Abstract
A novel Gram-strain positive, aerobic, actinobacterial strain, designated CF11/1T, was isolated from a sand sample obtained in the Sahara Desert, Chad. The black-pigmented isolate was aerobic and exhibited optimal growth from 25 to 35 °C at pH 6.0–8.0 and with 0–8 % (w/v) NaCl, indicating that it is a halotolerant mesophile. Chemotaxonomic and molecular characteristics of the isolate matched those described for members of the genus Geodermatophilus. The G+C content in the genome was 74.4 mol%. The peptidoglycan contained meso-diaminopimelic acid as diagnostic diaminoacid. The main phospholipids were diphosphatidylglycerol, phosphatidylcholine, phosphatidylethanolamine, phosphatidylinositol and a minor fraction of phosphatidylglycerol; MK-9(H4) was the dominant menaquinone, and galactose was detected as a diagnostic sugar. The major cellular fatty acid was branched-chain saturated acid iso-C16:0. Analysis of 16S rRNA gene sequences showed 95.3–98.6 % pairwise sequence identity with the members of the genus Geodermatophilus. Based on phenotypic and chemotaxonomic properties, as well as phylogenetic distinctiveness, the isolate represents a novel species, Geodermatophilus africanus, with the type strain CF11/1T (DSM 45422 = CCUG 62969 = MTCC 11556).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The family Geodermatophilaceae was originally proposed by Normand et al. (1996), but a formal description of the family name was only published after a decade (Normand 2006). Today, the family comprises the genera Blastococcus, Modestobacter and Geodermatophilus (as type genus). Geodermatophilus was first proposed by Luedemann (1968) and accepted in the approved lists of bacterial names by Skerman et al. (1980). The members of this genus are frequently isolated from arid soils (Urzì et al. 2001), although some have also been isolated from rhizosphere soil (Zhang et al. 2011; Jin et al. 2013). Nevertheless, this genus was for a long time poorly studied and sampled due to challenges in culturing (Urzì et al. 2004). Eleven named species have been classified in the genus Geodermatophilus: Geodermatophilus obscurus (Luedemann 1968), Geodermatophilus ruber (Zhang et al. 2011), Geodermatophilus nigrescens (Nie et al. 2012), Geodermatophilus arenarius (Montero-Calasanz et al. 2012; Validation list no. 150 March 2013), Geodermatophilus siccatus (Montero-Calasanz et al. 2013a; Validation list no. 151 May 2013), Geodermatophilus saharensis (Montero-Calasanz et al. 2013b; Validation list no. 151 May 2013), Geodermatophilus tzadiensis (Montero-Calasanz et al. 2013c), Geodermatophilus telluris (Montero-Calasanz et al. 2013d), ‘Geodermatophilus soli’, ‘Geodermatophilus terrae’ (Jin et al. 2013) and ‘Geodermatophilus normandii’ (Montero-Calasanz et al. 2013e). The genomes of only one type strain, G. obscurus G-20T has been sequenced so far (Ivanova et al. 2010). Moreover, four named subspecies have been identified but with their names not yet validly published: ‘G. obscurus subsp. amargosae’, ‘G. obscurus subsp. utahensis’, ‘G. obscurus subsp. dictyosporus’ (Luedemann 1968) and ‘G. obscurus subsp. everesti’ (Ishiguro and Fletcher 1975; Normand and Benson 2012). This study describes the taxonomic position of a novel halotolerant species in the genus Geodermatophilus based on a polyphasic approach.
Materials and methods
Sample collection and culture conditions
During an environmental screening of arid surface soils in the Sahara Desert (Republic of Chad) in 2007, representative red sand samples (sand grain diameter 1–2 mm) were collected near Ourba (for details see Favet et al. 2013). Portions of sand were suspended in physiological saline, shaken for 1 h at 26 °C and kept overnight at 4 °C then shaken for an additional 2 h before being streaked out on R2A (DSMZ medium 830) and trypticase soy broth (TSB; DSMZ medium 535) plates and incubated at 25 °C for 3–10 days (for details see Giongo et al. 2013). Purified strain CF11/1T was stored in Microbank™ Blue Colour Beads (Pro-Lab Diagnostics, Richmond, Canada) before accession into the DSMZ open collection.
