Abstract
The toxicity of n-butanol in microbial fermentations limits its formation. The stress response of Clostridium acetobutylicum involves various stress proteins and therefore, over-expression of genes encoding stress proteins constitutes an option to improve solvent tolerance. Over-expression of groESL, grpE and htpG, significantly improved butanol tolerance of C. acetobutylicum. Whereas the wild type and vector control strain did not survive 2 % (v/v) butanol for 2 h, the recombinant strains showed 45 % (groESL), 25 % (grpE) and 56 % (htpG), respectively, of the initial c.f.u. after 2 h of butanol exposure. As previously, over-expression of groESL led to higher butanol production rates, but the novel strains over-expressing grpE or htpG produced only 51 and 68 %, respectively, of the wild type butanol concentrations after 72 h clearly differentiating butanol tolerance and production. Not only butanol tolerance but also the adaptation to butanol in successive stress experiments was significantly facilitated by increased levels of GroESL, GrpE and HtpG. Re-transformation and sequence analyses of the plasmids confirmed that not the plasmids, but the host cells evolved to a more robust phenotype.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Clostridial acetone/butanol/ethanol (ABE) fermentation has regained much interest in the past years because butanol represents a superior biofuel with several advantages over ethanol: butanol has a higher energy content, is less hydroscopic, less volatile and less corrosive, which makes it perfectly compatible with the current gasoline infrastructure (Lee et al. 2008; Green 2011). Since the maximum butanol tolerance is 20 g l−1 or less for clostridial strains, product removal by different techniques, such as gas stripping or pervoration, have been considered to economically improve ABE fermentation (Ezeji et al. 2005, 2007). The major cellular impact of high butanol concentrations is the destruction of the cytoplasmic membrane and various macromolecules, accompanied by the loss of vital functions (Bowles and Ellefson 1985; Taylor et al. 2008; Liu and Qureshi 2010). On the molecular level, butanol causes a stress response which is not yet fully understood; the most important issues and experimental approaches have been reviewed in detail recently (Ezeji et al. 2010; Nicolaou et al. 2010).
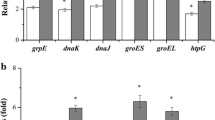
Referring to the development of suitable genomic tools, clostridial metabolite and solvent stresses have been studied in a global approach using DNA microarrays (Alsaker et al. 2004, 2010; Tomas et al. 2004; Alsa; Heluane et al. 2011; Hönicke et al. 2012) and proteome analyses (Terracciano et al. 1988; Mao et al. 2010, 2011). The results clearly showed that different heat shock proteins (HSPs) are involved in Clostridium acetobutylicum’s stress response to butanol. These molecular chaperones play important roles in protein biosynthesis and degradation and are highly upregulated under various stress conditions (Narberhaus and Bahl 1992; Narberhaus et al. 1992; Bahl et al. 1995; Rüngeling et al. 1999). Transcriptome data revealed that dnaK, dnaJ, grpE, groES, groEL, clpP, clpC, htpG and hsp18 were significantly induced after butanol stress (Tomas et al. 2004). Interestingly, the same HSPs were generally upregulated in C. acetobutylicum Rh8, a mutant strain with improved butanol tolerance and production (Mao et al. 2010). An increased abundance of HSPs was not only detected after external butanol exposure, but also during the life cycle-associated solventogenic metabolism (Alsaker and Papoutsakis 2005; Jones et al. 2008). Taking high ATP consumption rates into account, the protective function of multiple HSPs against solvent stress is primarily based on fast refolding and/or degradation of denaturated proteins, although the regulatory circuits are not well understood (Nicolaou et al. 2010). The second known defense mechanism of C. acetobutylicum is a higher content of saturated fatty acids in the cell membrane to counteract the increasing fluidity caused by butanol (Vollherbst-Schneck et al. 1984; Baer et al. 1987, 1989; Taylor et al. 2008; Ezeji et al. 2010).
Regarding microbial solvent tolerance, very few bacteria exhibit a similar or better performance than species of Clostridium, e.g., Pseudomonas putida or lactic acid bacteria (Fischer et al. 2008; Knoshaug and Zhang 2009; Rühl et al. 2009; Li et al. 2010; Winkler et al. 2010). Therefore, it seems more reasonable to engineer solventogenic clostridia with naturally high butanol production capacities, although a major metabolic breakthrough for recombinant butanol production was described recently (Bond-Watts et al. 2011; Shen et al. 2011).
