Abstract
Planctomycetes are part of the PVC superphylum together with Verrucomicrobia, Chlamydiae and others. They are budding bacteria with very distinctive characteristics, such as a remarkable morphology and cell biology. Planctomycetes can be found in almost all habitats, and seem to have a preference for marine biotic and abiotic surfaces, on which they frequently occur in biofilm-forming communities. To extend the number of axenic cultures of planctomycetal strains, we isolated Pan97T from a biofilm in a volcanic site close to the Italian island Panarea in the Thyrrhenian Sea. The physiology, genome and morphology of the novel strain were characterised revealing typical planctomycetal characteristics, such as, division by polar budding and presence of crateriform structures. The strain shows pear-shaped cells of 1.5 ± 0.3 µm × 0.8 ± 0.2 µm and forms white- to cream-coloured colonies on solid medium. Strain Pan97T is mesophilic and neutrophilic, since growth was observed at a pH range of 5.5–9.5 with optimal growth at pH 7.0 and at a temperature range of 15–40 °C with a maximal growth rate at 36 °C. Pan97T has a genome size of 6,496,182 bp with a G + C content of 56.2%. 5264 protein-coding genes were identified, of which 2141 genes (41%) encode hypothetical proteins. Based on the phylogenetic analysis, we suggest that Pan97T (DSM 101992T = LMG 29460T) represents a novel species of a novel genus within the family Planctomycetaceae, for which we propose the name Bremerella gen. nov., with strain Pan97T classified as Bremerella volcania sp. nov. Based on our analysis, we also propose the reclassification of Blastopirellula cremea Lee et al. 2013 as Bremerella cremea comb. nov., as this species is considered to be the type species of the novel genus Bremerella.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Planctomycetes were first discovered in 1924 and were initially classified as eukaryotes (Gimesi 1924). Subsequently, they were acknowledged to be bacteria (Hirsch 1972). Phylogenetically, they are classified as a phylum, and together with Verrucomicrobia and Chlamydiae, they are part of the so-called PVC superphylum (Wagner and Horn 2006). Planctomycetes were considered exceptional due to their compartmentalised cell plan (Lindsay et al. 1997), nucleus-like structure (Fuerst and Webb 1991), endocytosis (Lonhienne et al. 2010) and the lack of a peptidoglycan cell wall (König et al. 1984). However, Planctomycetes faced a paradigm shift resulting from new insights into their cell biology (Wiegand et al. 2018). Before their recognition as Gram-negative bacteria, they were seen as microorganisms beyond the bacterial cell plan (Devos et al. 2013; Fuerst and Sagulenko 2011) and were postulated to link bacteria and eukaryotes (Devos and Reynaud 2010). The development of new microscopic techniques and genetic tools led to new insights into the cell organisation and physiology of the phylum Planctomycetes (Jogler and Jogler 2013; Overmann et al. 2017; Rivas-Marin et al. 2016). Several distinctive characteristics were discovered. These include an uncommon mechanism of cell division associated with the absence of canonical divisome proteins including FtsZ (Jogler et al. 2012; Pilhofer et al. 2008; Wiegand et al. 2018, 2019). A complex cytoplasmic membrane and an unusual macromolecule uptake system were found, which potentially might both take part in the digestion of internalised polysaccharides (Boedeker et al. 2017). Finally, it was found that Planctomycetes and also the closely related Verrucomicrobia (Boedeker et al. 2017) do possess a peptidoglycan cell wall (Jeske et al. 2015; van Teeseling et al. 2015). The planctomycetal cell envelope architecture is now considered Gram-negative (Devos 2014).
Planctomycetal species were found to divide either by budding or binary fission and some species even seem to use both mechanisms (Wiegand et al. 2019). Members of the class Phycisphaerae and the family Candidatus Brocadiaceae divide by binary fission while Planctomycetia divide by budding, which can be arbitrary budding in Gemmataceae and Isosphaeraceae or polar budding in Planctomycetaceae in association with a lifestyle switch (Wiegand et al. 2019). The lifecycle switch features the attachment of a swimming cell to a surface or to other cells. Subsequently, the sessile mother cell starts to divide by polar budding, thereby developing a swimming flagellated daughter cell (Jogler et al. 2011; Wiegand et al. 2018).
