Abstract
The taxonomic position of an aerobic actinobacterial strain, BMG841T, isolated from the Bulla Regia monument (Tunisia) and exhibiting a high resistance to gamma-radiation (D10 ~9 kGy) was determined using polyphasic approach. The optimal growth range was found to be 25–35 °C at pH of 7.0–8.5. The strain was observed to form black dry colonies. Chemotaxonomic characteristics of the isolate showed a cell wall type III, with galactose and glucose as diagnostic sugars; phosphatidylcholine, phosphatidylinositol, diphosphatidylglycerol, phosphatidylethanolamine and an unidentified glycolipid as main polar lipids; and MK-9(H4) as the predominant menaquinone. The major cellular fatty acids were identified as iso-C16:0 and iso-C15:0. Phylogenetic analysis indicated that strain BMG841T represents a novel member of the genus Geodermatophilus with high 16S rRNA gene sequence identity with Geodermatophilus saharensis (98.28 %). Based on phylogenetic and phenotypic analysis, strain BMG841T is proposed as the type strain (=DSM 46841T = CECT 8821T) of a novel species, Geodermatophilus bullaregiensis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Geodermatophilus was proposed by Luedemann (1968) to accommodate aerobic, Gram-positive actinomycetes with dl-2,6-diaminopimelic acid (dl-DAP) in the cell wall peptidoglycan and MK-9(H4) as the predominant menaquinone. Currently the genus comprises eighteen species, which have been isolated from Desert sands (Montero-Calasanz et al. 2012), soil (Luedemann 1968; Nie et al. 2012; Jin et al. 2013; Montero-Calasanz et al. 2014a; Bertazzo et al. 2014), rhizosphere (Zhang et al. 2011), sediment (Qu et al. 2013) and altered stones (Montero-Calasanz et al. 2014b; Hezbri et al. 2015). Members of the genus Geodermatophilus are well known as gamma-radiation resistant actinobacteria with LD10s around 8–9 kGy (Gtari et al. 2012; Montero-Calasanz et al. 2014b; Hezbri et al. 2015). In this paper, we describe the polyphasic characterisation of the type strain of a gamma radiation-resistant new species, Geodermatophilus bullaregiensis, isolated from the marble monument of Bulla Regia located in North Western Tunisia.
Materials and methods
Isolation and culture of strain
Dust sampled from crevices of a marble rock surface located in the ruin of Bulla Regia, a Roman City situated in North Western Tunisia, was suspended in physiological saline solution and shaken overnight at 28 °C before being streaked out on Luedemann medium (Luedemann 1968) and on R2A (DSMZ medium 830) supplemented with cycloheximide at 0.01 %. Strain BMG841T was isolated and maintained on Luedemann medium after 10 days incubation. Colonies and general cultural characteristics were observed from cultures growing at 28 °C for 7 days on different media: GYM Streptomyces medium (DSMZ medium 65), R2A medium (DSMZ medium 830) and Luedemann medium (DSMZ medium 877).
Phenotypic tests
Morphological characteristics were observed by using light microscopy (Zeiss AxioScope A1) and a field-emission scanning electron microscope (FE-SEM Merlin, Zeiss, Germany) after 7 days growth at 28 °C on GYM Streptomyces medium. Gram reaction was carried out by the standard Gram staining procedure (Gram 1884). Oxidase activity was analysed using filter-paper disks (Sartorius grade 388) impregnated with 1 % solution of N,N,N′,N′-tetramethyl-p-phenylenediamine (Sigma-Aldrich); a positive test was defined by the development of a blue-purple colour after applying biomass to the filter paper. Catalase activity was determined based on formation of bubbles following the addition of 1 drop of 3 % H2O2. Enzymatic activities were tested using API ZYM galleries according to the instructions of the manufacturer (bioMérieux). Oxidation of carbon and nitrogen compounds was tested using GEN III Microplates in an Omnilog device (BIOLOG Inc., Hayward, CA, USA) in comparison with the reference strains in parallel assays: Geodermatophilus africanus DSM 45422T (Montero-Calasanz et al. 2013d), G. amargosae DSM 46136T (Montero-Calasanz et al. 2014a), G. arenarius DSM 45418T (Montero-Calasanz et al. 2012), G. dictyosporus DSM 43161T (Montero-Calasanz et al. 2015), G. nigrescens DSM 45408T (Nie et al. 2012), G. normandii DSM 45417T (Montero-Calasanz et al. 2013b), G. saharensis DSM 45423T (Montero-Calasanz et al. 2013c), G. telluris DSM 45421T (Montero-Calasanz et al. 2013e) and G. tzadiensis DSM 45416T (Montero-Calasanz et al. 2013a). The GEN III microplates were inoculated with cells suspended in a viscous inoculating fluid (IF C) provided by the manufacturer at a cell density of at 83–84 % Transmittance (T) for strain BMG841T, at 70 % T for G. amargosae DSM 46136T, at 75–79 % T for G. africanus DSM 45422T, at 90 % T for G. arenarius DSM 45418T and at 80–83 % T for G. dictyosporus DSM 43161T and the remaining reference strains. The strains were studied in two independent technical replicates. Data were exported and analysed using the opm package for R (Vaas et al. 2012; Vaas et al. 2013) v.1.0.6.