Apart from traditional random mutagenesis, only two examples of targeted genetic engineering to increase butanol tolerance of C. acetobutylicum and other solventogenic clostridia have been published thus far. Based on early DNA microarray results which indicated the importance of stress proteins, the bicistronic groESL operon, coding for the class I (Hsp60) chaperonin GroEL and its co-chaperonin GroES, was homologously overexpressed. The recombinant C. acetobutylicum strain exhibited a significantly improved tolerance and also higher butanol production rates as compared to the wild type (Tomas et al. 2003). More recently, the gshAB genes from Escherichia coli were expressed in C. acetobutylicum to establish a heterologous glutathione biosynthetic pathway. The resulting strain exhibited an improved butanol tolerance and a slightly increased butanol titer as compared to the vector control (Zhu et al. 2011). In this study, two novel over-expression strains of C. acetobutylicum with enhanced butanol tolerance properties were generated. Interestingly, we observed that the HSP gene overexpressing strains revealed an enhanced adaptation to high butanol concentrations.
Materials and methods
Bacterial strains and cultivation conditions
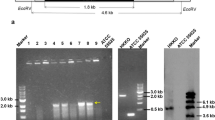
Strains, plasmids and oligonucleotides for PCR used in this study are listed in Supplementary Table 1. General recombinant DNA techniques were performed according to standard. Chromosomal DNA of C. acetobutylicum ATCC 824 (Fischer et al. 2006) was used as template and the oligonucleotides listed in Table 1 as PCR primers to amplify groESL (CAC2703-2704), grpE (CAC1281) and htpG (CAC3315). The DNA fragments were cloned via BamHI or BclI, respectively, and KasI restriction sites into the pTHydA plasmid as described by Girbal et al. (2005) and Hillmann et al. (2009). To generate the vector control pT, the hydA gene was excised from pTHydA by BamHI/KasI restriction and re-ligated after treatment with the Klenow fragment of DNA polymerase I (Fermentas GmbH, Germany). The resulting plasmids pT, pT::groESL, pT::grpE and pT::htpG, respectively, were transformed into E. coli DH5α and validated by DNA sequencing (LGC Genomics GmbH, Berlin, Germany). After in vivo methylation in E.coli ER2275 pAN2 (Mermelstein and Papoutsakis 1993), the plasmids were transformed into C. acetobutylicum ATCC 824 and selected on reinforced clostridial agar containing 40 μg erythromycin ml−1.
Clostridial strains were cultivated in clostridial growth medium (CGM) or mineral salts medium (MS-MES) with 40 μg erythromycin ml−1 for recombinant C. acetobutylicum under anaerobic conditions at 37 °C; media, conditions and fermentation product analyses were performed as described previously (Lehmann and Lütke-Eversloh 2011; Lehmann et al. 2012).
Butanol stress experiments
For butanol stress and adaptation experiments, 200 ml CGM in serum bottles were inoculated with 10 % (v/v) CGM precultures. When the cultures were grown to OD600 of 1, they were divided into 10 ml aliquots in Hungate tubes and subjected to 1 or 2 % (v/v) n-butanol. Respective controls were treated similarly but without butanol addition. To determine the c.f.u. values, samples were drawn regularly and serial dilutions were plated on RCA and incubated anaerobically for 48 h at 37 °C prior to colony counting.
Plasmid maintenance was controlled by colony PCR using plasmid- and gene-specific oligonucleotides (Supplementary Table 1). In addition, plasmids from three successive stress experiments were isolated and re-transformed into C. acetobutylicum ATCC 824 (Harris et al. 2000); furthermore, DNA sequence analyses of each plasmid was conducted (LGC Genomics GmbH, Berlin, Germany).
