Abstract
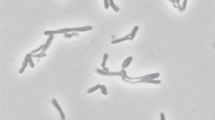
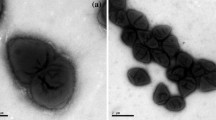
A psychrotolerant, obligate anaerobic, acetogenic bacterium designated strain SyrA5 was isolated from black anoxic sediment of a brackish fjord. Cells were Gram-positive, non-sporeforming rods. The isolate utilized H2/CO2, CO, fructose, glucose, ethanol, ethylene glycol, glycerol, pyruvate, lactate, betaine and the methyl-groups of several methoxylated benzoic derivatives such as syringate, trimethoxybenzoate and vallinate. The optimum temperature for growth was 29 °C, whilst slow growth occurred at 2 °C. The strain grew optimally with NaCl concentrations below 2.7% (w/v), but growth occurred up to 4.3% (w/v) NaCl. Growth was observed in the range from pH 5.9 to 8.5, optimum at pH 8. The G+C content was 44.1 mol%. Based upon 16S rRNA gene sequence analysis and DNA–DNA reassociation studies, the organism was classified in the genus Acetobacterium. Strain SyrA5 shared a 16S rRNA sequence similarity with A. carbinolicum of 100%, a fthfs gene (which codes for the N5,N10 tetrahydrofolate synthetase) sequence identity of 98.5–98.7% (amino acid sequence similarities were 99.4–100%) and a RNA–DNA hybridization homology of 64–68%. Despite a number of phenotypic differences between strain SyrA5 and A. carbinolicum we propose including strain SyrA5 as a subspecies of A. carbinolicum for which we propose the name Acetobacterium carbinolicum subspecies kysingense. The type strain is SyrA5 (=DSM 16427T, ATCC BAA-990).
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bache R. and Pfennig N. (1981). Selective isolation of Acetobacterium woodii on methoxylated aromatic acids and determination of growth yields. Arch. Microbiol. 130:255–261
Balch W.E., Schoberth S., Tanner R.S. and Wolfe R.S. (1977). Acetobacterium, a new genus of hydrogen-oxidizing, carbon dioxide-reducing, anaerobic bacteria. Int. J. Syst. Bacteriol. 27:355–361
Braun M. and Gottschalk G. (1982). Acetobacterium wieringae sp. nov., a new species producing acetic acid from molecular hydrogen and carbon dioxide. Zentralblatt fürBakteriologie Mikrobiologie und Hygiene I Abteilung Originale C-Allgemeine Angewandte Und Ökologische Mikrobiologie 3:368–376
Breznak J.A., Switzer J.M. and Seitz H.J. (1988). Sporomusa termitida sp. nov., an H2/CO2-utilizing acetogen isolated from termites. Arch. Microbiol. 150:282–288
Brioukhanov A.L., Thauer R.K. and Netrusov A.I. (2002). Catalase and Superoxide Dismutase in the Cells of Strictly Anaerobic Microorganisms. Microbiology 71:281–285
Brosius J., Dull T.J., Sleeter D.D. and Noller H.F. (1981). Gene organization and primary structure of a ribosomal-RNA operon from Escherichia coli. J. Mol. Biol. 148:107–127
Cashion P., Holder-Franklin M.A., McCully J. and Franklin M. (1977). A rapid method for the base ratio determination of bacterial DNA. Anal. Biochem. 81:461–466
Drake H.L. (ed.), 1994. Acetogenesis. In: Chapman and Hall Microbiology Series, Vol. 1. Chapman and Hall, New York
Drake H.L. and Daniel S.L. (2004). Physiology of the thermophilic acetogen Moorella thermoacetica. Res. Microbiol. 155:422–436
Eck R.V. and Dayhoff M.O. (1980). Atlas of protein sequence and structure 1966. Silver Spring, Maryland
Eichler B. and Schink B. (1984). Oxidation of primary aliphatic alcohols by Acetobacterium carbinolicum sp. nov., a homoacetogenic anaerobe. Arch. Microbiol. 140:147–152
