Abstract
Fungi belonging to the genus Cunninghamella have enzymes similar to those employed by mammals for the detoxification of xenobiotics, thus they are useful as models of mammalian drug metabolism, and as a source for drug metabolites. We report the transformation of the anti-cancer drug flutamide in Cunninghamella sp. The most predominant phase I metabolites present in the plasma of humans, 2-hydroxyflutamide and 4-nitro-3-(trifluoromethyl)aniline, were also produced in Cunninghamella cultures. Other phase I and phase II metabolites were also detected using a combination of HPLC, GC–MS and 19F-NMR.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Flutamide (2-methyl-N-[4-nitro-3-(trifluoromethyl)phenyl]propanamide is a non steroidal antiandrogen drug primarily used to treat prostate cancer and, like an increasing number of anti-cancer agents, it is fluorinated (Isanbor and O’Hagan 2006). In medicinal chemistry, the substitution of a fluorine atom is used to improve metabolic stability, bioavailability and interaction with the biological target (Purser et al. 2008). In particular the trifluoromethyl group, which is part of flutamide, is the most used fluorinated moiety in pharmaceuticals and agrochemicals. It is among the most lipophilic of all substituents and it is known to retard metabolic degradation by blocking cytochrome P450 enzymes (Park et al. 2001).
Some microorganisms have enzymes that are synonymous to those involved in xenobiotic detoxification in mammals (Prior et al. 2010; Zhang et al. 1996) and, as such, are useful models of drug metabolism (Smith and Rosazza 1974) and in the production of drug metabolites (Osorio-Lozada et al. 2008). Few studies have been published on the metabolism of fluorinated drugs by microorganisms despite the significance of this class of compounds. There are a few studies on the degradation of trifluoromethyl-containing xenobiotics, for example, Engesser et al. (1988 and 1990) reported the metabolism of 3- and 4-trifluoromethyl benzoate in Pseudomonas putida, via the corresponding catechols, to 2-hydroxy-6-oxo-7,7,7-trifluoro-hepta-2,4-dienoate. Herath and Khan (2010) examined the biotransformation of flutamide in 40 microorganisms, and demonstrated the production of three metabolites also found in mammals, which arose via nitroreduction, hydrolysis and acetylation. However, 2-hydroxyflutamide, which is a major metabolite in human plasma (Aizawa et al. 2003), was not observed in these fungal cultures, presumably because the enzyme required, cytochrome P450 (CYP1A2), was not produced by the fungi investigated.
In the present study, we investigated the biotransformation of flutamide in Cunninghamella species, which have been previously shown to be very effective models of mammalian drug metabolism (Amadio et al. 2010; Asha and Vidyavathi 2009), since they have cytochrome P450 activity and conjugation enzymes (Zhang et al. 2010).
Materials and methods
Materials
Flutamide, 2-hydroxyflutamide, 4-nitro-3-(trifluoromethyl)aniline, N-methyl-N-(trimethyl-silyl)trifluoroacetamide, sulfatase (from Helix pomatia type H-1), β-glucoronidase (from Escherichia coli) and β-glucosidase (from almonds) were purchased from Sigma.
Culture conditions
Cunninghamella elegans (strains DSM 1908, DSM 8217, DSM 63299), C. echinulata DSM 1905 and C. blakesleeana DSM 1906 (DSMZ, Germany) were selected for biotransformation of flutamide. Fungi were grown on Sabouraud dextrose/agar plates for 5 days at 26°C before being aseptically homogenized into 100 ml of sterile saline solution. The homogenate (10% v/v) was used to inoculate 50 ml of Sabouraud dextrose/broth in 250 ml Erlenmeyer flasks, which were incubated at 28°C with shaking at 150×g rpm. Control experiments were conducted in either the absence of flutamide or fungi. After 72 h, 5 mg flutamide dissolved in 20 μl dimethylformamide was added to the cultures and incubated for up to eight days. The entire cultures (supernatant and cells) were sonicated on ice (Sonicator U200S control, IKA Labortechnik) for 5 min at 50% amplitude, with intervals of 30 s after each minute to prevent overheating. Sonicates were centrifuged and the supernatant extracted with 50 ml ethyl acetate, and the extracts evaporated to dryness. Organic extracts were finally dissolved in 1 ml methanol and aqueous fractions were lyophilized and then reconstituted in 1 ml D2O prior to NMR analysis.