Besides cell biology, Planctomycetes have great ecological relevance as they are involved in global nitrogen and carbon cycles (Wiegand et al. 2018). Members of Candidatus Brocadiaceae are known for their performance of the anammox (anaerobic ammonium oxidation) reaction (Strous et al. 1999), in which ammonium is converted to dinitrogen gas (Peeters and van Niftrik 2019). Therefore, Planctomycetes can be used for the industrial removal of ammonium from wastewater (Kartal et al. 2010; Strous et al. 1999). Typically, Planctomycetes can be found in diverse habitats, especially on aquatic surfaces (Wiegand et al. 2018). They even make up the majority of cells in some biofilm communities (Webster and Negri 2006), in particular on marine algal surfaces (Bengtsson et al. 2012; Bondoso et al. 2014, 2015, 2017; Vollmers et al. 2017). In general, they often live associated with aquatic phototrophs such as diatoms, cyanobacteria and kelp, or in association with fungi, sponges or lichens (Jeske et al. 2016; Lage and Bondoso 2014). For example, Planctomycetes can make up 70% of the microbial community on the macroalgae Ecklonia radiata of the Australian shore (Wiegand et al. 2018). Also, in biofilms on surfaces of the kelp Laminaria hyperborea, Planctomycetes account for more than 50% of the bacterial biofilm (Bengtsson and Ovreas 2010). Given such high values and the fact that Planctomycetes are slow growers compared to other natural competitors in their ecological niche (Frank et al. 2015; Wiegand et al. 2018), it was hypothesised that their survival in such communities is mediated by the production of bioactive small molecules (Graça et al. 2016; Jeske et al. 2013). Also, the natural resistance of Planctomycetes to several classes of antibiotics (Cayrou et al. 2010)—often a result of the lack of targets due to their unusual cell division mechanism—could be associated with the production of bioactive compounds (Godinho et al. 2019; Graça et al. 2016; Jeske et al. 2013, 2016). Planctomycetal characteristics, including large genomes, the presence of giant genes (Kohn et al. 2016, 2019a; Wiegand et al. 2019) and their complex lifecycles has some similarities with other ‘talented’ small compound-producing bacteria, such as Actinobacteria, in particular members of the family Streptomycetaceae (Challis and Hopwood 2003).
To extend the collection of axenic cultures of Planctomycetes, we here sampled an active volcanic area in the Thyrrhenian Sea close to the Italian island Panarea (Fig. 1) and isolated the novel strain Pan97T from a red biofilm in the so called Hot Lake (Gugliandolo et al. 2015). Characterisation of its physiology and morphology, and analysis of the genome, suggest that strain Pan97T represents a novel species of a novel genus within the family Planctomycetaceae.
Sampling location of the novel strain Pan97T. The sampling spot is located in Italy, Europe (a) close to the island Panarea (b, red circle) which is part of the Aeolian Islands (b, white box). Pan97T was isolated from a hydrothermal vent (c, yellow dot), located close to Panarea. Satellite images were obtained using Google Earth Pro. (Color figure online)
Materials and methods
Sampling and cultivation
Strain Pan97T was isolated on the 9th of September 2013 from a red biofilm sampled in a hydrothermal vent close to the island Panarea in Italy (sampling site 38.5568 N, 15.1097 E) (Fig. 1). The samples were collected as previously described (Wiegand et al. 2019). Initial amplification and sequencing of the 16S rRNA gene was performed as described (Boedeker et al. 2017), which was intended to check that the strain is a Planctomycete. The strain was subsequently cultivated in M1 medium with HEPES as buffering agent and additionally supplemented with N-acetyl glucosamine (NAG) and artificial seawater (ASW). This medium, designated M1H NAG ASW, was prepared according to a protocol published earlier (Kallscheuer et al. 2019a).
Physiological analysis
Temperature and pH optima were determined by cultivating Pan97T in M1H NAG ASW medium. To this end, the strain was cultivated at temperatures of 10, 15, 20, 22, 24, 27, 30, 33, 36 and 40 °C in a shaking incubator at 110 rpm with an initial pH of 8.0. The growth rates were determined from optical density measurements at 600 nm (OD600) recorded with a UV–Vis spectrophotometer (Ultrospec II photometer LKB Biochrom). For determination of the pH optimum 100 mM HEPES was used as buffering agent for cultivations at pH 7.0, 7.5 and 8.0. For cultivations at pH 5.0 and 6.0 HEPES was replaced by 100 mM 2-(N-morpholino)ethanesulfonic acid (MES) and at pH 9.0 and 10.0 100 mM N-cyclohexyl-2-aminoethanesulfonic acid (CHES) was used as buffering agent. Cultivations for determination of the pH optimum were performed at 28 °C in a shaking incubator at 110 rpm. For determination of the salt tolerance of Pan97T different amounts of NaCl were added to M1H NAG ASW in order to yield a final NaCl concentration ranging from 0 to 2 M.
Microscopic techniques
Light microscopy and field emission scanning electron microscopy were performed as previously described (Kallscheuer et al. 2019a).
Genome information
Genome information of strain Pan97T is available from GenBank under accession number CP036289 (Wiegand et al. 2019) and the 16S rRNA gene sequence is available from GenBank under accession number MK554518.
Phylogenetic analysis
The 16S rRNA gene-based phylogenetic analysis of strain Pan97T was computed together with sequences of all described planctomycetal species (assessed in May 2019) and all isolates recently described (Boersma et al. 2019; Kallscheuer et al. 2019a, b, c, d; Kohn et al. 2019b; Peeters et al. 2019; Wiegand et al. 2019).
The 16S rRNA gene sequence alignment was made using SINA (Pruesse et al. 2012). The phylogenetic inference was calculated with RAxML (Stamatakis 2014) using the maximum likelihood method with 1000 bootstraps, nucleotide substitution model GTR, gamma distributed rate variation and estimation of proportion of invariable sites (GTRGAMMAI option) (Stamatakis 2014). In this phylogenetic analysis, three 16S rRNA genes of bacterial strains from the PVC superphylum (accession numbers AJ229235, CP010904.1 and NR_027571) were used as outgroup. The average nucleotide identity (ANI) was calculated using OrthoANI (Lee et al. 2016) and the average amino acid identity (AAI) was obtained using the aai.rb script of the enveomics collection (Rodriguez-R and Konstantinidis 2016). The percentage of conserved proteins (POCP) was calculated as described (Qin et al. 2014). The rpoB gene sequences were taken from publicly available genome annotations and sequence identities were determined as previously described (Bondoso et al. 2013). The alignment and matrix calculation was performed upon extracting only those parts of the sequence that would have been sequenced with the described primer set and the alignment and matrix calculation was done with Clustal Omega (Sievers et al. 2011).
