Abstract
Tumour necrosis factor related apoptosis inducing ligand (TRAIL) binds to death receptor 4 (DR4) activating the apoptotic signalling pathway. DNA damaging agents (genotoxins) such as etoposide increase DR4 expression and when combined with TRAIL induce a synergistic apoptotic response. The mechanism for up-regulation of DR4 expression following genotoxin treatment is not well understood. Herein, we determined that transcription factor NF-κB plays a role in genotoxin induced DR4 expression. Increased expression of DR4 following etoposide treatment is blocked by inhibition of the NF-κB pathway. Moreover, expression of the p65 subunit of NF-κB is sufficient to increase DR4 protein levels. Indeed, knockdown of p65 by RNA interference blocked etoposide up-regulation of DR4. We further identified a functional NF-κB binding site located in the DR4 promoter. Mutation of this site abrogates the induction of luciferase activity after p65 over-expression. Furthermore, electromobility shift assays and chromatin immunoprecipitaton suggest that NF-κB binds to this site upon etoposide treatment. MEK kinase 1 (MEKK1) is a serine threonine kinase that is activated following etoposide treatment and activates NF-κB. Expression of the kinase inactive MEKK1 (MEKK1-KM) abrogates the up-regulation of DR4 after etoposide treatment. Taken together, NF-κB plays a role in up-regulation of DR4 following etoposide treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tumour necrosis related apoptosis inducing ligand (TRAIL/Apo-2L) is a protein that binds to death receptor-4 (DR4, TRAIL-R1) and death receptor-5 (DR5, TRAIL-R2) to induce cell death. TRAIL induces apoptosis in a variety of cancer cells without inducing apoptosis in normal non-transformed cells. This gives TRAIL the potential to become a therapeutic agent in the treatment of cancers [1, 2]. Levels of DR4 and DR5 expression are thought to contribute at least in part to the extent of TRAIL induced apoptosis [3–5]. Unfortunately, many cancer cells are TRAIL resistant [6]. Hence, understanding the regulation of TRAIL receptors’ expression could provide insight into increasing the amount of TRAIL-induced apoptosis in TRAIL resistant cells.
We and others have reported that TRAIL in combination with DNA damaging agents including etoposide induce a synergistic apoptotic response in cancer cells [3, 7]. Etoposide treatment increases the levels of DR4 and DR5 expression in a variety of cancer cell lines. This increase in death receptors contributes to the synergistic apoptotic response when etoposide is combined with TRAIL. We have determined that up-regulation of DR5 following DNA damaging agent treatment is mediated by transcription factors NF-κB and p53 [8, 9]. The DR5 gene contains a p53 and a NF-κB cis-element within the first intronic region [10]. Similarly, an analogous p53 binding site has been reported in the first intron of the DR4 gene contributing to DR4 up-regulation following DNA damaging agent treatment, however an NF-κB binding site is absent [11]. Although a NF-κB site has not been reported in the DR4 gene, several reports link NF-κB activity with an increase in DR4 expression. Treatment with etoposide increased DR4 mRNA and protein levels in HEK 293 cells and was inhibited by blockage of the NF-κB activation [3]. Furthermore, over-expression of the NF-κB subunit c-Rel has been reported to increase DR4 mRNA and protein levels in CCR43 cells [12], and MDA-MB-231 cells [13]. The mechanism for NF-κB mediated DR4 expression is unknown.
MEK kinase 1 (MEKK1) is a serine/threonine kinase that participates in the induction of apoptosis [14–20]. MEKK1 has been found present in the IKK complex and activates the NF-κB signalling pathway [20, 21]. Over-expression of kinase inactive MEKK1 (MEKK1-KM) has been shown to block NF-κB activity and etoposide induced apoptosis [3]. In addition, over-expression of MEKK1 specifically elevates the levels of DR4 without affecting DR5 [15]. The role of MEKK1 in NF-κB mediated regulation of DR4 expression is unknown.
Herein, we report that etoposide increases the levels of DR4 protein and this increase is blocked by inhibition of the NF-κB activation. We further identified the NF-κB binding site in the DR4 promoter responsible for DR4 expression. In addition, we determine the involvement of MEKK1 in NF-κB activation and up-regulation of DR4 following etoposide treatment. This describes for the first time a mechanism for NF-κB up-regulation of DR4 following genotoxin treatment.
Material and methods
Cell culture
Cell lines were maintained in a humidified 5.0% CO2, 37° C incubator. The human embryonic kidney cell line (HEK 293) was grown in Dulbecco’s modified medium supplemented with 100 units/ml penicillin, 100 μg/ml streptomycin and 10% bovine calf serum (Invitrogen). The colorectal carcinoma cell line HT29 was maintained in RPMI medium 1640 supplemented with 100 units/ml penicillin, 100 μg/ml streptomycin, 1 mM HEPES and 10% fetal calf serum (Hyclone). HEK 293 cells stably transfected with empty vector pcDNA3 (293V), pcDNA3 expressing kinase inactive MEKK1 (MEKK1-KM), or pcDNA3 expressing ΔIκB were under selection with 1 mg/ml G418.
Transfections
HEK 293 cells were grown on six well plates and transfected with 4 μg of plasmid DNA with the exception of the p65 plasmid which used 8 μg DNA. Cells were treated with 6 μl of superfect reagent according to manufacturer’s specifications.
Western blotting
Cells were lysed as specified by Spector et al [22]. 50–100 μg of cell lysate protein were equally loaded and western blotted [22]. Blots were incubated with the appropriate antibody concentration overnight (1:200 dilution for rabbit anti-p65 (Santa Cruz), 1:200 for rabbit anti-IκB (Santa Cruz), 1:200 rabbit anti-DR4 (Santa Cruz), 1:600 for mouse anti-DR4 (Abcam), 1:1000 mouse anti-TRAIL (Sigma), 1:200 for rabbit anti-MEKK1 (Santa Cruz), 1:200 for rabbit anti-caspase-3 (Santa Cruz), 1:30,000 for mouse anti-tubulin (Sigma), and 1:500 for rabbit anti-actin (sigma)). After incubation with the appropriate secondary antibody conjugated to horseradish peroxidase, blots were visualized on X-ray film with enhanced Chemiluminescence reagents (Perkin Elmer Life Sciences). Signal density was obtained by scanning exposed X-ray films on a Fluor chem 9900 imaging system (Alpha Innotech) for 120 ms. Normalized density was obtained by dividing the rough density values of a sample band over loading control band (actin). Fold increase was obtained by dividing the normalized density over the normalized density of untreated cells (or untreated 293 V cells when dealing with stable transfections).
Flow cytometry
HEK 293 cells stably transfected with vector alone pcDNA3, ΔIκB, and MEKK1-KM expression vectors were treated for 24 h with 100 μM etoposide. HT29 cells were untreated or treated with 10 μM BAY11-7082 and 100 μM etoposide for 24 h. 1 × 106 cells were collected and blocked with 10% bovine calf serum dissolved in phosphate buffer saline (PBS) for 30 min. Cells were collected and incubated with 5 μg/ml mouse monoclonal antibody against DR4 conjugated with FITC (Axxora). Cells were washed 3 times before analyzing by flow cytometry.
