Abstract
Common fig (Ficus carica L.) is one of the most ancient domesticated species, originated, supposedly in Arabia, from where it diffused to the Middle East and Asia, and to the Mediterranean basin, where it greatly diffused. More than 600 fig varieties have been described, but it is conceivable that this number is underestimated. Along all the Italian territory, there is a rich germplasm of fig composed of a large number of varieties (approximately 300) of very not well defined origin. Effectively, during several centuries of cultivation and propagation by seed, a large number of genotypes appeared and were selected, leading to the generation of an uncountable number of genotypes, different in numerous traits, particularly in those related to leaves and fruits features. Unfortunately, the extensive existing fig genetic patrimony is facing genetic erosion; for this reason, it is extremely important to study and valorised it, in order to preserve the remaining biodiversity. The purpose of this study was to genetically characterize, with nSSR markers, 79 fig accessions, collected in several areas in Italy. The set of chosen markers resulted highly polymorphic, and allowed the characterization of all the studied accessions. Data were analysed by cluster analysis, and the results demonstrated a great genetic variability within the population. The nSSR used, moreover, allowed us to identify all accessions and to recognised possible homonyms and synonyms, and cases of intravarietal clones.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Common fig (Ficus carica L.), with olive tree, grape and palm, is one of the most ancient domesticated species (Mawa et al. 2013). Common fig has originated, supposedly, in Arabia (Storey 1976), from where it diffused to the Middle East and Asia, and to the Mediterranean basin, where it greatly diffused. Therefore, many authors consider the East Mediterranean region the area of the common fig domestication (Ercisli et al. 2012), and from there the cultivation spread to the West Mediterranean area, where fig populations were already present in natural habitats before domestication (Veberic and Mikulic-Petkovsek 2016). The process of domestication resulted in sweeter and bigger fruits (Falistocco 2009). Approximately, worldwide, more than 600 fig varieties have been described (Condit 1955), but it is conceivable that this number is underestimated. Unfortunately, changes in alimentary habits, the lack of intensive system of fig cultivation and the short fig shelf life, caused the downfall of fig cultivation. In the Mediterranean basin, fig biodiversity has been preserved, because its cultivation is merely based on the use of autochthonous genotypes. In Italy, fig germplasm consists of a large number of varieties (approximately 300) of not well defined origin (Grassi 1998; Barberis et al. 2001; Chessa et al. 2001; Minonne et al. 2001), diffused, primarily, in four regions of southern Italy (Campania, Calabria, Puglia and Sicily), and in Tuscany. Particularly, archeobotanical researches, combined with historical sources, date the presence of the fig tree from the Neolithic, through the Roman Empire to the Middle Ages (Turfa 2012; Mariotti Lippi et al. 2009; Rattighieri et al. 2013; Buonincontri et al. 2014; Mariotti Lippi et al. 2015) and further on; as a matter of fact, historical references, found in XIII–XVI century documents, report the names of some varieties of fig, Dottato, Verdino, Brogiotto Nero and Brogiotto Bianco, cultivated in Tuscany (Baldini 1953). Even though, in Italy, fig cultivation has a long tradition and its germplasm is still wide, with more than 90 genotypes (Baldini 1953; Basso 1960a, b), information on their genetic identity are scarce and dubious, making their cataloguing very difficult. Therefore, the establishment strategies for the preservation of the local fig germplasm is a necessity. For this reason, for example, in Tuscany, “Slow food”, a global organization to prevent the disappearance of local food cultures and traditions, individuated the “dry fig of Carmignano” as a typical regional production to be valorized and preserved. The scientific community, instead, is committed to characterize the existing fig germplasm. To assess the diversity among fig cultivars, different methods have been utilized, from morphological to biochemical and genetic characterizations. Morphological descriptors were used to evaluate and characterize fig cultivars in the Mediterranean area (Papadopoulou et al. 2002; Gaaliche et al. 2012; Çalişkan and Polat 2012; Giraldo et al. 2010) to determine diversity in different accessions of F. carica L.