Phenotypic procedures
Cultural characteristics were tested on GYM Streptomyces medium (DSMZ medium 65), TSB agar, GPHF medium (DSMZ medium 553), R2A medium, GEO medium (DSMZ medium 714), PYGV medium (DSMZ medium 621) and Luedemann medium (DSMZ medium 877) for 15 days. To determine its morphological characteristics, strain CF11/1T was cultivated on GYM Streptomyces medium. Colony features were observed at 4 and 15 days under a binocular microscope according to Pelczar (1957). Exponentially growing bacterial cultures were observed with an optical microscope (Zeiss AxioScope A1) with a 100-fold magnification and phase-contrast illumination. Micrographs of bacterial cells grown on GYM Streptomyces broth after 7 days were taken with a field-emission scanning electron microscope (FE-SEM Merlin, Zeiss, Germany). Gram reaction was performed using the KOH test described by Gregersen (1978). Cell motility was observed on modified ISP2 (Shirling and Gottlieb 1966) swarming agar (0.3 %, w/v) at pH 7.2 that contained (l−1) 4.0 g dextrin, 4.0 g yeast extract and 10.0 g malt extract. Oxidase activity was analysed using filter paper disks (Sartorius grade 388) soaked in a 1 % solution of N,N,N′,N′-tetramethyl-p-phenylenediamine (Sigma-Aldrich); a positive test was defined by the development of a blue–purple colour after applying biomass on the filter paper. Catalase activity was determined based on formation of bubbles following the addition of drops of 3 % H2O2 (1 drop). Growth rates were determined on plates of GYM medium for temperatures from 10 to 50 °C at 5 °C increments and for pH values 4.0–12.5 (in increments of 0.5 pH units) on modified ISP2 medium by adding NaOH or HCl, since the use of a buffer system inhibited growth of the cultures. Degradation of specific substrates was examined using agar plates with various basal media: casein degradation was tested on plates containing milk powder (5 % w/v), NaCl (0.5 %) and agarose (1 %); tyrosine degradation was investigated as previously described (Gordon and Smith 1955) on plates containing peptone (0.5 %), beef extract (0.3 %), l-tyrosine (0.5 %) and agarose (1.5 %); xanthine and hypoxanthine decomposition was tested by the same test, replacing l-tyrosine by hypoxanthine or xanthine (0.4 %); starch degradation was tested on plates containing nutrient broth (0.8 %), starch (1 %) and agarose (1.5 %), then developed by flooding in 1 % iodine solution. For all tests, a positive result was defined by the appearance of clear zones around the colonies. The utilization of carbon compounds and production of acid were tested using API 20 NE strips (bioMérieux) and GEN III Microplates in an Omnilog device (BIOLOG Inc., Hayward, CA, USA). The GEN III Microplates were inoculated with cells suspended in the viscous inoculating fluid (IF C) provided by the manufacturer at a cell density of 75–79 % T for strain CF11/1T, at 90 % T for G. arenarius CF5/4T and at 80–83 % T for the rest of reference strains. As growth rates were relatively slow, each plate was measured in three subsequent runs by restarting the OmniLog device twice, yielding a total running time of 10 days in Phenotype Microarray mode at 28 °C. Data was exported and analysed using the opm package for R (Vaas et al. 2012). Each strain was studied in two independent experiments, yielding a total of six recorded runs per strain. Reactions with a distinct behaviour between the two experiments were regarded as ambiguous. Enzymatic activity was screened using API ZYM galleries according to manufacturer instructions (bioMérieux). All physiological tests were performed at 28 °C using G. obscurus DSM 43160T, G. ruber DSM 45317T, G. nigrescens DSM 45408T, G. arenarius DSM 45418T, G. siccatus DSM 45419T, G. saharensis DSM 45423T, G. tzadiensis DSM 45416T, G. telluris DSM 45421T, G. soli DSM 45843T, G. terrae DSM 45844T and G. normandii DSM 45417T in parallel assays.
