-155 on proliferation. b, d The cell viability of HCT-116 and SW-620 at 72 h after cotransfection with siRNA and 155 M. e, f Knockdown of E2F2 in HCT-116 and SW-620 cells partially reverse the effect of miR-155 on cell cycle. g, h Knockdown of E2F2 partially reverse the effect of miR-155 on ratio of EdU-positive cells in HCT-116 and SW-620 cells (*P < 0.05, **P < 0.01, NS no significant difference)
Discussion
Changes in miRNA expression level frequently occur in cancer cells and these changes usually relate closely to the genesis and development of cancer. In many types of cancer, miR-155 promotes proliferation (Mattiske et al. 2012; Jiang et al. 2010; Sun et al. 2012; Xie et al. 2012; Li et al. 2012) whereas, in some types of cancer, miR-155 represses proliferation (Dai et al. 2012). miR-155 can act as either as an oncomiR or a tumor suppressor. Here we found that miR-155 acted as an oncomiR promoting the proliferation and cell cycle progress of colorectal carcinoma which had high miR-155 expression level. The high level of miR-155 may be one cause of colorectal carcinoma development. Consistently, miR-155 is associated with proliferation, metastasis and invasion of breast cancer (Mattiske et al. 2012; Jiang et al. 2010) which also has a high level of miR-155 which makes it a potential biomarker of breast cancer (Sun et al. 2012). Nanoparticle-based anti-miR-155 therapy has excellent therapeutic effects in miR-155-dependent mouse model of lymphoma (Babar et al. 2012). miR-155 is often correlated with poor prognosis and may act as a key target of colorectal carcinoma therapy.
To explore the mechanism underlying the function of miR-155, online software was used to predicted novel targets of miR-155. When we explored whether E2F2 was a direct target of miR-155, we used pMIR-reporter system which was a system to analyze miRNA function on predicted miRNA binding sites. However, only a DNA fragment at about 50–60 mer can be inserted into its multiclone site. Data of luciferase reporter assay showed that miR-155 could bind to the WT target sequence but not the MUT target sequence. To a certain extent, it also demonstrated that E2F2 was a direct target of miR-155 assisted by western blotting. However, there are still some limitations: the secondary structure of the entire 3′-UTR might also affect the binding of miR-155 and further studies needed.
siRNA technology was used to explore whether miR-155 regulated the proliferation and cell cycle of colorectal carcinoma cells by targeting E2F2. E2F2-siRNA can only partly reverse the function of miR-155, which suggested that E2F2 was not the only one target of miR-155 in colorectal carcinoma and thus there may be some other targets of miR-155.
E2F2 is a member of the E2F family which plays important roles in the regulation of cell cycle by repressing or activating cell cycle regulator including cyclin, cyclin dependent kinases (CDKs), checkpoint regulators (Timmers et al. 2006). E2F family members can be grouped into “activators” and “suppressors”. The so-called “activators” include E2F1, E2F2, E2F3. They have the same promotor binding element and they can promote cell proliferation (DeGregori 2002; Timmers et al. 2006; Wenzel et al. 2011; Rogoff and Kowalik 2004). However, when we explored the roles of E2F2 on the proliferation and cell cycle of colorectal carcinoma cells, we found that E2F2 suppressed the proliferation and cell cycle process of colorectal carcinoma cells. E2F “activators” also have tumor suppressive properties (Johnson and DeGregori 2006; Infante et al. 2008; Opavsky et al. 2007; Pusapati et al. 2010; Wong et al. 2003; Lee 2000). For example, E2F1, one of the classic “activator” E2Fs, is well studied. Ectopic expression of E2F1 promotes cell proliferation and tumorigenesis (Infante et al. 2008). However, over-expression of E2F1 also suppresses Ras-driven tumorigenesis and colony formation (Lee 2000). Johnson and DeGregori (2006) raised a hypothesis that E2F1 may function as a tumor suppressor when it is absent and as an oncogene when it is overexpressed. This hypothesis may be applied to E2F2 which has a very low expression level in colorectal carcinoma cells. There are a few studies about E2F2 “suppressor” capability: in mice, E2F2 repressed Myc-induced proliferation and tumorigenesis (Opavsky et al. 2007; Pusapati et al. 2010). In cultured cells, E2F2 represses cell cycle regulators to maintain quiescence (Infante et al. 2008). Nevertheless, little is known about roles of E2F2 in cancer cells, and here we validated for the first time that E2F2 acts as a tumor suppressor in colorectal carcinoma.
Data of western blotting demonstrated that E2F2 repressed the expression of survivin. Survivin is a multifunctional protein that controls cell proliferation, cell cycle and apoptosis. Survivin is expressed in most human cancers but is absent in normal and differentiated tissues. It has a close relationship with tumor survival (Jha et al. 2012). Repression of survivin by E2F2 is one evidence for the hypothesis that E2F2 acts as a tumor suppressor in colorectal carcinoma. However, the underlying mechanisms of how E2F2 acts as a tumor suppressor in colorectal carcinoma are still unknown. More exploration needs to be done.
In summary, miR-155, as an oncomiR, regulates proliferation and the cell cycle of colorectal carcinoma cells by targeting E2F2. E2F2 acts as a tumor suppressor in colorectal carcinoma. miR-155 plays an important role in colorectal carcinoma tumorigenesis and could be a potential therapeutic target for colorectal carcinoma treatment.