Biochemical markers, such as isozymes (Cabrita et al. 2001), and DNA markers, such as mitochondrial DNA-RFLP (Khadari et al. 2005), AFLP (Laddomada et al. 2008; Aradhya et al. 2010), and RAPD (Khadari et al. 1995; Chessa et al. 2001; De Masi et al. 2003; Ciarmiello et al. 2015) have been used to characterize fig varieties, revealing high polymorphism among fig accessions from different Mediterranean areas. Genomic microsatellite markers have been developed for common fig tree in recent years (Khadari et al. 2003; Giraldo et al. 2005, 2008; Achtak et al. 2009; Ferrara et al. 2016). Khadari et al. (2001), identified 8 SSR primers on different varieties of F. carica L. and other plants of the genus Ficus, showing the efficiency of the primers on fig varieties, but a partial transferability on other plants of the genus Ficus. The same primers were then utilized on 30 fig varieties from France and Morocco (Khadari et al. 2003), and 5 of the 8 primers were selected for the characterization of the entire population, considering their highest discrimination power. Giraldo et al. (2005), identified 26 new SSR primers and tested them on 15 fig cultivars from France and Spain; the primers were able to discriminate 9 different genotypes in the 15 accessions. Achtak et al. (2009) individuated, trough selection and validation, 6 SSR primers characterized by high discriminant capacity, useful for the identification of all the genotypes present in a Morocco’s fig collection. In a study of Abou-Ellail et al. (2014), seven fig cultivars were characterized using biochemical and microsatellite markers; the firsts showed different relation among the cultivars, but they were considered not very efficient, because their low polymorphism and because, in some cases, they were influenced by environmental factors; instead, the use of SSR allowed the discrimination of all the 7 cultivars, proving a higher efficiency. Recently, the analysis of simple sequence repeats (SSR or microsatellites) showed good efficiency in the identification and characterization of fig accessions of different origins (Baraket et al. 2011; Khadari 2012; Essid et al. 2015). Knap et al. (2016), developed 16 new SSR markers and evaluated their transferability on other species, with successful results.
The objective of the present work was to describe the genetic variability of common fig germplasm still present in Italy by identifying any genetic similarities, synonyms and homonyms, with the aid of a set of nuclear SSR (nSSR) markers. The study of genetic variability within the germplasm is the best condition for a correct preservation of genetic resources, and for the identification and valorisation of the most suitable genotypes, in view of a recovery of the fig industry, not only in Italy, but all over the world too.
Materials and methods
Plant material
The plant material used in this study was collected in ex situ fig germplasm private collection, located in Pescia (Pistoia, Tuscany, IT) (43°53′13″N, 10°41′18″E, 42 m a.s.l.). The plant collection, propagated by cuttings, is composed by 79 parthenocarpic fig accessions (Table 1). For each accession, leaves from healthy and actively growing shoots, were collected in August, and randomly sampled from different parts of each tree. Samples were placed in airtight plastic bags, immersed in liquid nitrogen, and frozen at − 80 °C, prior to subsequent analysis.
DNA extraction and molecular characterization
Genomic DNA was extracted following the CTAB (Cetyl Trimethylammonium Bromide) procedure (Doyle and Doyle 1987). After the spectrophotometer quantification, sample were analysed with nSSR markers.
For DNA amplification, 7 couples of nSSR dinucleotide primers (MFC and LMFC series), set already for fig characterization for their high level of polymorphism (Khadari et al. 2001; Giraldo et al. 2005), were used (Table 2).
PCR amplification was performed in a final volume of 20 μL containing: 1 × Reaction Buffer (International PBI, Milano, Italy), 0.2 mM dNTPs (Amersham Biosciences, Piscataway, USA), 2 mM MgCl2 (International PBI, Milano, Italy), 0.2 μM primer (MWG Biotech, Ebersberg), 20 ng genomic DNA and 1 U di Taq polymerase (Fisher Molecular Biology, Trevose, USA).
PCR amplification reaction was optimized in thermal cycler MJ PCT 100 Research (Watertown, Mass.), programming a first passage at 94 °C for 1 min followed by 35 cycles of 30 s at 94 °C, 30 s at the specific annealing temperature for each couple of primers (Table 2), and 1 min at 72 °C, for denaturation, annealing and primer extension, respectively; at the end of the cycles were allowed 8 min of incubation at 72 °C.
One of the two PCR primers in each reaction was end-labelled with a fluorescent dye (CY5, MWG Biotech, Ebersberg, Germany). The amplification products were separated with a CEQ 2000 Genetic Analysis System (Beckman Coulter, Inc.) sequencer on acrylamide gel CEQ Separation Gel LPA-1 (Beckman Coulter, Inc.). A marker CEQ DNA Size Standard kit 400 (Beckman Coulter, Inc.) was used to estimate the approximate molecular weight of the amplified products. Two reference samples were used in all runs.
Statistical analysis
Fragments were sized using a conservative binning approach (Kirby 1990), through the statistical R software (R Development Core Team 2005), which takes into account the type of replicate and compensates for the limits of fragment resolution. Genotypes showing a single allele in a given locus were indicated as homozygote.
