Abstract
The rpiB gene, encoding ribose-5-phosphate isomerase (RpiB) from Clostridium thermocellum, was cloned and expressed in Escherichia coli. RpiB converted d-psicose into d-allose but it did not convert d-xylose, l-rhamnose, d-altrose or d-galactose. The production of d-allose by RpiB was maximal at pH 7.5 and 65°C for 30 min. The half-lives of the enzyme at 50°C and 65°C were 96 h and 4.7 h, respectively. Under stable conditions of pH 7.5 and 50°C, 165 g d-allose l−1 was produced without by-products from 500 g d-psicose l−1 after 6 h.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
d-Allose, one of the rare aldohexoses, has attracted much attention as an inhibitor for ischemia/reperfusion injury, segmented neutrophil production, and lowered platelet count (Hossain et al. 2003, 2004, 2006). Moreover, a combination use of d-allose as a potent immunosuppressant with a low dose of FK506 significantly increased the rate of allograft survival with less tissue damage (Hossain et al. 2000). The biological manufacture of d-allose has been studied using l-rhamnose isomerase (l-RI) from Pseudomonas stutzeri, which is the only reported enzyme (Bhuiyan et al. 1998; Leang et al. 2004a; Morimoto et al. 2006; Menavuvu et al. 2006). Therefore, the investigation and further development for other sources of enzymes to produce d-allose should be greatly needed.
Ribose-5-phosphate isomerase (EC 5.3.1.6, RpiB) forms a homodimer and catalyzes the conversion of d-ribose-5-phosphate to d-ribulose-5-phosphate in the branch of the pentose phosphate pathway (Zhang et al. 2003). Moreover, this enzyme can take part in d-allose metabolism (Kim et al. 1997; Poulsen et al. 1999). However, d-allose production by the enzyme has not yet been reported.
In this study, the ribose-5-phosphate isomerase B (rpiB) gene from Clostridium thermocellum was cloned and expressed in E. coli. Furthermore, the production of d-allose from d-psicose by RpiB was investigated.
Materials and methods
Bacterial strains, plasmids, and culture conditions
The genomic DNA from Clostridium thermocellum was used as a source of rpiB gene. E. coli BL21(DE3) was used as a host. pBluescript II SK(+) (Stratagene, La Jolla, CA, USA) and pQE30 (Qiagen, Valencia, CA, USA) plasmids were used as cloning and expression vectors, respectively.
The recombinant E. coli for protein expression was cultivated in a Luria-Bertani (LB) medium containing 50 μg ampicillin ml−1 and 20 μg kanamycin ml−1 at 37°C with agitation at 200 rpm until the OD 600 reached 0.5. IPTG was added to the culture medium at 0.1 mM to induce RpiB and then the culture was grown at 16°C for 16 h.
Gene cloning
The rpiB gene (450 bp) was amplified from the genomic DNA of C. thermocellum by PCR using Pfu DNA polymerase (Daemyung Science, Seoul, Korea). The sequence of the oligonucleotide primers used for gene cloning was based on the DNA sequence of the C. thermocellum RpiB gene (GenBank Accession Number, ZP 00503831). Forward (RpiB-F, 5′-CCCATGGAGGAAAGTATGAAAATTGG-3′) and reverse primers (RpiB-R, 5′-CCTGCAGATCAACGGATGATCCATAAC-3′) were designed to introduce the NcoI and PstI restriction sites (underlined) for rpiB gene cloning. The primers were synthesized by Bioneer Co. (Daejon, Korea). The amplified DNA fragment obtained by PCR was purified and digested with both NcoI and PstI endonucleases (Promega, Madison, WI, USA). The digested DNA fragment was extracted, and then inserted into the pBluescript II SK(+) plasmid digested with the same restriction enzymes. The resultant plasmid, pBCTrpiB, was digested with both NcoI and PstI and the obtained fragment, contains rpiB gene, was treated by Klenow fragment (Takara, Shiga, Japan). Finally, the fragment was inserted into pQE30 plasmid digested to completion with SmaI and PstI and then the pQCTrpiB plasmid was obtained. E. coli BL21(DE3) strain was transformed with the plasmid and plated on LB agar containing 50 μg ampicillin ml−1 and 20 μg kanamycin ml−1 to select transformants. The expression of the gene encoding ribose-5-phosphate isomerase was determined by both SDS-PAGE and the assay of enzyme activity.