Engelmann T., Kaufmann F. and Diekert G. (2001). Isolation and characterization of a veratrol:corrinoid protein methyl transferase from Acetobacterium dehalogenans. Arch Microbiol 175:376–383
Escara J.F. and Hutton J.R. (1980). Thermal-stability and renaturation of DNA in dimethylsulfoxide solutions - acceleration of the renaturation rate. Biopolymers 19:1315–1327
Felsenstein J. (1985). Confidence-limits on phylogenies - an approach using the bootstrap. Evolution 39:783–791
Fitch W.M. (1971). Toward defining course of evolution - minimum change for a specific tree topology. Syst. Zool. 20:406–416
Fontaine F.E., Peterson W.H., McCoy E. and Johnson M.J. (1942). A new type of glucose fermentation by Clostridium thermoaceticum n. sp. J. Bacteriol. 43:701–715
Frazer A.C. and Young L.Y. (1986). Anaerobic C1-metabolism of the O-methyl-C14-labeled substituent of vanillate. Appl. Environ. Microbiol. 51:84–87
Friedrich M.W. (2002). Phylogenetic analysis reveals multiple lateral transfers of adenosine-5′-phosphosulfate reductase genes among sulfate-reducing microorganisms. J. Bacteriol. 184:278–289
Frings J. and Schink B. (1994). Fermentation of phenoxyethanol to phenol and acetate by a homoacetogenic bacterium. Arch. Microbiol. 162:199–204
Frings J., Wondrak C. and Schink B. (1994). Fermentative degradation of triethanolamine by a homoacetogenic bacterium. Arch. Microbiol. 162:103–107
Hall T.A. (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series 41:95–98
Heijthuijsen J.H.F.G. and Hansen T.A. (1990). One-carbon metabolism in anaerobic non-methanogenic bacteria. In: Codd G.A., Dijkhuizen L., Tabita F.R. (eds) Autotrophic Microbiology and One-Carbon Metabolism. Kluwer, Dordrecht, pp. 163–191
Henckel T., Friedrich M. and Conrad R. (1999). Molecular analyses of the methane-oxidizing microbial community in rice field soil by targeting the genes of the 16S rRNA, particulate methane monooxygenase, and methanol dehydrogenase. Appl. Environ. Microbiol. 65:1980–1990
Huss V.A.R., Festl H. and Schleifer K.H. (1983). Studies on the spectrophotometric determination of DNA hybridization from renaturation rates. Syst. Appl. Microbiol. 4:184–192
Jahnke K.D. (1992). Basic computer-program for evaluation of spectroscopic DNA renaturation data from Gilford-System-2600 spectrophotometer on a Pc/Xt/at type personal computer. J. Microbiol. Meth. 15:61–73
Jukes T.H. and Cantor C.R. (1969). Evolution of protein molecules. In: Munro H.N. (eds) Mammalian Protein Metabolism. Academic Press, New York, pp. 21–132
Kerby R., Niemczura W. and Zeikus J.G. (1983). Single-carbon catabolism in acetogens - analysis of carbon flow in Acetobacterium woodii and Butyribacterium methylotrophicum by fermentation and C13 nuclear magnetic resonance measurement. J.Bacteriol. 155:1208–1218
Kimura M. (1980). A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16:111–120
Klemm P., Christiansen G., Kreft B., Marre R. and Bergmans H. (1994). Reciprocal exchange of minor components of type 1 and F1C fimbriae results in hybrid organelles with changed receptor specificities. J.Bacteriol. 176:2227–2234
Kotsyurbenko O.R., Simankova M.V., Nozhevnikova A.N., Zhilina T.N., Bolotina N.P., Lysenko A.M. and Osipov G.A. (1995). New species of psychrophilic acetogens - Acetobacterium bakii sp. nov., A. paludosum sp. nov., A. fimetarium sp. nov. Arch. Microbiol. 163:29–34
