Abstract
The Lactobacillus plantarum group comprises five very closely related species. Some species of this group are considered to be probiotic and widely applied in the food industry. In this study, we compared the use of two different molecular markers, the 16S rRNA and dnaK gene, for discriminating phylogenetic relationships amongst L. plantarum strains using sequencing and DNA fingerprinting. The average sequence similarity for the dnaK gene (89.2%) among five type strains was significantly less than that for the 16S rRNA (99.4%). This result demonstrates that the dnaK gene sequence provided higher resolution than the 16S rRNA and suggests that the dnaK could be used as an additional phylogenetic marker for L. plantarum. Species-specific profiles of the Lactobacillus strains were obtained with RAPD and RFLP methods. Our data indicate that phylogenetic relationships between these strains are easily resolved using sequencing of the dnaK gene or DNA fingerprinting assays.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The current taxonomy of the L. plantarum group is comprised of five closely related species: L. plantarum subsp. plantarum, L. plantarum subsp. argentoratensis, L. paraplantarum , L. pentosus, and L. fabifermentans (De Bruyne et al. 2009). The L. plantarum and L. pentosus species have been shown to be probiotic and are widely used in the food and feed industries (Casey et al. 2007; Holzapfel et al. 2001; Merry et al. 1995; Oneca et al. 2003; Ruiz-Barba et al. 1994). Moreover, numerous studies have demonstrated that the L. plantarum subsp. plantarum strain provides beneficial effects to the immune system (Bujalance et al. 2007; Pathmakanthan et al. 2004), especially in treating inflammatory disease and mitigating pathogenic infection (Connelly 2008). Inconceivably, more than 28% of commercial probiotic products are mislabelled at the genus or species level (Huys et al. 2006). Thus, it is very important to accurately distinguish and identify the correct species designations for this probiotic bacterial species complex.
Traditionally, the methods used for L. plantarum group classification were dependent on phenotypic tests such as morphological, physiological and biochemical analyses (Abazinge et al. 1993; Petrovic et al. 2006). Currently, the identification of Lactobacillus strains relies mainly on API 50 CH carbohydrate fermentation strips (Boyd et al. 2005; Bringel et al. 2005; De Bruyne et al. 2009). However, this commercial identification kit may not always be adequate in reliably distinguishing closely related Lactobacillus species, especially in the L. casei and L. plantarum groups (Johansson et al. 1995). Therefore, there has been a shift towards molecular based techniques to improve species identification within the L. plantarum group. The 16S ribosome RNA (16S rRNA), an ~1,500 bp sequence that codes for a portion of the 30S ribosome, and comparative analysis of the sequences is a commonly used molecular method for bacterial identification (Petti 2007). Strains that generally show higher than 97% similarity of the 16S rRNA sequence are considered to be the same species (McCartney 2002; Vandamme et al. 1996). However, it may not always suitable to distinguish closely related species such as those of the L. plantarum group because the 16S rRNA sequence similarity of L. plantarum group strains reaches 98.9–99.9% (De Bruyne et al. 2009). Thus other phylogenetic markers with higher resolution should be identified.
The dnaK gene encodes the 70 kDa heat shock protein (HSP70), which plays a crucial role as a chaperone machine for protein folding and transport and in protecting organisms from heat- or stress-induced damage (Netzer and Hartl 1998). In fact, HSP70 is the most conserved protein found in all biota (Gupta and Golding 1993); thus, the dnaK sequence is well suited for examining deep-level phylogenies. In the present study, we analysed partial dnaK gene sequences and compared them to the 16S rRNA gene to evaluate the utility of dnaK as an alternative phylogenetic marker for L. plantarum group strains. In addition, the RAPD (random amplification of polymorphic DNA) and RFLP (restriction fragment length polymorphism) fingerprinting techniques were also evaluated for their ability to discriminate the L. plantarum group.
Materials and methods
Lactobacillus strains
The Lactobacillus type strains and isolates used in this study are listed in Table 1. They were obtained from the Bioresource Collection and Research Center (BCRC). All bacterial strains were incubated on Lactobacilli MRS Agar (LMRS agar, Difco) aerobically for 48 h at 37°C.