Chemotaxonomic analysis
Whole-cell amino acids and sugars were prepared according to Lechevalier and Lechevalier (1970), followed by thin layer chromatography (TLC) analysis (Staneck and Roberts 1974). Polar lipids were extracted, separated by two-dimensional TLC and identified according to procedures outlined by Minnikin et al. (1984) with modifications proposed by Kroppenstedt and Goodfellow (2006). Additionally, choline-containing lipids were detected by spraying with Dragendorff reagent (Merck) (Tindall 1990). Menaquinones (MK) were extracted from freeze-dried cell material using methanol as described by Collins et al. (1977) and analysed by high-performance liquid chromatography (HPLC) (Kroppenstedt 1982). For extraction and analysis of cellular fatty acids, the physiological age of each strain was standardised by consistently choosing the last quadrant streaked on GYM agar plates incubated at 28 °C for 4 days. Analysis was conducted using the microbial identification system (MIDI) Sherlock Version 4.5 (method TSBA40, TSBA6 database) as described by Sasser (1990). The composition of peptidoglycan hydrolysates (6 N HCl, 100 °C for 16 h) was examined by TLC as described by Schleifer and Kandler (1972). All chemotaxonomic tests were conducted with the same reference strains under standardised conditions.
Genetic and phylogenetic analysis
G+C content of chromosomal DNA was determined by HPLC according to Mesbah et al. (1989). Genomic DNA extraction, PCR-mediated amplification of the 16S rRNA gene and purification of the PCR product was carried out as described by Rainey et al. (1996). Phylogenetic analysis was based on an alignment inferred by POA version 2.0 (Lee et al. 2002) and filtered with GBLOCKS (Castresana 2000). Phylogenetic trees were inferred with maximum-likelihood (ML) and maximum-parsimony as optimality criteria using RAxML version 7.2.8 (Stamatakis et al. 2008) and PAUP* 4b10 (Swofford 2002), respectively. Bootstrap support values were calculated using the bootstopping criterion (Pattengale et al. 2009) as implemented in RAxML and 1000 replicates in the case of PAUP*. Rooting was done using the midpoint method (Hess and De Moraes Russo 2007) and checked for agreement with the phylogenetic classification. Pairwise similarities were calculated from exact pairwise sequence alignments using the Smith–Waterman algorithm within the European Molecular Biology Open Software (EMBOSS) suite (Rice et al. 2000). DNA–DNA hybridization tests were performed by double reciprocal analysis as described by De Ley et al. (1970) with the modifications suggested by Huss et al. (1983) using a Cary 100 Bio UV/VIS (Biotech).
Results and discussion
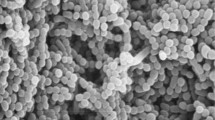
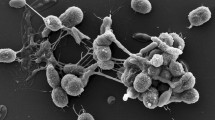
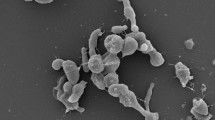
CF11/1T cells were Gram-type-positive, pleiotropic and with dried aspect. Individual cells, dimers and large aggregates were both observed, confirming reports by Ishiguro and Wolfe (1970) of synchronous morphogenesis on unspecific media. Motile zoospores were circular or elliptical; septated filaments from zoospore germination were observed (Fig. 1). Colonies were black-coloured, irregular, multilocular and opaque with a dry surface and an irregular margin. Similar appearances were observed in colonies of G. obscurus and G. telluris under the same growth conditions (Table 1). Moderate growth was observed on GYM Streptomyces medium and R2A medium, but not on TSB agar, GPHF, GEO, PYGV and Luedemann media. CF11/1T grew best at 25–35 °C; no growth was observed below 20 °C or above 37 °C. Growth was observed in the presence of 0–8 % NaCl, regarding as a halotolerant bacteria, and between pH 6.0–8.0. Results from phenotype microarray analysis are shown as a heatmap in the supplementary material (Fig. S1) in comparison to other type strains of the genus Geodermatophilus. A summary of select