All analyses were developed after removing duplicates, previously identified by using pair-wise comparisons among all genotypes, based on their multilocus nSSR profile, using an Excel spreadsheet (© Microsoft Corporation).
Data were processed using the software Identity 1.0 (Wagner and Sefc 1999), available at the web site http://www.boku.ac.at/zag/forsch/identity.htm. Identity 1.0 was used for the calculation of the number of alleles per locus (N), allele frequency, observed (Ho) and expected (He) heterozygosity, frequency of null alleles r = (He − Ho)/(1 + He) (Brookfield 1996), probability of genetic identity (PI) (Paetkau et al. 1995).
The number of effective alleles (Ne = 1/(1 − He), Brown and Weir 1983) was calculated using Microsoft Excel spreadsheet (© Microsoft Corporation).
The level of similarity/dissimilarity among examined accessions was obtained through the genetic similarity matrix utilizing Manhattan distance. Cluster analysis and construction of the dendrogram relative to genetic distances were obtained by using the unweighted pair-group method with arithmetic mean (UPGMA) algorithm, with XLSTAT 2009 software (Addinsoft™ 1995–2009).
Results
SSR markers polymorphism
The molecular analysis on 79 accessions was performed using seven SSR markers, belong to the series MFC and LMFC. For a better comprehension of markers characteristics, the number of amplified alleles per locus, frequency, heterozygosity, discrimination power and null allele frequency were observed (Table 3). From the 79 analysed accessions, SSR markers identified 43 alleles (Table 3). Allelic frequencies (f) varied from 0.006 and 0.651; the average expected heterozygosity (He) and observed heterozygosity (Ho) value were 0.664 and 0.670 respectively. The highest expected heterozygosity was reach by locus LMFC30 with 0.810, instead the higher observed heterozygosity was reach by locus LMFC4 with 0.798. The less observed and expected heterozygosity reached the values respectively of 0.456 and 0.520, both for the locus LMFC24.
The number of alleles (No) amplified, ranged from 3 alleles for the primer MFC4, to 9 alleles for LMFC30. The most frequent alleles in the examined population were alleles 278, for the locus LMFC24, with a frequency of 0.652 and 353 for the locus LMFC12, with a frequency of 0.513; the alleles less frequent were alleles 270 for the locus LMFC24 and allele 244 for LMFC30, with a frequency of 0.006.
Microsatellite LMFC30 showed the highest PIC (= 0.778) values (Table 3), reflecting the large number of alleles observed and similar allele frequency distribution in the population when compared to the other microsatellites (Table 3). The lowest PIC was found in microsatellite LMFC24 (= 0.466).
Identification of the examined population and internal relationship
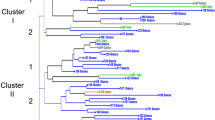
The utilization of the seven SSR markers, on 79 accessions allowed the discrimination and identification of 56 genotypes, of which 12 (named “Group”) consisted of a number of accessions from 2 to 6 (Fig. 1). Genetic profiles of fig trees are reported in Supplementary Data (Table S1). The relationship among all accessions was analysed and observed by cluster analysis (UPGMA) at Euclidean distances, by which dissimilarities between the studied figs are highlighted (Fig. 1). The dendrogram (Fig. 1) includes all the accessions and shows 56 different genotypes and differences can be observed between the accession with a dissimilarity index between 0 and 40,83. The examined population is divided in two main clusters (I and II); the firs cluster includes 1/3 of the identified genotypes, and contains two subgroups, I.1 and I.2. Subgroup I.1 is divided in two sets (I.1.A and I.1.B): set I.1.A is composed by five dark figs (“Pecciolo nero 1”, “Datterino”, “Sanguinella”, “Datto”, “Viola della Turchia”), and three light figs (“Perticone”, “Faraone”, “Cavallierino”); set.I.1.B, is mainly composed by dark figs (“Rocco nero”, “Dattero”, “Pecciolo nero 2”, “Cavaliere”, “Brogiotto nero 2”) and “Brogiotto nero 3”). In subgroup I.2, only the genotype “Corvo” is present.
Cluster II is divided in two subgroups, in which the majority of the studied accessions are included. Subgroup II.1 is composed by the sets II.1.A and II.1.B. Set II.1.A is split in two sub sets; in one sub set, there are two accessions of the genotype “Dottato” (2 and 1), that are positioned near each other (Table S1). In the other sub set (above), it is possible to observe the similarity and genetic nearness between “Dottato” accessions (3, 5, 6, 7, 8) (Group 1) (Fig. 1 and Table S1). Other interesting observations are synonymies, that are accessions named differently, but sharing the same alleles amplicons (Table S1), for example, “Troiano”, “Natalegna” and unknown (Group 2), and the accessions “San Pietro 1” and “Gentile Bianco” (Group 4).