DNA constructs and plasmids
Plasmids pGL3 DR4 −1773/+63, pGL3 DR4 −586/+63 were kind gifts from Dr. Shi-Yong Sun (Emory University). NF-κB sites I, II, or III on the pGL3 DR4 −1773/+63 plasmid were deleted or mutated using the QuikChange site-directed mutagenesis kit (Stratagene). Site I was deleted using primers with the forward sequence GGCTAATTTTTTTTCTATTTTTAGTAGAGACCTGTTTTGGCC and the reverse sequence GGCCAAAACAGGTCTCTACTAAAAATAGAAAAAAAATTAGCC to generate pGL3 DR4 −1773/+63 ΔNF-κB I. Site II was deleted using primers with the forward sequence CCGAGGGACCCAATTCTGCCTTTCTTCTAACCTGTTA and the reverse sequence CTTAACAGGTTAGAAGAAAGGCAGAATTGGGTCCCTC to create pGL3 DR4 −1773/+63 ΔNF-κB II. Important nucleotides for binding were mutated in NF-κB site III using the forward primer GAGACGGAGT GC CCCTCTTGTGGCCCAGGCTGG and the reverse primer CCAGCCTGGGCCACAAGAGGG GC ACTCCGTCTC to yield pGL3 DR4 −1773/+63 NF-κB III. Putative binding sites are underlined and point mutations are indicated as boldface. Nucleotides flanking the deletions are italized and underlined.
P65 and DR4 siRNA knockdown
HEK 293 cells were transfected with 100 nM human-specific p65 siRNA, or scrambled siRNA (Cell signalling). HEK 293 cells were transfected with 2.5 μg/ml of human specific DR4 siRNA, or non-specific control siRNA (Santa Cruz). Cells were transfected using RNAiFect reagent (Qiagen) according to the manufacturer’s specifications. Levels of p65 and DR4 were verified by western blotting 24 h after transfection.
Luciferase assays
HEK 293 cells were transfected with 0.4 μg β-GAL and cotransfected with 4 μg of luciferase vectors pGL3, pGL3 DR4 −1773/+63 (kind gift from Dr. S.-Y. Sun, Emory University), pGL3 DR4 −586/+63, pGL3 DR4-1773/+63 ΔNF-κB I, pGL3 DR4 −1773/+63 ΔNF-κB II, pGL3 DR4 −1773/+63 mNF-κB III, and the NF-κB reporter vector (Stratagene). When assays involved the transfection of additional plasmids; empty vector pcDNA3 was used to equalize the ammount of DNA transfected to cells. Cells analyzed for NF-κB activation were treated with 100 μM etoposide or 50 ng/ml of TNF-α. Cells were lysed with a cell culture lysis reagent (Promega). 20 μl of lysate was added to each of a 96-well plate before measurement. Luciferase activity was measured in an L-MAX 1.1 L luciferase plate reader (Molecular devices). The reaction consisted in the injection of 100 μL of luciferase assay substrate solution (Promega), the-post injection delay was set to 2 seconds, and the integration time was set to 10 seconds. To measure β-Gal activity, 50 μL of lysate was mixed with 50 μL of 200 mM sodium phosphate buffer, 2 mM magnesium chloride, 100 mM beta-mercapto-ethanol, and 1.22 mg/ml of Ortho-nitrophenyl-b-D-galactopyranoside (ONPG) in a 96-well plate, and incubated for 30 min. Plates were read in a Titertek Multiskan MCC/340 at a wavelength of 414 nm. Relative luciferase values were calculated by dividing luciferase reading over β-gal reading. Fold induction was obtained by dividing relative luciferase values against the untreated values for the −1776/+63 vector.
Chromatin immunoprecipitation
ChIP assay was performed as described by Spencer et al [23]. HEK 293 cells were treated with 100 μM etoposide for 24 h. The lysate was precipitated with 4 μg of a rabbit polyclonal antibody against p65 (Santa Cruz sc-109X) overnight at 4°C. DNA was extracted by using the QIAmp DNA mini kit and resuspended in 50 μl of doubly distilled water. Polymerase chain reaction was performed using the forward primer CTGCTTTTGGTTTCAGGTCTCCAT and the reverse primer TCTCCCGTGGTTTAAGGAGTTTCA. We performed thirty PCR cycles; each cycle consisted of temperatures of 95°C, 60°C, and 72°C; with 30 seconds at each temperature. The run was ended with 7 min at a temperature of 72°C. For each immunoprecipitation 1% of the chromatin was assayed for equal loading. Images were captured by a GeneGenious apparatus (Syngene).
Electromobility shift assays
HEK 293 cells were untreated or treated for 24 h with etoposide and nuclear extracts were prepared. Double stranded oligonucleotides with the NF-κB site III sequence TTGAGACGGAGTTTCCCTCTTGTG, consensus NF-κB sequence AGTTGAGGGGACTTTCCCAGG, or scrambled NF-κB site III sequence CCACAAGAGACATACGCGGTCTCAA were 5′ end labelled with [γ-32P]ATP (Perkin Elmer) and T4 polynucleotide kinase. Putative binding sites are underlined. A total volume of 20 μl of binding buffer contained 10 μg of nuclear extract and 0.5 ng of labelled probe. Competition assays were performed by adding an excess (2.5 ng) of unlabeled probe to the binding reaction. Where indicated, 4 μg of a rabbit polyclonal antibody against p65 (Santa Cruz sc-109X), or 4 μg of a mouse monoclonal antibody against p50 (Santa Cruz sc-8414X) were incubated with the binding mixtures. Mixtures were loaded into a 5% native polyacrylamide gel. The gel was dried and a film was exposed overnight.
Apoptotic assays
HT29 cells were plated in six-well plates and untreated or treated with 100 μM etoposide, 100 μg/ml HGS-ETR1 (Mapatumumab) (Human Genome Sciences), or 10 μM BAY11-7082. After the specified incubation period cells were collected by centrifugation and resuspended in DMEM with 100 ng/ml acridine orange stain with ethidium bromide. At least 200 cells were scored per sample. Alternatively, after treatment HT29 cells were suspended in hypotonic propidium iodide lysis buffer for 30 min at room temperature ( 50 μg/ml propidium iodide, 1 mg/ml sodium citrate, 10 μg/ml Rnase diluted in a 1:100 triton X-100 solution). Cells were washed with phosphate buffer saline (PBS) and sub G-1 peak was visualized using flow cytometry. We used immunoflourescence to determine apoptosis due to MEKK1 transfection. Cells were grown on coverslips and transfected with 2.5 μg/ml of siRNA, expression vectors, or treated with 100 ng/ml DR4:Fc. The slides were fixed and incubated with rabbit polyclonal anti-MEKK1 (Santa Cruz) antibody in a 1:500 dilution, and a secondary donkey anti-Rabbit antibody conjugated to Alexa Fluor 488 in a 1:2,000 dilution (Molecular Probes). Slides were mounted on 3 μl of a 1.5 μg/ml DAPI/Antifade solution (Molecular Probes). In cells positive for MEKK1, DAPI staining was indicative of apoptosis when chromatin was condensed; compared with the uniform and diffuse staining of DNA in normal cells. At least 200 cells were scored per sample. Fluorescence was visualized and captured using an Olympus BX51 fluorescent microscope equipped with a Coolsnap camera.
Caspase assays
Caspase-3 activity was determined by using a DEVD-AFC detection kit (Santa Cruz). DEVD is a synthetic tetrapeptide corresponding to the upstream amino acid sequence of the caspase-3 cleavage site in PARP; conjugated to the fluorophore AFC (7-amino-4trifluoromethyl coumarin). Cells were lysed using the lysis buffer as indicated in the manufacturer’s specifications; and the protein concentration was detected using a bradford assay. Protein assay concentration was 1 μg/μl diluted in reaction buffer containing 5 mM di-thiothreitol (DTT) and 5 μl of DEVD-AFC to a total volume of 100 μl. Samples were placed in a white 96 well plate with clear bottom (Costar). Fluorescence was detected using a Spectramax M5 (Molecular Devices) with an excitation wavelength of 400 nm, and an emission wavelenght of 505 nm.