Genomic DNA preparation
The genomic DNA was extracted using the DNeasy kit (Qiagen, Valencia, CA, USA) following the manufacturers’ instructions. The DNA concentration and purity was measured with a spectrophotometer and checked by agarose gel electrophoresis. Then, the genomic DNA was used as a template for PCR.
RAPD–PCR analysis
The RAPD profiles for the test strains were determined as described previously (Huang and Lee 2009). Forty random primers (Operon kits A and T series) were used for RAPD-PCR, and the amplification was carried out in a thermal cycler (GeneAmp® 2700, Applied Biosystems, CA, USA). The RAPD profiles were separated according to size by electrophoresis on 2% agarose gels.
PCR amplification of target gene
The partial dnaK fragments were amplified by PCR using primers Lpdnak-500F3 (5′-CCGTTCTTRTCRATRTCRAA-3′) and Lpdnak-1710R5 (5′-GAAAYYCAAGTYGGHGAAGT-3′), the 16S rRNA were amplified using the MicroSeq Full Gene 16S rRNA Bacterial Identification kit (Applied Biosystems, Foster City, CA, USA). PCR reactions were composed of 81 μL sterile MilliQ water, 10 μL 10× PCR buffer, 1.5 μL dNTPs (10 mM), 2.5 μL forward primer (10 mM), 2.5 μL reverse primer (10 mM), 2.5 U Taq DNA polymerase (DreamTaq, Fermentas) and 3 μL template DNA (100 ng/μL). The thermal protocol using the following conditions: initial strand denaturation at 94°C for 5 min, followed by 35 cycles of 94°C for 1 min, 58°C for 1 min and 72°C for 1.5 min, and a final extension step at 72°C for 7 min. The PCR products were visualized by electrophoresis on a 2% agarose gel with ethidium bromide staining.
DNA sequencing and RFLP analysis
The PCR products were purified using the QIA quick PCR purification Kit (Qiagen Inc., Valencia, CA, USA) and sequenced with the BigDye Terminator v3.1 cycle-sequencing kit on the 3730 DNA sequencer (Applied Biosystems and Hitachi, Foster City, CA, USA). The sequencing primers of dnaK gene were Lpdnak-500F3 and Lpdnak-1710R5, Lplntdnak-340r1 (5′-ATTCCAGCYGTTCAAGAAGC-3′) and Lpdnak-1340F9 (5′-AAMGTMCCACCACCAAGGTC-3′) designed from the conserved regions of the five type strains of the L. plantarum group. The sequencing primers of 16S rRNA gene were the MicroSeq Full Gene 16S rRNA Bacterial Identification kit. Twenty μl of the amplified target DNA of dnaK were digested with enzymes HaeIII, AluI, MspI and Tsp509I. The restriction profiles were separated according to size by electrophoresis on 2.5% agarose gels.
Phylogenetic data analysis
Sequence similarities were calculated using the programs of Wisconsin Package Version 10.1 (Accelrys Inc., San Diego, CA). The dnaK and 16S rRNA sequences were aligned using the Clustal X program, version 1.8 (Thompson et al. 1997). Phylogenetic trees were constructed with the PHYLIP computer program package (Felsenstein 1993) using the neighbor-joining method (Saitou and Nei 1987) with genetic distances computed by using Kimura’s 2-parameter model (Kimura 1980). The bootstrap values were based on 1,000 replications.
Results
RAPD fingerprinting profiles
Two series of random primers including OPA and OPT (Operon Alameda, CA, USA) were used during the RAPD fingerprinting analysis of strains from within the L. plantarum group. Each series consisted of 20 primers of random sequence. Among these 40 random primers, OPT-1 (5′-GGGCCACTC-3′) was the most noteworthy for distinguishing all Lactobacillus species within the L. plantarum group, producing unique DNA patterns for each species within the group (Fig. 1).
