Abstract
Fig, Ficus carica L., is a useful genetic resource for commercial cultivation. In this study, RAPD (60), ISSR (48), RAMPO (63), and SSR (34) markers were compared to detect polymorphism and to establish genetic relationships among Tunisian fig tree cultivars. The statistical procedures conducted on the combined data show considerable genetic diversity, and the tested markers discriminated all fig genotypes studied. The identification key established on the basis of SSR permitted the unambiguous discrimination of cultivars and confirmed the reliability of SSR for fingerprinting fig genotypes. The study findings are discussed in relation to the establishment of a national reference collection that will aid in the conservation of Tunisian fig resources.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Ficus is composed of about 700 species growing mainly in tropical and subtropical areas; it is classified into six subgenera (Berg 2003). The fig (Ficus carica L.) is a diploid species (2n = 26) (Weiblen 2000) that belongs to the family Moraceae. It is characterized by the presence of latex in all parts of the plant. The fig is probably the oldest cultivated crop (Zohary and Hopf 2000), being widely used as a fruit tree in the eastern Mediterranean regions of Europe and Africa and in southwestern regions of Asia (Storey 1975). In nature, this tree is present in two sexual forms, the caprifig (male tree or pollinator) and the domesticated fig (female tree and producer of the edible fruit). The biological cycle of Ficus carica is strongly tied to the pollinator wasp, a blastophage, Blastophaga psenes (Kjelberg et al. 1987).
In Tunisia, fig trees grow in all areas of the country, in close association with date palms in the south or olive trees in other regions. Previous studies have identified a large number of cultivars housed in collections throughout the country (Rhouma 1996; Mars et al. 1994). Despite these large resources, fig cultivation remains traditional, and fruit production is relatively low. In addition, the local germplasm is menaced by genetic erosion due to intensive urbanization, cultivation of monovarietal culture, and biotic stresses. Therefore, preservation and characterization of the Tunisian fig germplasm have become imperative.
Several studies have reported the use of morphometric and pomological parameters as well as isozyme markers to discriminate fig cultivars (Mars et al. 1998; Hedfi et al. 2003; Chatti et al. 2004a). Unfortunately, these parameters are highly influenced by environmental conditions and the stage of tissue development. To overcome these difficulties, molecular markers have been successfully designed to examine genetic diversity and to characterize fig cultivars. Among these markers, random amplified polymorphic DNA (RAPD), inter simple sequence repeats (ISSR), random amplified microsatellite polymorphism (RAMPO), and simple sequence repeats (SSR) have been proven to be suitable to survey molecular polymorphism and to examine relationships among fig cultivars (Salhi Hannachi et al. 2004, 2005, 2006; Chatti et al. 2004b, 2007; Khadari et al. 1995), but their utility and contribution to the objectives outlined have not been clearly established, taking into account the nature of each marker. Few comparative analyses of fig genetic diversity as inferred by RAPD and ISSR have been reported (Salhi Hannachi et al. 2005; Hidetoshi et al. 2009).
In this study, we demonstrate the production of suitable markers and the assessment of molecular polymorphism in a set of Tunisian fig cultivars using RAPD, ISSR, RAMPO, and SSR markers. The objectives of this study were (1) to compare the use of several kinds of molecular markers in the genetic characterization of fig trees; (2) to elucidate the genetic diversity and the genetic relationships among fig trees using pooled RAPD, ISSR, and RAMPO data in order to provide a strong valid molecular analysis; and (3) to establish an identification key of fig cultivars using SSR markers recognized as the most efficient for cultivar fingerprinting. Data are discussed in relation to conservation management of this crop and the establishment of a national reference collection of local fig germplasm. The identification of local fig resources is important in recognition of gene pools, identification of pitfalls in germplasm collections, and development of effective conservation and management strategies.
Materials and Methods
Plant Material
The 17 cultivars (15 common figs and 2 caprifigs) used in this study are cultivated mainly in the Sahel region of Tunisia and are housed in the collection of the Superior Institute of Agronomy, Chott Meriam (Table 1). The morphological and main phenotypic traits of these ecotypes have been previously reported (Chatti et al. 2004a). The plant material consisted of young leaves randomly sampled from adult trees and stored at −20°C until their use for DNA preparation.