Enzyme assay
The activity of RpiB was determined by measuring d-allose formation using d-psicose as a substrate. Unless otherwise stated, the reactions were carried out in a 50 mM Tris/HCl buffer (pH 7.5) containing 1% (w/v) d-psicose and 0.03 units enzyme ml−1 at 50°C for 30 min. For the investigation of substrate specificity, the reactions were performed with 1% (w/v) of each sugar for 16 h. One unit of enzyme activity was defined as the amount of enzyme required to produce 1 μmol d-allose per min at 50°C and pH 7.5.
Analytical methods
Protein concentrations were determined by a Bio-rad assay with bovine serum albumin as a standard. The concentrations of monosaccharides were analyzed by the cysteine-carbazole method (Dische and Borenfreund 1951) or HPLC with a refractive index detector (Shimadzu SCL-10A, Kyoto, Japan) using NH2P-50 4E (Showa Denko, Tokyo, Japan), BP-100 Ca2+ carbohydrate (Benson Ploymeric, Reno, NV, USA), and cosmosil (Nacalai Tesque., Kyoto, Japan) columns which were eluted at 30°C with 70% (v/v) acetonitrile at 0.8 ml min−1, at 80°C with water at 0.4 ml min−1, and at 35°C with 80% (v/v) acetonitrile at 1 ml min−1, respectively.
Results and discussion
Gene cloning
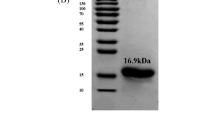
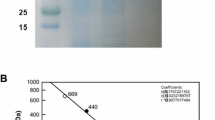
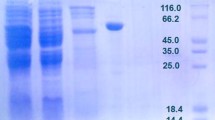
The rpiB genes based on the reported DNA sequence of RpiB was cloned into the pQE30 plasmid to produce pQCTrpiB and expressed in E. coli. The expressed pQCTrpiB was detected using SDS-PAGE. The molecular mass of the enzyme was estimated to be approximately 17 kDa by SDS-PAGE (data not shown) and this was consistent with the calculated value of 16,950 Da based on the 149-residue amino acid and six histidine sequences.
Although RpiB from C. thermocellum and l-RI from P. stutzeri can produce d-allose, the low homology between two enzymes indicated that two enzymes were not well aligned.
Substrate specificity
RpiB was active with d-allose, d-psicose, and d-ribose-5-phosphate but not other sugars (see Table 1). However, l-RI from Pseudomonas stutzeri converted l-rhamnose, d-xylose, d-altrose, and d-galactose (Leang et al. 2004b). These results suggest that RpiB from C. thermocellum is different from l-RI from P. stutzeri. Although d-ribose-5-phosphate is a substrate for RpiB, this enzyme had higher activities for d-allose and d-psicose.
pH, temperature, and metal ions effects
d-Allose production by RpiB was maximal at pH 7.5, whereas optimum pH for d-allose production by l-RI was at pH 9.0 (Bhuiyan et al. 1997). The activity of RpiB was maximal at 65°C for 30 min, while that of l-RI was shown at 60°C (Bhuiyan et al. 1997). Thermal inactivation of RpiB followed first-order kinetics with half-lives (t1/2) of 96 h, 18 h, 8.1 h, 4.7 h, 1.7 h, and 0.3 h at 50°C, 55°C, 60°C, 65°C, 70°C, and 75°C, respectively (Fig. 1). The results indicated RpiB from C. thermocellum has relatively unstable above 55°C. Therefore, the further reactions were performed at 50°C.
d-Allose-producing activity from d-psicose by RpiB was not activated by monovalent or divalent cations and not inhibited by EDTA because phosphosugar isomerases are metal-independent enzymes (Teplayakov et al. 1999). However, l-RI did not show any activity after its treatment with EDTA (Bhuiyan et al. 1997).
Bioconversion of d-psicose into d-allose
Using 500 g d-psicose l−1, 165 g d-allose l−1 was obtained with RpiB after 6 h at 50°C and pH 7.5, corresponding to the conversion yield of 33% (Fig. 2). A similar conversion occurred using d-psicose at 1.25, 2.5, and 5 g l−1 with 2.8 units enzyme ml−1 for 6 h. Previously, the l-RI from P. stutzeri converted 100 g d-psicose l−1 into 25 g d-allose l−1 with 8 g d-altrose l−1 as a by-product (Menavuvu et al. 2006) and an immobilized l-RI produced 150 g d-allose l−1 from 500 g d-psicose l−1 as a by-product (Morimoto et al. 2006). The yield and concentration of d-allose obtained with RpiB of C. thermocellum were therefore higher than those with l-RI of P. stutzeri which, unfortunately, produced a significant amount of d-altrose as a by-product, which is a serious problem in the purification of d-allose. However, the RpiB of C. thermocellum used in this study did not produce d-altrose (Fig. 3).
