Abstract
A metal-resistant and phosphate-solubilising bacterium, designated as strain D414T, was isolated from heavy metal (Pb, Cd, Cu and Zn)-polluted paddy soils at the surrounding area of Dabao Mountain Mine in Southeast China. The minimum inhibitory concentrations of heavy metals for strain D414T were 2000 mg L−1 (Cd), 800 mg L−1 (Pb), 150 mg L−1 (Cu) and 2500 mg L−1 (Zn). The strain possessed plant growth-promoting properties, such as 1-aminocyclopropane-1-carboxylate assimilation, indole production and phosphate solubilisation. Analysis of 16S rRNA gene sequence indicated that the isolate is a member of the genus Burkholderia where strain D414T formed a distinct phyletic line with validly described Burkholderia species. Strain D414T is closely related to Burkholderia tropica DSM 15359T, B. bannensis NBRC E25T and B. unamae DSM 17197T, with 98.5, 98.3 and 98.3 % sequence similarities, respectively. Furthermore, less than 34 % DNA–DNA relatedness was detected between strain D414T and the type strains of the phylogenetically closest species of Burkholderia. The dominant fatty acids of strain D414T were C14:0, C16:0, C17:0 cyclo and C18:1 ω7c. The DNA G+C content was 62.3 ± 0.5 mol%. On the basis of genotypic, phenotypic and phylogenetic data, strain D414T represents a novel species, for which the name Burkholderia metalliresistens sp. nov. is proposed, with D414T (=CICC 10561T = DSM 26823T) as the type strain
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The genus Burkholderia, with Burkholderia cepacia as the type species, was separated from the former Pseudomonas rRNA homology group II by Yabuuchi et al. 1992. More than 80 Burkholderia species with validly published names have been reported to date. Burkholderia species were isolated from highly diverse habitats (Coenye and Vandamme 2003; Caballero-Mellado et al. 2004; Reis et al. 2004; Chen et al. 2007). Burkholderia is a genus with a steady growth of species numbers. Previously described Burkholderia species were classified into two major species clusters and several subgroups on the basis of the phylogenetic analyses of 16S rRNA, recA, gyrB, rpoB and acdS gene sequences, as well as genome sequences (Estrada-de los Santos et al. 2013). Group A contains plant-associated species and some saprophytic bacterial species. Group B comprises members of the B. cepacia complex (opportunistic human pathogens), the B. pseudomallei subgroup (both human and animal pathogens) and the B. andropogonis subgroup (including B. andropogonis, B. rhizoxinica, B. endofungorum, B. caryophylli, B. symbiotica and B. soli) (Suárez-Moreno et al. 2012; Estrada-de los Santos et al. 2013; Martínez-Aguilar et al. 2013).
An increasing number of Burkholderia species with plant growth-promoting (PGP) traits and legume plant nodulation abilities in legumes have been isolated from soils and plant tissues. Burkholderia species with nitrogen-fixing ability were classified under the plant-associated group; these species include B. unamae, B. xenovorans, B. kururiensis, B. silvatlantica, B. tropica, B. ferrariae, B. bannensis, B. heleia, B. terrae and B. gisengisoli (Suárez-Moreno et al. 2012). The B. cepacia complex species B. vietnamiensis is also a nitrogen fixer. Moreover, B. sabiae, B. tuberum, B. phymatum, B. mimosarum and B. nodosa can fix nitrogen in symbiosis with legumes (Vandamme et al. 2002; Chen et al. 2006, 2007, 2008). Plant growth-promoting rhizobacteria (PGPR) with 1-aminocyclopropane-1-carboxylate (ACC) deaminase promotes plant growth by suppressing the increased level of ethylene content, which causes growth inhibition, in plant tissues under abiotic or biotic stress (Blaha et al. 2006). Plant-beneficial rhizobacteria produce indole-3-acetic acid (IAA) to promote growth. B. phytofirmans and B. unamae reportedly produce ACC deaminase and IAA (Caballero-Mellado et al. 2007; Onofre-Lemus et al. 2009). Furthermore, plant-associated Burkholderia species, such as B. tuberum STM678, solubilise phosphate and produce siderophores (Angus et al. 2013). The shoot lengths and dry weights of Macroptilium atropurpureum which were inoculated with strain STM678 significantly increased.