DNA Preparation
DNA was extracted from frozen leaves according to Dellaporta et al. (1984). After purification, the DNA concentration was estimated spectrophotometrically. DNA integrity was checked by analytical agarose gel electrophoresis according to Sambrook et al. (1989).
Primers and PCR Assays
Depending on the procedure, different primer sets were used in this study (Table 2). Universal oligonucleotides purchased from Operon Technologies (Alameda, CA, USA) were used to perform RAPD; the ISSR methodology utilized 21 primers that are complementary to microsatellite motives and arbitrarily defined. Six specific microsatellite primers identified by Khadari et al. (2001) were used to detect SSRs. Combined RAPD and ISSR primers were used for the RAMPO method (Chatti et al. 2007).
RAPD–PCR assays were carried out as described by Chatti et al. (2004b). ISSR–PCR was achieved according to Zehdi et al. (2004). RAMPO–PCR was performed according to Chatti et al. (2007). SSR–PCR amplification was performed according to Saddoud et al. (2007). All PCR reactions were carried out in a Crocodile III thermocycler (QBiogène, Illkirch, France).
The derived amplified products were electrophoresed on 1.5% agarose gel in 0.5× TBE buffer (pH 8.3), stained with ethidium bromide, and visualized under UV light (Sambrook et al. 1989). SSRs were first checked on 2% agarose gels and then resolved on 10% nondenaturing polyacrylamide gels, with bands revealed by ethidium bromide staining. The SSR size was estimated by comparison with a standard 100-bp ladder.
Data Analysis
The total number of amplified bands and polymorphic bands was scored for each primer. Only reproducible and unambiguous banding patterns were taken into account. The ability of the most informative primers to differentiate ecotypes was assessed by estimating their resolving power (Rp) as defined by Prevost and Wilkinson (1999). The Rp was described to correlate well with the primer’s efficiency in distinguishing between accessions according to the formula of Gilbert et al. (1999): Rp = ∑I b, where I b = 1 − (2 × ∣0.5 − p∣), where p is the proportion of the accessions containing I band.
In addition, pooled RAPD, ISSR, and RAMPO banding patterns were transformed into a binary character matrix, with each reproducible polymorphic DNA band scored as present (1) or absent (0). The derived data matrix was computed with the GeneDist program (version 3.572c) to produce a genetic distance matrix using the formula of Nei and Li (1979), which assesses the similarity between two individuals on the basis of the number of generated bands. The genetic distance matrix was computed with the Neighbor program to produce a tree file using the unweighted pair group method with arithmetic averaging (UPGMA) algorithm (Sneath and Sokal 1973). The TreeView program (Win32, version 1.5.2) was used to draw the dendrogram illustrating the relationships among ecotypes. All these analyses were conducted using appropriate programs in Phylip (Phylogeny Inference Package, version 3.5 c), kindly provided by Felsenstein (1995) and Page’s TreeView software (Page 1996).
Based on the SSR data, genetic polymorphism was assessed by estimates of the polymorphism level. Allele frequency and heterozygosity, observed (H obs) and expected (H exp) (Nei 1978), were estimated using the Genetix 4.01 program (Belkhir 2000). Wright’s inbreeding coefficient (F IS) was computed according to Weir and Cockerham (1984). In addition, shared allele distances, as genetic distances among cultivars, were calculated according to Chakraborty and Jin (1993) using the software MSA (Microsatellite Analyzer), version 3.14 (Dieringer and Schlötterer 2003). The multivariate correspondence factorial analysis was also performed using Genetix 4.01 (Belkhir 2000). Finally, genetic matrices were compared by computing Spearman’s rank correlations using the statistical package Statistica 6.0 (StatSoft 2001).
Identification Key
Multilocus genotypes were identified, taking advantage of SSR to fingerprint cultivars, and used to establish a cultivar identification key as described by Zehdi et al. (2002). Genotypes were hierarchically organized according to the greater number of alleles per locus. Ecotypes were then classified, and those with similar fingerprints were grouped.
Results
All four techniques discriminated successfully among the cultivars investigated (Fig. 1). The RAPD markers revealed 60 polymorphic bands (53.09%), ISSR revealed 48 (54.54%), and RAMPO revealed 63 (45.66%). This result suggests that the primers used are efficient for the detection of molecular polymorphism in the cultivars tested. The calculated resolving power rates scored using the tested primers were 27.13 for RAPD, 23.15 for ISSR, and 26.96 for RAMPO, confirming the efficiency of the primers.