Krumholz L.R., Harris S.H., Tay S.T. and Suflita J.M. (1999). Characterization of two subsurface H2-utilizing bacteria, Desulfomicrobium hypogeium sp. nov. and Acetobacterium psammolithicum sp. nov., and their ecological roles. Appl. Environ. Microbiol. 65:2300–2306
Kumar S., Tamura K., Jakobsen I.B. and Nei M. (2001). MEGA2: molecular evolutionary genetics analysis software. Bioinformatics 17:1244–1245
Lane D.J. (1991) 16S/23S rRNA sequencing. In: Stackebrandt E., Goodfellow M. (eds) Nucleic Acids Techniques in Bacterial Systematics. John Wiley, Chichester, pp. 115–175
Leaphart A.B. and Lovell C.R. (2001) Recovery and Analysis of Formyltetrahydrofolate Synthetase Gene Sequences from Natural Populations of Acetogenic Bacteria. Appl.Environ.Microbiol. 67:1392–1395
Ludwig W., Strunk O., Westram R., Richter L., Meier H., Yadhukumar, Buchner A., Lai T., Steppi S., Jobb G., Forster W., Brettske I., Gerber S., Ginhart A.W., Gross O., Grumann S., Hermann S., Jost R., Konig A., Liss T., Lussmann R., May M., Nonhoff B., Reichel B., Strehlow R., Stamatakis A., Stuckmann N., Vilbig A., Lenke M., Ludwig T., Bode A., Schleifer K.H. (2004). ARB: a software environment for sequence data. Nucleic Acids Res. 32:1363–1371
Lueders T., Manefield M. and Friedrich M.W. (2004). Enhanced sensitivity of DNA- and rRNA-based stable isotope probing by fractionation and quantitative analysis of isopycnic centrifugation gradients. Environ. Microbiol. 6:73–78
Madigan M.T., Martinko J.M. and Parker J. (2000). Microbial growth. In: Corey P.F. (eds) Brock Biology of Microorganisms. Prentice Hall, New Jersey, pp. 135–162
Maidak B.L., Cole J.R., Lilburn T.G., Parker C.T., Saxman P.R., Farris R.J., Garrity G.M., Olsen G.J., Schmidt T.M. and Tiedje J.M. (2001). The RDP-II (Ribosomal Database Project). Nucleic Acids Research 29:173–174
Mechichi T., Labat M., Patel B.K.C., Woo T.H.S., Thomas P. and Garcia J.L. (1999a). Clostridium methoxybenzovorans sp. nov., a new aromatic O-demethylating homoacetogen from an olive mill wastewater treatment digester. Int. J. Syst. Bacteriol. 49:1201–1209
Mesbah M., Premachandran U. and Whitman W.B. (1989). Precise measurement of the G+C content of deoxyribonucleic-acid by high-performance liquid-chromatography. Int. J. Syst. Bacteriol. 39:159–167
Mountfort D.O., Grant W.D., Clarke R. and Asher R.A. (1988). Eubacterium callanderi sp. nov. that demethoxylates O-methoxylated aromatic acids to volatile fatty acids. Int. J. Syst. Bacteriol. 38:254–258
Pfennig N. (1978). Rhodocyclus purpureus gen. nov. and sp. nov. a ring-shaped, vitamin B12-requiring member of family Rhodospirillaceae. Int. J. Syst. Bacteriol. 28:283–288
Rossello-Mora R. and Amann R. (2001). The species concept for prokaryotes. FEMS Microbiol. Rev. 25:39–67
Saitou N. and Nei M. (1987). The neighbor-joining method - a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4:406–425
Schuppert B. and Schink B. (1990). Fermentation of methoxyacetate to glycolate and acetate by newly isolated strains of Acetobacterium sp. Arch. Microbiol. 153:200–204
Seeley J.H.W., VanDemark P.J. and Lee J.J. (1991). Staining bacteria. In: Kennedy D., Park R.B. (eds) Microbes in Action, A Laboratory Manual of Microbiology. W. H. Freeman and Company, New York, pp. 71–90
Simankova M.V., Kotsyurbenko O.R., Stackebrandt E., Kostrikina N.A., Lysenko A.M., Osipov G.A. and Nozhevnikova A.N. (2000). Acetobacterium tundrae sp. nov., a new psychrophilic acetogenic bacterium from tundra soil. Arch. Microbiol. 174:440–447