differential phenotypic characteristics is presented in Table 1. Analysis of cell wall components revealed the presence of dl-diaminopimelic acid (cell wall type III), which is consistent with other species of the genus Geodermatophilus (Lechevalier and Lechevalier 1970; Montero-Calasanz et al. 2013e). Strain CF11/1T displayed primarily MK-9(H4) (87.2 %), in agreement with values reported for the family Geodermatophilaceae (Normand 2006), but also MK-8(H4) (4.5 %), MK-9(H0) (3.0 %), MK-9(H2) (2.1 %) and an unknown MK (3.2 %). Similar patterns were already observed for G. arenarius (Montero-Calasanz et al. 2012) and G. tzadiensis (Montero-Calasanz et al. 2013e). Most major fatty acids were saturated branched-chain acids: iso-C16:0 (36.2 %), anteiso-C17:0 (8.3 %), anteiso-C15:0 (7.7 %) and iso-C15:0 (7.1 %), complemented by the monounsaturated iso-H-C16:1 (7.9 %) and C16:1ω7c (7.2 %). The phospholipid pattern consisted of diphosphatidylglycerol (DPG), phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidylinositol (PI) and a small amount of phosphatidylglycerol (PG) (see Supplementary Fig. S2) and is in accordance with profiles obtained for the other Geodermatophilus species investigated in this study (Table 1). Whole-cell sugar analysis revealed galactose as diagnostic sugar (Lechevalier and Lechevalier 1970), but also glucose and traces of ribose. Genomic G+C content was 74.4 mol%.
The almost complete (1511 bp) 16S rRNA gene sequence of strain CF11/1T was determined. The 16S rRNA gene sequence showed the highest similarity with the homologous genes of G. siccatus (98.6), G. tzadiensis (98.3), ‘G. normandii’ (98.2), G. arenarius (97.8), G. nigrescens (97.5), G. saharensis (97.4), G. obscurus (97.2), G. ruber (97.0) and G. telluris (97.0 %) and all listed closely related type strains were placed within the same phylogenetic group by both maximum-likelihood and maximum-parsimony estimations (Fig. 2). The 16S rRNA gene sequences analysis thus strongly supports that strain CF11/1T belongs to the genus Geodermatophilus. However, similarities in 16S rRNA gene sequence between CF11/1T and closely related type strains indicated the need to prove the genomic distinctness of the type strain representing the novel species by DNA–DNA hybridizations. CF11/1T displayed a percentage of DNA–DNA relatedness of 32.3 ± 0.7 with G. siccatus, 23.5 ± 3.4 with G. tzadiensis, 19.3 ± 5.1 with G. normandii, 20.9 ± 0.5 with G. nigrescens, and 28.2 ± 1.0 with G. obscurus. DNA–DNA hybridizations of CF11/1T with the type strains of G. arenarius, G. saharensis, G. ruber and G. telluris were not conducted, because our hands-on experience from over 30 DDHs between pairs of strains related by 97–99 % 16S rRNA gene sequence identity in the genus Geodermatophilus clearly confirmed the observation reported by Stackebrandt and Ebers (2006) and Meier-Kolthoff et al. (2013) that such strains generally result in DNA–DNA hybridization values below the 70 % threshold recommended by Wayne et al. (1987) to confirm the species status of a novel strain.
Maximum likelihood phylogenetic tree inferred from 16S rRNA gene sequences, showing the phylogenetic position of strain CF11/1T relative to the type strains within the family Geodermatophilaceae. The branches are scaled in terms of the expected number of substitutions per site (see size bar). Support values from maximum-likelihood (left) and maximum-parsimony (right) bootstrapping are shown above the branches if equal to or larger than 60 %
Apart from the phylogenetic analysis based on 16S rRNA gene sequences, several phenotypic characteristics support the distinctiveness of strain CF11/1T from all other named Geodermatophilus species (Table 1). Based on the phenotypic and genotypic data presented, we propose that strain CF11/1T represents a novel species within the genus Geodermatophilus, with the name Geodermatophilus africanus sp. nov.