In set II.1.B, it is possible to observe similarities between two accessions named “Brogiotto bianco” (2 and 3), that have different geographic origins, and low genetic differences between “Brogiotto”, and “Brogiotto vero”, considered the same genotypes (Group 6). Moreover, three genotypes with different names (“Seccareccio”, “Paradiso 2”, “Verdino bianco”) share the same microsatellite amplicon (Table S1) and have genetic similarities with “Doro” and “Verdino” (Group 5). Other synonymies are observed in the accessions “Montalcino rosa” and “Cerreto” and “Trapani” and “Montecarlo” (Group 8 and 9) (Fig. 1 and Table S1).
Discussions
The seven primer pairs used in this study (Table 2) were selected from a set of 8 microsatellites identified and developed by Khadari et al. (2001) and from a set of 26 primers developed by Giraldo et al. (2005). For the selection of primers, the number of alleles amplified by the markers, the polymorphism and the results of amplification in other studies were considered (Saddoud et al. 2007; Giraldo and López-Corrales 2008; Aradhya et al. 2010; Saddoud et al. 2011; Abou-Ellail et al. 2014; Ferrara et al. 2016). Molecular analysis enables the identification of the genetic distances in the Tuscany fig population, allowing the observation of a rich biodiversity. SSR markers, together with other molecular methods, were previously used in the study of fig biodiversity in France (Khadari et al. 2001; Khadari 2012), Spain (Giraldo et al. 2005), Tunisia (Saddoud et al. 2007), in Brazil (do Val et al. 2013) and in different Mediterranean countries (Khadari 2012).
Giraldo et al. (2005) analysed 15 fig cultivars from different geographical areas, using 26 SSR markers, and obtained low value of observed heterozygosity (0.47); the data was endorsed by the easiness in agamic propagation of fig, by the narrow genetic base and by the small genetic variability in the studied population. However, results reported in this study on Italian fig biodiversity show a good level of polymorphism and genetic variability, with a higher average heterozygosity (0.67) (Table 3). The rescored value is, instead, in accord with a prior study of 72 Tunisian fig genotypes (Saddoud et al. 2007), where the observed heterozygosity reached the value of 0.7. The lower value of observed heterozygosity in the population studied in this research, compared to expected heterozygosity in four loci, is in accord with the study of Saddoud et al. (2007, 2011). In the set of primers used in this study, primers LMFC30 and MFC3 showed the best results in terms of polymorphism, amplifying 9 and 8 alleles respectively. These data are in accord with prior studies of Achtak et al. (2009) and Aradhya et al. (2010), in which the amplification of the same number of alleles for the considered primers is shown. As concern observed heterozygosity (Ho), the highest level was reached by the locus MFC4 with 0.798, instead the major expected heterozygosity (He) was found in locus LMFC30, with a value of 0.810 (data in partial accord with the study of Giraldo et al. 2005). Prior study (Khadari et al. 2003) showed low value of heterozygosity for the locus MFC4 (0.375 in 14 accessions analysed from Conservatoire Botanique National Méditerranéen de Porquerolles collection, and 0.560 in 16 Moroccan genotypes analysed); instead, locus MFC4, showed high level of heterozygosity in the study of Aradhya et al. (2010) on 194 fig genotypes, reaching the value of 0.818; the higher value registered in this study, was probably due to the wide number of fig samples analysed and to the worldwide origin of the accessions.
In this study, PIC value was higher than 0.5 for all loci analysed (data in accord with Ferrara et al. 2016), except for the locus LMFC24, that turned to be the less informative; instead, the locus LMFC30, with 0.7783 achieved the highest value. From the allelic analysis it is possible to observe the presence of unique alleles (f = 0.0063), thus typical of single genotypes, like allele 270 and 284 (locus LMFC24), specific for the genotype “Paradiso 1” and “Dottato 2” respectively; allele 401 (locus LMFC12) specific for “Corvo”; allele 244 (LMFC30), for “Siro” genotype; the presence of unique allele is not shown for the locus of the series MFC. The existence in the considered population of unique alleles could be a starting point for the univocal discrimination of genotypes (Jakše et al. 2004), originated in a restricted area of Italy.
