Abstract
During investigations of mycobiota in the coastal Arctic polythermal glaciers, different species of the ubiquitous genus Penicillium were isolated from the extreme subglacial environment. A group of Penicillium strains was obtained that did not belong to any known Penicillium species. This species was isolated in high numbers from the Kongsvegen subglacial ice and was not detected in the surrounding environment. A detailed analysis of secondary metabolite profiles, physiological and morphological characteristics, and partial β-tubulin gene sequences showed that the proposed new species Penicillium svalbardense is closely related but not identical to Penicillium piscarium and Penicillium simplicissimum. It differs in the production of secondary metabolites and in the morphological features of conidia and penicilli, and it is therefore described as a new species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The ubiquitous genus Penicillium contains mainly food, soil and airborne species (Pitt et al. 2000). This genus shows tolerance to cold environments, as demonstrated by the many species that grow on refrigerated food (Pitt and Hocking 1999) and that have been isolated from high mountain soils (Domsch et al. 1980; Petrovič et al. 2000). Penicillium spp. also inhabit extremely cold, polar areas of the world. They have been isolated from Arctic and Antarctic soils, permafrost, snow, sea ice and sea water (Vishniac 1993; McRae et al. 1999; Gunde-Cimerman et al. 2003; Frisvad 2004; Ivanushkina et al. 2005; Frisvad et al. 2006), and from glacial ice cores up to 38,600 years old (Abyzov 1993; Ma et al. 1999, 2000). Recent studies have revealed a new habitat for microorganisms within the Arctic polythermal glaciers. These are represented by bacteria (Foght et al. 2004) and fungi, primarily yeasts and the genus Penicillium (Sonjak et al. 2006). Among the Penicillium spp. detected, a group of biverticillate Penicillium strains was obtained that did not belong to any known Penicillium species. These strains were isolated exclusively from this extreme environment and were not detected in the neighbouring coastal areas (Sonjak et al. 2006). Here, we present the description of this new biverticillate Penicillium species, named Penicillium svalbardense, using morphological, physiological and molecular criteria.
Materials and methods
Site and sampling
In June and August 2001, fieldwork was undertaken in an area of Kongsfjorden, one of the largest fjords in west Spitsbergen, Svalbard (79°N, 12°E), Norway. Sediment-rich subglacial ice and the overlying clear glacial ice of the Kongsvegen polythermal glacier were sampled at the glacier margins where it was exposed. Samples were collected in sterile polypropylene bags using surface sterilized tools and transported to the laboratory, where they were processed as previously described (Sonjak et al. 2006). Aseptically melted inner layers of the glacial ice mass were filtered in aliquots of 100 ml. The membrane filters used (0.45 μm; Millipore) were placed on solid agar media. Additionally, mineral ice inclusions were collected aseptically, and 1 g of this sediment was spread directly over agar plates. Up to ten replicates of enumeration and different selective media were used (Sonjak et al. 2006). All of the plates were incubated for up to 14 weeks at 10°C and 24°C. After incubation, the colony forming units (CFU) were counted, with the mean CFUs calculated and expressed as CFU l−1 for the melt-water and CFU g−1 for the direct spreading of the sediments.
Strains examined
Strains of the proposed new Penicillium species are maintained in the EX-F culture collection of the Department of Biology, Biotechnical Faculty, University of Ljubljana, Slovenia and in the IBT culture collection of The Centre for Microbial Biotechnology, BioCentrum-DTU, Technical University of Denmark. The strains included in the analysis are listed in Table 1. Additional strains of related species, Penicillium simplicissimum, Penicillium piscarium and Penicillium cremeogriseum, were obtained from the IBT collection and the CBS culture collection of the Centraalbureau voor Schimmelcultures of the Royal Netherlands Academy of Arts and Sciences, Utrecht, The Netherlands (Table 1).