RAPD fingerprints of Lactobacillus plantarum group strains. Genomic DNA samples were amplified with random primers (OPT-1). M: 100 bp DNA ladder markers. Lanes: (1) L. plantarum subsp. plantarum BCRC 10069T; (2) L. plantarum subsp. plantarum BCRC 12251; (3) L. plantarum subsp. plantarum BCRC 14059; (4) L. plantarum subsp. argentoratensis BCRC 17638T; (5) L. plantarum subsp. argentoratensis BCRC 17640; (6) L. plantarum subsp. argentoratensis BCRC 12327; (7) L. paraplantarum BCRC 17178T; (8) L. paraplantarum BCRC 17970; (9) L. paraplantarum BCRC 17971; (10) L. pentosus BCRC 11053T; (11) L. pentosus BCRC 17972; (12) L. pentosus BCRC 17973; (13) L. fabifermentans BCRC 18841T
Target gene amplification and DNA sequencing
Approximately 1,500 and 1,100 bp of the 16S rRNA and dnaK genes, respectively, were amplified from all Lactobacillus strains. The amplified DNA fragments were used for sequencing and restriction analysis. To obtain the 1,100 bp dnaK fragment, sequencing primers Lplntdnak-340r1 and Lpdnak-1340F9 were designed to anneal to the conserved regions. The PCR product sequencing was repeated at least twice to confirm the reading and to resolve any ambiguity. The dnaK gene sequences of all Lactobacillus strains in this study were submitted to GenBank, and the accession numbers are listed in Table 1.
PCR–RFLP profiles
The dnaK amplicons were digested with the enzymes HaeIII, MspI, AluI and Tsp509I, respectively. Among the 22 Lactobacillus strains, digestions using Tsp509I produced five different restriction profiles and distinguished all L. plantarum species simultaneously. Conversely, digestions using the other three enzymes showed that some species produced the same restriction profiles (data not shown).
Phylogenetic tree based on 16S rRNA and dnaK gene sequences
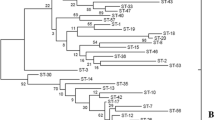
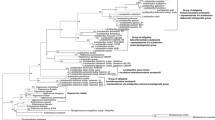
Similarities of the dnaK sequence ranged from 86.2 to 96.2%, compared to 98.9 to 99.9% for the 16S rRNA (Table 2). The nucleotide sequences of the dnaK and 16S rRNA genes from 22 Lactobacillus strains were determined, and phylogenetic trees were reconstructed using the neighbor-joining method. The bootstrap values at all nodes of the dnaK tree were significantly higher than those of 16S rRNA tree. Further, the topology of the dnaK tree showed five clearly separated groups (Fig. 3). In comparison, the 16S rRNA-based tree for all strains only showed two groups (Fig. 4) because the L. plantarum subsp. plantarum, L. plantarum subsp. argentoratensis, L. paraplantarum, and L. pentosus strains were indistinguishable from each other.
Discussion
Genomic DNA fingerprinting using the RAPD method has become widely accepted as a valid taxonomic and phylogenetic tool for a large range of organisms such as lactobacilli (Daud Khaled et al. 1997; Du Plessis and Dicks 1995; Huang and Lee 2009). Thus far, the RAPD technique was applied only to L. plantarum subsp. plantarum, L. paraplantarum, and L. pentosus; the other two Lactobacillus species were not checked (Torriani et al. 2001a). In this study, we used RAPD fingerprinting to classify full L. plantarum group strains, and the OPT-1 primer was used to simultaneously discriminate five members within the L. plantarum group, each of which formed their own specific-species DNA pattern (Fig. 1). PCR–RFLP technique is a simple, fast and low-cost way to identify bacteria (Olive and Bean 1999). The crucial element of this technique is the selection of the restriction enzymes. In this study, we also used a dnaK gene fragment in combination with PCR–RFLP to simultaneously discriminate between members of the L. plantarum group. Four restriction enzymes were tested, and the Tsp509I PCR–RFLP analysis was found to preliminary give species-specific discrimination (Fig. 2). The RFLP profiles of single species differed by two to three fragments, due to genetic drift. Thus, the other more reliable molecular technique should be identified.