Description of Geodermatophilus africanus sp. nov.
Geodermatophilus africanus (af.ri.ca’nus. L. masc. adj. africanus, of Africa)
Colonies are black-coloured, irregular, multilocular with a dry surface. Cells are Gram-strain positive, catalase positive and oxidase negative. No diffusible pigments are produced on any medium tested. Utilizes sodium lactate, d-serine, guanidine hydrochloride, d-galacturonic acid, d-glucuronic acid, glucuronamide, l-malic acid, bromo-succinic acid, potassium tellurite, β-hydroxy-butyric acid, acetoacetic acid and butyric acid as sole carbon source for energy and growth, but not d-maltose, β-gentiobiose, sucrose, d-turanose, stachyose, d-raffinose, α-d-lactose, d-melibiose, β-methyl-d-galactoside, d-salicin, N-acetyl-d-glucosamine, N-acetyl-β-d-mannosamine, N-acetyl-d-galactosamine, N-acetyl-neuraminic acid, d-mannose, d-fructose, d-galactose, 3-O-methyl-d-glucose, d-fucose, l-fucose, l-rhamnose, inosine, d-sorbitol, d-mannitol, d-arabitol, myo-inositol, glycerol, d-glucose-6-phosphate, d,l-aspartic acid, d-serine, gelatin, glycyl-l-proline, l-alanine, l-arginine, l-glutamic acid, l-histidine, l-pyroglutamic acid, l-serine, pectin, l-galactonic acid-ϒ-lactone, d-gluconic acid, mucic acid, quinic acid, d-saccharic acid, d-lactic acid methyl ester, l-lactic acid, citric acid, α-keto-glutaric acid, d-malic acid, α-keto-butyric acid, propionic acid, acetic acid and sodium formate, considering as ambiguous the utilization of dextrin, d-trehalose, d-cellobiose, d-glucose, d-fructose-6-phosphate, p-hydroxy-phenylacetic acid, methyl pyruvate, tween 40, ϒ-amino-N-butyric acid and α-hydroxy-butyric acid. Acid is produced from d-serine to guanidine hydrochloride and can be used as sole nitrogen sources, but not l-arginine, l-glutamic acid, l-serine, glycil-l-proline, l-alanine, l-histidine, l-pyroglutamic acid, inosine, N-acetyl-d-glucosamine, N-acetyl-β-mannosamine, N-acetyl-d-galactosamine and d,l-aspartic acid and ambiguous the utilization of ϒ-amino-N-butyric. Negative for the reduction of nitrate and denitrification, gelatine hydrolysis, indole production and degradation of casein, tyrosine, aesculin, starch, xanthine and hypoxanthine. Tests for alkaline phosphatase, esterase lipase (C8) and leucine arylamidase are positive. Tests are negative for acid phosphatase, Naphthol-AS-BI-phosphohydrolase, esterase (C4), lipase (C14), valine arylamidase, urease, cystine arylamidase, trypsin, α-chymotrypsin, α,ß-galactosidase, ß-glucuronidase, α,ß-glucosidase, N-acetyl-ß-glucosaminidase, α-mannosidase and α-fucosidase. NaCl tolerance ranges from 0 to 8 % (w/v). Cell growth ranges from 20 to 37 °C and pH 6.0–8.0. The peptidoglycan in the cell wall contains meso-diaminopimelic acid as diamino acid, with galactose as diagnostic sugar compounds. The predominant menaquinone is MK-9(H4). The main polar lipids are diphosphatidylglycerol, phosphatidylcholine, phosphatidylethanolamine, phosphatidylinositol with minor fraction of phosphatidylglycerol. Cellular fatty acids consist mainly of the branched-chain saturated acid iso-C16:0. The type strain has a genomic DNA G+C content of 74.4 mol%.