Results and discussion
Butanol tolerance
Homologous over-expression of the groESL operon in C. acetobutylicum leads to a significantly improved phenotype regarding butanol resistance and production (Tomas et al. 2003). In this study, two additional HSP encoding genes, i. e. grpE and htpG, were overexpressed in C. acetobutylicum and as a positive control, groESL over-expression was included. As shown in Fig. 1, all three over-expression strains exhibited enhanced butanol tolerance in comparison to the wild type and the vector control strain in the presence of 1 and 2 % (v/v) butanol. Tomas et al. (2003) reported that the growth inhibition by butanol of the groESL overexpressing strain was up to 85 % less than the vector control. This value is not directly comparable to the results presented here, because higher butanol concentrations and not the growth rates of challenged cultures, but survival rates according to c.f.u. (c.f.u.) counts after the butanol exposure were preferentially chosen as parameters. However, the highly improved butanol tolerance of C. acetobutylicum pT::groESL is in good agreement. Regarding the novel strains C. acetobutylicum pT::grpE and C. acetobutylicum pT::htpG, significantly increased butanol resistance can also be mediated by values of the stress proteins GrpE and HtpG. Whereas the wild type and the vector control had c.f.u. < 0.1 % after 6 h exposure to 1 % (v/v) butanol, referring to the initial c.f.u., the over-expressing strains still showed 12 % (groESL), 16 % (grpE) and 15 % (htpG), respectively, of the initial c.f.u.. In addition, the control strains did not survive 2 % (v/v) butanol for more than 2 h, but the three recombinant strains revealed 45 % (groESL), 25 % (grpE) and 56 % (htpG), respectively, of the initial c.f.u. after 2 h at 2 % (v/v) butanol (Fig. 1).
Increased butanol tolerance of C. acetobutylicum pT::groESL (c), pT::grpE (d) and pT::htpG (e) in comparison to the wild type (a) and vector control (b). The respective strains were subjected to 1 % (v/v) (diamonds) and 2 % (v/v) (circles) n-butanol for 8 h. Samples were drawn every 2 h and serial dilutions were plated onto RCA to determine the c.f.u. with respect to the controls without butanol (squares). The relative c.f.u. values are shown in percent of the initial sample (a–e). Panel (f) depicts the total c.f.u. counts of the 6 h samples of 1 % (v/v) (grey) and 2 % (v/v) (black) butanol: 1 C. acetobutylicum wild type, 2 vector control, 3 pT::groESL, 4 pT::grpE, 5 pT::htpG
Butanol production
Since genetic tools for clostridia became available only recently, traditional production strain-breeding was common (and quite successful) for a long time, using random mutagenesis and selection by survival of high butanol concentrations (Jones and Woods 1986; Papoutsakis 2008; Lütke-Eversloh and Bahl 2011). The phenotype of such production strains implements usually both enhanced butanol tolerance and production, justifying the relation between both features. This connection was confirmed by the groESL overexpressing C. acetobutylicum, which was reported to produce 40 % more butanol than the wild type (Tomas et al. 2003; Alsaker et al. 2010). Using mineral salts medium instead of CGM, C. acetobutylicum pT::groESL produced 130 % of the wild type butanol titer, validating the fact that higher GroES and GroEL protein levels improves solvent production.
Surprisingly, grpE and htpG over-expression did not enhanced butanol production, only 51 % (grpE) and 67 % (htpG) of the wild type butanol concentrations were detected in the culture supernatants (Table 1). Fermenation experiments in MS-MES medium further revealed clearly hampered growth of C. acetobutylicum pT::grpE, whereas the growth rates of C. acetobutylicum pT::groESL and C. acetobutylicum pT::htpG were only moderately reduced as compared to the wild type and the vector control (Table 1).