Analysis of fluorometabolites
Organic extracts in methanol were analysed by reversed phase HPLC using a Microsorb MV-100.5 C8 250 × 4.6 mm column. The eluate was monitored at 250 nm. Compounds were eluted with a gradient of acetonitrile/water (10–90% v/v acetonitrile) over 30 min (1 ml min−1 flow). The fluorometabolites were isolated by preparative reversed phase HPLC using a Zorbax SB-C18 9.4 mm × 25 cm column (Agilent Technologies) and a gradient of acetonitrile/water (20–60% v/v acetonitrile) over 30 min (3.5 ml min−1 flow).
GC–MS analysis was conducted on per-trimethylsilylated extracts and non-derivatised culture extracts that were dissolved in 1 ml ethyl acetate. Silylation was performed on 1 ml lyophilised extracts by adding 50 μl N-methyl-N-(trimethyl-silyl)trifluoroacetamide (MSTFA) and heating at 100°C for 1 h. Derivatised samples (1 μl) were injected in the splitless mode onto a HP-1 column (12 m × 0.25 mm × 0.33 μm) and the oven was held at 120°C for 2 min then raised to 300°C at 10°C min−1. Non-derivatized (1 μl) samples were injected in the splitless mode and the oven temperature held at 70°C for 3 min then raised to 250°C at 10°C min−1. The mass spectrometer was operated in the scan mode.
Fluorine-19 NMR (19F-NMR) spectroscopy was performed using a Varian 400 MHz spectrometer. Organic and aqueous extracts were dissolved in 800 μl of CDCl3 or D2O, respectively, to provide a lock signal.
Results
Analysis of phase I metabolites
The microbial biotransformation of flutamide was investigated in selected Cunnighamella species previously shown to transform xenobiotics (Asha and Vidyavathi 2009). Morphological growth as dispersed mycelia of Cunninghamella species in liquid cultures is a crucial condition for biotransformation (Amadio and Murphy 2010). C. elegans DSM 8217, DSM 63299 and C. echinulata DSM 1905 grew forming clumps which resulted in negligible biotransformation. Interestingly, when the three species were mixed together as single inoculum the most favourable growth morphology of substrate was achieved, but it was not possible to determine the predominant species involved in the metabolic process. C. elegans DSM 1908 and C. blakesleeana DSM 1906 both had reproducible morphological growth as homogeneously dispersed mycelia and biotransformed flutamide to the same type of metabolites and with comparable yield. For the purpose of this study C. elegans 1908 strain was chosen for more detailed investigation.
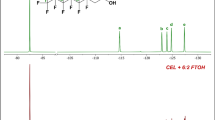
Organic extracts analyzed by 19F-NMR showed the presence of four fluorometabolites (Supplementary Fig. 1) with resonances at −60.22 (2-hydroxyflutamide), −60.21 (flutamide), −60.47 and −62.95 ppm (all singlets). Signals of flutamide and 2-hydroxyflutamide were assigned by comparison with authentic standards. The most predominant metabolite was the 2-hydroxyflutamide that it is also known to be the major pharmacologically active metabolite in mammals (Kostrubsky et al. 2007). HPLC analysis of the organic extracts (Fig. 1) showed that approx. 50% of flutamide was degraded to a variety of more polar metabolites by Cunninghamella species over eight days. No metabolite peaks were found in the control experiments and no disappearance of flutamide was observed in uninoculated control flasks. Flutamide had a retention time (tR) of 27.5 min and was metabolised to four major metabolites with tR 20.8, 20, 19.5 and 17.7 min (peaks M1, M2, M3, M4, respectively). The chromatographic profile was comparable to that reported by (Kang et al. 2008) using human liver microsomes, and the tR of M1 and M2 were identical to those of 2-hydroxyflutamide and 4-nitro-3-(trifluoromethyl)aniline standards, respectively. In an attempt to increase the degree of biotransformation, a longer period of incubation (up to 14 days) and different flutamide starting concentrations (0.01–1 mg/ml) were investigated; in each case there was no difference in the extent of flutamide biotransformation (data not shown).