Identification of potential NF-κB binding sites
We used the on-line program TFSEARCH (TRANSFAC databases) to analyze the sequence of a region 2800 base pairs ahead of the transcription start site in the DR4 promoter [24]. We set the threshold limit to 85.5 and searched for NF-κB binding sites.
Results
Etoposide treatment increases DR4 expression in an NF-κB dependent manner
DNA damaging agents have been reported to increase DR4 expression. To determine whether DNA damaging agents induce the up-regulation of DR4, we treated human embryonic kidney (HEK) 293 cells and the colorectal cancer cell line, HT29 with 100 μM of the topoisomerase II inhibitor etoposide. Increased DR4 expression was detected 3 h after treatment in HEK 293 cells (Fig. 1A, B) and 6 h after treatment in HT29 cells (Fig. 1C). In HEK 293 cells a 2.5 fold peak expression of DR4 was observed at 6 h (Fig. 1B). Flow cytometry for DR4 shows that treatment with 100 μM etoposide increases DR4 expression at the cell surface at 24 h (Fig. 1D, E). DNA damaging agent treatment has also been reported to activate the NF-κB pathway [3]. We transfected HEK 293 cells with a NF-κB luciferase reporter vector and treated these cells with 100 μM etoposide. Addition of etoposide increased the levels of NF-κB activity to 5-fold when compared to untreated cells (Fig. 1F). As positive control, these cells were treated with 50 ng/ml of tumour necrosis factor-α (TNF-α) a strong activator of NF-κB; this increased the activity up to 16 fold (Fig. 1F). To determine whether NF-κB plays a role in DR4 regulation, we inhibited the NF-κB pathway by treating with BAY11-7082, an inhibitor that blocks IκBα degradation [25] or by over-expressing the super repressor ΔIκB. The repressor protein IκB inhibits nuclear localization of NF-κB and the activation of target genes [26]. HEK 293 cells were stably transfected with a ΔIκB expression vector. This effectively reduced levels of DR4 to lower than basal and blocked up-regulation of DR4 after etoposide treatment (Fig. 1A, B). Furthermore, flow cytometry shows that DR4 surface expression in response to etoposide is blocked by the expression of the ΔIκB protein (Fig. 1D). Finally, etoposide-induced expression of DR4 in HT29 cells was blocked upon treatment with 10 μM BAY11-7082 (Fig. 1C, E). These results suggest that NF-κB signalling plays a role in up-regulation of DR4 in response to etoposide.
Etoposide increases DR4 levels in an NF-κB dependent manner: (A) Etoposide through the mediation of NF-κB increases the levels of DR4. HEK 293 cells stably transfected with empty vector pcDNA3 (vector alone) or a ΔIκB expression vector were treated with 100 μM etoposide; cells were lysed at 3, 6, 12 or 24 h. Levels of DR4 protein were detected by western blotting. Blot is representative of three independent experiments. (B) Densiomentry fold increase for DR4 western blots. Fold increase was calculated with the density of 0 h 293V cells as a reference. Averages for 293V cells are representative of 6 independent experiments; averages for ΔIκB cells are representative of 3 independent experiments. Error bars represent standard deviation. (C) HT29 cells were treated with 100 μM etoposide alone or in combination with 10 μM BAY11-7086 and harvested at 0, 3, 6, 12 and 24 h after treatment. DR4 levels were detected by western blotting. (D) Inhibition of NF-κB blocks DR4 surface expression. Vector alone and ΔIκB cells were untreated or treated with etoposide for 24 h and labelled with a DR4 antibody conjugated with FITC. As negative controls, unlabeled cells (Blank) and cells incubated with a non-specific FITC antibody (FITC control Ab) were used. Flow cytometry was used to detect FITC fluorescence in cells. (E) HT29 cells were untreated or treated with 100 μM etoposide alone or in combination with 10 μM BAY11-7086 and labelled with a DR4 antibody conjugated with FITC. Flow cytometry was used to detect DR4 surface expression. (F) Etoposide activates the NF-κB pathway. HEK 293 cells were transfected with a NF-κB luciferase reporter vector and treated with 100 μM etoposide, or with 50 ng/ml TNF-α as a positive control. Cells were harvested at 0, 3, 6, 12 and 24 h after treatment and luciferase assays were performed. Results are the average of three independent experiments; error bars represent standard deviation
p65 Subunit of NF-κB regulates DR4 expression
We have previously shown that the p65 subunit of NF-κB is involved in the up-regulation of DR5. Over-expression of p65 activates a luciferase vector containing the DR5 promoter, and it increases the levels of DR5 protein [9]. To determine the involvement of p65 in the up-regulation of DR4, we transfected HEK 293 cells with a p65 expression vector. Western blotting was used to detect DR4 protein levels 24 h after transfection. Over-expression of p65 effectively increased the levels of DR4 1.6 fold (Fig. 2A, B). To determine the role of NF-κB in the transcriptional activation of DR4, we obtained a luciferase vector containing the 1.7 kb DR4 promoter region and transfected the promoter construct into HEK 293 cells. Cells were co-transfected with the p65 expression vector and treated or untreated with 10 μM BAY11-7082. Luciferase assays indicate that expression of p65 effectively increases the activity of the DR4 promoter six-fold. This was abrogated by treatment with BAY11-7082, which reduced the luciferase activity to basal levels (Fig. 2C). This suggests that p65 regulates the expression of DR4 through its promoter. To further determine the role of p65 in the regulation of DR4 we transfected p65 siRNA or scrambled siRNA to HEK 293 cells. These cells were treated with etoposide for 24, 48, and 72 h. Etoposide treatment increased DR4 levels in cells transfected with scrambled siRNA. This up-regulation was blocked in cells transfected with p65 siRNA and levels of DR4 reduced to levels lower than basal (Fig. 2D). By western blotting we showed that p65 siRNA effectively lowers the levels of p65 when compared with untransfected cells (Fig. 2E). These results suggest that the NF-κB subunit p65 participates in the up-regulation of DR4 in response to etoposide.