Results and discussion
Sampling and cultivation
After sampling an active volcanic area close to the island Panarea in Italy, a novel Planctomycetes strain, designated Pan97T, was isolated from a red biofilm from the so-called Hot Lake (Gugliandolo et al. 2015; Sieland et al. 2009), a submarine depression of the seafloor at a water depth of 19.1 m (Fig. 1). Panarea is one out of seven islands of the volcanic archipelago of the Aeolian Islands (Raab et al. 2017), which have a quaternary active volcanic structure and are characterised by a volcanic landform derived from repeated episodes of volcanic activity and volcano-tectonic collapse (Lucchi et al. 2017). Increased temperatures and the presence of different sources of carbon, nitrogen and sulphur render this area an interesting location for the isolation of novel planctomycetal strains. At the time of sampling, the sampling site was filled with water at pH 5.62 and 27.6 °C, compared to the surface water at pH 8.06 and 20.0 °C.
Phylogenetic analysis
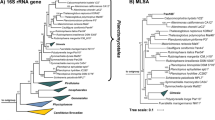
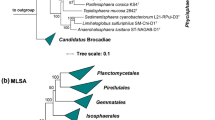
The phylogenetic analysis of Pan97T showed that this strain belongs to the Pirellula clade (Wiegand et al. 2019). Upon 16S rRNA gene sequence-based phylogenetic inference (Fig. 2), Pan97T clustered with Blastopirellula cremea LHWP2T (Lee et al. 2013), the second described species of the genus Blastopirellula. The two strains show a 16S rRNA gene identity of 99.0%. As this value is above the proposed species threshold of 98.7% (Stackebrandt and Ebers 2006), both strains may belong to the same species. However, it has been shown previously that Planctomycetes tend to have high 16S rRNA sequence identities despite belonging to different species (Bondoso et al. 2013; Kohn et al. 2019b) and higher thresholds can be applied to particular taxonomic groups (Meier-Kolthoff et al. 2013). As there is no genome available for B. cremea LHWP2T, genome-based analysis of these strains was not possible. To come to an educated decision, we compared all genomes from species annotated as belonging to B. cremea in NCBI (Table 1). By 16S rRNA gene analysis only, all of the three genomes may also belong to the species B. cremea. However, when comparing genome-based markers, this picture changed distinctly. For rpoB sequence identity, the species threshold is 95.5% (Bondoso et al. 2013) and for ANI a threshold of 95–96% is accepted (Kim et al. 2014). The ANI values as well as the rpoB sequence identities for the analysed strains are significantly below these thresholds (Table 1). We therefore assume that 16S rRNA gene identity comparisons in this clade do not follow the species threshold of 98.7% (Stackebrandt and Ebers 2006) and conclude that Pan97T does not belong to the species B. cremea.
Phylogenetic tree of the phylum Planctomycetes and of strain Pan97T. Phylogeny was computed with 16S rRNA genes using a maximum likelihood approach. At the nodes the bootstrap values after 1000 re-samplings are given in %. The outgroup contains three 16S rRNA genes from the PVC superphylum (accession numbers AJ229235, CP010904.1 and NR_027571). The Gimesia clade includes species of the genera Gimesia, Planctopirus, Fuerstiella, Schlesneria, Rubinisphaera and Planctomicrobium, while the thermophilic clade includes species of the genera Thermostilla, Thermogutta and Thermopirellula
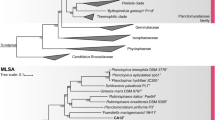
When assessing the values for species differentiation it became evident that Pan97T and B. cremea do not belong to the same genus as Blastopirellula marina. Their 16S rRNA gene identities are below the genus threshold of 94.5% (Yarza et al. 2014) and the rpoB sequence identities are below the corresponding genus threshold of 75.5–78.0% (Kallscheuer et al. 2019d) (Fig. 3 and Table 1). The same is true for AAI values, which were also below the genus threshold of 60–80% (Luo et al. 2014). Only the POCP is slightly above the proposed genus threshold of 50% (Qin et al. 2014).
Similarity values of the novel isolate Pan97T with the two described Blastopirellula species B. cremea and B. marina. The black font gives the 16S rRNA gene identity, the orange font gives the rpoB sequence identity, the turquoise font the whole genome-based average nucleotide identity (ANI) and the red font the percentage of conserved proteins (POCP). The colour of the circles discriminates between strains that are concluded to form different genera
Taken together, we propose that the genus Blastopirellula genus of the family Planctomycetaceae is divided into Blastopirellula and the novel genus Bremerella. As B. marina is the type species of the genus Blastopirellula (Schlesner et al. 2004), we propose the reclassification of B. cremea (Lee et al. 2013) as Bremerella cremea comb. nov., the type species of the genus Bremerella gen. nov. We further propose that strain Pan97T represents a novel species within this novel genus named Bremerella volcania sp. nov.