Dendrogram observations, based on the differences in their allelic profile, allowed the separation of the analysed accessions in different groups (Fig. 1). Genetic structure does not show particular grouping, probably due to narrow genetic base from which the fig were domesticated. From the analysis of the population, it was not possible to discriminate between common fig (partenocarpic) and “San Pedro” fig type, as was already partially observed by Ferrara et al. (2016). Instead, our results are in accord with other studies (Saddoud et al. 2007; Baraket et al. 2011) in which correlation was not highlighted. Nevertheless, it is possible to observe in the I cluster, the presence almost exclusive of dark figs, thus showing not only phenotypic but also genetic similarity, even if some other dark varieties are distributed along the dendrogram.
The observation of the allelic profile and the dendrogram, allowed the identification of cases of homonymy, synonymy and possible denomination mistakes that caused varietal confusion (Giraldo and López-Corrales 2008). Of the four accessions named “Brogiotto nero”, only two have relevant similarities (“Brogiotto nero” 2 and 3) (cluster I dendrogram), instead, the other two accessions possess different allelic profiles and are clustered in genetically distant groups; for this reason, this can be considered wrong denomination. The same result is shown for the two “Paradiso” accessions analysed: “Paradiso 2” showed the same allelic profile of the accession “Seccareccio” (case of synonymy) (Table S1), instead, “Paradiso 1” is situated in a different cluster in the dendrogram, showing important genetic differences if compared with “Paradiso 2” (Table S1). Another particular case concerns the 7 analysed accessions of “Dottato” cultivar (Fig. 1), coming from different cultivated areas: five of them showed genetic proximity (II.1.A in dendrogram, Group 1), instead, the other two accessions (“Dottato” 1 and 2), are positioned in another subgroup, showing an additional case of wrong nomination. Case of synonymy is shown for the cultivars “San Pietro 1” and “Gentile bianco” (Group 4), and homonymy is shown between San Piero 1 and “San Piero 2” (Fig. 1 and Table S1). The unknown cultivar was probably recognized: it possess the same allelic profile of the two synonymy “Natalegna” and “Troiano” (Group 2). The reason of the varietal confusion and the presence of homonymies and synonymies is probably due to the ancient origin of fig cultivars and migrations, so that, during century figs cultivar were renamed differently (Aradhya et al. 2010); nevertheless, wrong naming by growers, mistake during propagation or errors in labelling, are other possible hypothesis. Denomination errors and varietal confusion were yet observed in fig population by Aradhya et al. (2010) and Giraldo and López-Corrales (2008), where fig coming from germplasm banks were studied and characterized. Moreover it is known that in fig, as in other ancient species, the lack of varietal standards has led, with the time, to the formation of a heterogeneous population of clones, that increased the confusion in the classification within the species (Condit 1955). In our study, some accessions showed a less than 10% genetic dissimilarity (group 1, 5, 10, 11 and 12), due to minor genetic differences between the allelic profiles; in accord to several Authors (Cipriani et al. 2002; Hocquigny et al. 2004; Beghé et al. 2013), such small genetic discrepancies might have been originated by somatic mutations, which frequently occur in long vegetatively propagated species. Accessions with such characteristics, therefore, might be considered as heterogeneous clones of the same cultivar (polyclonal cultivar).
Conclusion
Seventy-nine Italian accessions, collected in a collection field in Tuscany, were studied, using SSR markers, allowing the individuation of 56 different genotypes. Moreover, thanking the allelic analysis, homonymies and synonymies were individuated, making a step forward in the characterization of the germplasm collection. In the present study, it was possible to identify a cluster, in which the majority of dark figs are contained, but the primers set utilized does not recognized all dark figs accessions; however, phenotypic characters, such as peel pigmentation, are probably controlled by simple Mendelian gene and may be unrelated to molecular markers.
The present study permit the detection of identity errors in figs denomination and allowed the creation of a databank in which, Tuscany and Italian fig genotypes are included. This result will help a univocal varietal identification and the evaluation of the real fig biodiversity. In addition to the genetic analyses, to better characterize and valorise Italian fig germplasm, morphological and agronomical analyses could be integrate in further studies.