Tsp509I RFLP profiles of the dnaK gene PCR products. M: 100 bp DNA ladder markers. (1) L. plantarum subsp. plantarum BCRC 10069T; (2) L. plantarum subsp. plantarum BCRC 12251; (3) L. plantarum subsp. plantarum BCRC 14059; (4) L. plantarum subsp. argentoratensis BCRC 17638T; (5) L. plantarum subsp. argentoratensis BCRC 17640; (6) L. plantarum subsp. argentoratensis BCRC 12327; (7) L. paraplantarum BCRC 17178T; (8) L. paraplantarum BCRC 17970; (9) L. paraplantarum BCRC 17971; (10) L. pentosus BCRC 11053T; (11) L. pentosus BCRC 17972; (12) L. pentosus BCRC 17973; (13) L. fabifermentans BCRC 18841T
Whole genome DNA–DNA hybridization is a gold standard assay and has long been applied to bacterial species delineation (Stackebrandt et al. 2002; Wayne et al. 1987). However, this technique is time consuming, expensive, not always repeatable and difficult to apply to a large number of bacteria (Mehlen et al. 2004; Pontes et al. 2007). In contrast, DNA sequences of protein-encoding genes seem to be more effective than the 16S rRNA gene and may replace DNA–DNA hybridization for species identification (Stackebrandt et al. 2002; Zeigler 2003). In this study, we used the 16S rRNA and partial dnaK gene sequences to classify L. plantarum group strains. A reliable phylogenetic tree based on dnaK clearly showed five species groups. The bootstrap values at all nodes of the dnaK tree were significantly higher than those of the 16S rRNA tree. In addition, ten nodes were observed where bootstrap values reached ≥80% (Fig. 3), whereas only one such node existed on the 16S rRNA tree (Fig. 4). The average nucleotide sequence similarity of dnaK between the L. plantarum group type strains was significantly less than that of 16S rRNA (89.2 and 99.4%, respectively). At present, several phylogenetic targets of protein-encoding genes have been exploited for the differentiation of L. plantarum group species (atpA, tuf, recA, hsp60, pheS and rpoA) (Blaiotta et al. 2008; Chavagnat et al. 2002; Naser et al. 2007; Torriani et al. 2001b). However, only two targets (recA and pheS) showed good resolution at a high discrimination level. In the present study, we found that the phylogenetic information in the dnaK gene was compatible with that from other protein-encoding genes in distinguishing phenotypically closely related species L. plantarum subsp. plantarum, L. plantarum subsp. argentoratensis, L. paraplantarum, L. pentosus and L. fabifermentans.
Phylogenetic tree of 22 Lactobacillus strains based on dnaK sequences. The tree was constructed with the neighbour-joining method. Genetic distances were computed by Kimura’s two-parameter model. L. pseudoficulneum was included as an outgroup. Only bootstrap percentages above 70% are shown (based on 1,000 replications). The scale bar represents 0.02% sequence divergence
Phylogenetic tree of 22 Lactobacillus strains based on 16S rRNA sequences. The tree was constructed with the neighbour-joining method. Genetic distances were computed by Kimura’s two-parameter model. L. pseudoficulneum was included as an outgroup. Only bootstrap percentages above 70% are shown (based on 1,000 replications). The scale bar represents 0.01% sequence divergence
Our data confirm that the sequence of the dnaK gene is significantly more polymorphic than that of the 16S rRNA gene. Thus, we propose that dnaK should complement the 16S rRNA for classification of the L. plantarum group. Furthermore, the DNA fingerprinting profiles obtained with the RAPD and RFLP methods might also be useful for species discrimination between L. plantarum group strains. In conclusion, the dnaK target gene appears to be an ideal phylogenetic marker for accurate and rapid discrimination and classification of L. plantarum group species, as well as subspecies, using direct sequencing and DNA fingerprinting techniques.