The type strain, CF11/1T = DSM 45422 = CCUG 62969 = MTCC 11556 was isolated in 2007 from sand of the Sahara Desert collected in Ourba (N15.23.905, E22.42.297), Republic of Chad (N15.23.905). The INSDC accession number for the 16S rRNA gene sequence of strain CF11/1T is HE654550.
References
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Favet J, Lapange A, Giongo A, Kennedy S, Aung Y–Y, Cattaneo A, Davis-Richarson AG, Brown CT, Kort R, Brumsack H-J, Schnetger B, Chappell A, Kroijenga J, Beck A, Schwibbert K, Mohamed AH, Kirchner T, Dorr de Quadros P, Tripplett EW, Broughton WJ, Gorbushina AA (2013) Microbial hitchhikers on intercontinental dust: catching a lift in Chad. ISME J 7:850–867
Giongo A, Favet J, Lapanje A, Gano KA et al (2013) Microbial hitchhikers on intercontinental dust: high-throughput sequencing to catalogue microbes in small sand samples. Aerobiologia 29:71–84
Gordon RE, Smith MM (1955) Proposed group of characters for the separation of Streptomyces and Nocardia. J Bacteriol 69:147–150
Gregersen T (1978) Rapid method for distinction of gram-negative from positive bacteria. Appl Microbiol Biotechnol 5:123–127
Hess PN, De Moraes Russo CA (2007) An empirical test of the midpoint rooting method. Biol J Linn Soc 92:669–674
Huss VAR, Fest H, Schleifer KH (1983) Studies on the spectrophotometric determination of DNA hybridization from renaturation rates. Syst Appl Microbiol 4:184–192
Ishiguro EE, Fletcher DW (1975) Characterization of Geodermatophilus strains isolated from high altitude Mount Everest soils. Mikrobiologika 12:99–108
Ishiguro EE, Wolfe RS (1970) Control of morphogenesis in Geodermatophilus: ultrastructural studies. J Bacteriol 104:566–580
Ivanova N, Sikorski J, Jando M, Munk C, Lapidus A, Glavina Del Rio T, Copeland A, Tice H, Cheng JF, Lucas S et al (2010) Complete genome sequence of Geodermatophilus obscurus type strain (G-20T). Stand Genomic Sci 2:158–167
Jin L, Lee HG, Kim HS, Ahn CY, Oh HM (2013) Geodermatophilus soli sp. nov. and Geodermatophilus terrae sp. nov., two novel actinobacteria isolated from grass soil. Int J Syst Bacteriol. doi:10.1099/ijs.0.048892-0
Kroppenstedt RM (1982) Separation of bacterial menaquinones by HPLC using reverse phase (RP18) and a silver loaded ion exchanger. J Liq Chromatogr 5:2359–2387
Kroppenstedt RM, Goodfellow M (2006) The family Thermomonosporaceae: Actinocorallia, Actinomadura, Spirillispora and Thermomonospora. In: Dworkin M, Falkow S, Schleifer KH, Stackebrandt E (eds) The prokaryotes. Archaea and Bacteria: Firmicutes, Actinomycetes, 3rd edn, vol 3, Springer, New York, pp. 682–724
Lechevalier MP, Lechevalier HA (1970) Chemical composition as a criterion in the classification of aerobic actinomycetes. Int J Syst Bacteriol 20:435–443
Lee C, Grasso C, Sharlow MF (2002) Multiple sequence alignment using partial order graphs. Bioinformatics 18:452–464
Luedemann GM (1968) Geodermatophilus, a new genus of the Dermatophilaceae (Actinomycetales). J Bacteriol 96:1848–1858
Meier-Kolthoff JP, Göker M, Spröer C, Klenk HP (2013) When should a DDH experiment be mandatory in microbial taxonomy? Arch Microbiol. doi:10.1007/s00203-013-0888-4
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G+C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Syst Bacteriol 39:159–167
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal K, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Montero-Calasanz MC, Göker M, Pötter G, Rohde M, Spröer C, Schumann P, Gorbushina AA, Klenk HP (2012) Geodermatophilus arenarius sp. nov., a xerophilic actinomycete isolated from Saharan desert sand in Chad. Extremophiles 16:903–909