Many Burkholderia species play important roles in pollution remediation (Lim et al. 2008; Otsuka et al. 2011; Lu et al. 2012). Jiang et al. (2008) isolated the heavy metal-tolerant Burkholderia sp. J62 from heavy metal-contaminated soils and found that this strain promotes growth through IAA production, siderophore synthesis and phosphate solubilisation. Moreover, the Cd and Pb contents in the biomasses and tissues of Zea mays L. and Lycopersicon esculentum increase upon inoculation with Burkholderia sp. J62 in heavy metal-contaminated soils. Some scholars isolated Burkholderia species which exhibit heavy metal resistance, promote plant growth and enhance metal uptake from heavy metal-polluted soils (Li et al. 2007; Jiang et al. 2008; Guo et al. 2011; Martínez-Aguilar et al. 2013). The present study aims to investigate the taxonomy of an isolate which belongs to a new Burkholderia species. The name Burkholderia metalliresistens sp. nov. is proposed for this new species.
Materials and methods
Isolation and screening of isolate
Strain D414T was isolated from arable layers (0–20 cm) of heavy metal-polluted paddy soils at the surrounding area of Dabao Mountain Mine, Guangdong, Southeast China. The basic properties of the soil samples are shown in Supplementary Table 1. Ten grams of soils that were collected from the heavy metal-polluted paddy fields were added to 100 ml sterile modified SMN mediums without agar in a clean bench. The enrichment mediums were incubated at 200 rpm for 72 h at 30 °C in a rotary shaker. The selective medium used was modified SMN medium [2 g L−1 (NH4)2SO4, 5 g L−1 mannitol, 0.5 g L−1 K2HPO4, 0.5 g L−1 MgSO4, 0.1 g L−1 NaCl, 0.05 g L−1 FeSO4, 0.05 g L−1 MnSO4, 20 g L−1 agar and 1000 mL of distilled water; pH 6.8 ± 0.2] supplemented with 50 mg L−1 Cd as CdCl2. Single colonies which grew well were selected and streaked on other modified SMN agar plates supplemented with 0.5 % ACC as the sole carbon source. The isolates were purified by streaking in three other modified SMN agar plates. Finally, the pure isolates were stored in 30 % v/v glycerol at −80 °C for long-term storage and on modified SMN slants for short-term storage.
Phenotypic characteristics
Morphological properties were examined by light microscopy (Carl Zeiss Axioskop, Germany) after 3 days of culture on modified SMN agar at 30 °C. After serial dehydration in 50, 60, 70, 80, 90 and 100 % ethanol solutions (three times for 10 min at each stage), the critical-point dried samples were sputter-coated with gold and viewed under a scanning electron microscope (FEI Quanta 200, Netherlands). The Gram reaction was performed following the method described by Palinska and Marquardt (2008).
Strain D414T and reference type strains were inoculated with a suspension of late exponential growth phase cells in 0.7 % (w/v) YNB minimal growth medium (Difco) adjusted to pH 7.0, and their utilisation of different substrates and enzyme activities was determined with the API 20NE and API 50CH microtest systems in accordance with the manufacturers’ instructions (bioMérieux, France). The effects of temperature and pH on the growth of strain D414T were determined in modified SMN medium. The growth of the strain on LB broth (Difco), yeast nitrogen base agar (Difco) and trypticase soy agar (Difco) was also evaluated.