Peaks M1 and M2 had the same retention times as 2-hydroxyflutamide and 4-nitro-3-(trifluoromethyl)aniline, respectively, by comparison with authentic standards. To identify the other compounds the metabolites were isolated by preparative reversed phase HPLC and then re-analyzed by HPLC to confirm their retention times. Owing to the small amounts recovered the metabolites were analysed by GC–MS (Table 1). The fractions containing the compounds M1 and M2 had mass spectra identical to the authentic standards of 2-hydroxyflutamide and 4-nitro-3-(trifluoromethyl)aniline, respectively, as anticipated from the HPLC analyses. The compounds M3 and M4 required silylation prior to GC–MS analysis; M3 had a similar mass spectrum to 2-hydroxyflutamide, thus is probably another mono-hydroxylated metabolite, and the molecular ion of M4 was identical to silylated 2-methyl-N-[4-amino-3-(trifluoromethyl)phenyl]propanamide. In the recent study by (Herath and Khan 2010), the yeast Rhodotorula mucilaginosa (and other fungi that they investigated) produced two of the same metabolites that were identified here, M2 and M4, and these authors suggested that fungal metabolism of flutamide occurs exclusively via amide bond cleavage. Here, we have shown that in Cunninghamella sp. the main pathway of biotransformation is oxidation to hydroxyflutamide, presumably via cytochrome P450 activity (Fig. 2). Shet et al. (1997) demonstrated that the inhibition of flutamide metabolism in humans is caused by its own main metabolite, 2-hydroxyflutamide, which possibly accounts for the incomplete catabolism of flutamide in C. elegans.
Analysis of phase II metabolites
Several flutamide conjugates such as sulfate, glucuronide and glutathione-related adducts have been recovered in urine from rats and humans (Tevell et al. 2006). In our study, examination of aqueous extracts by 19F-NMR showed six signals (Fig. 3a) with resonances at −50.16, −59.99, −60.05, 60.72, −60.78 and −75.43 ppm. C. elegans has the ability to form conjugates with aromatic hydrocarbons (Cerniglia et al. 1982) and possible these are possibly phase II metabolites. The aqueous fraction was treated with the sequential addition of deconjugation enzymes (added in the order: sulfatase, β-glucuronidase and β-glucosidase) and the reactions monitored by 19F-NMR spectroscopy between each addition, demonstrating that a sulfated metabolite was present (Fig. 3b). Subsequent treatment with other deconjugative enzymes did not result in any further changes to the spectrum. HPLC analysis of the aqueous extract revealed the presence of flutamide and 2-hydroxyflutamide, which were not completely removed by ethyl acetate extraction. It was not possible to determine the identity of the other compounds present, although other conjugation reactions, e.g. glutathione-S-transferase and glucuronsyl transferase, are known in C. elegans (Zhang et al. 1996).
Discussion
We have determined the major biotransformation products of the anticancer drug flutamide in the fungus C. elegans. The work demonstrates for the first time using microorganisms that 2-hydroxyflutamide is the most predominant metabolite, which is similar to previous in vivo and in vitro studies conducted in mammals. The results support the potential of using the fungus Cunninghamella, which can transform a broad range of xenobiotic compounds in an analogous fashion to mammals and is therefore a model of mammalian drug metabolism to generate primary and conjugated metabolites of flutamide. Considering that promoting alternatives to animal testing is one of the objectives of the recent European Union legislation governing the testing and evaluation of chemicals, REACH (Regulation, Evaluation, Authorisation and Restriction of Chemicals), and the US Food and Drug Administration’s guidelines for testing drug metabolites (Guengerich 2009), the use of microbial systems is becoming an important tool for testing drugs and for production of valuable drug metabolites.