Morphology, physiology and extracellular enzymatic activity
For the determination of the morphological characteristics, the isolates were inoculated as three-point cultures on Czapek yeast autolysate agar (CYA), CYA with 5% NaCl (CYAS), malt extract agar (MEA), yeast extract sucrose agar (YES) and creatine sucrose agar (CREA), and grown for seven days at 25°C (CYA also at 15°C) in the dark (Frisvad and Samson 2004). For the determination of micro-morphological characteristics, microscope slides were prepared from MEA medium. Water solution of 60% (v/v) lactic acid without a colour dye was used as the mounting medium. The slides were examined under oil immersion with a BX51 microscope (Olympus, Japan) by differential interference contrast (DIC), at up to 1000× magnification. Digital micrographs were taken with DP12 digital camera and analysed using the DPSOFT 3.2 application software (Olympus, Japan). For the selected strains (Table 1), the following extracellular enzymatic activities were tested: fatty acid esterase (Tween 80 medium), protease (casein hydrolysis medium, gelatine hydrolysis medium), amylase (starch agar), β-glucosidase (aesculin agar) and urease (urease test agar) (Paterson and Bridge 1994).
Secondary metabolite analysis
Agar plugs (6 mm in diameter) were cut out of seven-day-old cultures growing on CYA and YES media. According to the method of Smedsgaard (1997), the cultures were extracted ultrasonically for 60 min with 500 μl of the solvent mixture of methanol/dichloromethane/ethyl acetate (3:2:1) containing 0.5% (v/v) formic acid. The organic solvent was transferred to a clean vial and evaporated by vacuum centrifugation. The residues were re-dissolved in 500 μl methanol, filtered through 0.45 μm Minisart filters, and analysed by high-performance liquid chromatograph (HPLC) (A1100; Agilent, Germany) with diode array detection at 210 nm, with 5 μl injections (Frisvad and Thrane 1987, 1993; Smedsgaard 1997). Separation was obtained with a 2 × 100 mm Luna2 OOD-4251-BO-C18 column (Phenomenex, Germany). The elution gradient was initially linear, from 85% water/15% acetonitrile to 100% acetonitrile over 20 min; 100% acetonitrile was then maintained for 5 min. A flow-rate of 0.4 ml min−1 was used. Both eluents contained 0.005% (v/v) trifluoroacetic acid. An alkylphenone analytical standard was used to define the retention time and an index was calculated for each peak detected. The secondary metabolites were identified by comparison with standard and by their characteristic UV spectra.
DNA isolation, amplification and sequence analysis
For selected isolates (Table 1), DNA was extracted following mechanical lysis (Gerrits van den Ende and de Hoog 1999), from ∼200 mg of four-day-old cultures grown on complete yeast extract medium (CYM; Raper et al. 1972), at 25°C in the dark. Amplification of the partial β-tubulin gene was carried out as described by O’Donnell and Cigelnik (1997) using the T1 and T22 primers. PCR was performed in a 50 μl reaction mixture with ∼10 ng of genomic DNA (GeneAmp PCR system 2400; Perkin Elmer, USA). The PCR products were separated electrophoretically on 1% agarose gels, and the expected bands were excised and purified with the DNA Gel Extraction Kit (Promega, USA), following the manufacturer protocol. The PCR fragments were sequenced with the T1 and T22 primers using a BigDyeTM terminator Ready Reaction Cycle Sequencing Kit on an ABI 3730xl DNA Analyser (Applied Biosystems, USA), as provided by the Macrogen Company (Korea). The partial β-tubulin gene sequences were aligned using CLUSTAL W (Thompson et al. 1994). For the phylogenetic analysis, the neighbor-joining (NJ) method (Saitou and Nei 1987) was used. The data were first analysed using the Tamura-Nei parameter distance calculation model (Tamura and Nei 1993), which was then used to construct the NJ tree using MEGA3.1 software (Kumar et al. 2004). To determine the support for each clade, bootstrap analysis was performed with 1000 replications. All of the sequences generated in this study have been deposited at GenBank; their accession numbers are given in Table 1.