Morphological and physiological analyses
The morphology of strain Pan97T was characterised from cells harvested during the exponential growth phase. The strain shows pear-shaped cells of 1.5 ± 0.3 µm × 0.8 ± 0.2 µm (Fig. 4a–c) and forms white- to cream-coloured colonies on solid medium indicating a lack of carotenoid biosynthesis. Pan97T cells are arranged in rosettes and usually form aggregates (Fig. 4d). The cells divide by polar budding (Fig. 4a) with the daughter cells having the same shape as the mother cell. The surface of the cell shows overall crateriform structures and fibre-like structures occur at the division pole regions of the cells (Fig. 4e). No holdfast structures or flagella were observed during microscopic analyses. The phenotypic features show similarities between Pan97T and the close relatives B. cremea and B. marina in terms of cell size, aggregate formation and cell division mechanism (Table 2). Daughter cells of Pan97T have a pear-like shape, whereas they have bean-like shapes in case of B. cremea and B. marina.
Morphological characteristics of strain Pan97T. Morphologic characteristics were analysed by phase contrast (a, b) and scanning electron microscopy (d, e). The scale bar is 1 µm. Pan97T cells divide by polar budding (a) and grow in rosettes and usually form aggregates (b). Average cell size (c) was determined based on phase contrast images
In the physiological analyses, strain Pan97T was found to grow at a pH range of 5.5–9.5 with optimal growth at pH 7.0 (Fig. 5a) and at a temperature range of 15–40 °C with a maximal growth rate at 36 °C (Fig. 5b). Based on these values, strain Pan97T is mesophilic and neutrophilic. In our experiments, a maximal growth rate of 0.079 h−1 was observed, which corresponds to a generation time of 9 h. The in vitro determined optimum growth conditions of Pan97T do not fit to the environmental conditions found at the sampling location in the sampling year (pH 5.62 and 27.6 °C), indicating that this might not be the preferred location of the strain. However, the temperature optimum of 36 °C is high compared to the close relatives B. cremea and B. marina, both showing optimal growth at 30 °C, which might in some way reflect the higher temperatures at the sampling location in comparison to other locations from which we have isolated strains (e.g. the North Sea or the Baltic Sea; Wiegand et al. 2019). In cultivations with different NaCl concentrations, Pan97T showed the highest growth rate of 0.035 h−1 at an NaCl concentration of 0.1 M, but was also capable of growth at the highest tested concentration of 2 M (with 13% of the growth rate observed at 0.1 M NaCl) (Fig. 5c).
Temperature and pH optimum and NaCl tolerance of strain Pan97T. The figure shows the growth rates calculated from cultivations performed at different temperatures (a), different pH (b) and different NaCl concentrations (c). The growth rates were obtained from the slope of the plot of ln(OD600) against the cultivation time for each tested condition
Genome characteristics
The phenotypic and genomic characteristics of Pan97T and its close relatives B. cremea LHWP2T (Lee et al. 2013) and B. marina DSM 3645T (Schlesner et al. 2004) are summarised in Table 2. Pan97T has a genome size of 6,496,182 bp with a G + C content of 56.2%. 5264 protein-coding genes were identified, of which 2141 genes (41%) encode hypothetical proteins. The values correspond to 820 protein-coding genes per Mb and a coding density of 88.5%. Compared to B. marina, with a genome size of 6,663,851 bp and a G + C content of 57.4%, strain Pan97T has a slightly smaller genome and G + C content, but a slightly larger coding density. Pan97T harbours two copies of the 16S rRNA gene, while the gene is present in single copy in B. marina DSM 3645T (Table 2).
Genome-based analysis of the central carbon metabolism
As the central carbon metabolism of Planctomycetes has not been investigated in detail, we performed a genome-based analysis in order to check for presence of key metabolic enzymes participating in the central carbon metabolism in Pan97T. We included genes coding for enzymes involved in glycolytic pathways, gluconeogenesis, the tricarboxylic acid (TCA) cycle and anaplerotic reactions and compared the data obtained for Pan97T to B. marina DSM 3645T. Both strains harbour genes coding for enzymes of the Embden–Meyerhof–Parnas (EMP) pathway (the most common glycolytic pathway) as well as for the Entner–Doudoroff pathway (Table 3). From this data alone, we could not ultimately decide which pathway(s) both strains actually follow for sugar degradation. It should be noted that we were not able to identify a gene coding for phosphoglycerate mutase in B. marina DSM 3645T, however, as all other genes belonging to the EMP pathway could be identified, we assume that this glycolytic route is functional in B. marina. Pan97T and B. marina DSM 3645T harbour a complete gene set required for a functional pentose phosphate pathway and TCA cycle (Table 2), which leads to the assumption that the central carbon metabolism of both strains is similar to most heterotrophic bacteria. For the gluconeogenesis, we could only identify a partial set of genes in both strains, with only two genes, ppdK (encoding pyruvate phosphate dikinase) and pckA (encoding phosphoenolpyruvate carboxykinase), present in both strains. It thus remains to be elucidated if the two strains are capable of de novo sugar biosynthesis. Additionally, both strains lack the glyoxylate shunt, which is typically required as the anaplerotic pathway during growth either with acetate or with compounds that are degraded to acetate or acetyl-CoA (e.g. fatty acids). Absence of the glyoxylate shunt suggests that both strains are either unable to use such compounds as sole carbon and energy source or that they follow other pathways with a related function.