References
Aizawa Y, Ikemoto I, Kishimoto K, Wada T, Yamazaki H, Ohishi Y, Kiyota H, Furuta N, Suzuki H, Ueda M (2003) Flutamide-induced hepatic dysfunction in relation to steady-state plasma concentrations of flutamide and its metabolites. Mol Cell Biochem 252:149–156
Amadio J, Murphy CD (2010) Biotransformation of fluorobiphenyl by Cunninghamella elegans. Appl Microbiol Biotechnol 86:345–351
Amadio J, Gordon K, Murphy CD (2010) Biotransformation of flurbiprofen by Cunninghamella species. Appl Environ Microbiol 76:6299–6303
Asha S, Vidyavathi M (2009) Cunninghamella—a microbial model for drug metabolism studies—a review. Biotechnol Adv 27:16–29
Cerniglia CE, Freeman JP, Mitchum RK (1982) Glucuronide and sulfate conjugation in the fungal metabolism of aromatic-hydrocarbons. Appl Environ Microbiol 43:1070–1075
Engesser KH, Cain RB, Knackmuss HJ (1988) Bacterial metabolism of side-chain fluorinated aromatics—co-metabolism of 3-trifluoromethyl(tfm)-benzoate by Pseudomonas putida (arvilla) mt-2 and Rhodococcus rubropertinctus n657. Arch Microbiol 149:188–197
Engesser KH, Rubio MA, Knackmuss HJ (1990) Bacterial metabolism of side-chain-fluorinated aromatics—unproductive meta-cleavage of 3-trifluoromethylcatechol. Appl Microbiol Biotechnol 32:600–608
Guengerich FP (2009) Introduction: human metabolites in Safety Testing (MIST) Issue. Chem Res Toxicol 22:237–238
Herath W, Khan IA (2010) Microbial metabolism. Part 11. Metabolites of flutamide. Chem Pharm Bull 58:562–564
Isanbor C, O’Hagan D (2006) Fluorine in medicinal chemistry: a review of anti-cancer agents. J Fluorine Chem 127:303–319
Kang P, Dalvie D, Smith E, Zhou S, Deese A, Nieman JA (2008) Bioactivation of flutamide metabolites by human liver microsomes. Drug Metab Dispos 36(7):1425–1437
Kostrubsky SE, Strom SC, Ellis E, Nelson SD, Mutlib AE (2007) Transport, metabolism, and hepatotoxicity of flutamide, drug-drug interaction with acetaminophen involving phase I and phase II metabolites. Chem Res Toxicol 20:1503–1512
Osorio-Lozada A, Surapaneni S, Skiles GL, Subramanian R (2008) Biosynthesis of drug metabolites using microbes in hollow fiber cartridge reactors: case study of diclofenac metabolism by Actinoplanes species. Drug Metab Dispos 36:234–240
Park BK, Kitteringham NR, O’Neill PM (2001) Metabolism of fluorine-containing drugs. Ann Rev Pharmacol Toxicol 41:443–470
Prior JE, Shokati T, Christians U, Gill RT (2010) Identification and characterization of a bacterial cytochrome p450 for the metabolism of diclofenac. Appl Microbiol Biotechnol 85:625–633
Purser S, Moore PR, Swallow S, Gouverneur V (2008) Fluorine in medicinal chemistry. Chem Soc Rev 37:320–330
Shet MS, McPhaul M, Fisher CW, Stallings NR, Estabrook RW (1997) Metabolism of the antiandrogenic drug (flutamide) by human CYP1A2. Drug Metab Dispos 25:1298–1303
Smith RV, Rosazza JP (1974) Microbial models of mammalian metabolism—aromatic hydroxylation. Arch Biochem Biophys 161:551–558
Tevell A, Lennernas H, Jonsson M, Norlin M, Lennernas B, Bondesson U, Hedeland M (2006) Flutamide metabolism in four different species in vitro and identification of flutamide metabolites in human patient urine by high performance liquid chromatography/tandem mass spectrometry. Drug Metab Dispos 34:984–992
Zhang DL, Yang YF, Leakey JEA, Cerniglia CE (1996) Phase I and phase II enzymes produced by Cunninghamella elegans for the metabolism of xenobiotics. FEMS Microbiol Lett 138:221–226
Acknowledgments
The authors acknowledge financial assistance from the Environmental Protection Agency STRIVE Programme.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Amadio, J., Murphy, C.D. Production of human metabolites of the anti-cancer drug flutamide via biotransformation in Cunninghamella species. Biotechnol Lett 33, 321–326 (2011). https://doi.org/10.1007/s10529-010-0425-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-010-0425-3