Chemotaxonomic properties
For chemotaxonomic property studies, strain D414T and reference type strains (B. tropica DSM 15359T, B. bannensis NBRC E25T and B. unamae DSM 17197T) were cultured in modified SMN medium at 30 °C for 48 h. The strains in the late exponential growth phase were harvested by centrifugation at 10,000×g for 10 min and washed with distilled water several times for further study. The whole-cell fatty acids were saponified, methylated and then extracted using the standard protocol of the MIDI Sherlock Microbial Identification System (version 6.0). Fatty acids were analysed by GC (Agilent Technologies 6850) and identified using the TSBA6 database of the Microbial Identification System (Sasser 1990).
The growth and nitrogen-fixing ability of strain D414T were evaluated in Ashby’s nitrogen-free medium and serials of other test media (Jones 1970). The nitrogenase activity of strain D414T was determined by acetylene reduction assay (ARA) (Hardy et al. 1968). Flasks which contain 20 mL of Ashby’s nitrogen-free semi-solid medium (with 5 g L−1 agar) were inoculated with the isolate at a concentration of 1 × 108 cfu mL−1. After incubation at 30 °C for 48 h, 10 % of the gas volume in each flask was replaced with acetylene. The flasks were incubated for 1 h at 30 °C and analysed for ethylene production by GC with an FID detector (Agilent 7890A). B. tropica DSM 15359T and B. unamae DSM 17197T served as positive controls. Uninoculated flasks served as negative controls. The experiment was carried out thrice.
Whole-cell protein SDS-PAGE analyses were performed as described by Laemmli (1970). Bacterial cultures (1.0 mL) in the late exponential growth phase were harvested and washed as described above. The pellets were resuspended in 100 μL of 0.125 M Tris–HCl, 4 % SDS, 20 % glycerol and 10 % mercaptoethanol at pH 6.8. The mixtures were heated at 100 °C for 4 min. The samples were cooled on ice and centrifuged at 10,000×g for 10 min, and the concentration was recovered and stored at −20 °C until use. The lysate solutions (10 µL) were subjected to SDS-PAGE electrophoresis on vertical slabs at 50 mA and 120 V. The samples were fixed by immersing the gels in a 10 % (v/v) trichloroacetic solution for 1 h and then stained overnight with Coomassie blue [0.25 % (w/v) Coomassie blue R-250, 50 % (v/v) methanol and 10 % (v/v) acetic acid]. Excess stain was removed by rinsing with a solution of 25 % (v/v) methanol and 10 % (v/v) acetic acid (Ghazi et al. 2009).
Phylogenetic analysis
Genomic DNA extraction and 16S rRNA gene PCR were carried out following the procedures by Chun and Goodfellow (1995). The universal bacterial 16S rRNA gene primers (the forward primer P1and the reverse primer P6) were used (Wei et al. 2002). The PCR product was purified and directly sequenced by an automated DNA Sequencing System (ABI 3730XL, USA) with the forward primer P1 and the reverse primer P6 as described above. Two sequencing reactions with the forward primer P1 and the reverse primer P6 were carried out. Sequence fragments were then assembled using Contig Express (Vector NTI Suite 8; InforMax, Bethesda, MD, USA). The almost complete 16S rRNA gene sequence of the strain was aligned by ClustalX version 1.8 (Thompson et al. 1997) with almost complete 16S rRNA gene sequences of validly described type strains obtained from the GenBank/EMBL/DDBJ databases and EzTaxon-e (Kim et al. 2012). A phylogenetic tree was constructed by neighbour-joining (Saitou and Nei, 1987), minimum-evolution (Rzhetsky and Nei 1993) and maximum-parsimony (Fitch 1971) in TREECON 1.3b (Van de Peer and De Wachter. 1994) and MEGA 3.1 (Kumar et al. 2004). The genetic distance matrices were estimated by the Kimura two-parameter model (Kimura 1980). The topology of the tree was evaluated by bootstrap analysis based on 1000 replicates (Felsenstein 1985).
DNA–DNA hybridisation and G+C content
DNA–DNA relatedness between strain D414T and its most closely related type strain described by De Ley et al. (1970) was determined by thermal denaturation with a spectrophotometer (Beckman DU 800, USA). The DNA G+C content was determined through HPLC (Mesbah et al. 1989). All analyses on DNA–DNA relatedness and G+C content were carried out in triplicates.