p65 plays an important role in the induction of DR4: (A) p65 over-expression increases the levels of DR4; this is inhibited by blockage of NF-κB signalling. HEK 293 cells were transiently transfected with empty vector pcDNA3 or with a p65 expression vectors. In cells were transfected with etoposide. After 24 h cells were lysed and DR4 and p65 levels were detected by western blotting. (B) Densiomentry fold increase for p65 western blots. Fold increase was calculated with the density of untreated HEK 293 cells as a reference. Averages are representative of 3 independent experiments. Error bars represent standard deviation. Statistical significance was calculated by a Student’s t test. (C) p65 increases transcriptional activity of DR4 promoter. HEK 293 cells were transfected with the luciferase vector pGL3 containing the DR4 promoter region (DR4 promoter) or empty pGL3 vector. Cells were co-transfected with an expression vector for p65 and treated or untreated with 10 μM of the inhibitor BAY11-7082. Cells were lysed after 24 h and luciferase assays were performed. Experiment was repeated 3 times; error bars represent standard deviation. Statistical significance was calculated by a Student’s t test. (D) p65 knockdown blocks the up-regulation of DR4 in response to etoposide. HEK 293 cells were transfected with siRNA specific for p65 or with a control siRNA containing the same nucleotides in a different order (scrambled). Cells were treated with etoposide and lysed at 24, 48, and 72 h. DR4 levels were detected by western blotting. (E) p65 siRNA lowers the levels of p65 protein. Cells were untransfected or transfected with p65 siRNA. 24 h after transfection cells were untreated or treated with lysed and western blotted for p65
Identification of an NF-κB binding site in DR4 promoter responsible for the up-regulation of DR4
Although there are several reports linking NF-κB to the up-regulation of DR4, a NF-κB binding site has not been identified. We analyzed the sequence of the DR4 promoter starting −2,800 base pairs before the transcriptional start site with the TFSEARCH tool (TRANSFAC databases) [24]. We identified three potential NF-κB binding sites; A site located between 563 and 553 base-pairs from of the transcription start site (−563/553) was denoted site I. Site II was located at −776/−766, and site III was located at −1366/1356 (Fig. 3A). To determine the importance of these sites we obtained a truncated luciferase vector that contains a truncated promoter region that lacks sites II and III (Fig. 3A). Luciferase assays in HEK 293 cells indicate that this truncated promoter lacks responsiveness to p65 over-expression. While p65 induces a 6 fold increase in activity, the elimination of sites II and III decreases luciferase activity to basal levels (Fig. 3B). This suggests that site II, site III, or both sites are important for the expression of DR4. To elucidate the importance of these potential NF-κB binding sites, we generated mutants for each site and evaluated their responsiveness to p65 expression. Through site-directed mutagenesis, we mutated highly conserved sites in the sequence of site III and deleted site I and site II (Fig. 3C). Luciferase assays indicate that deletion of site I and II do not suppress the activation of the DR4 promoter in response to p65 expression. In cells transfected a vector containing the wild type promoter (pGL3 DR4 (−1773/+63), a two-fold increase of luciferase activity was observed (Fig. 3D). In cells transfected with a promoter lacking NF-κB site I (pGL3 DR4 ΔNF-κB I), over-expression of p65 resulted in a 1.5-fold increase in luciferase activity (Fig. 3D). In cells transfected with a promoter lacking NF-κB site II (pGL3 DR4 ΔNF-κB II), over-expression of p65 resulted in a 2-fold increase in luciferase activity (Fig. 3D). In contrast, mutation of site III abrogated responsiveness to p65. In cells transfected with the pGL3 DR4 mNFκB III vector, there was no increase in luciferase activity in response to p65 (Fig. 3D). This suggests that NF-κB site III is important for its up-regulation in response to p65 over-expression. To determine whether etoposide treatment resulted in p65 binding to this NF-κB binding site in the DR4 promoter, we used electromobility shift assay (EMSA) with a probe specific to site III in the DR4 promoter. Etoposide treatment caused retardation in mobility of a probe containing the sequence of site III; no shift was observed in untreated cells (Fig. 3E lanes 2 and 3). Addition of a competing unlabeled probe caused the disappearance of this band; while addition of a scrambled unlabeled probe did not have an effect (Fig 3E lanes 4 and 5). Etoposide caused the shift of an identical band when a NF-κB consensus probe was used (Fig. 3E lanes 8 and 9). Addition of p65 specific antibodies caused a decreased intensity of this band; however the band was not affected when a control non-specific antibody was added (Fig. 3E lanes 6 and 7). When probes corresponding to site I and site II were used, no shifted band was observed following etoposide treatment (data not shown). To determine whether p65 binds directly to the DR4 promoter following etoposide treatment, chromatin immunoprecipitation (ChIP) assay was performed. We used a set of primers specific for site III and a p65 specific antibody. Binding of p65 was detected in this region and was increased following etoposide treatment in HEK293 cells. The ChIP assay showed that untreated cells have an intensity volume of 2,818 relative units; while cells treated with etoposide have 5404 relative units (Fig. 3F). Input values of 10,602 relative units for untreated and 7,229 relative units were obtained. This means that 74% of promoter is bound to NF-κB in response to etoposide treatment, while 25% of promoter is bound in untreated cells. Using primers specific for regions corresponding to sites I and II, p65 binding failed to be detected (data not shown). These results suggest that site III in the DR4 promoter binds p65 in response to etoposide treatment regulating DR4 expression.
An NF-κB binding site located in the promoter of DR4 is responsible for its up-regulation in response to etoposide. (A) TFSEARCH found 3 potential NF-κB binding sites in the DR4 promoter. The location of the potential NF-κB sites in these vectors is outlined in the figure. (B) Elimination of NF-κB sites II and III abrogates DR4 up-regulation. We obtained 2 luciferace vectors; one containing the entire DR4 promoter pGL3 DR4 −1773/+63 and a truncated vector pGL3 DR4 −586/+63; HEK 293 cells were transfected with these vectors and co-transfected with p65. 24 h after transfections cells were lysed and luciferase assays were performed. The results are the average of three independent experiments; error bars represent standard deviation. Statistical significance was calculated by using a Student’s t test (C) Site directed mutagenesis was performed on the pGL3 DR4 −1773/+63 plasmid. Three different plasmids were generated: pGL3 DR4 −1773/+63 ΔNF-κB I containing deletion of site I, pGL3 DR4 −1773/+63 ΔNF-κB II containing a deletion of site II, and pGL3 DR4 −1773/+63 mNF-κB III containing point mutation of critical sites for NF-κB binding in site III. The wild type and mutated sequences of these sites are listed. (D) NF-κB site III is crucial for DR4 up-regulation in response to p65. HEK 293 cells were transfected with luciferase vector containing one of the ΔNF-κB I, ΔNF-κB II, or mNF-κB III mutations; where indicated cells were co-transfected with a p65 expression vector. After 24 h cells were lysed and luciferase assays were performed. Experiments were performed in triplicate; error bars represent standard deviation. Statistical significance was calculated by a Student’s t test. (E) Etoposide treatment causes p65 to bind to a probe containing the NF-κB site III sequence. HEK 293 cells were treated with etoposide and nuclear lysates were isolated. EMSA assays were performed using a probe containing NF-κB site III sequence, an NF-κB consensus sequence, and a scrambled site III probe. Competitor probe consisted of an unlabeled NF-κB site III sequence, while non competitor consisted of a scrambled NF-κB site III sequence (N.S. comp.). In addition we added antibodies against p65 and FLAG tag as a negative control (ctrl. Ab.). (F) Etoposide treatment recruits p65 to the DR4 promoter. HEK 293 cells were untreated or treated with etoposide for 24 h; chromatin immunoprecipitation assays were performed using antibodies specific for p65. PCR was performed using primers specific for the NF-κB site III region. Intensity volumes for each band are specified under the image. Percentage of Bound promoter was calculated by dividing ChIP relative units by input relative units for each treatment and multiplying by 100
Etoposide-induced increase in DR4 protein levels leads to a sensitization of cells to TRAIL induced cell death