Results and discussion
Polythermal glaciers are characterized by areas of massive surface ablation (80–100 m) that are drained by a stable, open-channel system with ice temperatures below zero and with cold (subfreezing) ice at the surface, margins and terminus of the glacier. The glaciers are exposed to rapid movements (Ekström et al. 2003; Fahnestock 2003), and consequently to frictional and geothermal melting of the ice at their base. Together with groundwater and seasonal supraglacial water, the water arising at the base contributes to the subglacial waters, which interact with rocks and sediments beneath the glacial ice (Foght et al. 2004). These processes create subglacial environments that have until recently been considered abiotic. However, a very high number of Penicillium species, at up to 13,000 CFU l−1, was recently isolated from the subglacial ice of this extreme environment (Gunde-Cimerman et al. 2003).
One of the Penicillium species isolated almost exclusively from the Kongsvegen glacier (Gunde-Cimerman et al. 2003; Sonjak et al. 2006) has characteristics that are different from known species and was therefore acknowledged as a new species; this was named P. svalbardense. It was isolated from subglacial ice in total counts of 200 CFU l−1, with the highest proportions obtained on a medium used for the detection of moderate xerophiles, dichloran 18% glycerol agar (DG18; Hocking and Pitt 1980), at 10°C, followed by malt extract agar with 5% NaCl (MEA5NaCl; Gunde-Cimerman et al. 2003) and malt extract agar with 15% NaCl (MEA15NaCl; Gunde-Cimerman et al. 2003), at 24°C. It was isolated from clear glacial ice at lower total counts, of up to 43 CFU l−1, again with the highest proportion obtained on DG18 at 10°C. The species was also isolated from Kongsvegen subglacial mineral ice inclusions, with the highest number (10 CFU g−1) obtained on MEA5NaCl at 10°C (Table 2).
To date, the genus Penicillium comprises approximately 225 described species that show a high diversity of morphology and secondary metabolites (Pitt et al. 2000). A combination of micromorphological, macromorphological and physiological characters is therefore needed to achieve satisfactory identification and classification (Frisvad and Samson 2004).
The proposed new species of P. svalbardense is characterized by its biverticillate penicilli, rough-walled stipes, globose to subglobose, smooth to slightly rough-walled conidia, good growth on CYA at 25°C and 30°C, and moderate to good growth on CREA. The morphology of their penicilli resembles most those of P. simplicissimum and P. piscarium from the subgenus Furcatum; however, there are differences in size, ornamentation and shape of the conidia (Table 3), which are among the most stable morphological characters in the genus Penicillium (Frisvad and Samson 2004).
Since Penicillium species produce several mycotoxins and other secondary metabolites that are species specific and usually very consistently expressed, chemotaxonomic studies are often used for their identification and classification (Frisvad et al. 1998). Thus, secondary metabolite production has been used extensively to distinguish between Penicillium species in the subgenus Furcatum (Frisvad and Filtenborg 1990). On the basis of secondary metabolites, P. svalbardense is phenotypically most closely related to P. piscarium, since they both produce indole diterpenes. However, P. piscarium does not produce xanthoepocin, which is found in P. svalbardense and was reported to be produced by P. simplicissimum (Igarashi et al. 2000). Both P. piscarium and P. simplicissimum also produce additional secondary metabolites that are not found in P. svalbardense (Tuthill et al. 2001) (Table 3).