Banding patterns generated in Tunisian Ficus carica cultivars using RAPD (a), ISSR (b), RAMPO (c), and SSR (d). Lane L: Standard molecular size (1 kb) ladder, Gibco-BRL for RAPD, ISSR, and RAMPO; 100 bp for SSR. Lane T: Negative control. Lanes 1–17: Fig cultivars as numbered in Table 1
Genetic Diversity and Phylogenetic Relationships as Assessed by Aggregated Patterns of ISSR, RAPD, and RAMPO Markers
Cultivar polymorphism was detected using combined data of RAPD, ISSR, and RAMPO markers. A genetic distance matrix was established among cultivars for each kind of marker (Fig. 2). The derived genetic distance matrix generated by the combined data exhibited distances ranging from 0.05 to 0.61, with a mean of 0.41. Thus, it may be assumed that the ecotypes studied are highly divergent at the DNA level. The smallest genetic distance (0.05) was scored between the Soltani2 and Kahli1 cultivars, suggesting that these cultivars are similar and strongly clustered. The Kahli2, Baghali, and Bither Abiadh2 cultivars, displaying the maximum distance value of 0.61, seem to be the most divergent. All the remaining ecotypes exhibited intermediate levels of similarity.
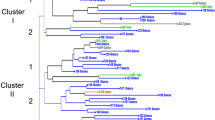
Based on the RAPD, ISSR, and RAMPO data, the UPGMA dendrogram shows the ecotypes clustered into two main groups (Fig. 3). The first cluster (a) is composed of Bither Abiadh2, Bither Abiadh3, and Besbessi, cultivars characterized by green fruits and two crops in a year. All the remaining cultivars are clustered in the second group (b) which exhibits secondary tree branching. In addition, the male trees (Jrani and Assafri) are grouped in the same subcluster, with no significant divergence from the common type.
SSR Analysis
This study demonstrated a total of 34 alleles and 31 genotypes. The allele number varied from four alleles for MFC5 and MFC7 to 7 for the MFC3, MFC6, and MFC8 loci, with a mean value of 5.67 for the collection (Tables 3, 4).
Allele Frequency
For each locus, allele frequency was calculated for the two caprifigs, for the domesticated fig trees, and for all cultivars, ranging from 0.029 for the 176-bp allele of MFC3 and 188-bp allele of MFC6 to 0.588 for the 132-bp allele of MFC7 (Table 4). For the MFC2 locus, five alleles were obtained. The most frequent allele was 174, with a frequency of 0.382 in the entire sample. Allele 176 of the MFC2 locus is present only in caprifig trees; it can be characteristic for pollinators and can have implications in sex determination in Ficus carica. A probability test (P = 0.00002) indicates significant differences between frequencies calculated for caprifigs and domesticated fig trees at this locus.
The MFC3 locus had seven alleles, with a size range of 124–194 bp. The 131-bp allele is the main allele in all cultivars (0.324). Pollinators are represented only by the two alleles 131 and 134. A probability test showed no significant differences (P = 0.40009) between domesticated fig and caprifig samples for this locus.
Four specific alleles were generated for the MFC5 locus, varying from 118 to 140 bp. The most frequent allele was 138 bp (0.441). The differences between pollinators and domesticated fig trees are not significant (P = 0.26950). The amplimers obtained for the MFC6 locus revealed seven alleles, from 166 to 188 bp, the most frequent of which was 186 bp in all cultivars (0.353). Domesticated fig tree profiles show all detected alleles; on the other hand, patterns of caprifigs presented only the MFC6 alleles of 166 and 184 bp. A probability test for assessment divergence between pollinator and domesticated fig trees showed a significant difference (P = 0.01968) (Table 4).
Four alleles were obtained by means of MFC7 specific primers, from 132 to 148 bp. The greatest allele frequency is noted for the 132-bp allele. The difference between pollinators and domesticated fig trees was significant (P = 0.01968) at this locus.
MFC8-specific primers generated seven alleles, from 145 to 202 bp. It is important to note that the allele number is relatively low, compared with the possible number of 30 alleles. A low allele frequency of 0.059 was obtained for allele 164, and the highest frequency (0.353) was estimated for allele 160. This is the most important allele in all the samples. The difference between pollinators and domesticated fig trees is significant (P = 0.04249).