It has been reported that combined treatment of TRAIL with DNA damaging agents synergistically induces apoptosis in a variety of cell lines; however TRAIL is able to activate both DR4 and DR5. Mapatumumab is a monoclonal antibody specific for DR4 that is able to induce its trimerization and activation of the apoptotic signalling pathway [27]. Acridine orange staining shows that treatment of HT29 cells with a combination of 100 μg/ml Mapatumumab and 100 μM etoposide increased cell death synergistically when compared with cells treated with Mapatumumab or etoposide alone. Only 2% of untreated HT29 cells underwent apoptosis; treatment with etoposide alone for 48 h induced 30% cell death; while treatment with Mapatumumab alone induced 10% apoptosis at 48 h (Fig. 4A). Indeed, HT29 cells have been shown to be apoptotic resistant when TRAIL is used as a single agent [28, 29]. Combination treatment of etoposide and Mapatumumab effectively increased cell death to 70% (Fig. 4A). To determine whether NF-κB is involved in this synergistic increase in apoptosis, we inhibited NF-κB activation with BAY11-7082. Blockage of NF-κB decreased apoptosis after combination treatment; addition of etoposide, Mapatumumab, and BAY11-7082 induced apoptosis in only 40% of the cells (Fig. 4A). To determine whether this synergistic increase in cell death is a result of apoptosis, we examined the activation of caspase-3. To confirm these results we used sub-G1 peak assays to measure chromatin degradation (a hallmark of apoptosis). HT29 cells were treated as described above and 48 h after treatment cells were suspended in hypotonic PI lysis buffer. After 30 min sub-G1 peak was measured through flow cytometry. Only 18% of untreated cells displayed chromatin degradation; etoposide treatment increased this number to 41%. Mapatumumab treatment induced chromatin degradation in 30% of the cells; while a combination treatment of mapatumumab and etoposide induced it in 54% of the cells. Treatment with BAY 11-7082, etoposide, and mapatumumab reduced the number of cells displaying chromatin degradation to 36% (Fig. 4B). Although combination treatment did not result in a synergistic but an additive increase in cells undergoing chromatin degradation; blockage of NF-κB by BAY 11-7082 reduced the number of cells displaying a sub-G1 peak. To determine whether chromatin degradation was a result of apoptosis; we inhibited caspases by adding 20 μM of the pan-caspase inhibitor z-VAD-FMK. This treatment effectively reduced chromatin degradation induced by etoposide, mapatumumab alone or in combination (Fig. 4B). Furthermore, caspase-3 activation in these cells was measured using a fluorimetric caspase assay using DEVD-AFC. Our results show that combination treatment of mapatumumab and etoposide show the highest caspase-3 activity. Addition of BAY 11-782 to cells treated with mapatumumab and etoposide reduced the activity level to 74% when compared to cells treated with mapatumumab and etoposide (Fig. 4C). Etoposide treatment only induced 60% of the activity and mapatumumab only induced 50% of the activity. Untreated cells had 32% the levels of caspase activity displayed by cells treated with mapatumumab and etoposide (Fig. 4C). These results suggest that in response to Mapatumumab and etoposide cells undergo apoptosis. To determine the degree of NF-κB activation under these treatments, we performed a western blot for IκB. Degradation of IκB is indicative of NF-κB activation. Treatment with etoposide decreased the levels of IκB 50% when compared to untreated cells. Similarly Mapatumumab alone decreased the levels of IκB to 75%, and combination of Mapatumumab and etoposide decreased the IκB levels to 50% when compared to untreated cells (Fig. 4B, D). BAY 11-7082 is an inhibitor of IκBα degradation. When cells were treated with BAY 11-7082 alone a 2-fold increase in IκB were observed (Fig. 4E, F). Furthermore, when cells were treated with Mapatumumab and etoposide, BAY11-7082 restores IκB back to basal levels (Fig. 4E, F). These experiments demonstrate that BAY11-7082 treatment inhibits the degradation of IκB; this implies that NF-κB is inhibited in these cells. Taken together, these results suggest that the apoptotic synergy observed upon etoposide and Mapatumumab treatment is mediated through the NF-κB pathway.
Mapatumumab and etoposide synergize to induce apoptosis through the mediation of NF-κB. (A) Inhibition of NF-κB disrupts synergy between Mapatumumab and etoposide. HT29 cells were untreated, treated with 100 μM etoposide, 10 μM BAY 11-7082, or 100 μg/ml of Mapatumumab alone or in combination. 48 h after treatment cells were collected and cell death was assessed by acridine orange staining. Error bars represent the standard deviation of three independent experiments. Statistical significance was calculated using a Student’s t test. (B) Combination treatment between Mapatumumab and Etoposide increases DNA fragmentation; this is disrupted by NF-κB blockage. Cells were treated with 100 μM etoposide, 10 μM BAY 11-7082, 100 μg/ml of Mapatumumab, or 20 μM of z-VAD-FMK alone or in combination. 48 h after transfection cells were collected and stained with propidium iodide staining buffer. Flow cytometry was used to visualized the sub-G1 peak, indicative of DNA fragmentation. Flow data is representative of 3 independent experiments. (C) NF-κB blockage lowers caspase-3 activation in cells treated with mapatumumab and etoposide. Cells were treated as indicated in Fig. 4A; 12 h after treatment cells were lysed. Caspase-3 activity was measured after addition of DEVD-AFC substrate to 100 μg of protein. Caspase activity was measured by fluorescence at 400 nm excitation and 205 nm detection. Percent caspase-3 activity was determined in relation to the value of Mapatumumab + Etoposide which was set to 100%. (D) Combination treatment of Etoposide and Mapatumumab leads to the degradation of IκB. HT29 cells were treated as described in Fig. 4A and lysed at 24 h. (E) Densiomentry fold increase for IκB western blots. Fold increase was calculated with the density of untreated cells as a reference. Averages are representative of 4 independent experiments. Error bars represent standard deviation
MEKK1 activates NF-κB and increases DR4 expression
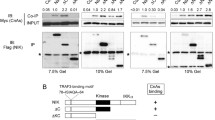
The serine/threonine kinase MEKK1 has been reported to play an important role in death receptor up-regulation and apoptosis induction. Over-expression of MEKK1 is enough to specifically up-regulate DR4 but not DR5 in a variety of cell lines contributing to apoptosis [15]. Expression of kinase-inactive MEKK1 (MEKK1-KM) blocks etoposide induced apoptosis and death-receptor up-regulation [15]. We stably transfected HEK 293 cells with an expression vector for MEKK1-KM or with empty vector pcDNA3. Etoposide treatment increased DR4 protein levels with a peak of 2.3 fold after 6 h of treatment (Fig. 5A, B). This was inhibited by the expression of MEKK1-KM which lowered the DR4 levels to a 1.3 fold increase at 6 h (Fig. 5A, B). In addition, expression of MEKK1-KM blocked cell surface expression of DR4 in response to etoposide treatment at 24 h as determined by flow cytometry (Fig. 5C). To determine whether MEKK1 has the ability to activate NF-κB, we co-transfected HEK 293 cells with a NF-κB luciferase vector and a expression vector for MEKK1 or kinase inactive MEKK1-KM. Transfection with MEKK1 increases NF-κB luciferase reporter activity whereas MEKK1-KM fails to activate NF-κB luciferase reporter activity (Fig. 5D). Cells transfected with the MEKK1 expression vector were taken as a reference (100% activity). Transfection with empty vector pcDNA3 only induces 33% of the activity induced by MEKK1. Transfection of MEKK1-KM reduced luciferase activity to 42% compared to MEKK1 luciferase activity (Fig. 5D). This suggests that MEKK1 plays a role in the activation of NF-κB. This experiment was repeated in cells expressing the ΔIκB super-repressor. In these cells expression of MEKK1 did not increase luciferase activity above pcDNA3 transfected cells (Fig. 5D). This suggests that NF-κB signalling is downstream of MEKK1. To elucidate whether MEKK1 participates in the transcriptional regulation of DR4, we co-transfected the DR4 luciferase plasmid into HEK-293 cells stably transfected with either empty vector or a expression vector for MEKK1-KM. MEKK1-KM significantly reduced basal luciferase of the DR4 promoter. Over-expression of MEKK1-KM reduced luciferase activity in half when compared with cells transfected with vector alone (Fig. 5E). This effect does not depend upon the activation of the JNK pathway since addition of the JNK inhibitor SP600125 [30] did not lower the activity of the DR4 promoter (Fig. 5E). A similar down-regulation in basal levels was observed when these cells were co-transfected with ΔIκB but no further reduction was observed when MEKK1-KM and ΔIκB were co-transfected (Fig. 5E). Transfection of a siRNA vector specific for human DR4 effectively blocked DR4 up-regulation in response to etoposide; however etoposide treatment of cells transfected with control siRNA lead to an increase of DR4 levels (Fig. 5F). This shows that DR4 siRNA can effectively block the up-regulation of DR4. To determine whether DR4 plays a role in MEKK1 induced apoptosis, we transfected cells with DR4 siRNA or non-specific control siRNA. HEK 293 cells were grown on glass coverslips and transfected with siRNA; 24 h later they were co-transfected with a MEKK1 expression vector or empty vector pcDNA3. Cells were fluorescently labelled for MEKK1 and cell death was determined as described in Material and Methods section. This indicates that MEKK1 expression resulted in 53% apoptosis in cells treated with control siRNA; in contrast, DR4 siRNA reduced MEKK1 induced apoptosis to 40%. Apoptosis in cells transfected with empty vector pcDNA3 was 8%. To further investigate the role of TRAIL in MEKK1 induced apoptosis, we treated cells with 100 ng/ml DR4:Fc a soluble protein that competes with death receptors for TRAIL binding. Treatment with DR4:Fc significantly reduced the apoptotic response of HEK293 cells in response to MEKK1 expression. Transfection of MEKK1 induced 35% cell death in HEK293 cells; addition of DR4:Fc reduced apoptosis to 24% (Fig. 5D). We also observed that over-expression of dominant-negative FADD that blocks death receptor activation abrogated MEKK1 induced apoptosis in HEK 293 cells (data not shown) [15]. These results indicate that DR4 expression and TRAIL binding are important for the cell death response in response to MEKK1; further suggesting that MEKK1 is involved in the regulation of DR4 and TRAIL induced apoptosis.