References
Abazinge MDA, Fontenot JP, Allen G, Flick GJ (1993) Ensiling characteristics of crab waste and wheat straw treated with different additives. J Agric Food Chem 41:657–661
Blaiotta G, Fusco V, Ercolini D, Aponte M, Pepe O, Villani F (2008) Lactobacillus strain diversity based on partial hsp60 gene sequences and design of PCR-restriction fragment length polymorphism assays for species identification and differentiation. Appl Environ Microbiol 74:208–215
Boyd MA, Antonio MAD, Hillier SL (2005) Comparison of API 50 CH strips to whole-chromosomal DNA probes for identification of Lactobacillus species. J Clin Microbiol 43:5309–5311
Bringel F, Castioni A, Olukoya DK, Felis GE, Torriani S, Dellaglio F (2005) Lactobacillus plantarum subsp. argentoratensis subsp. nov., isolated from vegetable matrices. Int J Syst Evol Microbiol 55:1629–1634
Bujalance C, Moreno E, Jimenez-Valera M, Ruiz-Bravo A (2007) A probiotic strain of Lactobacillus plantarum stimulates lymphocyte responses in immunologically intact and immunocompromised mice. Int J Food Microbiol 113:28–34
Casey PG, Gardiner GE, Casey G, Bradshaw B, Lawlor PG, Lynch PB, Leonard FC, Stanton C, Ross PP, Fitzgerald GF, Hill C (2007) A five-strain probiotic combination reduces pathogen shedding and alleviates disease signs in pigs challenged with Salmonella enterica serovar Typhimurium. Appl Environ Microbiol 73:1858–1863
Chavagnat F, Haueter M, Jimeno J, Casey MG (2002) Comparison of partial tuf gene sequences for the identification of lactobacilli. FEMS Microbiol Lett 217:177–183
Connelly P (2008) Lactobacillus plantarum—a literature review of therapeutic benefits. J Aust Tradit Med Soc 14:79–82
Daud Khaled AK, Neilan BA, Henriksson A, Conway PL (1997) Identification and phylogenetic analysis of Lactobacillus using multiplex RAPD-PCR. FEMS Microbiol Lett 153:191–197
De Bruyne K, Camu N, De Vuyst L, Vandamme P (2009) Lactobacillus fabifermentans sp. nov. and Lactobacillus cacaonum sp. nov., isolated from Ghanaian cocoa fermentations. Int J Syst Evol Microbiol 59:7–12
Du Plessis EM, Dicks LMT (1995) Evaluation of random amplified polymorphic DNA (RAPD)-PCR as a method to differentiate Lactobacillus acidophilus, Lactobacillus crispatus, Lactobacillus amylovorus, Lactobacillus gallinarum, Lactobacillus gasseri, and Lactobacillus johnsonii. Curr Microbiol 31:114–118
Felsenstein J (1993) PHYLIP (phylogeny inference package), version 3.5c. Department of Genetics, University of Washington, Seattle
Gupta RS, Golding GB (1993) Evolution of HSP70 gene and its implications regarding relationships between archaebacteria, eubacteria, and eukaryotes. J Mol Evol 37:573–582
Holzapfel WH, Haberer P, Geisen R, Bjorkroth J, Schillinger U (2001) Taxonomy and important features of probiotic microorganisms in food and nutrition. Am J Clin Nutr 73:365S–373S
Huang CC, Lee FL (2009) Development of novel species-specific primers for species identification of the Lactobacillus casei group based on the RAPD fingerprints. J Sci Food Agric 89:1831–1837
Huys G, Vancanneyt M, D’Haene K, Vankerckhoven V, Goossens H, Swings J (2006) Accuracy of species identity of commercial bacterial cultures intended for probiotic or nutritional use. Res Microbiol 157:803–810
Johansson ML, Quednau M, Ahrne S, Molin G (1995) Classification of lactobacillus plantarum by restriction endonuclease analysis of total chromosomal DNA using conventional agarose gel electrophoresis. Int J Syst Bacteriol 45:670–675
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
McCartney AL (2002) Application of molecular biological methods for studying probiotics and the gut flora. Br J Nutr 88:S29–S37