The temperature range and optimum for growth were tested at 5–45 °C on plates of GYM Streptomyces medium and at 50–60 °C on modified Streptomyces medium by adding MgCl2 and substituting agar with GELRITE (Sigma-Aldrich) (Shungu et al. 1983). The pH range was investigated between pH 4.0 and 12.5 at intervals of 0.5 pH units. Degradation of specific substrates was examined using agar plates with various basal media: casein degradation was tested on plates containing milk powder (5 % w/v), NaCl (0.5 %) and agarose (1 %); tyrosine degradation was determined as previously described by Gordon and Smith (1955) on plates containing peptone (0.5 %), beef extract (0.3 %), l-tyrosine (0.5 %) and agarose (1.5 %); xanthine and hypoxanthine decomposition (0.4 %) were examined using the same basal medium; starch degradation was tested on plates containing nutrient broth (0.8 %), starch (1 %) and agarose (1.5 %), then developed by flooding in 1 % iodine solution. For all tests, a positive result was defined by the appearance of clear zones around the colonies.
Chemotaxonomic analyses
Biomass for chemotaxonomic and genotypic studies was obtained by cultivation in shaken flasks (~150 rpm) containing GYM Streptomyces broth at 28 °C for 7 days. The diagnostic isomer of diaminopimelic acid in whole-cell hydrolysates (6 N HCl, 100 °C for 16 h) of strain BMG841T was identified by TLC on cellulose plates using the solvent system of Schleifer and Kandler (1972). Sugar analysis of whole-cell hydrolysates (1 N H2SO4, 95 °C for 2 h) was carried out as described by Staneck and Roberts (1974). About 100 mg of freeze-dried cells were used for the extraction of polar lipids followed by two-dimensional TLC separation and identification according to Minnikin et al. (1984) with modifications of Kroppenstedt and Goodfellow (2006). Choline-containing lipids were detected by spraying with Dragendorff’s reagent (Merck) (Tindall 1990). Menaquinones (MK) were extracted from about 300 mg of freeze-dried cell material using the method of Collins et al. (1977) and analysed by high-performance liquid chromatography (HPLC) (Groth et al. 1997). Cellular fatty acids were prepared by harvesting 40 mg of bacterial cells from the third quadrant of the streaked plate followed by saponification and methylation of the cells. The extraction and analysis of cellular fatty acids was conducted using the Microbial Identification System (MIDI) Sherlock Version 6.1 (method TSBA40, ACTIN6 database) as described by Sasser (1990). All chemotaxonomic analyses were conducted under standardized conditions with strain BMG841T and cultures of the same set of reference strains as listed in Table 1.
Phylogenetic analyses
Genomic DNA extraction, PCR-mediated amplification of the 16S rRNA gene and purification of the PCR product was carried out as described by Rainey et al. (1996). The identification of strain BMG841T based on the 16S rRNA gene sequence was performed using Ez-Taxon (Kim et al. 2012) and RDP-II (Maidak et al. 2001) servers. Pairwise similarities were calculated as recommended by Meier-Kolthoff et al. (2013). Sequence analyses and phylogenetic reconstruction were performed using MEGA 6.0 (Tamura et al. 2013). The stability of relationships was assessed by 1000 replicates through bootstrap analysis (Felsenstein 2005).