Heavy metal resistance and phosphate solubilisation properties
The minimum inhibitory concentrations (MICs) of the metals (Pb, Cd, Cu and Zn) for the strain D414T were determined by plate dilution as described by Aleem et al. (2003). The modified SMN agar without the metals served as the control. The cultures were incubated at 30 °C for 7 days, and the experiments were carried out in triplicate.
The phosphate solubilisation ability of strain D414T was determined in Pikovskays’s medium (Zaidi et al. 2006) with 0.5 % tricalcium phosphate. When the bacterial suspension reached 1.0 (1 × 108 cfu mL−1) of optical density (600 nm), 1 mL of the suspension was added into 250 mL conical flasks which contain 100 mL of the medium. After incubation at 200 rpm for 72 h at 30 °C in a rotary shaker, the supernatants were collected by centrifugation at 8000×g for 10 min. The soluble phosphate in the supernatants was estimated by Mo-blue (Watanabe and Olsen 1965). Each experiment was performed in triplicates.
Results and discussion
In the present study, strain D414T with the ability to assimilate ACC was isolated from a heavy metal-polluted paddy soil. Glick et al. (2007) reported that the plant-associated strains with the ability to assimilate ACC can prevent ethylene-induced growth inhibition by decreasing the amount of ACC, the immediate precursor of ethylene, through hydrolysis. ACC assimilation is a key plant-beneficial trait of PGPR (Blaha et al. 2006). Strain D414T was found to be 0.5–0.7 µm × 0.8–2.0 µm in cell size, Gram negative, aerobic and non-motile (Fig. 1). After 3 days of cultivation at 30 ± 1 °C, milk-like, yellowish-white, convex and circular colonies with clear margins were observed on the modified SMN agar plate. A diffusible pigment was not observed in the modified SMN medium. Strain D414T grew within the temperature range of 10–50 °C (optimum 30 °C) and pH range of 3.0 to 8.0 (optimum 7.0) (Table 1). The strain grew well on LB broth (Difco), yeast nitrogen base agar (Difco) and trypticase soy agar (Difco) but weakly on Ashby’s nitrogen-free medium. Furthermore, the result of the ARA showed that the nitrogenase activity of strain D414 was ranged from 87.2 to 105.6 nmol C2H4 h−1 mL−1.
The major fatty acids of strain D414T were C14: 0, C16: 0, C17: 0 cyclo and C18: 1 ω7c (Supplementary Table 2). The results demonstrate that the fatty acid characteristics of strain D414T are consistent with those of the genus Burkholderia (Coenye et al. 2000). However, strain D414T differed from closely related Burkholderia species, namely, B. tropica DSM 15359T, B. bannensis NBRC E25T and B. unamae DSM 17197T, by having higher concentrations of C16: 0 (25.1 %), C17: 0 cyclo (25.2 %) and C18: 1 ω7c (6.1 %) and lower concentration of the Sum In Feature 2 (C14:0 3-OH and/or iso-C16:1 I; 8.3 %) (Reis et al. 2004; Caballero-Mellado et al. 2004; Aizawa et al. 2011). Moreover, the results of API 20NE and API 50CH microtests showed that the physiological characters of type strain D414T can be distinguished from closely related type strains of Burkholderia species, such as B. tropica DSM 15359 T, B. bannensis NBRC E25T and B. unamae DSM 17197T (Table 1). Furthermore, bacteria with highly similar protein patterns possess high genome similarity (Vandamme et al. 1996). In this study, the SDS-PAGE results strongly suggest that strain D414T can be distinguished from its closely related type strains as shown in Supplementary Fig. 1.