References
Babar IA, Cheng CJ, Booth CJ et al (2012) Nanoparticle-based therapy in an in vivo microRNA-155 (miR-155) dependent mouse model of lymphoma. Proc Natl Acad Sci USA 109:E1695–E1704
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Calame K (2007) microRNA-155 function in B cells. Immunity 27:825–827
Carrington JC, Ambros V (2003) Role of microRNAs in plant and animal development. Science 301:336–338
Dai Y, Qiu Z, Diao Z et al (2012) MicroRNA-155 inhibits proliferation and migration of human extravillous trophoblast derived HTR-8/SVneo cells via down-regulating cyclin D1. Placenta 33:824–829
DeGregori J (2002) The genetics of the E2F family of transcription factors shared functions and unique roles. Biochim Biophys Acta 1602:131–150
Dimova DK, Dyson NJ (2005) The E2F transcriptional network: old acquaintances with new faces. Oncogene 24:2810–2826
Faraoni I, Antonetti FR, Cardone J et al (2009) miR-155 gene: a typical multifunctional microRNA. Biochim Biophys Acta 1792:497–505
Foshay KM (2007) Small RNAs, big potential: the role of MicroRNAs in stem cell function. Curr Stem Cell Res Ther 2:264–271
Infante A, Laresgoiti U, Fernández-Rueda J et al (2008) E2F2 represses cell cycle regulators to maintain quiescence. Cell Cycle 7:3915–3927
Jha K, Shukla M, Pandey M (2012) Survivin expression and targeting in breast cancer. Surg Oncol 21:125–131
Jiang S, Zhang HW, Lu MH et al (2010) MicroRNA-155 functions as an oncomiR in breast cancer by targeting the suppressor of cytokine signaling 1 gene. Cancer Res 70:3119–3127
Johnson DG, DeGregori J (2006) Putting the oncogenic and tumor suppressive activities of E2F into context. Curr Mol Med 6:731–738
Lagos-Quintana M, Rauhut R, Lendeckel W et al (2001) Identification of novel genes coding for small expressed RNAs. Science 294:853–858
Lee TAFP (2000) Exogenous E2F expression is growth inhibitory before, during, and after cellular transformation. Oncogene 19:2257–2268
Li S, Chen T, Zhong Z et al (2012) microRNA-155 silencing inhibits proliferation and migration and induces apoptosis by upregulating BACH1 in renal cancer cells. Mol Med Rep 5:949–954
Mattiske S, Suetani RJ, Neilsen PM et al (2012) The oncogenic role of miR-155 in breast cancer. Cancer Epidemiol Biomarkers Prev 21:1236–1243
McClellan KA, Ruzhynsky VA, Douda DN et al (2007) Unique requirement for Rb/E2F3 in neuronal migration evidence for cell cycle-independent functions. Mol Cell Biol 27:4823–4825
Meltzer PS (2005) Cancer genomics: small RNAs with big impacts. Nature 435:745–746
O’Connell RM, Kahn D, Gibson WSJ et al (2010) MicroRNA-155 promotes autoimmune inflammation by enhancing inflammatory T cell development. Immunity 33:607–619
Opavsky R, Tsai S-Y, Guimond M et al (2007) Specific tumor suppressor function for E2F2 in Myc-induced T cell lymphomagenesis. Proc Natl Acad Sci USA 104:15400–15405
Pusapati RV, Weaks RL, Rounbehler RJ et al (2010) E2F2 suppresses Myc-induced proliferation and tumorigenesis. Mol Carcinog 49:152–156
Rogoff HA, Kowalik TF (2004) Life, death and E2F. Cell Cycle 3:845–846
Sun Y, Wang M, Lin G et al (2012) Serum microRNA-155 as a potential biomarker to track disease in breast cancer. PLoS ONE 7:e47003
Timmers C, Sharma N, Opavsky R et al (2006) E2f1, E2f2, and E2f3 control E2F target expression and cellular proliferation via a p53-dependent negative feedback loop. Mol Cell Biol 27:65–78
Vigorito E, Perks KL, Abreu-Goodger C et al (2007) MicroRNA-155 regulates the generation of immunoglobulin class-switched plasma cells. Immunity 27:847–859
Volinia S, Calin GA, Liu CG et al (2006) A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA 103:2257–2261
Wenzel PL, Chong J-L, Sáenz-Robles MT et al (2011) Cell proliferation in the absence of E2F1-3. Dev Biol 351:35–45
Wong CF, Barnes LM, Dahle AL et al (2003) E2F modulates keratinocyte squamous differentiation. J Biol Chem 278:28516–28522
Xanthoulis A, Tiniakos DG (2013) E2F transcription factors and digestive system malignancies: how much do we know. World J Gastroenterol 19:3189–3198
Xie Q, Chen X, Lu F et al (2012) Aberrant expression of microRNA 155 may accelerate cell proliferation by targeting sex-determining region Y box 6 in hepatocellular carcinoma. Cancer 118:2431–2442
Zhang G, Xiao G, Tian H et al (2013) Upregulation of microRNA-155 promotes the migration and invasion of colorectal cancer cells through the regulation of claudin-1 expression. Int J Mol Med 31:1375–1380
Acknowledgments
This study was supported by grants from National Natural Science Foundation of China (No. 81372331). We thank Drs. Yihua Ma, Meijuan Xie and Yan Wang for helpful comments on the manuscript and all members of the Tao Xi laboratory for helpful discussion.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, T., Yang, J., Lv, X. et al. miR-155 regulates the proliferation and cell cycle of colorectal carcinoma cells by targeting E2F2. Biotechnol Lett 36, 1743–1752 (2014). https://doi.org/10.1007/s10529-014-1540-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-014-1540-3