Genes involved in biosynthesis of flagella
As we were not able to confirm presence of flagella in Pan97T during microscopic analyses (Fig. 4), we searched for genes coding for enzymes involved in the biosynthesis of flagella in the genomes of Pan97T and B. marina DSM 3645T. Indeed, the same set of genes required for assembly of flagella is present in both species (Table 4), suggesting no major differences in mechanism of motility of the two related species. The analysis also showed that proteins with regulatory activity, e.g. transcriptional regulators (FlhC, FlhD, FliT) and (anti-)sigma factors (FlgM, FliA) are probably absent in both strains (Table 4). This indicates that regulation of flagella biosynthesis might involve other regulatory proteins distinct from those of model organisms in which flagella biosynthesis was investigated before.
Putative gene clusters involved in secondary metabolite production
Species of the family Planctomycetaceae typically have complex lifestyles and are often found in higher abundance in biofilms on nutrient-rich biotic surfaces in marine environments. Such a lifestyle is believed to require an additional level of regulation/interaction by production of secondary metabolites, which may allow its producer to cope with abiotic (e.g. UV light) or biotic stresses (competing bacteria, etc.). To obtain a first insight into the secondary metabolism of strain Pan97T we performed an AntiSMASH analysis based on the genome sequence (Blin et al. 2019). Five clusters coding for enzymes known to produce secondary metabolites were identified. One cluster is associated with terpene production, two with polyketide biosynthesis (one putative type I polyketide synthase and one putative mixed type I polyketide synthase-non-ribosomal peptide synthetase). In addition, one putative cluster each was found to code for putative bacteriocin- and resorcinol biosynthetic proteins. The number of biosynthetic clusters is in the middle range, as 3–13 putative biosynthetic clusters were found for the planctomycetal genomes analysed so far (Wiegand et al. 2019), thereby suggesting some potential of Pan97T as a source of novel bioactive compounds.
Description of Bremerella gen. nov.
Bremerella (Bre.me.rel’la. N.L. dim. fem. n. Bremerella; in honour of the German microbiologist Erhard Bremer).
Species belonging to this genus have pear- to ovoid shaped cells, form aggregates and divide by polar budding. The type species is Bremerella cremea comb. nov. (basonym: Blastopirellula cremea).
Description of Bremerella cremea comb. nov.
Bremerella cremea (cre.me’a. N.L. fem. adj. cremea cream–white).
Basonym: Blastopirellula cremea Lee et al. 2013.
Characteristics are as described previously (Lee et al. 2013). The type strain is LHWP2T ( KACC 15559T = JCM 17758T).
Description of Bremerella volcania sp. nov.
Bremerella volcania (vol.ca’ni.a. L. fem. adj. volcania corresponding to the volcano area in which the type strain was found).
Cells are pear-shaped (1.5 ± 0.3 µm × 0.8 ± 0.2 µm) and form white- to cream-coloured colonies on solid medium. Cells are arranged in rosettes and usually form aggregates. Division is performed by polar budding with the daughter cell having the same shape as the mother cell. Cells of the type strain grow at a temperature range of 15–40 °C and a pH range of 5.5–9.5. Genome information of the type strain is available from GenBank under accession number CP036289 and the 16S rRNA gene sequence is available from GenBank accession number MK554518. The type strain is Pan97T (DSM 101992T = LMG 29460T) isolated from a red biofilm obtained from a hydrothermal vent close to the island Panarea in Italy in September 2013.
References
Bengtsson MM, Ovreas L (2010) Planctomycetes dominate biofilms on surfaces of the kelp Laminaria hyperborea. BMC Microbiol 10:261
Bengtsson MM, Sjotun K, Lanzen A, Ovreas L (2012) Bacterial diversity in relation to secondary production and succession on surfaces of the kelp Laminaria hyperborea. ISME J 6:2188–2198
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T (2019) AntiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:W81–W87
Boedeker C, Schuler M, Reintjes G, Jeske O, van Teeseling MC, Jogler M, Rast P, Borchert D, Devos DP, Kucklick M, Schaffer M, Kolter R, van Niftrik L, Engelmann S, Amann R, Rohde M, Engelhardt H, Jogler C (2017) Determining the bacterial cell biology of planctomycetes. Nat Commun 8:14853
Boersma A, Kallscheuer N, Wiegand S, Rast R, Peeters S, Mesman R, Heuer A, Boedeker C, Jetten M, Rohde M, Jogler M, Jogler C (2019) Alienimonas californiensis gen. nov. sp. nov., a novel Planctomycete isolated from the kelp forest in Monterey Bay. Antonie Van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01367-4