Butanol adaptation
As depicted in Fig. 1, we observed an unexpected phenomenon in the repeatedly conducted butanol stress experiments: in contrast to the control strains, i. e. wild type and vector control, the stress protein overexpressing C. acetobutylicum strains harboring plasmids pT::groESL, pT::grpE, or pT::htpG, respectively, always revealed several colonies on the c.f.u. counting plates after exposure to high butanol concentrations (Fig. 1). This led to the hypothesis that the HSP overexpressing strains must have a mechanism to adapt to butanol stress more efficiently, and the question was whether the plasmids mutated or the whole cells evolved, enabled by improved tolerance mechanisms as well as a simple selection pressure. Therefore, consecutive butanol stress experiments were conducted and are summarized in Figs. 2 and 3. Plasmids were isolated after each stress experiment and sequence analyses did not show any amino acid exchange of the respective stress proteins. Moreover, re-transformation of the isolated plasmids into C. acetobutylicum ATCC 824 did not alter the original phenotype (data not shown). Following the successive challenges at 2 % (v/v) butanol for 6 h, improved butanol resistance was noticed for all over-expression strains subsequently monitored for their resistance to both 1 and 2 % (v/v) butanol concentrations (Fig. 2). The absolute c.f.u. counts for the over-expression strains are shown in Fig. 3, exhibiting a better performance for all three overexpession strains, but particularly for C. acetobutylicum pT::grpE. The difference between butanol-stressed cells and the control cultures clearly demonstrated the positive influence of increased HSP abundance for enhanced butanol tolerance and adaptation.
Butanol stress adaptation of C. acetobutylicum pT::groESL, pT::grpE and pT::htpG. The experimental scheme is shown on the left: after each butanol stress experiment, two colonies of cultures exposed to 2 % (v/v) butanol for 6 h were used to inoculate fresh CGM, grown overnight and subjected again to 1 % (v/v) (diamonds) and 2 % (v/v) (circles) n-butanol for 8 h in comparison to the control without butanol (squares). The c.f.u. (c.f.u.) were determined as described in Fig. 1, the average data of two similar replicates each are shown
Improved survival rates of C. acetobutylicum pT::groESL, pT::grpE and pT::htpG after butanol exposure. The strains were exposed to 2 % (v/v) butanol for 6 h and spread on RCA plates (grey), the respective controls (black) were treated similarly without butanol exposure prior to plating. From each strain, two randomly-picked colonies were used for a subsequent stress experiment employing 6 h of incubation in the presence of 1 % (v/v) (a) and 2 % (v/v) (b) butanol. The c.f.u. were determined from serial dilutions, average c.f.u. values from two independent experiments are shown
References
Alsaker KV, Papoutsakis ET (2005) Transcriptional program of early sporulation and stationary-phase events in Clostridium acetobutylicum. J Bacteriol 187:7103–7118
Alsaker KV, Spitzer TR, Papoutsakis ET (2004) Transcriptional analysis of spo0A over-expression in Clostridium acetobutylicum and its effect on the cell’s response to butanol stress. J Bacteriol 186:1959–1971
Alsaker KV, Paredes C, Papoutsakis ET (2010) Metabolite stress and tolerance in the production of biofuels and chemicals: gene-expression-based systems analysis of butanol, butyrate, and acetate stresses in the anaerobe Clostridium acetobutylicum. Biotechnol Bioeng 105:1131–1147
Baer SH, Blaschek HP, Smith TL (1987) Effect of butanol challenge and temperature on lipid composition and membrane fluidity of butanol-tolerant Clostridium acetobutylicum. Appl Environ Microbiol 53:2854–2861
Baer SH, Bryant DL, Blaschek HP (1989) Electron spin resonance analysis of the effect of butanol on the membrane fluidity of intact cells of Clostridium acetobutylicum. Appl Environ Microbiol 55:2729–2731