The results show that all alleles were found in domesticated fig trees, except for alleles 134 and 176 of the MFC7 and MFC2 loci, which are present only in the pollinator genotypes (Table 4).
Heterozygosity and F-Statistics
For the MFC2, MFC5, and MFC6 loci, observed heterozygosity is greater than expected (Table 3). A hypothesis of heterozygote excess can be formulated; however, a heterozygote deficit is observed in the MFC3 and MFC8 loci (H obs < H exp). For the MFC7 locus, observed and calculated heterozygosity are similar. Taking into account all the loci tested, the cultivars are in Hardy–Weinberg equilibrium, and values for observed and expected heterozygosity are similar (0.706 and 0.721, respectively).
To confirm our previous assumptions, a statistical test was applied and the Hardy–Weinberg equilibrium hypothesis tested for every locus using F-statistics. The null hypothesis is accepted only for the MFC7 locus (F IS = −0.308; P > 0.05) and discarded for all others. This deviation from the equilibrium for the majority of loci can be explained by the vegetative propagation system of Ficus carica.
The fig samples have a significant heterozygote excess for the MFC2 (F IS = −0.469; P < 0.05) and MFC6 loci (F IS = −0.296; P < 0.05), but not for the MFC5 locus (F IS = −0.389; P > 0.05). A significant deficit is observed for the MFC3 (F IS = +0.356; P < 0.05) and MFC8 loci (F IS = +0.503; P < 0.05). Hardy–Weinberg equilibrium (F IS = −0.0361; P > 0.05) is maintained over all loci analyzed (Table 3).
Genetic Diversity Analysis
The scored genetic distances, based on Chakraborty and Jin (1993) (DAS), varied from 0.00 to 1.00, with a mean of 0.59 (Fig. 2). The smallest value was observed between the Soltani2 and Kahli1 cultivars which have the same alleles for all tested loci, showing a maximum of similarity at the DNA level.
The dendrogram based on the scored genetic distances (Fig. 3) clustered the genotypes into two main groups. The first (a) has two subgroups: a-1 is composed of two pollinators, and a-2 includes the Besbessi and Goutti cultivars. The second group (b) also has two subgroups: b-1 is composed of Bither Abiadh2 and Bither Abiadh3, and b-2 is composed of the remaining cultivars in this study. The observed tree branches grouped cultivars with the same name (e.g., Hemri1, Hemri2, Bidhi1, Bidhi2, Bither Abiadh2, Bither Abiadh3, Soltani1, and Soltani2). The strong association between the Soltani 2 and Kahli1 cultivars, however, suggests that they are characterized by large similarities at the DNA level and might correspond to the same genotype based on SSR. It is important to note that these cultivars are clearly discriminated by the combined data (Fig. 3).
In addition, correspondence factorial analysis was used to refine details of the genetic diversity. The distribution of fig cultivars in plan 1–2 (40.29% of the global genetic diversity) showed agglomeration of cultivars. Nevertheless, we observed significant detachment of some cultivars: pollinators Jrani and Assafri, Bither Abiadh2, Bither Abiadh3, Besbessi, and Goutti. In addition, the Soltani2 and Kahli1 cultivars are very close (Fig. 4).
Biplot of correspondence factorial analysis based on SSR markers. Axis 1, first dimension. Axis 2, second dimension. Data points (+) represent fig cultivars as numbered in Table 1
Identification Key
To establish a cultivar identification key, we took into account each cultivar’s multilocus fingerprint. In total, 31 genotypes were identified in our study. It is important to note that these loci were hierarchically classified according to the provided genotypes. The precise diagram was then drawn to discriminate 15 cultivars, among 17 (88%), according to their SSR profiles (Fig. 5).
Correlation Coefficient Between Matrices
Positive and negative correlations were estimated for genetic matrix combinations. Low correlations calculated between SSR and RAMPO, SSR and ISSR, and RAPD and ISSR matrices support the utility of these markers (Table 5; Fig. 6).