Bondoso J, Harder J, Lage OM (2013) rpoB gene as a novel molecular marker to infer phylogeny in Planctomycetales. Antonie Van Leeuwenhoek 104:477–488
Bondoso J, Balague V, Gasol JM, Lage OM (2014) Community composition of the Planctomycetes associated with different macroalgae. FEMS Microbiol Ecol 88:445–456
Bondoso J, Albuquerque L, Nobre MF, Lobo-da-Cunha A, da Costa MS, Lage OM (2015) Roseimaritima ulvae gen. nov., sp. nov. and Rubripirellula obstinata gen. nov., sp. nov. two novel planctomycetes isolated from the epiphytic community of macroalgae. Syst Appl Microbiol 38:8–15
Bondoso J, Godoy-Vitorino F, Balague V, Gasol JM, Harder J, Lage OM (2017) Epiphytic Planctomycetes communities associated with three main groups of macroalgae. FEMS Microbiol Ecol 93:fiw255
Cayrou C, Raoult D, Drancourt M (2010) Broad-spectrum antibiotic resistance of Planctomycetes organisms determined by Etest. J Antimicrob Chemother 65:2119–2122
Challis GL, Hopwood DA (2003) Synergy and contingency as driving forces for the evolution of multiple secondary metabolite production by Streptomyces species. Proc Natl Acad Sci USA 100:14555–14561
Devos DP (2014) Re-interpretation of the evidence for the PVC cell plan supports a Gram-negative origin. Antonie Van Leeuwenhoek 105:271–274
Devos DP, Reynaud EG (2010) Evolution. Intermediate steps. Science 330:1187–1188
Devos DP, Jogler C, Fuerst JA (2013) The 1st EMBO workshop on PVC bacteria—Planctomycetes–Verrucomicrobia–Chlamydiae superphylum: exceptions to the bacterial definition? Antonie Van Leeuwenhoek 104:443–449
Frank O, Michael V, Pauker O, Boedeker C, Jogler C, Rohde M, Petersen J (2015) Plasmid curing and the loss of grip–the 65-kb replicon of Phaeobacter inhibens DSM 17395 is required for biofilm formation, motility and the colonization of marine algae. Syst Appl Microbiol 38:120–127
Fuerst JA, Sagulenko E (2011) Beyond the bacterium: planctomycetes challenge our concepts of microbial structure and function. Nat Rev Microbiol 9:403–413
Fuerst JA, Webb RI (1991) Membrane-bounded nucleoid in the eubacterium Gemmata obscuriglobus. Proc Natl Acad Sci USA 88:8184–8188
Gimesi N (1924) Hydrobiologiai Tanulmanyok (Hydrobiologische Studien). I: Planctomyces bekefii gim. nov. gen. et sp. Kiadja a Magyar Ciszterci Rend Budapest, pp 1–8
Godinho O, Calisto R, Ovreas L, Quinteira S, Lage OM (2019) Antibiotic susceptibility of marine Planctomycetes. Antonie Van Leeuwenhoek 112:1273–1280
Graça AP, Calisto R, Lage OM (2016) Planctomycetes as novel source of bioactive molecules. Front Microbiol 7:1241
Gugliandolo C, Lentini V, Bunk B, Overmann J, Italiano F, Maugeri TL (2015) Changes in prokaryotic community composition accompanying a pronounced temperature shift of a shallow marine thermal brine pool (Panarea Island, Italy). Extremophiles 19:547–559
Hirsch P (1972) Two identical genera of budding and stalked bacteria: Planctomyces Gimesi 1924 and Blastocaulis Henrici and Johnson 1935. Int J Syst Evol Microbiol 22:107–111
Jeske O, Jogler M, Petersen J, Sikorski J, Jogler C (2013) From genome mining to phenotypic microarrays: Planctomycetes as source for novel bioactive molecules. Antonie Van Leeuwenhoek 104:551–567
Jeske O, Schuler M, Schumann P, Schneider A, Boedeker C, Jogler M, Bollschweiler D, Rohde M, Mayer C, Engelhardt H, Spring S, Jogler C (2015) Planctomycetes do possess a peptidoglycan cell wall. Nat Commun 6:7116
Jeske O, Surup F, Ketteniss M, Rast P, Forster B, Jogler M, Wink J, Jogler C (2016) Developing techniques for the utilization of planctomycetes as producers of bioactive molecules. Front Microbiol 7:1242
Jogler M, Jogler C (2013) Toward the development of genetic tools for Planctomycetes. In: Fuerst JA (ed) Planctomycetes: cell structure, origins and biology. Humana Press, Totowa, pp 141–164
Jogler C, Glockner FO, Kolter R (2011) Characterization of Planctomyces limnophilus and development of genetic tools for its manipulation establish it as a model species for the phylum Planctomycetes. Appl Environ Microbiol 77:5826–5829
Jogler C, Waldmann J, Huang X, Jogler M, Glöckner FO, Mascher T, Kolter R (2012) Identification of proteins likely to be involved in morphogenesis, cell division, and signal transduction in Planctomycetes by comparative genomics. J Bacteriol 194:6419–6430
Kallscheuer N, Jogler M, Wiegand S, Peeters S, Heuer A, Boedeker C, Jetten M, Rohde M, Jogler C (2019a) Rubinisphaera italica sp. nov. isolated from a hydrothermal area in the Tyrrhenian Sea close to the volcanic island Panarea. Antonie Van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01329-w
Kallscheuer N, Jogler M, Wiegand S, Peeters S, Heuer A, Boedeker C, Jetten M, Rohde M, Jogler C (2019b) Three novel Rubripirellula species isolated from artificial plastic surfaces submerged in the German part of the Baltic Sea and the estuary of the river Warnow. Antonie Van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01368-3