Bahl H, Müller H, Behrens S, Joseph H, Narberhaus F (1995) Expression of heat shock genes in Clostridium acetobutylicum. FEMS Microbiol Rev 17:341–348
Bond-Watts BB, Bellerose RJ, Chang MCY (2011) Enzyme mechanism as a kinetic control element for designing synthetic biofuel pathways. Nat Chem Biol 7:222–227
Bowles LK, Ellefson WL (1985) Effects of butanol on Clostridium acetobutylicum. Appl Environ Microbiol 50:1165–1170
Ezeji TC, Karcher PM, Qureshi N, Blaschek HP (2005) Improving performance of a gas stripping-based recovery system to remove butanol from Clostridium beijerinckii fermentation. Bioprocess Biosyst Eng 27:207–214
Ezeji TC, Qureshi N, Blaschek HP (2007) Bioproduction of butanol from biomass: from genes to bioreactors. Curr Opin Biotechnol 18:220–227
Ezeji T, Milne C, Price ND, Blaschek HP (2010) Achievements and perspectives to overcome the poor solvent resistance in acetone and butanol-producing microorganisms. Appl Microbiol Biotechnol 85:1697–1712
Fischer RJ, Oehmcke S, Meyer U, Mix M, Schwarz K, Fiedler T, Bahl H (2006) Transcription of the pst operon of Clostridium acetobutylicum is dependent on phosphate concentration and pH. J Bacteriol 188:5469–5478
Fischer CR, Klein-Marcuschamer D, Stephanopoulos G (2008) Selection and optimization of microbial hosts for biofuels production. Metab Eng 10:295–304
Girbal L, von Abendroth G, Winkler M, Benton PM, Meynial-Salles I, Croux C, Peters JW, Happe T, Soucaille P (2005) Homologous and heterologous over-expression in Clostridium acetobutylicum and characterization of purified clostridial and algal Fe-only hydrogenases with high specific activities. Appl Environ Microbiol 71:2777–2781
Grant SG, Jessee J, Bloom FR, Hanahan D (1990) Differential plasmid rescue from transgenic mouse DNAs into Escherichia coli methylation-restriction mutants. Proc Natl Acad Sci U S A 87:4645–4649
Green EM (2011) Fermentative production of butanol—the industrial perspective. Curr Opin Biotechnol 22:337–343
Harris LM, Desai RP, Welker NE, Papoutsakis ET (2000) Characterization of recombinant strains of the Clostridium acetobutylicum butyrate kinase inactivation mutant: need for new phenomenological models for solventogenesis and butanol inhibition? Biotechnol Bioeng 67:1–11
Heap JT, Pennington OJ, Cartman ST, Carter GP, Minton NP (2007) The ClosTron: a universal gene knock-out system for the genus Clostridium. J Microbiol Methods 70:452–464
Heluane H, Dagher MRE, Bruno-Bárcena JM (2011) Meta-analysis and functional validation of nutritional requirements of solventogenic clostridia growing under butanol stress conditions and coutilization of d-glucose and d-xylose. Appl Environ Microbiol 77:4473–4485
Hillmann F, Döring C, Riebe O, Ehrenreich A, Fischer RJ, Bahl H (2009) The role of PerR in O2-affected gene expression of Clostridium acetobutylicum. J Bacteriol 191:6082–6093
Hönicke D, Janssen H, Grimmler C, Ehrenreich A, Lütke-Eversloh T (2012) Global transcriptional changes of Clostridium acetobutylicum cultures with increased butanol:acetone ratios. N Biotechnol. doi:10.1016/j.nbt.2012.01.001
Jones DT, Woods DR (1986) Acetone-butanol fermentation revisited. Microbiol Rev 50:484–524
Jones SW, Paredes CJ, Tracy B, Cheng N, Sillers R, Senger RS, Papoutsakis ET (2008) The transcriptional program underlying the physiology of clostridial sporulation. Genome Biol 9:R114
Knoshaug EP, Zhang M (2009) Butanol tolerance in a selection of microorganisms. Appl Biochem Biotechnol 153:13–20
Lee SY, Park JH, Jang SH, Nielsen LK, Kim J, Jung KS (2008) Fermentative butanol production by clostridia. Biotechnol Bioeng 101:209–228
Lehmann D, Lütke-Eversloh T (2011) Switching Clostridium acetobutylicum to an ethanol producer by disruption of the butyrate/butanol fermentative pathway. Metab Eng 13:464–473
Lehmann D, Hönicke D, Ehrenreich A, Schmidt M, Weuster-Botz D, Bahl H, Lütke-Eversloh T (2012) Modifying the product pattern of Clostridium acetobutylicum: physiological effects of disrupting the acetate and acetone formation pathways. Appl Microbiol Biotechnol 94:743–754