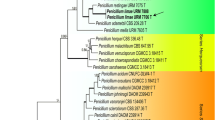
Molecular phylogenetic analyses based on ITS (Peterson 2000; Tuthill 2001) and recently particularly of the partial β-tubulin gene sequences (Seifert and Louis-Seize, 2000; Samson et al. 2004; Frisvad et al. 2006) have been used for Penicillium species delimitation. Skouboe et al. (1996) showed that ribosomal internal transcribed spacer (ITS) sequences were to invariant to provide sufficiently resolved phylogram. Therefore, in the case of P. svalbardense, partial β-tubulin gene sequencing was performed. We used T1 and T22 primers to amplify about 1500 bp of the β-tubulin gene. The smaller portion of about 700 bp that included the bt2a–bt2b region (Glass and Donaldson 1995) was sequenced. The sequences obtained were compared to the available sequences of the National Center for Biotechnology Informatic (NCBI) by using blast program BLAST-n (Altschul et al. 1990, 1997). Due to the lack of the Penicillium β-tubulin sequences in gene banks no close matches were obtained. The phylogenetic relationships of the partial β-tubulin gene sequences of ten strains belonging to different species from the subgenus Furcatum and additionally of the P. janthinellum strain used as an outgroup were inferred from NJ analysis. The tree produced is shown in Fig. 1. The P. svalbardense strains included in the analysis were grouped into a cluster with a 100% bootstrap value. The alignment of these strains showed no differences in base composition. P. svalbardense is phylogenetically most closely related to P. piscarium. The tree topology indicates on the intra-specific variation of β-tubulin sequences in single species P. piscarium and therefore on possible existence of two races, one being P. svalbardense and the other the two P. piscarium strains IBT 21815 and IBT 21002. However, the genetic dissimilarity between P. svalbardense and the closest P. piscarium strain, and particularly additional differences in morphology and secondary metabolite profiles clearly point to a new species.
Penicillium svalbardense Frisvad, Sonjak & Gunde-Cimerman, sp. nov.
Coloniae in agaro CYA 29–36 mm diam. post 7 dies 25°C, 30°C 40–46 mm, planae. Conidiophora mononematosa, biverticillata; stipites 250–500 μm longi, 2.5–3.5(−4) μm lati, asperulati; metulae leves, 13–25 × 2.5–3.5 μm; phialides lageniformes, collulo brevi praeditae, 9–13 × 2.3–3.5 μm. Conidia globosa vel subglobosa, leves vel asperulata, 2.7–3.4 μm diam, acervata caeruleo-grisea. Metabolitum: xanthoepocinum.
On CYA at 25°C after 7 days (Fig. 2a): colonies 29–36 mm in diameter, low, plane, with few light radial and annular zones, mononematous; conidiogenesis moderate to good, bluish-grey; mycelium white at margins; creamy-rose-coloured exudate, occasionally present; reverse light brown in centre.
On MEA at 25°C after 7 days (Fig. 2b): colonies 35–39 mm in diameter, low, plane, mononematous; conidiogenesis good, bluish-grey; mycelium white at margins; no exudate and soluble pigments; reverse uncoloured.
On YES at 25°C after 7 days (Fig. 2c): colonies 37–39 mm in diameter, low, umbonate in centre, with radial and annular zones; conidiogenesis weak to moderate, bluish green to brown; mycelium white; no exudate and soluble pigments; reverse yellow brown.
On CREA at 25°C after 7 days: colonies 22–25 mm in diameter, good growth, no acid or low acid production.
On CYA at 30°C after 7 days: colonies 40–46 mm in diameter.
On CYA at 15°C after 7 days: colonies 18–25 mm in diameter.
On CYAS at 25°C after 7 days: colonies 5–8 mm in diameter.
Extracellular enzymatic activities: positive fatty acid esterase, β-glucosidase and urease activities.
Conidiophores (Figs. 2d–h, 3) biverticillate, stipes 250–500 μm × 2.5–3.5 (−4) μm, rough-walled; metulae smooth-walled, 13–25 μm × 2.5–3.5 μm; phialides flask-shaped with a short neck 9–13 μm × 2.3–3.5 μm; conidia (Figs. 2i, 3) globose to subglobose, smooth to slightly rough-walled, 2.7–3.4 μm in diameter.
Etymology: the species is named after its origin, Svalbard archipelago.
GenBank accession numbers of partial β-tubulin gene sequences
EX-F 1227—DQ834933, EX-F 1307—DQ486644, EX-F 1319—DQ486643, CBS 372.48—DQ486650, IBT 13051—EF123659; IBT 15303—DQ834935, IBT 21815—DQ486648, IBT 21002—DQ486649, IBT 12452—DQ834934, CBS 223.66—DQ834936, IBT 15467—DQ486651.