References
Abou-Ellail M, Mahfouze SA, El-Enay MAM, Mustafa NSA (2014) Using biochemical and simple sequence repeats (SSR) markers to characterize (Ficus carica L.) cultivars. World Appl Sci J 29(3):313–321. https://doi.org/10.5829/idosi.wasj.2014.29.03.13835
Achtak H, Oukabli A, Ater M, Santoni S, Kjellberg F, Khadari B (2009) Microsatellite markers as reliable tools for fig cultivar identification. J Am Soc Hortic Sci 134(6):624–631. https://doi.org/10.1007/s10681-010-0286-9
Aradhya MK, Stover E, Velasco D, Koehmstedt A (2010) Genetic structure and differentiation in cultivated fig (Ficus carica L.). Genetica 138:681–694. https://doi.org/10.1007/s10709-010-9442-3
Baldini E (1953) Alcuni aspetti della coltura del fico nella provincia di Firenze. Riv Ortoflorfrutt Ital 7–8:185–203
Baraket G, Chatti K, Saddoud O, Abdelkarim AB, Mars M, Trifi M, Hannachi AS (2011) Comparative assessment of SSR and AFLP markers for evaluation of genetic diversity and conservation of fig, Ficus carica L., genetic resources in Tunisia. Plant Mol Biol Rep 29(1):171–184. https://doi.org/10.1007/s11105-010-0217-x
Barberis A, Chessa I, Nieddu G (2001) Analisi multivariata dei descrittori primari del germoplasma di fico della Sardegna. Italus Hortus 8(5):12–15
Basso M (1960a) Contributo allo studio della coltura del fico nella provincia di Pisa. Riv Ortoflorfrutt Ital 3–4:1–17
Basso M (1960b) Contributo allo studio della coltura del fico nella provincia di Livorno. Riv Ortoflorfrutt Ital 5–6:194–208
Beghé D, Ganino T, Dall’Asta C, Silvanini A, Cirlini M, Fabbri A (2013) Identification and characterization of ancient Italian chestnut using nuclear microsatellite markers. Sci Hortic 164:50–57. https://doi.org/10.1016/j.scienta.2013.09.009
Brookfield JFY (1996) A simple new method for estimating null allele frequency from heterozygote deficiency. Molecular Ecology 5(3):453-455. https://doi.org/10.1046/j.1365-294X.1996.00098x
Brown AHD, Weir BS (1983) Measuring genetic variability in plant populations. In: Tanksley SD, Orton PA (eds) Isozymes in plant genetics and breeding. Elsevier Science Publications, Amsterdam
Buonincontri M, Moser D, Allevato E, Basile B, Di Pasquale G (2014) Farming in a rural settlement in central Italy: cultural and environmental implications of crop production through the transition from Lombard to Frankish influence (8th–11th centuries A.D.). Veg Hist Archaeobot 23:775–788. https://doi.org/10.1007/s00334-013-0429-8
Cabrita LF, Aksoy U, Hepaksoy S, Leitão JM (2001) Suitability of isozyme, RAPD and AFLP markers to assess genetic differences and relatedness among fig (Ficus carica L.) clones. Sci Hortic 87:261–273. https://doi.org/10.1016/S0304-4238(00)00181-3
Çalişkan O, Polat AA (2012) Morphological diversity among fig (Ficus carica L.) accessions sampled from the Eastern Mediterranean Region of Turkey. Turk J Agric For 36:179–193. https://doi.org/10.3906/tar-1102-33
Chessa I, Erre P, Nieddu M, Satta D, Nieddu G (2001) Applicazione di marcatori molecolari RAPD in una collezione di germoplasma sardo di fico (Ficus carica L.). Italus Hortus 8(5):16–19
Ciarmiello LF, Piccirillo P, Carillo P, De Luca A, Woodrow P (2015) Determination of the genetic relatedness of fig (Ficus carica L.) accessions using RAPD fingerprint and their agro-morphological characterization. S Afr J Bot 97:40–47. https://doi.org/10.1016/j.sajb.2014.11.012
Cipriani G, Marrazzo MT, Marconi R, Cimato A, Testolin R (2002) Microsatellite markers isolated in olive (Olea europaea L.) are suitable for individual fingerprinting and reveal polymorphism within ancient cultivars. Theor Appl Genet 104(2–3):223–228. https://doi.org/10.1007/s001220100685
Condit IJ (1955) Fig varieties: a monograph. Calif Agric 23(11):323–538
De Masi L, Cipollaro M, Di Bernardo G, Galderisi U, Galano G, Pavone E, Grassi G, Simeone A, Cascino A (2003) Clonal selection and molecular characterization by RAPD analysis of the fig (Ficus carica L.) “Dottato” and “Bianco del Cilento” cultivars in Italy. Acta Hortic 605:65–68