Li J, Zhao JB, Zhao M, Yang YL, Jiang WH, Yang S (2010) Screening and characterization of butanol-tolerant microorganisms. Lett Appl Microbiol 50:373–379
Liu S, Qureshi N (2010) How microbes tolerate ethanol and butanol. N Biotechnol 26:117–121
Lütke-Eversloh T, Bahl H (2011) Metabolic engineering of Clostridium acetobutylicum: recent advances to improve butanol production. Curr Opin Biotechnol 22:634–647
Mao S, Luo Y, Zhang T, Li J, Bao G, Zhu Y, Chen Z, Zhang Y, Li Y, Ma Y (2010) Proteome reference map and comparative proteomic analysis between a wild type Clostridium acetobutylicum DSM 1731 and its mutant with enhanced butanol tolerance and butanol yield. J Proteome Res 9:3046–3061
Mao S, Luo Y, Bao G, Zhang Y, Li Y, Ma Y (2011) Comparative analysis on the membrane proteome of Clostridium acetobutylicum wild type strain and its butanol-tolerant mutant. Mol Biosyst 7:1660–1677
Mermelstein LD, Papoutsakis ET (1993) In vivo methylation in Escherichia coli by the Bacillus subtilis phage phi 3T I methyltransferase to protect plasmids from restriction upon transformation of Clostridium acetobutylicum ATCC 824. Appl Environ Microbiol 59:1077–1081
Narberhaus F, Bahl H (1992) Cloning, sequencing, and molecular analysis of the groESL operon of Clostridium acetobutylicum. J Bacteriol 174:3282–3289
Narberhaus F, Giebeler K, Bahl H (1992) Molecular characterization of the dnaK gene region of Clostridium acetobutylicum, including grpE, dnaJ, and a new heat shock gene. J Bacteriol 174:3290–3299
Nicolaou SA, Gaida SM, Papoutsakis ET (2010) A comparative view of metabolite and substrate stress and tolerance in microbial bioprocessing: from biofuels and chemicals, to biocatalysis and bioremediation. Metab Eng 12:307–331
Papoutsakis ET (2008) Engineering solventogenic clostridia. Curr Opin Biotechnol 19:420–429
Rühl J, Schmid A, Blank LM (2009) Selected Pseudomonas putida strains able to grow in the presence of high butanol concentrations. Appl Environ Microbiol 75:4653–4656
Rüngeling E, Laufen T, Bahl H (1999) Functional characterisation of the chaperones DnaK, DnaJ and GrpE from Clostridium acetobutylicum. FEMS Microbiol Lett 170:119–123
Shen CR, Lan EI, Dekishima Y, Baez A, Cho KM, Liao JC (2011) Driving forces enable high-titer anaerobic 1-butanol synthesis in Escherichia coli. Appl Environ Microbiol 77:2905–2915
Taylor M, Tuffin M, Burton S, Eley K, Cowan D (2008) Microbial responses to solvent and alcohol stress. Biotechnol J 3:1388–1397
Terracciano JS, Rapaport E, Kashket ER (1988) Stress- and growth phase-associated proteins of Clostridium acetobutylicum. Appl Environ Microbiol 54:1989–1995
Tomas CA, Welker NE, Papoutsakis ET (2003) Over-expression of groESL in Clostridium acetobutylicum results in increased solvent production and tolerance, prolonged metabolism, and changes in the cell’s transcriptional program. Appl Environ Microbiol 69:4951–4965
Tomas CA, Beamish J, Papoutsakis ET (2004) Transcriptional analysis of butanol stress and tolerance in Clostridium acetobutylicum. J Bacteriol 186:2006–2018
Vollherbst-Schneck K, Sands JA, Montenecourt BS (1984) Effect of butanol on lipid composition and fluidity of Clostridium acetobutylicum ATCC 824. Appl Environ Microbiol 47:193–194
Winkler J, Rehmann M, Kao KC (2010) Novel Escherichia coli hybrids with enhanced butanol tolerance. Biotechnol Lett 32:915–920
Zhu L, Dong H, Zhang Y, Li Y (2011) Engineering the robustness of Clostridium acetobutylicum by introducing glutathione biosynthetic capability. Metab Eng 13:426–434
Acknowledgments
The authors thank the Süd-Chemie AG and the Federal Ministry of Education and Research, Germany (Grant no. 0315419A) for financial support.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mann, M.S., Dragovic, Z., Schirrmacher, G. et al. Over-expression of stress protein-encoding genes helps Clostridium acetobutylicum to rapidly adapt to butanol stress. Biotechnol Lett 34, 1643–1649 (2012). https://doi.org/10.1007/s10529-012-0951-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-012-0951-2