Montero-Calasanz MC, Göker M, Rohde M, Schumann P, Pötter G, Spröer C, Gorbushina AA, Klenk HP (2013a) Geodermatophilus siccatus sp. nov., isolated from arid sand of the Saharan desert in Chad. Antonie Van Leeuwenhoek 103:449–456
Montero-Calasanz MC, Göker M, Pötter G, Rohde M, Spröer C, Schumann P, Gorbushina AA, Klenk HP (2013b) Geodermatophilus saharensis sp. nov., isolated from sand of the Saharan desert in Chad. Arch Microbiol 195:153–159
Montero-Calasanz MC, Göker M, Broughtonc WJ, Cattaneoc A, Favetc J, Pötter G, Rohde M, Spröer C, Schumann P, Klenk H-P, Gorbushina AA (2013c) Geodermatophilus tzadiensis sp. nov., a UV radiation-resistant bacterium isolated from sand of the Saharan desert. Syst Appl Microbiol 36:177–182
Montero-Calasanz MC, Göker M, Pötter G, Rohde M, Spröer C, Schumann P, Klenk HP, Gorbushina AA (2013d) Geodermatophilus telluris sp. nov., a novel actinomycete isolated from Saharan desert sand in Chad. Int J Syst Evol Microbiol 63:2254–2259
Montero-Calasanz MC, Göker M, Pötter G, Rohde M, Spröer C, Schumann P, Gorbushina AA, Klenk HP (2013e) Geodermatophilus normandii sp. nov., isolated from Saharan desert sand. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.051201-0
Nie GX, Ming H, Li S, Zhou EM, Cheng J, Yu TT, Zhang J, Feng HG, Tang SK, Li WJ (2012) Geodermatophilus nigrescens sp. nov., isolated from a dry-hot valley. Antonie Van Leeuwenhoek 101:811–817
Normand P (2006) Geodermatophilaceae fam. nov., a formal description. Int J Syst Evol Microbiol 56:2277–2278
Normand P, Benson DR (2012). Genus I. Geodermatophilus Luedemann 1968. 1994. In: Goodfellow M, Kämpfer P, Busse HJ, Trujillo ME, Suzuki KI, Ludwig W, Whitman WB (eds) Bergey’s Manual of Systematic Bacteriology, 2nd edn, vol 5, The Actinobacteria Part 1, Springer, New York, pp 528–530
Normand P, Orso S, Cournoyer B, Jeannin P, Chapelon C, Dawson J, Evtushenko L, Misra AK (1996) Molecular phylogeny of the genus Frankia and related genera and emendation of the family Frankinaceae. Int J Syst Bacteriol 46:1–9
Pattengale ND, Alipour M, Bininda-Emonds ORP, Moret BME, Stamatakis A (2009) How many bootstrap replicates are necessary? Lect Notes Comput Sci 5541:184–200
Pelczar MJ Jr (ed) (1957) Manual of microbiological methods. McGraw-Hill Book Co., New York
Rainey FA, Ward-Rainey N, Kroppenstedt RM, Stackebrandt E (1996) The genus Nocardiopsis represents a phylogenetically coherent taxon and a distinct actinomycete lineage: proposal of Nocardiopsiaceae fam. nov. Int J Syst Bacteriol 46:28–96
Rice P, Longden I, Bleasby A (2000) EMBOSS: the European molecular biology open software suite. Trends Genet 16:276–277
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. USFCC Newsl 20:16
Schleifer KH, Kandler O (1972) Peptidoglycan types of bacterial cell walls and their taxonomic implications. Bacteriol Rev 36:407–477
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Skerman VBD, McGowan V, Sneath PHA (1980) Approved lists of bacterial names. Int J Syst Bacteriol 30:225–420
Stackebrandt E, Ebers J (2006) Taxonomic parameter revisited: tarnished gold standards. Microbiol Today 33:152–155
Stamatakis A, Hoover P, Rougemont J (2008) A rapid bootstrap algorithm for the RAxML web servers. Syst Biol 57:758–771
Staneck JL, Roberts GD (1974) Simplified approach to identification of aerobic actinomycetes by thin-layer chromatography. Appl Microbiol 28:226–231
Swofford DL (2002) PAUP*: Phylogenetic Analysis Using Parsimony (*and Other Methods), Version 4.0 b10. Sinauer Associates, Sunderland