Discussion
Our results demonstrate that it is feasible to discriminate Tunisian fig cultivars using molecular markers, to detect problems of appellation (homonymy and synonymy), and to characterize the genetic diversity of local fig resources. All the techniques used here successfully discriminated between the cultivars maintained ex situ and housed in the Chott Meriam collection. Clustering and multivariate analyses suggest that a typically continuous genetic diversity characterizes the Tunisian fig germplasm with a common genetic basis. This finding confirms results obtained by Chatti et al. (2007), Saddoud et al. (2007), and Salhi Hannachi et al. (2004, 2005, 2006) using dominant and codominant markers found in other fig cultivars kept in situ and ex situ in north and south Tunisia.
The identification key permitted the unambiguous discrimination of the cultivars and confirmed the reliability of microsatellites for fingerprinting fig genotypes, except for the Soltani2 and Kahli1 cultivars. A similar result was obtained for north and south Tunisian figs, conserved ex situ and in situ. Microsatellite multilocus genotyping can distinguish unambiguously 70 well-defined cultivars with a resolving power of 97% (Saddoud et al. 2007). Taking into consideration all the tested loci, significant differences are detected between pollinators and domesticated fig trees. Our results showed that all the alleles are current in domesticated fig tree patterns, except for alleles 134 and 176 of the MFC7 and MFC2 loci, respectively, which are present only in the pollinator genotypes.
In addition, the mean genetic distances were relatively high, suggesting a large variability at the DNA level of the Tunisian fig germplasm. Therefore, the topology of the UPGMA dendrograms shows clustering independent of the origin of cultivars and of their botanical types.
On the other hand, positive and negative correlations were estimated for genetic matrix combinations. Low correlations were calculated between SSR and RAMPO, SSR and ISSR, and RAPD and ISSR matrices, demonstrating the utility of these markers. A low correlation (s = 0.0186) was also found between RAPD and ISSR genetic distances by Salhi Hannachi et al. (2005) for fig cultivars from south Tunisia. As suggested by Fig. 5, SSR markers generate the largest genetic distances in comparison with other markers used. Our results demonstrate that these markers are useful to describe diversity. The applied assays are informative in genotyping and discrimination of fig cultivars. These markers have a higher effective resolving power that revealed their higher relative information content. This result suggests that the tested markers are the best choice for the evaluation of diversity and for assessing genetic relationships between fig cultivars. Obviously the number of primer combinations or the cultivars must be increased to gain a deeper insight into genetic diversity in this fruit crop.
In addition, the methods used are suitable for discrimination of ecotypes and, therefore, for the resolution of appellation problems. This assumption is strongly supported since cultivars with similar names are characterized by a great divergence, exemplified by the case of Soltani, Hemri, and Bidhi cultivars. On the other hand, SSR markers permit the resolution of the problem of synonymy not illuminated by RAMPO (Chatti et al. 2007) or the RAPD technique. In this study, the combination of RAPD, ISSR, and RAMPO markers was able to discriminate between the closely clustered Soltani2 and Kahli1 cultivars using SSR.
Several studies have used molecular markers to measure genetic diversity and to characterize fig cultivars from Mediterranean regions. RAPD markers were shown to be useful for distinguishing between Turkish cultivated fig clones, as reported by Cabrita et al. (2001). Akbulut et al. (2009) demonstrated the utility of RAPD markers in the analysis of genetic variation and genetic relationships among Turkish wild fig. As reported in another study, RAPD markers were used to analyze genetic diversity in the Jordan fig, and the data illustrated variability in the genetic pool of the local common fig, as a valuable source for incorporation into potential breeding programs (Sadder and Ateyyeh 2006). Compared with the results obtained here, an earlier study showed RAPD markers to be powerful enough to differentiate even between clones of the same fig cultivars (Papadopoulou et al. 2002). European and Asian fig varieties were fingerprinted by ISSR, RAPD, and SSR markers, suggesting that the genetic diversity of the fig population is low and that the use of multiple markers is critical to estimate the relatedness of figs at the variety level (Hidetoshi et al. 2009). Similar results were obtained with complementary SSR and ISSR approaches, using combined data to discriminate among five denominations of Morocco fig accessions (Oukabli and Khadari 2005). The results appear to be useful for the establishment of a reference collection, and the authors suggest that among the identified polyclonal fig varieties, a reference genotype must have the most desirable pomological traits. The establishment of a reference collection can overcome varietal heterogeneity. ISSR and SSR markers constitute the first database for a reference collection in northern Morocco. Both molecular markers and pomological traits distinguished genotypes with the same name (Khadari et al. 2005), since cases of mislabeling and homonymy were detected. Similarly, Khadari et al. (2003) described genetic parameters of SSR loci and characterized Mediterranean fig cultivars to establish a representative collection of cultivated fig genetic variability in the Mediterranean area.