do Val ADB, Souza CS, Ferreira EA, Salgado SML, Pasqual M, Cançado GMA (2013) Evaluation of genetic diversity in fig accessions by using microsatellite markers. Genet Mol Res 12(2):1383–1391. https://doi.org/10.4238/2013.April.25.9
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Ercisli S, Tosun M, Karlidag H, Dzubur A, Hadziabulic S, Aliman Y (2012) Color and antioxidant characteristics of some fresh fig (Ficus carica L.) genotypes from Northeastern Turkey. Plant Foods Hum Nutr 67:271–276. https://doi.org/10.1007/s11130-012-0292-2
Essid A, Aljane F, Ferchichi A, Hormaza JI (2015) Analysis of genetic diversity of Tunisian caprifig (Ficus carica L.) accessions using simple sequence repeat (SSR) markers. Hereditas 152:1. https://doi.org/10.1186/s41065-015-0002-9
Falistocco E (2009) Presence of triploid cytotypes in the common fig (Ficus carica L.). Genome 52:919–925. https://doi.org/10.1139/g09-068
Ferrara G, Mazzeo A, Pacucci C, Mataresse AMS, Tarantino A, Crisosto C, Incerti O, Marcotuli I, Nigro D, Blanco A, Gadaleta A (2016) Characterization of edible fig germplasm from Puglia, southeastern Italy: is the distinction of three fig types (Smyrna, San Pedro and Common) still valid? Sci Hortic 205:52–58. https://doi.org/10.1016/j.scienta.2016.04.016
Gaaliche B, Saddoud O, Mars M (2012) Morphological and pomological diversity of fig (Ficus carica L.) cultivars in Northwest of Tunisia. ISRN Agron 2012, Article ID 326461. https://doi.org/10.5402/2012/326461
Giraldo E, López-Corrales M (2008) Optimization of the management of an ex situ germplasm bank in common fig with SSRs. J Am Soc Hortic Sci 133(1):69–77
Giraldo E, Viruel MA, Lòpez-Corrales M, Hormaza JI (2005) Characterisation and cross-species transferability of microsatellites in the common fig (Ficus carica L.). J Hortic Sci Biotechnol 80:217–224. https://doi.org/10.1080/14620316.2005.11511920
Giraldo E, López-Corrales M, Roger JP, Khadari B, Hochu I, Santoni S, Hormaza JI (2008) Standardization of experimental protocols and SSR markers for the management of fig germplasm collections. Acta Hortic (ISHS) 798:213–216. https://doi.org/10.17660/ActaHortic.2008.798.29
Giraldo E, López-Corrales M, Hormaza JI (2010) Selection of the most discriminating morphological qualitative variables for characterization of fig germplasm. J Am Soc Hortic Sci 135(3):240–249
Grassi G (1998) Studies of Italian fig germplasm. Acta Hortic (ISHS) 480:97–102
Hocquigny S, Pelsy F, Dumas V, Kindt S, Heloir MC, Merdinoglu D (2004) Diversification within grapevine cultivars goes through chimeric states. Genome 47(3):579–589. https://doi.org/10.1139/g04-006
Jakše J, Satovic Z, Javornik B (2004) Microsatellite variability among wild and cultivated hops (Humulus lupulus L.). Genome 47:889–899. https://doi.org/10.1139/g04-054
Khadari B (2012) Ex situ management of fig (Ficus carica L.) genetic resources: towards the establishment of the Mediterranean reference collection. Acta Hortic (ISHS) 940:67–74. http://www.actahort.org/books/940/940_7.htm
Khadari B, Lashermes P, Kjellberg F (1995) RAPD fingerprints for identification and genetic characterization of fig (Ficus cairica L.) genotypes. J Genet Breed 49:77–86
Khadari B, Hochu I, Santoni S, Kjellberg F (2001) Identification and characterisation of microsatellite loci in the common fig (Ficus carica L.) and representative species of genus Ficus. Mol Ecol Notes 1:191–193. https://doi.org/10.1046/j.1471-8278.2001.00072.x
Khadari B, Hochu I, Bouzid L, Roger JP, Kjellberg F (2003) The use of microsatellite markers for identification and genetic diversity evaluation of fig collection in CBNMP. Acta Hortic 605:77–86
Khadari B, Grout C, Santoni S, Kjellberg F (2005) Contrasted genetic diversity and differentiation among Mediterranean populations of Ficus carica L.: a study using mtDNA RFLP. Genet Resour Crop Evol 52:97–109. https://doi.org/10.1007/s10722-005-0290-4
Kirby LT (1990) DNA fingerprinting. An introduction. Stockton Press, New York