MEKK1 regulates etoposide induced up-regulation of DR4. (A) Kinase inactive MEKK1 blocks DR4 up-regulation. HEK 293 cells stably transfected with empty vector pcDNA3 (Vector alone), or an expression vector for MEKK1-KM were treated with etoposide for 0, 3, 6, 12 and 24 h. Cells were lysed and DR4 levels were assessed by western blotting. (B) Densiomentry fold increase for western blots. Fold increase was calculated with the density of 0 h 293V (vector alone) cells as a reference. Averages for 293V cells are representative of 6 independent experiments; averages for MEKK1-KM cells are representative of 3 independent experiments. Error bars represent standard deviation. (C) Dominant negative MEKK1-KM inhibits surface expression of DR4. Vector alone and MEKK1-KM cells were untreated or treated with etoposide for 24 h and labelled with a DR4 antibody conjugated with FITC. As negative controls, unlabeled cells (blank) and cells incubated with a non-specific FITC antibody (FITC control Ab) were used. Flow cytometry was used to detect FITC fluorescence in cells. (D) MEKK1 induces NF-κB activation. HEK 293 cells were co-transfected with empty luciferase vector pGL3 or the NF-κB reported plasmid. These cells were co-transfected with empty vector pcDNA3, a expression vector for MEKK1, or a expression vector for MEKK1-KM. Expression levels of MEKK1 and MEKK1-KM were detected by western blotting for antibodies against MEKK1. As a control NF-κB was inhibited by expressing ΔIκB. Values were calculated as percentages with respect to luciferase activity in response to MEKK1 expression. Experiments were repeated three times; error bars represent standard deviation. Statistical significance was calculated using a Student’s t test. (E) Expression of kinase inactive-MEKK1 reduces basal activity levels of the DR4 promoter. HEK293 cells stably transfected with empty vector pcDNA3, ΔIκB or MEKK1-KM were transfected with the pGL3 DR4-promoter vector. Additionally, a set of cells expressing empty vector pcDNA3 was treated with JNK inhibitor SP600125. Cells were lysed 24 h after treatment and luciferase assays were performed. Experiments were repeated 3 times; error bars represent standard deviation. Statistical significance was calculated using a Student’s t test. (F) DR4 siRNA inhibits up-regulation of DR4. HEK 293 cells were transfected with siRNA specific for DR4 or non-specific control siRNA. 24 h after transfections cells were untreated or treated with etoposide for an additional 24 h. Cells were lysed and immunoblotted for DR4. (G) Blockage of DR4 expression blocks MEKK1 induced apoptosis. HEK 293 cells were grown on glass cover-slips and transfected with control siRNA or DR4 siRNA. 24 h after siRNA transfections the cells were co-transfected with an expression vector for MEKK1 or empty vector pcDNA3. 24 h after MEKK1 transfections cells were fixed and labelled for with a MEKK1 specific antibody, and a FITC secondary antibody. Cells were stained with DAPI in order to visualize cell nuclei. MEKK1 positive cells were visualized by immunofluorescence and cell death was determined by counting MEKK1 positive cells displaying nuclear fragmentation. Bars represent the average of three independent experiments; standard deviation is represented by error bars. Statistical significance was calculated using a Student’s t test. (H) MEKK1 induced apoptosis is inhibited by blockage of the TRAIL pathway. HEK 293 cells were transfected with empty vector pcDNA3 or with an expression vector for MEKK1. Cells were treated with 100 ng/ml DR4:Fc. After 24 h Cells were stained for MEKK1 and the nucleus was stained using DAPI; cells were visualized by immunofluorescence. Apoptosis was assessed by chromatin condensation in cells positive for MEKK1 over-expression. Experiment was done in triplicates; error bars represent standard deviation. Statistical significance was calculated using a Student’s t test
Discussion
TRAIL’s ability to induce apoptosis in cancer cells but not in normal cells has potential as a treatment strategy against cancer. Currently, there are phase I and II clinical trials involving recombinant human TRAIL or agonistic antibodies against DR4 and DR5 [5]. Insight into the regulation of TRAIL death receptor expression could increase the effectiveness of these agents and provide rationale for combinational therapies in cancers. Indeed, treatments with DNA damaging agents [3], histone deacetylase (HDAC) inhibitors [4], or proteasome inhibitors [31] increase the levels of DR4 and DR5 expression and contribute to their cytotoxicity. In all instances addition of TRAIL synergistically increases the apoptotic response compared to using these drugs alone [5]. Herein we report that combined treatment between etoposide and the agonistic antibody Mapatumumab results in an increase in apoptosis and blocking NF-κB mediated DR4 expression inhibits this increased apoptosis (Fig. 6).