The number of polymorphisms was 23.15 for the 10 ISSR primers, 27.13 for the 15 RAPD primers, and 26.96 for the 18 RAMPO combined primers, which was lower than described for Asian figs (Rp 18.74 in 13 ISSR primers, Rp 48.42 in 19 RAPD primers, and H obs 0.44 in SSR) and for other Tunisian cultivars (Rp 29.65 in 4 ISSR primers, Salhi Hannachi et al. 2004; Rp 62.28 in 7 RAPD primers, Salhi Hannachi et al. 2005). The observed heterozygosity calculated in SSR (0.706) is higher overall than that obtained for Asian (0.44) and Spanish (0.47) figs (Ikegami et al. 2009; Giraldo et al. 2005). A similar value (0.7) was calculated for Tunisian figs of south and north Tunisia by Saddoud et al. (2007). Three DNA markers (RAPD, ISSR, and SSR) were applied for a more comprehensive and precise estimation of the unknown genetic background, structure, relationships, and diversity of 19 fig varieties distributed in Japan by Ikegami et al. (2009); the dendrogram topologies showed no correlation with geographic origin. Similar results were obtained for Turkish figs by Ikten et al. (2010); geographic- and color-based groups were not genetically distinct among the Turkish figs. No correlations were found between similarity matrices, and results demonstrate that each method is useful and informative for evaluating genetic diversity. In fact, each marker represents only a part of the genome in the incongruence of the dendrogram. These results suggest that the use of multiple markers is critical to estimate the relatedness of figs at the variety level, because different markers engender different classification results based on their respective characters, as reported by Ikegami et al. (2009). On the other hand, Rout and Aparajita (2009) reported that 107 ISSR markers were used to characterize and assess phylogenetic relationships among 23 Ficus accessions in eastern India. Results indicate that the genetic relationships among the Ficus accessions were in accordance with their morphological characters and confirm the presence of accessions in the natural distribution of parental species.
In view of the economic importance of the fig, fig cultivars have been the subject of many studies, particularly evaluations of genetic diversity. This study is an important step that ensures the conservation and the development of a national database. On the whole, our data concur with results based on morphological and molecular analysis reported in fig germplasm. Consequently, it may be assumed that this study provides evidence of the effectiveness of the SSR procedure, either for the identification of cultivars or for the assessment of genetic diversity. In addition, ISSR and RAPD markers are useful tools to detect polymorphism and to discriminate fig cultivars, since a low and negative correlation was found. Thus, all tested genotypes were significantly divergent at the DNA level and ought to be preserved in a reference collection. The aggregated molecular markers can also be used to discriminate fig cultivars in the case of genotypes sharing a narrow genetic basis and to verify the distinctness between the genetically close fig accessions. Based on the scored multilocus genotypes, a cultivar identification key was established that should be useful in the account, listing, and certification of trees as well as in the rational management of Tunisian fig germplasm and ex situ conservation strategies. These results are of particular interest given the current and future use of local cultivars in improving production, diversification, and food security.