Mehlen AM, Goeldner S, Ried S, Stindl S, Ludwig W, Schleifer KH (2004) Development of a fast DNA–DNA hybridization method based on melting profiles in microplates. Syst Appl Microbiol 27:689–695
Merry RJ, Dhanoa MS, Theodorou MK (1995) Use of freshly cultured lactic acid bacteria as silage inoculants. Grass For Sci 50:112–123
Naser SM, Dawyndt P, Hoste B, Gevers D, Vandemeulebroecke K, Cleenwerck I, Vancanneyt M, Swings J (2007) Identification of lactobacilli by pheS and rpoA gene sequence analyses. Int J Syst Evol Microbiol 57:2777–2789
Netzer WJ, Hartl FU (1998) Protein folding in the cytosol: chaperonin-dependent and -independent mechanisms. Trends Biochem Sci 23:68–73
Olive DM, Bean P (1999) Principles and applications of methods for DNA-based typing of microbial organisms. J Clin Microbiol 37:1661–1669
Oneca M, Irigoyen A, Ortigosa M, Torre P (2003) PCR and RAPD identification of L. plantarum strains isolated from ovine milk and cheese. Geographical distribution of strains. FEMS Microbiol Lett 227:271–277
Pathmakanthan S, Li CK, Cowie J, Hawkey CJ (2004) Lactobacillus plantarum 299: beneficial in vitro immunomodulation in cells extracted from inflamed human colon. J Gastroenterol Hepatol 19:166–173
Petrovic T, Niksic M, Bringel F (2006) Strain typing with ISLpl1 in lactobacilli. FEMS Microbiol Lett 255:1–10
Petti CA (2007) Detection and identification of microorganisms by gene amplification and sequencing. Clin Infect Dis 44:1108–1114
Pontes SP, Lima-Bittencourt CI, Chartone-Souza E, Amaral Nascimento AM (2007) Molecular approaches: advantages and artifacts in assessing bacterial diversity. J Ind Microbiol Biotechnol 34:463–473
Ruiz-Barba JL, Cathcart DP, Warner PJ, Jiménez-Díaz R (1994) Use of Lactobacillus plantarum LPCO10, a bacteriocin producer, as a starter culture in spanish-style green olive fermentations. Appl Environ Microbiol 60:2059–2064
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Stackebrandt E, Frederiksen W, Garrity GM, Grimont PA, Kampfer P, Maiden MC, Nesme X, Rossello-Mora R, Swings J, Truper HG, Vauterin L, Ward AC, Whitman WB (2002) Report of the ad hoc committee for the re-evaluation of the species definition in bacteriology. Int J Syst Evol Microbiol 52:1043–1047
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Torriani S, Clementi F, Vancanneyt M, Hoste B, Dellaglio F, Kersters K (2001a) Differentiation of Lactobacillus plantarum, L. pentosus and L. paraplantarum species by RAPD-PCR and AFLP. Syst Appl Microbiol 24:554–560
Torriani S, Felis GE, Dellaglio F (2001b) Differentiation of Lactobacillus plantarum, L. pentosus, and L. paraplantarum by recA gene sequence analysis and multiplex PCR assay with recA gene-derived primers. Appl Environ Microbiol 67:3450–3454
Vandamme P, Pot B, Gillis M, Devos P, Kersters K, Swings J (1996) Polyphasic taxonomy, a consensus approach to bacterial systematics. Microbiol Rev 60:407–438
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE, Stackebrandt E, Starr MP, Truper HG (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Zeigler DR (2003) Gene sequences useful for predicting relatedness of whole genomes in bacteria. Int J Syst Evol Microbiol 53:1893–1900
Acknowledgments
We thank S. K. Chen, C. C. Liao and G. F. Yuan (Food Industry Research and Development Institute, Taiwan, ROC) for their encouragement. This research was supported by Ministry of Economic Affairs, ROC (project no. 98-EC-17-A-01-04-0525).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huang, CH., Lee, FL. & Liou, JS. Rapid discrimination and classification of the Lactobacillus plantarum group based on a partial dnaK sequence and DNA fingerprinting techniques. Antonie van Leeuwenhoek 97, 289–296 (2010). https://doi.org/10.1007/s10482-009-9409-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-009-9409-5