Kallscheuer N, Wiegand S, Jogler M, Boedeker C, Peeters S, Rast P, Heuer A, Jetten M, Rohde MCJ (2019c) Rhodopirellula heiligendammensis sp. nov., Rhodopirellula pilleata sp. nov., and Rhodopirellula solitaria sp. nov. isolated from natural or artificial marine surfaces in Northern Germany and California, USA. Antonie Van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01366-5
Kallscheuer N, Wiegand S, Peeters SH, Jogler M, Boedeker C, Heuer A, Rast P, Jetten MSM, Rohde M, Jogler C (2019d) Description of three bacterial strains belonging to the new genus Novipirellula gen. nov., reclassificiation of Rhodopirellula rosea and Rhodopirellula caenicola and readjustment of the genus threshold of the phylogenetic marker rpoB for Planctomycetaceae. Antonie van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01374-5
Kartal B, Kuenen JG, van Loosdrecht MC (2010) Sewage treatment with anammox. Science 328:702–703
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351
Kohn T, Heuer A, Jogler M, Vollmers J, Boedeker C, Bunk B, Rast P, Borchert D, Glockner I, Freese HM, Klenk HP, Overmann J, Kaster AK, Rohde M, Wiegand S, Jogler C (2016) Fuerstia marisgermanicae gen. nov., sp. nov., an Unusual Member of the Phylum Planctomycetes from the German Wadden Sea. Front Microbiol 7:2079
Kohn T, Heuer A, Jogler M, Vollmers J, Boedeker C, Bunk B, Rast P, Borchert D, Glockner I, Freese HM, Klenk HP, Overmann J, Kaster AK, Rohde M, Wiegand S, Jogler C (2019a) Corrigendum: Fuerstia marisgermanicae gen. nov., sp. nov., an unusual member of the phylum Planctomycetes from the German Wadden Sea. Front Microbiol 10:1029
Kohn T, Wiegand S, Boedeker C, Rast P, Heuer A, Schüler M, Rohde C, Müller R-W, Brümmer F, Rohde M, Engelhardt H, Jogler M, Jogler C (2019b) Planctopirus ephydatiae, a novel planctomycetal species isolated from the freshwater sponge Ephydatia fluviatilis. Syst Appl Microbiol. https://doi.org/10.1016/j.syapm.2019.126022
König E, Schlesner H, Hirsch P (1984) Cell wall studies on budding bacteria of the Planctomyces/Pasteuria group and on a Prosthecomicrobium sp. Arch Microbiol 138:200–205
Lage OM, Bondoso J (2014) Planctomycetes and macroalgae, a striking association. Front Microbiol 5:267
Lee HW, Roh SW, Shin NR, Lee J, Whon TW, Jung MJ, Yun JH, Kim MS, Hyun DW, Kim D, Bae JW (2013) Blastopirellula cremea sp. nov., isolated from a dead ark clam. Int J Syst Evol Microbiol 63:2314–2319
Lee I, Ouk Kim Y, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103
Lindsay MR, Webb RI, Fuerst JA (1997) Pirellulosomes: a new type of membrane-bounded cell compartment in planctomycete bacteria of the genus Pirellula. Microbiol UK 143:739–748
Lonhienne TG, Sagulenko E, Webb RI, Lee KC, Franke J, Devos DP, Nouwens A, Carroll BJ, Fuerst JA (2010) Endocytosis-like protein uptake in the bacterium Gemmata obscuriglobus. Proc Natl Acad Sci USA 107:12883–12888
Lucchi F, Romagnoli C, Tranne CA (2017) Volcanic landforms and landscapes of the Aeolian Islands (Southern Tyrrhenian Sea, Sicily): implications for hazard evaluation. In: Soldati M, Marchetti M (eds) Landscapes and landforms of Italy. Springer, Cham, pp 443–453
Luo C, Rodriguez RL, Konstantinidis KT (2014) MyTaxa: an advanced taxonomic classifier for genomic and metagenomic sequences. Nucleic Acids Res 42:e73
Meier-Kolthoff JP, Göker M, Spröer C, Klenk HP (2013) When should a DDH experiment be mandatory in microbial taxonomy? Arch Microbiol 195:413–418
Overmann J, Abt B, Sikorski J (2017) Present and future of culturing bacteria. Annu Rev Microbiol 71:711–730
Peeters SH, van Niftrik L (2019) Trending topics and open questions in anaerobic ammonium oxidation. Curr Opin Chem Biol 49:45–52
Peeters S, Wiegand S, Kallscheuer N, Jogler M, Heuer A, Jetten M, Rast P, Boedeker C, Rohde M, Jogler C (2019) Three marine strains constitute the novel genus and species Crateriforma conspicua gen. nov. sp. nov. in the phylum Planctomycetes. Antonie van Leeuwenhoek. Accepted manuscript (ANTO-D-19-00328)
Pilhofer M, Rappl K, Eckl C, Bauer AP, Ludwig W, Schleifer KH, Petroni G (2008) Characterization and evolution of cell division and cell wall synthesis genes in the bacterial phyla Verrucomicrobia, Lentisphaerae, Chlamydiae, and Planctomycetes and phylogenetic comparison with rRNA genes. J Bacteriol 190:3192–3202
Pruesse E, Peplies J, Glöckner FO (2012) SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28:1823–1829
Qin Q-L, Xie B-B, Zhang X-Y, Chen X-L, Zhou B-C, Zhou J, Oren A, Zhang Y-Z (2014) A proposed genus boundary for the prokaryotes based on genomic insights. J Bacteriol 196:2210–2215
Raab G, Halpern D, Scarciglia F, Raimondi S, Norton K, Pettke T, Hermann J, de Castro Portes R, Aguilar Sanchez AM, Egli M (2017) Linking tephrochronology and soil characteristics in the Sila and Nebrodi mountains, Italy. CATENA 158:266–285