References
Akbulut M, Ercisli S, Karlidag H (2009) RAPD-based study of genetic variation and relationships among wild fig genotypes in Turkey. Genet Mol Res 8(3):1109–1115
Belkhir K (2000). Genetix version 4.01. Laboratoire Génome et Populations, CNRS UPR 9060, Université de Montpellier
Berg CC (2003) Flora melesiana precursor for the treatment of Moraceae 1: the main subdivision of Ficus: the subgenera. Blumea 48:176–178
Cabrita LF, Aksoy U, Hepaksoy S, Leitão JM (2001) Suitability of isozyme, RAPD and AFLP markers to assess genetic differences and relatedness among fig (Ficus carica L.) clones. Sci Hortic 87:261–273
Chakraborty R, Jin L (1993) A unified approach to study hypervariable polymorphisms: statistical considerations of determining relatedness and population distances. In: Chakraborty R, Epplen JT, Jefferys AJ (eds) DNA fingerprinting: state of the science. Birkhäuser Verlag, Basel, pp 153–175
Chatti K, Salhi Hannachi A, Mars M, Marrakchi M, Trifi M, (2004a) Analyse de la diversité génétique de cultivars tunisiens de figuier (Ficus carica L.) à l’aide de caractères morphologiques. Fruits 59(1):49–61
Chatti K, Salhi Hannachi A, Mars M, Marrakchi M, Trifi M (2004b) Genetic diversity and phylogenic relationships in Tunisian fig (Ficus carica L.) cultivars mediated by RAPD. Biologia Tunisie 1(2):1–4
Chatti K, Saddoud O, Salhi Hannachi A, Mars M, Marrakchi M, Trifi M (2007) Inferring of genetic diversity and relationships in a Tunisian fig (Ficus carica L.) germplasm collection by random amplified microsatellite polymorphisms. J Integr Plant Biol 49:386–391
Dellaporta SL, Wood J, Hicks JB (1984) Maize DNA miniprep. In Malmberg R, Messing J, Sussex I (eds) Molecular biology of plants. Cold Spring Harbor Laboratory, Cold Spring Harbor Press, New York, pp 36–38
Dieringer D, Schlötterer C (2003) Microsatellite analyzer (MSA): a platform independent analysis tool for large microsatellite data sets. Mol Ecol Notes 2:1–3
Felsenstein J (1995) Phylip (Phylogeny Interference Package) version 3.5 c. Department of Genetics, University of Washington, Seattle
Gilbert JE, Lewis RV, Wilkinson MJ, Caligari PDS (1999) Developing an appropriate strategy to assess genetic variability in plant germplasm collections. Theor Appl Genet 98:1125–1131
Giraldo E, Viruel MA, López-Corrales M, Hormaza JI (2005) Characterization and cross-species transferability of microsatellites in the common fig (Ficus carica L.). J Hortic Sci Biotechnol 80:217–224
Hedfi J, Trifi M, Salhi Hannachi A, Rhouma A, Marrakchi M (2003) Morphological and isoenzymatic polymorphism in Tunisian fig (Ficus carica L.) collection. Acta Hortic 605:319–325
Hidetoshi I, Hitoshi N, Keita H, Mitsuo A, Takao N (2009) Analysis of genetic diversity among European and Asian fig varieties (Ficus carica L.) using ISSR, RAPD, and SSR markers. Genet Resour Crop Evol 56(2):201–209
Ikegami H, Nogata H, Hirashima K, Awamura M, Nakahara T (2009) Analysis of genetic diversity among European and Asian fig varieties (Ficus carica L.) using ISSR, RAPD, and SSR markers. Genet Resour Crop Evol 56:201–209
Ikten H, Mutlu N, Gulsen O, Kocatas H, Aksoy U (2010) Elucidating genetic relationships, diversity and population structure among the Turkish female figs. Genetica 138:169–177
Khadari B, Lashermes PH, Kjellberg F (1995) RAPD fingerprints for identification and genetic characterization of fig (Ficus carica L.) genotypes. J Genet Breed 49:77–86
Khadari B, Hochu I, Santoni S, Kjelberg F (2001) Identification and characterization of microsatellite loci in the common fig (Ficus carica) and representative species of the genus Ficus. Mol Ecol Notes 1:191–193
Khadari B, Hochu I, Bouzid L, Santoni S, Roger JP, Kjelberg F (2003) The use of microsatellite markers for identification and genetic diversity evaluation of the fig collection in CBNMP. Acta Hortic 605:77–86
Khadari B, Oukabli A, Ater M, Mamouni A, Roger JP, Kjellberg F (2005) Molecular characterization of Moroccan fig germplsam using intersimple sequence repeat and simple sequence repeat markers to establish a reference collection. Hortscience 40:29–32