Tindall BJ (1990) A comparative study of the lipid composition of Halobacterium saccharovorum from various sources. Syst Appl Microbiol 13:128–130
Urzì C, Brusetti L, Salamone P, Sorlini C, Stackebrandt E, Daffonchio D (2001) Biodiversity of Geodermatophilaceae isolated from stones and monuments in the Mediterranean basin. Environ Microbiol 3:471–479
Urzì C, La Cono V, Stackebrandt E (2004) Design and application of two oligonucleotide probes for the identification of Geodermatophilaceae strains using fluorescence in situ hybridization (FISH). Environ Microbiol 6:678–685
Vaas LAI, Sikorski J, Michael V, Göker M, Klenk HP (2012) Visualization and curve-parameter estimation strategies for efficient exploration of phenotype microarray kinetics. PLoS ONE 7(4):e34846
Validation List no. 150 List of new names and new combinations previously effectively, but not validly published. Int J Syst Bacteriol March 2013
Validation List no. 151 List of new names and new combinations previously effectively, but not validly published. Int J Syst Bacteriol May 2013
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Trüper HG (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Zhang Y-Q, Chen J, Liu H-Y, Zhang Y-Q, Li W-J, Yu L-Y (2011) Geodermatophilus ruber sp. nov., isolated from rhizosphere soil of a medical plant. Int J Syst Evol Microbiol 61:190–193
Acknowledgments
We would like to gratefully acknowledge the help of Agathe Stricker of the International Committee of the Red Cross in Geneva, Switzerland, for organizing the collection of the sand samples from which strain CF11/1T was isolated. Jocelyne Favet, Arlette Cattaneo and William J. Broughton of the Laboratoire de Biologie Moléculaire de Plantes Supérieures (University of Geneva, Switzerland) are acknowledged for their contribution to the isolation of the strain, Bettina Sträubler and Birgit Grün (DSMZ, Braunschweig), for their help in DNA–DNA hybridization analysis and Brian J. Tindall (DSMZ, Braunschweig) for his guidance in chemotaxonomic analyses. M.C. Montero-Calasanz is the recipient of a postdoctoral contract from European Social Fund Operational Programme (2007–2013) for Andalusia and a DSMZ postdoctoral fellowship.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
The parameter “Maximum Height” estimated from the respiration curves as measured by an OmniLog phenotyping device and discretized and visualized as a heatmap using the opm package. Plates and substrates are rearranged according to their overall similarity (as depicted using the row and column dendrograms). Orange colour indicates positive reaction; purple colour indicate negative reaction; white colour indicate ambiguous reaction. Letters (A/B) indicate each replicate of experiment (PDF 61 kb)
Fig. S2
Polar lipids profile of Geodermatophilus africanus sp. nov. CF11/1T, after separation by two-dimensional TLC. Plate was sprayed with molydatophosphoric acid for detection of total polar lipid. DPG, diphosphadidylglycerol; PG, phosphadidylglycerol; PE, phosphatidethanolamine; PC, phosphatidylcholine; PI, phosphatidylinositol; GL, unknown glycolipid; L, unknown lipid (TIFF 277 kb)
Rights and permissions
About this article
Cite this article
Montero-Calasanz, M.C., Göker, M., Pötter, G. et al. Geodermatophilus africanus sp. nov., a halotolerant actinomycete isolated from Saharan desert sand. Antonie van Leeuwenhoek 104, 207–216 (2013). https://doi.org/10.1007/s10482-013-9939-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-013-9939-8