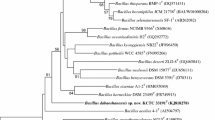
Based on 16S rRNA gene sequence analysis, the strain D414T separated from other closely-related species of the Burkholderia in the phylogenetic tree (Supplementary Fig. 2). The results revealed that the novel strain D414T was integrated into the group A (plant-associated and saprophytic Burkholderia lineage) described by Suárez-Moreno et al. (2012) and Estrada-de los Santos et al. (2013). The strain is closely related to B. tropica DSM 15359T, B. bannensis NBRC E25T and B. unamae DSM 17197T with similarities of 98.5, 98.3 and 98.3 %, respectively (Fig. 2).
Neighbour-joining phylogenetic tree based on almost complete 16S rRNA gene sequences showing relationships between Burkholderia metalliresistens sp. nov. strain D414T and the most closely related type strains of the genus Burkholderia. Numbers at branching nodes indicate bootstrap percentages (based on 1000 replications). Bar 0.01 substitutions per nucleotide position
The DNA–DNA relatedness values of strain D414T with the three aforementioned reference type strains were 22.7 to 33.9 %, 25.4 to 27.2 % and 32.9 to 33.7 %, respectively. These values are significantly lower than the 70 % cut-off point recommended for the delineation of genomic species (Wayne et al. 1987). The results of the DNA–DNA relatedness study confirm that strain D414T belongs to a novel Burkholderia species. The DNA G+C content is range from 61.8 to 62.8 %, which is within the G+C content range of group A (61.6 to 64.2 %) but different from the G+C content of group B (66.1 to 68.1 %), B. andropogonis (59.0 %) and B. rhizoxinica/B. endofungorum group (60.7 %) as described by Estrada-de los Santos et al. (2013).
Strain D414T showed high MIC values for Cd (2000 mg L−1), Pb (800 mg L−1), Cu (150 mg L−1) and Zn (2500 mg L−1). The order of toxicity of metals to strain D414 was Cu > Pb > Cd > Zn. Rhizosphere bacteria and endophytes, which were isolated from heavy metal-contaminated soils or plants, often exhibit considerably high heavy metal resistance to adapt to such environments (Pal et al. 2005). In the present study, the average content of soluble phosphate was estimated to be ranged from 154.3 to 161.0 mg L−1. Phosphorus is an essential macronutrient for plants; thus, phosphate solubilisation is an important trait of PGPR (Guo et al. 2011). Therefore, Burkholderia species could potentially be employed for the phytoremediation of multiple metal-contaminated soils.
Based on a combination of phenotypic and genotypic studies, strain D414T represents a novel species of the genus Burkholderia, for which the name Burkholderia metalliresistens sp. nov. is proposed.
Description of Burkholderia metalliresistens sp. nov
Burkholderia metalliresistens (metalliresistens: me.tal.li.re.sis’tens.L. n. metallum, metal; L. part. adj. resistens, resisting; N.L. part. adj. metalliresistens, metal resisting, referring to the ability of the organism to resist metal).
The cells of this species are 0.5–0.7 µm (width) × 0.8–2.0 µm (length) in size, Gram negative, aerobic and non-motile. The colonies are milk-like, yellowish-white, convex and circular with clear margins on modified SMN agar after 3 days of cultivation at 30 °C. Grows within the temperature range of 10–50 °C (optimum 30 °C) and pH range of 3.0–8.0 (optimum 7.0). Grows well on LB broth (Difco), yeast nitrogen base agar (Difco) and trypticase soy agar (Difco) but weakly on Ashby’s nitrogen-free medium. Positive for the production of indole; activity of urease, oxidase, catalase and nitro-d-methyl galactose; and assimilation of ACC, esculine, glucose, arabinose, mannose, mannitol, N-acetylglucosamine, malate, gluconate, capric acid, adipate, citrate, phenylacetate, d-galactose, d-ribose, d-xylose, d-adonitol, d-fructose, l-rhamnose, inositol, sorbitol, salicin, d-cellobiose, d-melibiose, d-gentiobiose, d-lyxose, fucose and d-arabitol. Negative for nitrate reduction, glucose fermentation, assimilation of arginine and gelatin hydrolysis, erythritol, l-xylose, methyl-β-d-pyranoid xyloside, l-sorbose, dulcitol, methyl-α-d-pyranoid mannoside, methyl-α-d-pyranoid glucoside, amygdalin, arbutin, d-lactose, d-saccharose, inulin, d-melezitose, d-raffinose, amylum, glycogen, xylitol, d-tagatose, l-arabitol, potassium gluconate, 2-ketogluconate and 5-ketogluconate. The tricalcium phosphate solubilisation ability is positive in Pikovskays’s medium. The major fatty acids are C14: 0, C16: 0, C17:0 cyclo and C18: 1 ω7c. The DNA G+C content of the type strain is 62.3 ± 0.5 mol%.