Abbreviations
- CFU:
-
colony forming units
- DIC:
-
differential interference contrast
- HPLC:
-
high-performance liquid chromatography
- PCR:
-
polymerase chain reaction
- ITS:
-
internal transcribed spacer
References
Abyzov SS (1993) Microorganisms in the Antarctic ice. In: Friemann EI (ed) Antarctic microbiology. Wiley-Liss, Inc., New York, pp. 265–285
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Domsch KH, Gams W, Anderson TH (1980) Compendium of soil fungi. Academic Press, London
Ekström G, Nettles M, Abers GA (2003) Glacial earthquakes. Science 302:622–624
Fahnestock M (2003) Geophysics: glacial flow goes seismic. Science 302:578–579
Foght J, Aislabie J, Turner S, Brown CE, Ryburn J, Saul DJ, Lawson W (2004) Culturable bacteria in subglacial sediments and ice from two southern hemisphere glaciers. Microb Ecol 47:329–340
Frisvad JC (2004) Chemical diversity of psychrotolerant fungi. In: Watanabe MM, Suzuki K, Seki T (eds) Innovative roles of biological resources centers. Society for Culture Colections and Worlds Federation of Culture Colections, Tsukuba, pp. 165–167
Frisvad JC, Filtenborg O (1990) Revision of Penicillium subgenus Furcatum based on secondary metabolites and conventional characters. In: Samson RA, Pitt JI (eds) Modern concepts in Penicillium and Aspergillus classification. Plenum Press, New York, pp. 159–170
Frisvad JC, Larsen TO, Dalsgaard PW, Seifert KA, Louis-Seize G, Lyhne EK, Jarvis BB, Fettinger JC, Overy DP (2006) Four psychrotolerant species with high chemical diversity consistently producing cycloaspeptide A, Penicillium jamesonlandense sp. nov., Penicillium ribium sp. nov., Penicillium soppii and Penicillium lanosum. Int J Syst Evol Microbiol 56:1427–1437
Frisvad JC, Samson RA (2004) Polyphasic taxonomy of Penicillium subgenus Penicillium. A guide to identification of food and air-borne terverticillate Penicillia and their mycotoxins. Stud Mycol 49:1–173
Frisvad JC, Thrane U (1987) Standardized high-performance liquid chromatography of 182 mycotoxins and other fungal metabolites based on alkylphenone retention indices and UV-VIS spectra (diode array detection). J Chromatogr 404:195–214
Frisvad JC, Thrane U (1993) Liquid column chromatography of mycotoxins. J Chromatogr Library 54:253–372
Frisvad JC, Thrane U, Filtenborg O (1998) Role and use of secondary metabolites in fungal taxonomy. In: Frisvad JC, Bridge PD, Arora DK (Eds) Chemical fungal taxonomy. Marcel Dekker, New York, pp. 289–319
Gerrits van den Ende AHG, de Hoog GS (1999) Variability and molecular diagnostics of the neurotropic species Cladophialophora bantiana. Stud Mycol 43:151–162
Glass NL, Donaldson GC (1995) Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 61:1323–1330
Gunde-Cimerman N, Sonjak S, Zalar P, Frisvad JC, Diderichsen B, Plemenitaš A (2003) Extremophilic fungi in Arctic ice: a relationship between adaptation to low temperature and water activity. Phys Chem Earth pt B 28:1273–1278
Hocking AD, Pitt JI (1980) Dichloran—glycerol medium for enumeration of xerophilic fungi from low—moisture foods. Appl Environ Microbiol 39:488–492
Igarashi Y, Kuwamori Y, Takagi K, Ando T, Fudou R, Furumai T, Oki T (2000) Xanthoepocin, a new antibiotic from Penicillium simplicissimum IFO5762. J Antibiot (Tokyo) 53:928–933
Ivanushkina NE, Kochkina GA, Ozerskaya SM (2005) Fungi in ancient permafrost sediments of the Arctic and Antarctic regions. In: Castello JD, Rogers SO (eds) Life in ancient ice. Princeton University Press, Princeton, pp. 127–139