Proposed pathway for etoposide induced up-regulation of DR4, and synergy with Mapatumumab. The DNA damaging agent etoposide leads to the activation of MEKK1 and NF-κB. Activated MEKK1 has been shown by us and others to phosphorylate the inhibitor IκB leading to its degradation and subsequent activation of NF-κB. An active NF-κB transcription factor which includes the p65 subunit moves to the nucleus to activate transcription of the DR4 gene. The DR4 protein localizes to the plasma membrane; upon binding to TRAIL or Mapatumumab it forms the death inducing signalling complex (DISC) and activates caspase-8. Caspase-8 is an apical caspase that activates the death signalling program resulting in apoptosis. In our model, treatment with etoposide leads to the increase of DR4 in the cell surface. This increase in death receptors sensitizes cells to TRAIL or Mapatumumab. Our model provides a mechanism for the synergistic increase in apoptosis when cells are treated with a combination etoposide and activators of the TRAIL pathway
The transcriptional regulation of DR4 and DR5 is only beginning to be understood. Following etoposide treatment, the DR5 gene is activated through the cooperation of transcription factors NF-κB and p53 [8, 9]. These proteins bind to the DR5 gene at two binding sites located in the first intronic region [9, 10]. An equivalent p53 binding site is present in the first intronic region of DR4 [11]. This site was found to increase promoter activity in response to DNA damaging agents; however an equivalent NF-κB site is not present. We have now defined the existence of a NF-κB binding site in the DR4 promoter that regulates DR4 expression. Besides NF-κB, an AP-1 binding site was identified within the DR4 promoter also contributing to DR4 expression [32]. A similar site has not been identified for the DR5 promoter suggesting there are differences in the regulation of the DR4 and DR5 promoters.
Depending on the stimulus and cellular context, NF-κB can mediate both survival and apoptotic responses. Pro-apoptotic proteins expression such as Fas, Fas ligand, TRAIL, TNFα, p53, Bax, and Bcl-XS are increased mediated by NF-κB activation [33]. In contrast, anti-apoptotic proteins expression such as c-IAP1, c-IAP2, XIAP, survivin, Bcl-2, Bcl-XL, c-FLIP, and decoy receptor 1, are increased by NF-κB activation [34]. Whether NF-κB favours life or death could be a function of stimuli used [9]. Indeed, we observed different levels of NF-κB activation in response to pro-survival TNF-α and pro-apoptotic etoposide; TNF-α activated NF-κB to much higher levels than etoposide. The survival responses also seem to be dependent on the specific NF-κB subunits being activated, as well as the cell line being used. In mouse embryonic fibroblasts, the p65 subunit opposes apoptosis by inhibiting DR4 and DR5 expression. In these cells the c-Rel subunit was reported to increase the expression of DR4 and DR5 [12, 13]. Contrary to these findings, we found that the p65 subunit of NF-κB up-regulates DR4 following etoposide treatment contributing to apoptosis. Similarly, we previously reported that the p65 subunit plays a pro-apoptotic role in the induction of the DR5 gene [9]. Moreover, in HEK 293 cells c-Rel has been reported to play a minor role in the apoptotic process [9, 13]. Thus, depending upon the cell type and stimuli, NF-κB could contribute to both survival and apoptotic signalling.
MEKK1 is a serine threonine kinase that is involved in the mitogen activated protein kinase (MAPK) pathway [35]. Similarly to NF-κB, MEKK1 is involved in the activation of pro-apoptotic and pro-survival pathways. MEKK1 activates the c-Jun N-terminal kinase (JNK); a cytoprotective pathway activated by cellular stresses such as cold shock and UV radiation [17, 36, 37]. In response to DNA damaging agents, MEKK1 is activated and plays a role in the induction of apoptosis [14–16, 18]. Over-expression of kinase inactive MEKK1 blocks etoposide induced up-regulation of DR4, and DR5 and blocks etoposide induced apoptosis [3]. Furthermore, expression of kinase inactive MEKK1 inhibits etoposide induced NF-κB activation [3]. Indeed, MEKK1 has been reported to bind and activate IKK leading to the degradation of IκB and the activation of NF-κB [21, 38, 39]. We have determined that blockage of MEKK1 lowers etoposide induced DR4 expression. This effect does not seem to be dependent on the activation of the JNK pathway. Furthermore, MEKK1 induced apoptosis is blocked by inhibition of the TRAIL apoptotic pathway suggesting MEKK1 induction of DR4 contributes to the activation of the TRAIL apoptotic pathway (Fig. 6).
Although TRAIL effectively kills a wide variety of cancer cells, there are number of malignant tumours that are resistant [6]. These include pancreatic cancer [40], melanoma [41], and neuroblastoma [42]. In addition, repeated treatment with TRAIL results in acquired resistance in some cancer cell lines [6, 43]. Herein we show that a TRAIL resistant cell line HT29 can be sensitized by increasing levels of DR4 after etoposide treatment. TRAIL resistance has been linked to lowered DR4 levels [44], and mutations in the DR4 gene [6]. In TRAIL resistant leukemia cells, activation of DR4 seems to be preferred to DR5 when HDAC inhibitors are combined with agonist antibodies against DR4 and DR5 [45]. By increasing DR4 expression through combination treatments such as DNA damaging agents partially mediated by NF-κB activation, TRAIL resistance might be eliminated.
References
Walczak H, Miller RE, Ariail K et al (1999) Tumoricidal activity of tumor necrosis factor-related apoptosis-inducing ligand in vivo. Nat Med 5:157–163
Griffith TS, Chin WA, Jackson GC, Lynch DH, Kubin MZ (1998) Intracellular regulation of TRAIL-induced apoptosis in human melanoma cells. J Immunol 161:2833–2840
Gibson SB, Oyer R, Spalding AC, Anderson SM, Johnson GL (2000) Increased expression of death receptors 4 and 5 synergizes the apoptosis response to combined treatment with etoposide and TRAIL. Mol Cell Biol 20:205–212
Singh TR, Shankar S, Srivastava RK (2005) HDAC inhibitors enhance the apoptosis-inducing potential of TRAIL in breast carcinoma. Oncogene 24:4609–4623
Duiker EW, Mom CH, de Jong S et al (2006) The clinical trail of TRAIL. Eur J Cancer 42:2233–2240
Zhang L, Fang B (2005) Mechanisms of resistance to TRAIL-induced apoptosis in cancer. Cancer Gene Ther 12:228–237
Keane MM, Ettenberg SA, Nau MM, Russell EK, Lipkowitz S (1999) Chemotherapy augments TRAIL-induced apoptosis in breast cell lines. Cancer Res 59:734–741
Shetty S, Gladden JB, Henson ES et al (2002) Tumor necrosis factor-related apoptosis inducing ligand (TRAIL) up-regulates death receptor 5 (DR5) mediated by NFκB activation in epithelial derived cell lines. Apoptosis 7:413–420
Shetty S, Graham BA, Brown JG et al (2005) Transcription factor NF-κB differentially regulates death receptor 5 expression involving histone deacetylase 1. Mol Cell Biol 25:5404–5416
Yoshida T, Maeda A, Tani N, Sakai T (2001) Promoter structure and transcription initiation sites of the human death receptor 5/TRAIL-R2 gene. FEBS Lett 507:381–385