The type strain D414T (=CICC 10561T = DSM 26823T) was isolated from multiple metal-polluted paddy soil at the surrounding area of Dabao Mountain Mine, Guangdong, Southeast China.
Abbreviations
- MIC:
-
Minimum inhibitory concentration
References
Aizawa T, Vijarnsorn P, Nakajima M, Sunairi M (2011) Burkholderia bannensis sp. nov., an acid-neutralizing bacterium isolated from torpedo grass (Panicum repens) growing in highly acidic swamps. Int J Syst Evol Microbiol 61:1645–1650
Aleem A, Isar J, Malik A (2003) Impact of long term application of industrial wastewater on the emergence of resistance traits in Azotobacter chroococcum isolated from rhizosphere soil. Bioresour Technol 86:7–13
Angus AA, Lee A, Lum MR, Shehayeb M, Hessabi R, Fujishige NA, Yerrapragada S, Kano S, Song N, Yang P et al (2013) Nodulation and effective nitrogen fixation of Macroptilium atropurpureum (siratro) by Burkholderia tuberum a nodulating and plant growth promitng beta-proteobacterium, are influenced by environmental factors. Plant Soil 369:543–562
Blaha D, Pringet-Combaret C, Mirza MS, Moe¨nne-Loccoz Y (2006) Phylogeny of the 1-aminocyclopropane-1-carboxylic acid deaminase-encoding gene acdS in phytobeneficial and pathogenic proteobacteria and relation with strain biogeography. FEMS Microbiol Ecol 56:455–470
Caballero-Mellado J, Martinez-Aguilar L, Paredes-Valdez G, de Los Estrada, Santos P (2004) Burkholderia unamae sp. nov., an N2-fixing rhizospheric and endophytic species. Int J Syst Evol Microbiol 54:1165–1172
Caballero-Mellado J, Onofre-Lemus J, Estrada-de Los Santos P, Martinez-Aguilar L (2007) The tomato rhizosphere, an environment rich in nitrogen-fixing Burkholderia species with capabilities of interest for agriculture and bioremediation. Appl Environ Microbiol 73:5308–5319
Chen WM, James EK, Coenye T, Chou JH, Barrios E, de Faria SM, Elliott GN, Sheu SY, Sprent JI, Vandamme P (2006) Burkholderia mimosarum sp. nov., isolated from root nodules of Mimosa spp. from Taiwan and South America. Int J Syst Evol Microbiol 56:1847–1851
Chen WM, de Faria SM, James EK, Elliott GN, Lin KY, Chou JH, Sheu SY, Cnockaert M, Sprent JI, Vandamme P (2007) Burkholderia nodosa sp. nov., isolated from root nodules of the woody Brazilian legumes Mimosa bimucronata and Mimosa scabrella. Int J Syst Evol Microbiol 57:1055–1059
Chen WM, de Faria SM, Chou J-H, James EK, Elliott GN, Sprent JI, Bontemps C, Young JPW, Vandamme P (2008) Burkholderia sabiae sp. nov., isolated from root nodules of Mimosa caesalpiniifolia. Int J Syst Evol Microbiol 58:2174–2179
Chun J, Goodfellow M (1995) A phylogenetic analysis of the genus Nocardia with 16S rRNA gene sequences. Int J Syst Bacteriol 45:240–245
Coenye T, Vandamme P (2003) Diversity and significance of Burkholderia species occupying diverse ecological niches. Environ Microbiol 5:719–729
Coenye T, Gillis M, Vandamme P (2000) Pseudomonas antimicrobica Attafuah and Bradbury 1990 is a junior synonym of Burkholderia gladioli (Severini 1913) Yabuuchi et al. 1993. Int J Syst Evol Microbiol 50:2135–2139
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Estrada-de los Santos P, Martı´nez-Aguilar L, Vinuesa P, Hirsch AM, Caballero-Mellado J (2013) Phylogenetic analysis of Burkholderia species by Multilocus sequence analysis. Curr Microbiol 67:51–60