Radiation resistance assay
Ionizing-radiation resistance analysis was performed on strains BMG841T and Geodermatophilus obscurus DSM 43160T, used as a control, following the protocol outlined by Gtari et al. (2012). Non-sporulating cultures were obtained by growth in Luedemann medium containing tryptose (Difco, Detroit, USA) for 5 days at 28 °C (Ishiguro and Wolfe 1970). Collected cells were then washed twice with 0.9 % NaCl, homogenized and subsequently re-suspended in saline solution. One milliliter of cell suspension of each strain was introduced into separate sterile 1.5-mL Eppendorf tubes and subsequently exposed to 1–10 kGy, at a dose rate of 63,319 Gy min−1, in a 60Co irradiator. Subsequently, the cell suspensions were cooled directly on ice, tenfold dilution series were prepared and plated in triplicate on solid Luedemann medium, and incubated at 28 °C. After a 2-week period, CFUs were counted and the survival fractions were calculated based on a non-irradiated control by using the R software packages mgcv (Wood 2014) and lethal (Hofner 2014) as described by Montero-Calasanz et al. (2014b).
Results and discussion
Strain BMG841T was observed to exhibit good growth on Luedemann and GYM Streptomyces media and moderate growth on R2A medium. No diffusible pigments were observed. Multilocular colonies were observed with a maximum diameter of 2.3 mm and intense black pigmentation with very dry surfaces and irregular margins. Cells of strain BMG841T were observed to be pleiomorphic and Gram stain-positive. Individual and cells aggregated in cauliflower-like clumps, together with groups of cuboid cells (Fig. 1), were observed as described by Ishiguro and Wolfe (1970). Strain BMG841T was found to grow in the presence of up to 4 % NaCl but not with 8 % NaCl. The temperature range for growth was found to range from 10 to 40 °C (with 25 to 35 °C as optimum) and the pH range from 6.5 to 10.5 (with 7.0–8.5 as optimum). Results from phenotype microarray analysis are shown as a heatmap in the supplementary material (Fig. S1) in comparison to the reference type strains of the genus Geodermatophilus. A summary of selected differential phenotypic characteristics is presented in Table 1.
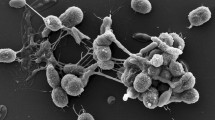
Scanning electron micrograph of strain BMG841T grown for 7 days at 28 °C on GYM Streptomyces medium and showing masses of cuboid and elliptical cells together with short septated filaments. Hezbri et al. (2015)
Chemotaxonomic data of strain BMG841T are in agreement with those previously described for the members of the genus Geodermatophilus (Hezbri et al. 2015). The cell wall peptidoglycan was found to be type III, containing DL-DAP as the diagnostic diamino acid (Lechevalier and Lechevalier 1970; Montero-Calasanz et al. 2015). The whole-cell sugar analysis revealed the presence of glucose and galactose as diagnostic sugars (Lechevalier and Lechevalier 1970), along with traces of ribose and mannose. The predominant menaquinone was identified as MK-9(H4) (71.5 %) as reported by Normand (2006) for all the members of the family Geodermatophilaceae. However, MK-9(H0) and MK-9(H2) were also present as minor components (17.2 and 8.6 %, respectively). These results are in accordance with those found for the strain Geodermatophilus soli DSM 45843T (Jin et al. 2013). The major fatty acids were identified as iso-C16:0 (38.6 %) and iso-C15:0 (20.4 %), consistent with the profiles of strains G. dictyosporus DSM 43161T (Montero-Calasanz et al. 2015), G. nigrescens DSM 45408T (Nie et al. 2012), G. arenarius DSM 45418T (Montero-Calasanz et al. 2012), G. tzadiensis DSM 45416T (Montero-Calasanz et al. 2013a), G. telluris DSM 45421T (Montero-Calasanz et al. 2013e), G. normandii DSM 45417T (Montero-Calasanz et al. 2013b) and G. amargosae DSM 46136T (Montero-Calasanz et al. 2014a) (Table 1). The polar lipid profile was found to contained phosphatidylcholine, phosphatidylinositol, diphosphatidylglycerol, phosphatidylethanolamine and an unidentified glycolipid (Supplementary Fig. S2), similar to the pattern obtained by Montero-Calasanz et al. (2015) for G. dictyosporus DSM 43161T.
Analysis of the 16S rRNA gene sequence of strain BMG841T (1458 bp; GenBank/EMBL/DDBJ accession number LN626271) supports the affiliation of strain BMG841T to the genus Geodermatophilus (Fig. 2), with the strain being closely related to the type strain of G. saharensis (98.3 % sequence similarity). Meier-Kolthoff et al. (2013) reported that an Actinobacterial-specific 16S rRNA threshold of 99.0 % with a maximum probability of error of 1.0 % can be assumed to correspond to DNA–DNA hybridization values below the recommended 70 % threshold recommended to assign a given strain to a new prokaryotic species (Wayne et al. 1987). Therefore, DNA–DNA hybridizations with the two closely related neighbours, G. saharensis and G. dictyosporus appear to be dispensable based on the 16S rRNA sequence similarity, and strain BMG841T can be proposed as the type strain of a novel species of the genus Geodermatophilus.