Liu X, Yue P, Khuri FR, Sun SY (2004) p53 Upregulates death receptor 4 expression through an intronic p53 binding site. Cancer Res 64:5078–5083
Ravi R, Bedi GC, Engstrom LW et al (2001) Regulation of death receptor expression and TRAIL/Apo2L-induced apoptosis by NF-κB. Nat Cell Biol 3:409–416
Chen X, Kandasamy K, Srivastava RK (2003) Differential roles of RelA (p65) and c-Rel subunits of nuclear factor κB in tumor necrosis factor-related apoptosis-inducing ligand signaling. Cancer Res 63:1059–1066
Mendoza FJ, Henson ES, Gibson SB (2005) MEKK1-induced apoptosis is mediated by Smac/Diablo release from the mitochondria. Biochem Biophys Res Commun 331:1089–1098
Bild AH, Mendoza FJ, Gibson EM et al (2002) MEKK1-induced apoptosis requires TRAIL death receptor activation and is inhibited by AKT/PKB through inhibition of MEKK1 cleavage. Oncogene 21:6649–6656
Gibson EM, Henson ES, Villanueva J, Gibson SB (2002) MEK kinase 1 induces mitochondrial permeability transition leading to apoptosis independent of cytochrome c release. J Biol Chem 277:10573–10580
Yujiri T, Fanger GR, Garrington TP, Schlesinger TK, Gibson S, Johnson GL (1999) MEK kinase 1 (MEKK1) transduces c-Jun NH2-terminal kinase activation in response to changes in the microtubule cytoskeleton. J Biol Chem 274:12605–12610
Gibson S, Widmann C, Johnson GL (1999) Differential involvement of MEK kinase 1 (MEKK1) in the induction of apoptosis in response to microtubule-targeted drugs versus DNA damaging agents. J Biol Chem 274:10916–10922
Widmann C, Gibson S, Johnson GL (1998) Caspase-dependent cleavage of signaling proteins during apoptosis. A turn-off mechanism for anti-apoptotic signals. J Biol Chem 273:7141–7147
Hirano M, Osada S, Aoki T et al (1996) MEK kinase is involved in tumor necrosis factor alpha-induced NF-κB activation and degradation of IκB-alpha. J Biol Chem 271:13234–13238
Lee FS, Peters RT, Dang LC, Maniatis T (1998) MEKK1 activates both IκB kinase alpha and IκB kinase beta. Proc Natl Acad Sci USA 95:9319–9324
David L (1998) Spector RDG, Leslie A. Leinwand. Culture and Biochemical analysis of Cells. Cold Spring Harbor Laboratory Press, New York
Spencer VA, Sun JM, Li L, Davie JR (2003) Chromatin immunoprecipitation: a tool for studying histone acetylation and transcription factor binding. Methods 31:67–75
Heinemeyer T, Wingender E, Reuter I et al (1998) Databases on transcriptional regulation: TRANSFAC, TRRD and COMPEL. Nucleic Acids Res 26:362–367
Pierce JW, Schoenleber R, Jesmok G et al (1997) Novel inhibitors of cytokine-induced IκBalpha phosphorylation and endothelial cell adhesion molecule expression show anti-inflammatory effects in vivo. J Biol Chem 272:21096–21103
Brockman JA, Scherer DC, McKinsey TA et al (1995) Coupling of a signal response domain in I κB alpha to multiple pathways for NF-κB activation. Mol Cell Biol 15:2809–2818
Pukac L, Kanakaraj P, Humphreys R et al (2005) HGS-ETR1, a fully human TRAIL-receptor 1 monoclonal antibody, induces cell death in multiple tumour types in vitro and in vivo. Br J Cancer 92:1430–1441
Vasilevskaya IA, O’Dwyer PJ (2005) 17-Allylamino-17-demethoxygeldanamycin overcomes TRAIL resistance in colon cancer cell lines. Biochem Pharmacol 70:580–589
Izeradjene K, Douglas L, Tillman DM, Delaney AB, Houghton JA (2005) Reactive oxygen species regulate caspase activation in tumor necrosis factor-related apoptosis-inducing ligand-resistant human colon carcinoma cell lines. Cancer Res 65:7436–7445
Bennett BL, Sasaki DT, Murray BW et al (2001) SP600125, an anthrapyrazolone inhibitor of Jun N-terminal kinase. Proc Natl Acad Sci USA 98:13681–13686
Sayers TJ, Murphy WJ (2006) Combining proteasome inhibition with TNF-related apoptosis-inducing ligand (Apo2L/TRAIL) for cancer therapy. Cancer Immunol Immunother 55:76–84
Guan B, Yue P, Lotan R, Sun SY (2002) Evidence that the human death receptor 4 is regulated by activator protein 1. Oncogene 21:3121–3129
Kucharczak J, Simmons MJ, Fan Y, Gelinas C (2003) To be, or not to be: NF-κB is the answer–role of Rel/NF-κB in the regulation of apoptosis. Oncogene 22:8961–8982
Dutta J, Fan Y, Gupta N, Fan G, Gelinas C (2006) Current insights into the regulation of programmed cell death by NF-κB. Oncogene 25:6800–6816
Widmann C, Gibson S, Jarpe MB, Johnson GL (1999) Mitogen-activated protein kinase: conservation of a three-kinase module from yeast to human. Physiol Rev 79:143–180
Xia Y, Makris C, Su B et al (2000) MEK kinase 1 is critically required for c-Jun N-terminal kinase activation by proinflammatory stimuli and growth factor-induced cell migration. Proc Natl Acad Sci USA 97:5243–5248
Xu S, Cobb MH (1997) MEKK1 binds directly to the c-Jun N-terminal kinases/stress-activated protein kinases. J Biol Chem 272:32056–32060
Nakano H, Shindo M, Sakon S et al (1998) Differential regulation of IκB kinase alpha and beta by two upstream kinases, NF-κB-inducing kinase and mitogen-activated protein kinase/ERK kinase kinase-1. Proc Natl Acad Sci USA 95:3537–3542
Nemoto S, DiDonato JA, Lin A (1998) Coordinate regulation of IκB kinases by mitogen-activated protein kinase kinase kinase 1 and NF-κB-inducing kinase. Mol Cell Biol 18:7336–7343
Hinz S, Trauzold A, Boenicke L et al (2000) Bcl-XL protects pancreatic adenocarcinoma cells against CD95- and TRAIL-receptor-mediated apoptosis. Oncogene 19:5477–5486
Fulda S, Kufer MU, Meyer E, van Valen F, Dockhorn-Dworniczak B, Debatin KM (2001) Sensitization for death receptor- or drug-induced apoptosis by re-expression of caspase-8 through demethylation or gene transfer. Oncogene 20:5865–5877
Eggert A, Grotzer MA, Zuzak TJ et al (2001) Resistance to tumor necrosis factor-related apoptosis-inducing ligand (TRAIL)-induced apoptosis in neuroblastoma cells correlates with a loss of caspase-8 expression. Cancer Res 61:1314–1319
Zhang L, Gu J, Lin T, Huang X, Roth JA, Fang B (2002) Mechanisms involved in development of resistance to adenovirus-mediated proapoptotic gene therapy in DLD1 human colon cancer cell line. Gene Ther 9:1262–1270
Kurbanov BM, Fecker LF, Geilen CC, Sterry W, Eberle J (2006) Resistance of melanoma cells to TRAIL does not result from upregulation of antiapoptotic proteins by NF-κB but is related to downregulation of initiator caspases and DR4. Oncogene
Kabore AF, Sun J, Hu X, McCrea K, Johnston JB, Gibson SB (2006) The TRAIL apoptotic pathway mediates proteasome inhibitor induced apoptosis in primary chronic lymphocytic leukemia cells. Apoptosis 11:1175–1193
Acknowledgements
We would like to thank Dr. Shi-Yong Sun (Emory University) for the kind gift of DR4 (−1773/+63) and DR4 (−586/+63) luciferase plasmid. Furthermore we would like to thank Human Genome Science for the kind gift of Matatumumab monoclonal antibody. The work is supported by a grant from Canadian Institutes of Health Research (MOP 68872). Mr. Mendoza is supported by a Natural Sciences and Engineering Research Council of Canada doctoral post graduate scholarship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mendoza, F.J., Ishdorj, G., Hu, X. et al. Death receptor-4 (DR4) expression is regulated by transcription factor NF-κB in response to etoposide treatment. Apoptosis 13, 756–770 (2008). https://doi.org/10.1007/s10495-008-0210-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10495-008-0210-0