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Ghazi F, Henni DE, Benmechernene Z, Kihal M (2009) Phenotypic and whole cell protein analysis by SDS-PAGE for identification of dominants lactic acid bacteria isolated from algerian raw milk. World J Dairy Food Sci 4:78–87
Glick BR, Cheng Z, Czarny J, Duan J (2007) Promotion of plant growth by ACC deaminase-producing soil bacteria. Eur J Plant Pathol 119:329–339
Guo JK, Tang SR, Ju XH, Ding YZ, Liao SQ, Song NN (2011) Effects of inoculation of a plant growth promoting rhizobacterium Burkholderia sp. D54 on plant growth and metal uptake by a hyperaccumulator Sedum alfredii Hance grown on multiple metal contaminated soil. World J Microbiol Biotechnol 27:2835–2844
Hardy RW, Holsten RD, Jackson EK, Burns RC (1968) The acetylene-ethylene assay for N2 fixation: laboratory and field evaluation. Plant Physiol 43:1185–1207
Jiang CY, Sheng XF, Qian M, Wang QY (2008) Isolation and characterization of a heavy metal-resistant Burkholderia sp. from heavy metal-contaminated paddy field soil and its potential in promoting plant growth and heavy metal accumulation in metal-polluted soil. Chemosphere 72:157–164
Jones K (1970) Nitrogen fixation in the phyllosphere of the Douglas fir, Pseudotsuga douglasii. Ann Bot 34:239–244
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H et al (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Li WC, Ye ZH, Wong MH (2007) Effects of bacteria on enhanced metal uptake of the Cd/Zn-hyperaccumulating plant, Sedum alfredii. J Exp Bot 58:4173–4182
Lim JH, Baek SH, Lee ST (2008) Burkholderia sediminicola sp. nov., isolated from freshwater sediment. Int J Syst Evol Microbiol 58:565–569
Lu P, Zheng LQ, Sun JJ, Liu HM, Li SP, Hong Q, Li WJ (2012) Burkholderia zhejiangensis sp. nov., a methyl-parathion-degrading bacterium isolated from a wastewater-treatment system. Int J Syst Evol Microbiol 62:1337–1341
Martı´nez-Aguilar L, Salazar-Salazar C, Me´ndez RD, Caballero-Mellado J, Hirsch AM, Va´squez-Murrieta M, Estrada-de los SP (2013) Burkholderia caballeronis sp. nov., a nitrogen fixing species isolated from tomato (Lycopersicon esculentum) with the ability to effectively nodulate Phaseolus vulgaris. Antonie van Leeuwenhoek doi: 10.1007/s10482-013-0028-9
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G+C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Syst Bacteriol 39:159–167
Onofre-Lemus J, Hernandez-Lucas I, Girard L, CaballeroMellado J (2009) ACC (1-aminocyclopropane-1-carboxylate) deaminase activity, a widespread trait in Burkholderia species, and its growth-promoting effect on tomato plants. Appl Environ Microbiol 75:6581–6590
Otsuka Y, Muramatsu Y, Nakagawa Y, Matsuda M, Nakamura M, Murata H (2011) Burkholderia oxyphila sp. nov., a bacterium isolated from acidic forest soil that catabolizes (+)-catechin and its putative aromatic derivatives. Int J Syst Evol Microbiol 61:249–254
Pal A, Dutta S, Mukherjee PK, Paul AK (2005). Occurrence of heavy metal resistance in microflora from serpentine soil of Andaman. J Basic Microbiol 45:207–218
Palinska KA, Marquardt J (2008) Genotypic and phenotypic analysis of strains assigned to the widespread cyanobacterial morphospecies Phormidium autumnale (Oscillatoriales). Arch Microbiol 189:325–335