Phylogenetic position of strain BMG841T within the family Geodermatophilaceae based on 16S rRNA gene sequences. Only bootstrap values higher than 50 % are shown above the branches. Hezbri et al. (2015)
With an LD10 around 9 KGy (Fig. 3), strain BMG841T showed only a minor difference in inactivation kinetics compared to the reference strain G. obscurus DSM 43160T used as a positive control (Gtari et al. 2012). Compared to the model radio-resistant bacteria Deinococcus radiodurans (LD10 ~10 kGy) and the actinobacterial species Rubrobacter xylanophilus (LD10 ~5.5 kGy), Kineococcus radiotolerans (LD10 ~2 kGy) (Sghaier et al. 2008), Geodermatophilus poikilotrophi (Montero-Calasanz et al. 2014b), G. dictyosporus (Montero-Calasanz et al. 2015), G. aquaeductis (Hezbri et al. 2015) and G. obscurus DSM 43160T (Gtari et al. 2012), strain BMG841T can be considered as a new highly radio-resistant representative of the actinobacteria.
Survival of strains BMG841T and G. obscurus DSM 43160T following gamma-radiation exposure and estimation from mean c.f.u. ml−1. LD50 and LD10 values are indicated in the upper panel of each figure. Y-axis is on a logarithmic scale. Hezbri et al. (2015)
On the basis of the genotypic and phenotypic data, strain BMG841T can be clearly distinguished from its phylogenetically close relatives. Therefore, strain BMG841T represents a novel species of the genus Geodermatophilus, for which we propose the name Geodermatophilus bullaregiensis sp. nov.
Description of Geodermatophilus bullaregiensis sp. nov.
Geodermatophilus bullaregiensis (bul.la.re.gi.en’sis. L. n. Bulla Regia, a Roman town in Northern Africa, today North-Western Tunisia; N.L. masc. adj. bullaregiensis, derived from Bulla Regia referring to the origin of isolation).
Colonies are black-coloured, irregular, multilocular with a dry surface. Cells are Gram-stain positive, catalase positive and oxidase negative. No diffusible pigments are produced on any of the tested media. Temperature range for growth ranges from 10 to 40 °C and the pH range from 6.5 to 10.5. Can oxidize several carbon and nitrogen sources (Table 1), degrades aesculin but is negative for nitrate reduction and denitrification, indole production and degradation of casein, tyrosine, starch, xanthine, gelatine and hypoxanthine. Tests for alkaline phosphatase and leucine arylamidase are positive but those for esterase (C4), esterase lipase (C8), valine arylamidase, ß-galactosidase and α- and ß-glucosidase acid phosphatase, Naphthol-AS-BI-phosphohydrolase, lipase (C14), cystine arylamidase, trypsin, α-chymotrypsin, α-galactosidase, ß-glucuronidase, N-acetyl-ß-glucosamidase, α-mannosidase, urease, and α-fucosidase are negative. The peptidoglycan in the cell wall contains meso-diaminopimelic acid as the diamino acid, with glucose and galactose as diagnostic sugars. The predominant menaquinone is MK-9(H4). The main polar lipids are phosphatidylcholine, phosphatidylinositol, diphosphatidylglycerol, phosphatidylethanolamine and an unidentified glycolipid. Cellular fatty acids consist mainly of the branched-chain saturated acids iso-C16:0 and iso-C15:0.
The type strain is BMG841T (=DSM 46841T = CECT 8821T). The INSDC accession number of the 16S rRNA gene sequence of the type strain is LN626271.