The use of RpiB has advantages of higher yield and no by-product formation. Moreover, the metal-independent RpiB dose not require metal ions in the reaction, thus it is probably more suitable for diverse molecular evolution than metal-dependent l-RI. Therefore, RpiB of C. thermocellum is a more proper enzyme to produce d-allose from d-psicose than l-RI of P. stutzeri, which has been known to the only one enzyme for the d-allose production.
References
Bhuiyan HS, Itami Y, Izumori K (1997) Isolation of an l-rhamnose isomerase-constitutive mutant of Pseudomonas sp. strain LL172: purification and characterization of the enzyme. J Ferment Bioeng 84:319–323
Bhuiyan HS, Itami Y, Rokui Y, Katayama T, Izumori K (1998) d-Allose production from d-psicose using immobilized l-rhamnose isomerase. J Ferment Bioeng 85:539–541
Dische Z, Borenfreund E (1951) A new spectrophotometric method for the detection and determination of keto sugars and trioses. J Biol Chem 192:583–587
Hossain MA, Wakabayashi H, Goda F, Kobayashi S, Maeba T, Maeta H (2000) Effect of the immunosuppressants FK 506 and d-allose on allogenic orthotopic liver transplantation in rats. Transplant Proc 32:2021–2023
Hossain MA, Izuishi K, Maeta H (2003) Protective effects of d-allose against ischemia reperfusion injury of the rat river. J Hepatobiliary Pancreat Surg 10:218–225
Hossain MA, Izuishi K, Tokuda M, Izumori K, Maeta H (2004) d-Allose has a strong suppressive effect against ischemia/reperfusion injury: a comparative study with allopurinol and superoxide dismutase. J Hepatobiliary Pancreat Surg 11:181–189
Hossain MA, Wakabayashi H, Izuishi K, Okano K, Yachida S, Tokuda M, Izumori K, Maeta H (2006) Improved microcirculatory effect of d-allose on hepatic ischemia reperfusion following partial hepatectomy in cirrhotic rat liver. J Biosci Bioeng 101:369–371
Kim C, Song S, Park C (1997) The d-allose operon of Escherichia coli K-12. J Bacteriol 179:7631–7637
Leang K, Takada G, Ishimura A, Okita M, Izumori K (2004) Cloning, nucleotide sequence, and overexpression of the l-rhamnose isomerase gene from Pseudomonas stutzeri in Escherichia coli. Appl Environ Microbiol 70:3298–3304
Leang K, Takada G, Fukai Y, Morimoto K, Granström TB, Izumori K (2004) Novel reactions of l-rhamnose isomerase from Pseudomonas stutzeri and its relation with d-xylose isomerase via substrate specificity. Biochim Biophys Acta 1674:68–77
Menavuvu BT, Poonperm W, Leang K, Noguchi N, Okada H, Morimoto K, Granström TB, Takada G, Izumori K (2006) Efficient biosynthesis of d-allose from d-psicose by cross-linked recombinant l-rhamnose isomerase: separation of product by ethanol crystallization. J Biosci Bioeng 101:340–345
Morimoto K, Park CS, Ozaki M, Takeshita K, Shimonishi T, Granström TB, Takata G., Tokuda M, Izumori K (2006) Large scale production of d-allose from d-psicose using continuous bioreactor and separation system. Enzyme Microb Technol 38:855–859
Poulsen TS, Chang YY, Hove-Jensen B (1999) d-Allose catabolism of Escherichia coli: involvement of alsI and regulation of als regulon expression by allose and ribose. J Bacteriol 181:7126–7130
Teplyakov A, Obmolova G, Badet-Denisot MA, Badet B (1999) The mechanism of sugar phosphate isomerization by glucosamine 6-phosphate synthase. Protein Sci 8:596–602
Zhang RG, Andersson CE, Sakarina T, Evdokimova E, Edwards AM, Joachimiak A, Savchenko A, Mowbray SL (2003) The 2.2 Å resolution structure of RpiB/AlsB from Escherichia coli illustrates a new approach to the ribose-5-phosphate isomerase reaction. J Mol Biol 332:1083–1094
Acknowledgments
This study was supported by the Second Phase of Brain Korea 21 Project (Ministry of Education & Human Resources Development) and by Grant R01-2004-000-10012-0 from the Basic Research Program of the Korea Science & Engineering Foundation (KOSEF).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Park, CS., Yeom, SJ., Kim, HJ. et al. Characterization of ribose-5-phosphate isomerase of Clostridium thermocellum producing d-allose from d-psicose. Biotechnol Lett 29, 1387–1391 (2007). https://doi.org/10.1007/s10529-007-9393-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-007-9393-7