Kjelberg F, Gouyon PH, Ibrahim M, Raumond M, Valdeyron G (1987) The stability of the symbiosis between dioecious figs and theirs pollinators: a study of Ficus carica L. and Blastophaga Psenes L. Int J Org Evol 41(4):693–704
Mars M, Carraut A, Marrakchi M, Gouiaa M, Gaaliche F (1994) Ressources génétiques fruitières en Tunisie (poirier, oranger, figuier, grenadier). Plant Genet Resour Newsl 100:14–17
Mars M, Marrakchi M, Chelbi T (1998) Multivariate analysis of fig (Ficus carica L.) germplasm southern Tunisia. Acta Hortic 480:75–81
Nei M (1978) Estimation of average heterozygosity and genetic distance from small number of individuals. Genetics 89:583–590
Nei M, Li WH (1979) Mathematical model for studying genetical variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 74:5267–5273
Oukabli A, Khadari B (2005) Caractérisation des variétés polyclonales marocaines de figuiers, Ficus carica L. Fruits 60:47–54
Page RDM (1996) Treeview: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Papadopoulou K, Ehaliotis C, Tourna M, Kastanis P, Karydis I, Zervadis G (2002) Genetic relatedness among dioecious Ficus carica L. cultivars by random amplified polymorphic DNA analysis, and evaluation of agronomic and morphological characters. Genetica 114:183–194
Prevost A, Wilkinson MJ (1999) A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98:107–112
Rhouma A (1996). Les ressources phytogénétiques oasiennes: le figuier (Ficus carica L.). In: Proceedings of the 3èmes Journées Nationales sur les Acquis de la Recherche Agronomique, Vétérinaire et Halieutique, Nabeul, p 29
Rout GR, Aparajita S (2009) Genetic relationships among 23 Ficus accessions using inter simple sequence repeat markers. J Crop Sci Biotechnol 12:91–96
Sadder MT, Ateyyeh AF (2006) Molecular assessment of polymorphism among local Jordanian genotypes of the common fig (Ficus carica L.). Sci Hortic 107:347–351
Saddoud O, Chatti K, Salhi Hannachi A, Mars M, Rhouma A, Marrakchi M, Trifi M (2007) Genetic diversity of Tunisian figs (Ficus carica L.) as revealed by nuclear microsatellites. Hereditas 144:149–157
Salhi Hannachi A, Trifi M, Zehdi S, Hedfi J, Mars M, Rhouma A, Marrakchi M (2004) Inter simple sequence repeat fingerprints to assess genetic diversity in Tunisian fig (Ficus carica L.) germplasm. Genet Resour Crop Evol 51(3):269–272
Salhi Hannachi A, Chatti K, Mars M, Marrakchi M, Trifi M (2005) Comparative analysis of genetic diversity in two Tunisian collections of fig cultivars based on random amplified polymorphic DNA and inter simple sequence repeats fingerprints. Genet Resour Crop Evol 52(5):563–573
Salhi Hannachi A, Chatti K, Saddoud O, Mars M, Rhouma A, Marrakchi M, Trifi M (2006) Genetic diversity of different Tunisian fig (Ficus carica L.) collections revealed by RAPD fingerprints. Hereditas 143:15–22
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor, New York
Sneath PMA, Sokal RR (1973) Numerical taxonomy. W.H. Freeman and Company, San Francisco
StatSoft, Inc (2001) Statistica (data analysis software systems), version 6. StatSoft, Inc., Tulsa. www.statsoft.com
Storey WB (1975) Figs. In: Janick J, Moore J (eds) Advances in fruit breeding. Purdue University Press, Indiana, pp 568–589
Weiblen GD (2000) Phylogenetic relationships of functionally diocious Ficus (Moraceae) based on ribosomal DNA sequences and morphology. Am J Bot 87(9):1342–1357
Weir BS, Cockerham C (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Zehdi S, Trifi M, Ould Mohamed Salem A, Rhouma A, Marrakchi M (2002) Survey of inter simple sequence repeat (ISSR) in Tunisian date-palms (Phoenix dactylifera L.). J Genet Breed 56:77–83
Zehdi S, Trifi M, Billotte N, Marrakchi M, Pintaud JC (2004) Genetic diversity of Tunisian date palms (Phoenix dactylifera L.) revealed by nuclear microsatellite polymorphism. Hereditas 141(3):278–287
Zohary D, Hopf M (2000) Domestication of plants in the old world, 3rd edn. Oxford University Press, Oxford
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chatti, K., Baraket, G., Abdelkrim, A.B. et al. Development of Molecular Tools for Characterization and Genetic Diversity Analysis in Tunisian Fig (Ficus carica) Cultivars. Biochem Genet 48, 789–806 (2010). https://doi.org/10.1007/s10528-010-9360-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-010-9360-1