References
Bertazzo M, Montero-Calasanz MC, Martinez-Garcia M et al (2014) Geodermatophilus brasiliensis sp. nov., isolated from Brazilian soil. Int J Syst Evol Microbiol 64:2841–2848
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230
Felsenstein J (2005) Using the quantitative genetic threshold model for inferences between and within species. Philos Trans R Soc Lond B Biol Sci 360:1427–1434
Gordon RE, Smith MM (1955) Proposed group of characters for the separation of Streptomyces and Nocardia. J Bacteriol 69:147–150
Gram H (1884) Über die isolierte Färbung der Schizomyceten in Schnitt- und Trockenpräparaten. Fortschr Med 2:185–189
Groth I, Schumann P, Rainey FA et al (1997) Demetria terragena gen. nov., sp. nov., a new genus of actinomycetes isolated from compost soil. Int J Syst Bacteriol 47:1129–1133
Gtari M, Essoussi I, Maaoui R et al (2012) Contrasted resistance of stone-dwelling Geodermatophilaceae species to stresses known to give rise to reactive oxygen species. FEMS Microbiol Ecol 80:566–577
Hezbri K, Ghodhbane-Gtari F, Montero-Calasanz MC et al (2015) Geodermatophilus aquaeductus sp. nov., isolated from the ruins of Hadrian’s aqueduct. Antonie Van Leeuwenhoek 107(1):291–296
Hofner B (2014) Lethal: compute lethal doses (LD) with confidence intervals. R package version 0.5. http://r-forge.r-project.org/projects/lethal/
Ishiguro EE, Wolfe RS (1970) Control of Morphogenesis in Geodermatophilus: ultrastructural Studies. J Bacteriol 104:566–580
Jin L, Lee H-G, Kim H-S et al (2013) Geodermatophilus soli sp. nov. and Geodermatophilus terrae sp. nov., two actinobacteria isolated from grass soil. Int J Syst Evol Microbiol 63:2625–2629
Kim O-S, Cho Y-J, Lee K et al (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kroppenstedt R, Goodfellow M (2006) The family Thermomonosporaceae: Actinocorallia, Actinomadura, Spirillispora and Thermomonospora. In: Falkow S, Schleifer KHS, Dworkin m (eds) The prokaryotes, 3rd edn. Springer, New York, pp 682–724
Lechevalier MP, Lechevalier H (1970) Chemical composition as a criterion in the classification of aerobic actinomycetes. Int J Syst Bacteriol 20:435–443
Luedemann GM (1968) Geodermatophilus, a new genus of the Dermatophilaceae (Actinomycetales). J Bacteriol 96:1848–1858
Maidak BL, Cole JR, Lilburn TG et al (2001) The RDP-II (ribosomal database project). Nucleic Acids Res 29:173–174
Meier-Kolthoff JP, Göker M, Spröer C, Klenk H-P (2013) When should a DDH experiment be mandatory in microbial taxonomy? Arch Microbiol 195:413–418
Minnikin DE, O’Donnell AG, Goodfellow M et al (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Montero-Calasanz MC, Göker M, Pötter G et al (2012) Geodermatophilus arenarius sp. nov., a xerophilic actinomycete isolated from Saharan desert sand in Chad. Extremophiles 16:903–909
Montero-Calasanz MC, Göker M, Broughton WJ et al (2013a) Geodermatophilus tzadiensis sp. nov., a UV radiation-resistant bacterium isolated from sand of the Saharan desert. Syst Appl Microbiol 36:177–182
Montero-Calasanz MC, Göker M, Pötter G et al (2013b) Geodermatophilus normandii sp. nov., isolated from Saharan desert sand. Int J Syst Evol Microbiol 63:3437–3443
Montero-Calasanz MC, Göker M, Pötter G et al (2013c) Geodermatophilus saharensis sp. nov., isolated from sand of the Saharan desert in Chad. Arch Microbiol 195:153–159
Montero-Calasanz MC, Göker M, Pötter G et al (2013d) Geodermatophilus africanus sp. nov., a halotolerant actinomycete isolated from Saharan desert sand. Antonie Van Leeuwenhoek 104:207–216
Montero-Calasanz MC, Göker M, Pötter G et al (2013e) Geodermatophilus telluris sp. nov., an actinomycete isolated from Saharan desert sand. Int J Syst Evol Microbiol 63:2254–2259
Montero-Calasanz MC, Göker M, Rohde M et al (2014a) Description of Geodermatophilus amargosae sp. nov., to accommodate the not validly named Geodermatophilus obscurus subsp. amargosae (Luedemann, 1968). Curr Microbiol 68:365–371
Montero-Calasanz MC, Hofner B, Göker M et al (2014b) Geodermatophilus poikilotrophi sp. nov.: a multitolerant actinomycete isolated from dolomitic marble. Biomed Res Int 20:914767. doi:10.1155/2014/914767