Reis VM, Estrada-de los Santos P, Tenorio-Salgado S, Vogel J, Stoffels M, Guyon S, Mavingui P, Baldani VLD, Schmid M et al (2004) Burkholderia tropica sp. nov., a novel nitrogen-fixing, plant-associated bacterium. Int J Syst Evol Microbiol 54:2155–2162
Rzhetsky A, Nei M (1993) Theoretical foundation of the minimum-evolution method of phylogenetic inference. Mol Biol Evol 10:1073–1095
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. USFCC Newsl 20:1–6
Suárez-Moreno ZR, Caballero-Mellado J, Coutinho BG, Mendonc¸a-Previato L, James EK, Venturi V (2012) Common features of environmetal and potentially beneficial plant associated Burkholderia. Microb Ecol 63:249–266
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Van de Peer Y, De Wachter R (1994) TREECON for Windows: a software package for the construction and drawing of evolutionary trees for the Microsoft Windows environment. Comput Appl Biosci 10:569–570
Vandamme P, Pot B, Gillis M, De Vos P, Kersters K, Swings J (1996) Polyphasic taxonomy, a consensus approach to bacterial systematics. Microbiol Rev 60:407–438
Vandamme P, Goris J, Chen WM, De Vos P, Willems A (2002) Burkholderia tuberum sp. nov. and Burkholderia phymatum sp. nov. nodulate the roots of tropical legumes. Syst Appl Microbiol 25:507–512
Watanabe FS, Olsen SR (1965) Test of an ascorbic acid method for determining phosphorous in water and NaHCO3 extracts from soil. Soil Sci Soc Am Proc 29:677–678
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE et al (1987) International committee on systematic bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Wei GH, Wang ET, Tan ZY, Zhu ME, Chen WX (2002) Rhizobium indigoferae sp. nov. and Sinorhizobium kummerowiae sp. nov., respectively isolated from Indigofera spp. and Kummerowia stipulacea. Int J Syst Evol Microbiol 52:2231–2239
Yabuuchi E, Kosako Y, Oyaizu H, Yano I, Hotta H, Hashimoto Y, Ezaki T, Arakawa M (1992) Proposal of Burkholderia gen. nov. and transfer of seven species of the genus Pseudomonas homology group II to the new genus, with the type species Burkholderia cepacia (Palleroni and Holmes 1981) comb. nov. Microbiol Immunol 36:1251–1275
Zaidi S, Usmani S, Singh BR, Musarrat J (2006) Significance of Bacillus subtilis strain SJ-101 as a bioinoculant for concurrent plant growth promotion and nickel accumulation in Brassica juncea. Chemosphere 64:991–997
Acknowledgments
We wish to thank J.P. Euzéby for the helpful advice on etymology of bacterial names. We also acknowledge JinXia Li at the CICC and Dr. Sabine Gronow at the DSMZ for deposit of the isolates. This research was financially supported by the Central Public Research Institutes Basic Funds for Research and Development (Agro-Environmental Protection Institute, Ministry of Agriculture) and National Natural Science Foundation of China (41473115).
Author information
Authors and Affiliations
Corresponding authors
Additional information
The GenBank/EMBL/DDBJ accession numbers for the 16S rRNA gene sequences of strain D414T is KF601211.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Guo, J.K., Ding, Y.Z., Feng, R.W. et al. Burkholderia metalliresistens sp. nov., a multiple metal-resistant and phosphate-solubilising species isolated from heavy metal-polluted soil in Southeast China. Antonie van Leeuwenhoek 107, 1591–1598 (2015). https://doi.org/10.1007/s10482-015-0453-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-015-0453-z