Kumar S, Tamura K, Nei M (2004) MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Ma L, Catranis CM, Starmer WT, Rogers SO (1999) Revival and characterization of fungi from ancient polar ice. Mycologist 13:70–73
Ma L, Rogers SO, Catranis CM, Starmer WT (2000) Detection and characterization of ancient fungi entrapped in glacial ice. Mycologia 92:286–295
McRae CF, Hocking AD, Seppelt RD (1999) Penicillium species from terrestrial habitats in the Windmill Islands, East Antarctica, including a new species, Penicillium antarcticum. Polar Biol 21:97–111
O’Donnell K, Cigelnik E (1997) Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol Phylogenet Evol 7:103–116
Paterson RRM, Bridge PD (1994) Biochemical techniques for filamentous fungi. IMI Technical Handbooks 1:19–26
Peterson SW (2000) Phylogenetic analysis of Penicillium species based on ITS and LSU-rDNA nucleotide sequences. In: Samson RA, Pitt JI (eds) Integration of modern taxonomic methods for Penicillium and Aspergillus classification. Harwood Academic Publishers, Amsterdam, pp. 163–178
Petrovič U, Gunde-Cimerman N, Zalar P (2000) Xerotolerant mycobiota from high altitude Anapurna soil, Nepal. FEMS Microbiol Lett 182:339–342
Pitt JI, Hocking AD (1999) Fungi and Food Spoilage. Aspen Publishers, Inc., Gaithersburg
Pitt JI, Samson RA, Frisvad JC (2000) List of accepted species and their synonyms in the family Trichocomaceae. In: Samson RA, Pitt JI (eds) Integration of modern taxonomic methods for Penicillium and Aspergillus classification. Harwood Academic Publishers, Amsterdam, pp. 9–47
Raper CA, Raper JR, Miller RE (1972) Genetic analysis of the life cycle of Agaricus bisporus. Mycologia 64:1088–1117
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Samson RA, Seifert KA, Kuijpers AFA, Houbraken JAMP, Frisvad JC (2004) Phylogenetic analysis of Penicillium subgenus Penicillium using partial β-tubulin sequences. Stud Mycol 49:175–200
Seifert KA, Louis-Seize G (2000) Phylogeny and species concepts in the Penicillium aurantiogriseum complex as inferred from the β-tubulin gene DNA sequences. In: Samson RA, Pitt JI (eds) Integration of modern taxonomic methods for Penicillium and Aspergillus classification. Harwood Academic Publishers, Amsterdam, pp. 189–198
Skouboe P, Boysen M, Pedersen LH, Frisvad JC, Rossen L (1996) Identification of Penicillium species using the internal transcribed spacer (ITS) regions. In: Rossen L, Rubio V, Dawson MT, Frisvad JC (eds) Fungal identification techniques. European Commission, Brussels, pp. 160–164
Smedsgaard J (1997) Micro-scale extraction procedure for standardized screening of fungal metabolite production in cultures. J Chromatogr 760:264–270
Sonjak S, Frisvad JC, Gunde-Cimerman N (2006) Penicillium mycobiota in Arctic subglacial ice. Microb Ecol 52:207–216
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tuthill DE, Frisvad JC, Christensen M (2001) Systematics of Penicillium simplicissimum based on rDNA sequences, morphology and secondary metabolites. Mycologia 93:298–308
Vishniac HS (1993) The microbiology of Antarctic soils. In: Friedman EI (ed) Antarctic microbiology. Wiley-Liss, Inc., New York, pp. 297–341
Acknowledgements
We thank Polona Zalar for the illustration of the P. svalbardense micromorphological features and Dr. Walter Gams for the latin diagnosis.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sonjak, S., Uršič, V., Frisvad, J.C. et al. Penicillium svalbardense, a new species from Arctic glacial ice. Antonie van Leeuwenhoek 92, 43–51 (2007). https://doi.org/10.1007/s10482-006-9133-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-006-9133-3