Rivas-Marin E, Canosa I, Santero E, Devos DP (2016) Development of genetic tools for the manipulation of the Planctomycetes. Front Microbiol 7:914
Rodriguez-R LM, Konstantinidis KT (2016) The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. PeerJ Preprints 4:e1900v1
Schlesner H, Rensmann C, Tindall BJ, Gade D, Rabus R, Pfeiffer S, Hirsch P (2004) Taxonomic heterogeneity within the Planctomycetales as derived by DNA-DNA hybridization, description of Rhodopirellula baltica gen. nov., sp. nov., transfer of Pirellula marina to the genus Blastopirellula gen. nov. as Blastopirellula marina comb. nov. and emended description of the genus Pirellula. Int J Syst Evol Microbiol 54:1567–1580
Sieland R, Steinbrückner D, Hamel M, Merkel B, Schipek M (2009) Geochemical investigations and gas quantification of submarine fluid discharges in the hydrothermal system of Panarea (Aeolian Islands, Italy). In: 1st International workshop research in shallow marine and fresh water systems, pp 87
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Stackebrandt E, Ebers J (2006) Taxonomic parameter revisited: tarnished gold standards. Microbiol Today 33:152–155
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Strous M, Fuerst JA, Kramer EH, Logemann S, Muyzer G, van de Pas-Schoonen KT, Webb R, Kuenen JG, Jetten MS (1999) Missing lithotroph identified as new planctomycete. Nature 400:446–449
van Teeseling MC, Mesman RJ, Kuru E, Espaillat A, Cava F, Brun YV, VanNieuwenhze MS, Kartal B, van Niftrik L (2015) Anammox Planctomycetes have a peptidoglycan cell wall. Nat Commun 6:6878
Vollmers J, Frentrup M, Rast P, Jogler C, Kaster AK (2017) Untangling genomes of novel planctomycetal and verrucomicrobial species from Monterey Bay kelp forest metagenomes by refined binning. Front Microbiol 8:472
Wagner M, Horn M (2006) The Planctomycetes, Verrucomicrobia, Chlamydiae and sister phyla comprise a superphylum with biotechnological and medical relevance. Curr Opion Biotechnol 17:241–249
Webster NS, Negri AP (2006) Site-specific variation in Antarctic marine biofilms established on artificial surfaces. Environ Microbiol 8:1177–1190
Wiegand S, Jogler M, Jogler C (2018) On the maverick Planctomycetes. FEMS Microbiol Rev 42:739–760
Wiegand S, Jogler M, Boedeker C, Pinto D, Vollmers J, Rivas-Marín E, Kohn T, Peeters SH, Heuer A, Rast P, Oberbeckmann S, Bunk B, Jeske O, Meyerdierks A, Storesund JE, Kallscheuer N, Lücker S, Lage OM, Pohl T, Merkel BJ, Hornburger P, Müller R-W, Brümmer F, Labrenz M, Spormann AM, Op den Camp H, Overmann J, Amann R, Jetten MSM, Mascher T, Medema MH, Devos DP, Kaster A-K, Øvreås L, Rohde M, Galperin MY, Jogler C (2019) Cultivation and functional characterization of 79 Planctomycetes uncovers their unique biology. Nat Microbiol. https://doi.org/10.1038/s41564-019-0588-1
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer KH, Whitman WB, Euzeby J, Amann R, Rossello-Mora R (2014) Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol 12:635–645
Acknowledgements
Part of this research was funded by the Deutsche Forschungsgemeinschaft Grants KA 4967/1-1 and JO 893/4-1, Grant ALWOP.308 of the Nederlandse Organisatie voor Wetenschappelijk Onderzoek (NWO), SIAM (Soehngen Institute for Anaerobic Microbiology) Grant No. 024002002 and the Radboud Excellence fellowship. We thank Ina Schleicher for skillful technical assistance. Brian Tindall and Regine Fähnrich from the DSMZ as well as the BCCM/LMG Bacteria collection we thank for support during strain deposition. We thank the Scientific Diving Center of the Bergakademie Freiberg, Germany, Thomas Pohl, Peter Hornburger and all participants of the 2013 Panarea Expedition for sampling support.
Author information
Authors and Affiliations
Contributions
SR wrote the manuscript, analysed the data and prepared the figures, SW and MJ performed the genomic and phylogenetic analysis, AH and PR isolated the strains and performed the cultivation and strain deposition, SHP and CB performed the light microscopic analysis, MSMJ and NK contributed to text preparation and revised the manuscript, MR performed the electron microscopic analysis, CJ took the samples and supervised the study. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical statement
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rensink, S., Wiegand, S., Kallscheuer, N. et al. Description of the novel planctomycetal genus Bremerella, containing Bremerella volcania sp. nov., isolated from an active volcanic site, and reclassification of Blastopirellula cremea as Bremerella cremea comb. nov.. Antonie van Leeuwenhoek 113, 1823–1837 (2020). https://doi.org/10.1007/s10482-019-01378-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-019-01378-1