Knap T, Jakše J, Cregeen S, Javornik B, Hladnik M, Bandelj D (2016) Characterization and defining of a core set of novel microsatellite markers for use in genotyping and diversity study of Adriatic fig (Ficus carica L.) germplasm. Braz J Bot 39(4):1095–1102. https://doi.org/10.1007/s40415-016-0299-2
Laddomada B, Gerardi C, Mita G, Lumare D, Minonne F, Marchiori S, Fiocchetti F (2008) Molecular characterization of Apulian fig (Ficus carica L.) germplasm collection using fluorescence-based AFLP markers. Acta Hortic 798:205–212
Mariotti Lippi M, Bellini C, Mori Secci M, Gonnelli T (2009) Comparing seeds/fruits and pollen from a Middle Bronze age pit in Florence (Italy). J Archaeol Sci 36(5):1135–1141. https://doi.org/10.1016/j.jas.2008.12.017
Mariotti Lippi M, Bellini C, Mori Secci M, Gonnelli T, Pallecchi P (2015) Archaeobotany in Florence (Italy): landscape and urban development from the late Roman to the Middle Ages. Plant Biosyst 149(1):216–227. https://doi.org/10.1080/11263504.2013.822433
Mawa S, Husain K, Jantan I (2013). Ficus carica L. (Moraceae): phytochemistry, traditional uses and biological activities. Evid Based Complement Altern Med, Article ID 974256. https://doi.org/10.1155/2013/974256
Minonne F, Ippolito I, Marchiori S (2001) L’attività dell’orto botanico di Lecce nel reperimento e la propagazione delle vecchie cultivar di Ficus carica L. Italus Hortus 8(5):30–33
Paetkau D, Calvert W, Stirling I, Stroberck C (1995) Microsatellite analysis of population structure in Canadian polar bears. Mol Ecol 4:347–354. https://doi.org/10.1111/j.1365-294X.1995.tb00227.x
Papadopoulou K, Ehaliotis C, Tourna M, Kastanis P, Karydis I, Zervakis G (2002) Genetic relatedness among dioecious Ficus carica L. cultivars by random amplified polymorphic DNA analysis, and evaluation of agronomic and morphological characters. Genetica 114:183–194. https://doi.org/10.1023/A:1015126319534
R Development Core Team (2005) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. http://www.R-project.org
Rattighieri E, Rinaldi R, Bowes K, Mercuri AM (2013) Land use from seasonal archaeological sites: the archaeobotanical evidence of small Roman farmhouses in Cinigiano, South-Eastern Tuscany-Central Italy. Ann Bot 3:207–215. https://doi.org/10.4462/annbotrm-10267
Saddoud O, Chatti K, Salhi-Hannachi A, Mars M, Rhouma A, Marrakchi M, Trifi M (2007) Genetic diversity of Tunisian figs (Ficus carica L.) as revealed by nuclear microsatellites. Hereditas 144(4):149–157. https://doi.org/10.1111/j.2007.0018-0661.01967.x
Saddoud O, Baraket G, Chatti K, Trifi M (2011) Using morphological characters and simple sequence repeat (SSR) markers to characterize tunisian fig (Ficus carica L.) cultivars. Acta Biol Crac 53:7–14. https://doi.org/10.2478/v10182-011-0019-y
Storey WB (1976) Fig Ficus carica (Moraceae). In: Simmonds NW (ed) Evolution of crop plants, 2nd edn. Wiley-Blackwell, London, pp 205–208
Turfa JM (2012) Background on agriculture, agronomy in prehistoric Italy. In: Turfa JM (ed) Divining the etruscan world: the brontoscopic calendar and religious practice. Cambridge University Press, Cambridge, pp 152–153
UPOV (2010) Guidelines for the conduct of tests for distinctness, homogeneity and stability, fig edn. International union for the protection of new varieties of plants, Genéve
Veberic R, Mikulic-Petkovsek M (2016) Phytochemical composition of common fig (Ficus carica L.) cultivars. In: Simmons MSJ, Preedy VR (eds) Nutritional composition of fruit cultivars. Elsevier, Amsterdam, pp 235–255
Wagner HW, Sefc KM (1999) Identity 1.0: freeware program for the analysis of microsatellite data. Vienna. http://www.boku.ac.at/zag/forsch/identity.htm
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rodolfi, M., Ganino, T., Chiancone, B. et al. Identification and characterization of Italian common figs (Ficus carica) using nuclear microsatellite markers. Genet Resour Crop Evol 65, 1337–1348 (2018). https://doi.org/10.1007/s10722-018-0617-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-018-0617-6