Montero-Calasanz MC, Hezbri K, Göker M et al (2015) Description of gamma radiation-resistant Geodermatophilus dictyosporus sp. nov. to accommodate the not validly named Geodermatophilus obscurus subsp. dictyosporus (Luedemann, 1968). Extremophiles 19:77–85
Nie G-X, Ming H, Li S et al (2012) Geodermatophilus nigrescens sp. nov., isolated from a dry-hot valley. Antonie Van Leeuwenhoek 101:811–817
Normand P (2006) Geodermatophilaceae fam. nov., a formal description. Int J Syst Evol Microbiol 56:2277–2278
Qu J-H, Hui M, Qu J-Y et al (2013) Geodermatophilus taihuensis sp. nov., isolated from the interfacial sediment of a eutrophic lake. Int J Syst Evol Microbiol 63:4108–4112
Rainey FA, Ward-Rainey N, Kroppenstedt RM, Stackebrandt E (1996) The genus Nocardiopsis represents a phylogenetically coherent taxon and a distinct actinomycete lineage: proposal of Nocardiopsaceae fam. nov. Int J Syst Bacteriol 46:1088–1092
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. USFCC Newsl 20:16
Schleifer KH, Kandler O (1972) Peptidoglycan types of bacterial cell walls and their taxonomic implications. Bacteriol Rev 36:407–477
Sghaier H, Ghedira K, Benkahla A, Barkallah I (2008) Basal DNA repair machinery is subject to positive selection in ionizing-radiation-resistant bacteria. BMC Genom 9:297
Shungu D, Valiant M, Tutlane V et al (1983) GELRITE as an agar substitute in bacteriological media. Appl Environ Microbiol 46:840–845
Staneck JL, Roberts GD (1974) Simplified approach to identification of aerobic actinomycetes by thin-layer chromatography. Appl Microbiol 28:226–231
Tamura K, Stecher G, Peterson D et al (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tindall BJ (1990) A comparative study of the lipid composition of Halobacterium saccharovorum from various sources. Syst Appl Microbiol 13:128–130
Vaas LAI, Sikorski J, Michael V et al (2012) Visualization and curve-parameter estimation strategies for efficient exploration of phenotype microarray kinetics. PLoS One 7:e34846
Vaas LAI, Sikorski J, Hofner B et al (2013) Opm: an R package for analysing OmniLog(R) phenotype microarray data. Bioinformatics 29:1823–1824
Wayne LG, Brenner DJ, Colwell RR et al (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Wood S (2014) mgcv: mixed GAM computation vehicle with GCV/AIC/REML smoothness estimation, R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/
Zhang Y-Q, Chen J, Liu H-Y et al (2011) Geodermatophilus ruber sp. nov., isolated from rhizosphere soil of a medicinal plant. Int J Syst Evol Microbiol 61:190–193
Acknowledgments
This research was supported by The Ministère de l’Enseignement Supérieur et de la Recherche Scientifique, Tunisia (LR03ES03). Chemotaxonomic work was done at the DSMZ (German Collection of Microorganisms and Cell Cultures) Braunschweig. The authors would like to thank Professor Bernhard Schink for his advice in naming the new species.
Conflict of interest
Authors disclose that there are no conflicts of interest. No research involving human participants and/or animals was performed. No financial interests tied directly or indirectly to this research exist that may be important to readers need to be disclosed.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 1 (PPT 205 kb)
Supplementary Fig. S1. The parameter “Maximum Height” based on respiration curves generated with the OmniLog phenotyping illustrated as a heatmap using the opm package. Plates and substrates are rearranged according to their overall similarity (as depicted using the row and column dendrograms). Ochre colour indicates positive reaction; purple colour indicates negative reaction; white colour indicates ambiguous reaction. Letters (A/B) correspond to the two experimental replicates
Supplementary material 2 (PPT 314 kb)
Supplementary Fig. S2. Two-dimensional TLC polar lipids profiling of strain BMG841T after revelation with molydatophosphoric acid spraying. DPG, diphosphadidylglycerol; PE, phosphatidethanolamine; PC, phosphatidylcholine; PI, phosphatidylinositol; GL, unidentified glycolipid
Rights and permissions
About this article
Cite this article
Hezbri, K., Ghodhbane-Gtari, F., del Carmen Montero-Calasanz, M. et al. Description of Geodermatophilus bullaregiensis sp. nov.. Antonie van Leeuwenhoek 108, 415–425 (2015). https://doi.org/10.1007/s10482-015-0494-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-015-0494-3