Smibert R.M. and Krieg N.R. (1994). Phenotypic characterization. In: Gerhardt P., Murray R.G.E., Wood W.A., Krieg N.R. (eds) Methods for General and Molecular Bacteriology. American Society for Microbiology, Washington, D. C., pp. 607–655
Soerensen A.H., Winthernielsen M. and Ahring B.K. (1991). Kinetics of lactate, acetate and propionate in unadapted and lactate-adapted thermophilic, anaerobic sewage-sludge - the influence of sludge adaptation for start-up of thermophilic uasb-reactors. Appl. Microbiol. Biotechnol. 34:823–827
Tamaoka J. and Komagata K. (1984). Determination of DNA-base composition by reversed-phase high-performance liquid-chromatography. FEMS Microbiol. Lett. 25:125–128
Tanaka K. and Pfennig N. (1988). Fermentation of 2-methoxyethanol by Acetobacterium malicum sp. nov. and Pelobacter venetianus. Arch. Microbiol. 149:181–187
Tindall B.J. (1990a). A comparative study of the lipid composition of Halobacterium saccharovorum from various sources. Syst. Appl. Microbiol. 13:128–130
Tindall B.J. (1990b). Lipid composition of Halobacterium lacusprofundi. FEMS Microbiol. Lett. 66:199–202
Traunecker J., Preuss A. and Diekert G. (1991). Isolation and characterization of a methyl chloride utilizing, strictly anaerobic bacterium. Arch. Microbiol. 156:416–421
Tschech A. and Pfennig N. (1984). Growth-yield increase linked to caffeate reduction in Acetobacterium woodii. Arch. Microbiol. 137:163–167
Visuvanathan S., Moss M.T., Stanford J.L., Hermontaylor J. and Mcfadden J.J. (1989). Simple enzymic method for isolation of DNA from diverse bacteria. J. Microbiol. Met. 10:59–64
Wayne L.G., Brenner D.J. and Colwell R.R., et al. 1987. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int. J. Syst. Bacteriol. 37: 463–464
Wheeler D.L., Church D.M., Lash A.E., Leipe D.D., Madden T.L., Pontius J.U., Schuler G.D., Schriml L.M., Tatusova T.A., Wagner L. and Rapp B.A. (2002). Database resources of the National Center for Biotechnology Information: 2002 update. Nucleic Acids Research 30:13–16
Widdel F., Kohring G.W. and Mayer F. (1983). Studies on dissimilatory sulfate-reducing bacteria that decompose fatty-acids .3. characterization of the filamentous gliding Desulfonema limicola gen. nov., sp. nov., and Desulfonema magnum sp. nov. Arch. Microbiol. 134:286–294
Widdel F. and Bak F. (1992). Gram-negative mesophilic sulfate-reducing bacteria. In: Balows H., Trüper H.G., Dworkin M., Harder W., Schleifer K.H. (eds) The prokaryotes. 2 ed., Springer-Verlag, New York
Willems A., Collins M.D. (1996) Phylogenetic relationships of the genera Acetobacterium and Eubacterium sensu stricto and reclassification of Eubacterium alactolyticum as Pseudoramibacter alactolyticus gen nov, comb nov. Int. J. System. Bacteriol 46:1088–1087
Zeikus J.G. (1983). Metabolism of one-carbon compounds by chemotropic anaerobes. Ad. Micro. Physiol. 24:215–299
Zeikus J.G., Lynd L.H., Thompson T.E., Krzycki J.A., Weimer P.J. and Hegge P.W. (1980). Isolation and characterization of a new, methylotrophic, acidogenic anaerobe, the Marburg Strain. Curr Microbiol 6:381–386
Acknowledgements
The study was supported by NSF grant No. 21-00-0309. The authors thank Peter Westermann for providing help with the measurement of acetate.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Paarup, M., Friedrich, M., Tindall, B. et al. Characterization of the psychrotolerant acetogen strain SyrA5 and the emended description of the species Acetobacterium carbinolicum . Antonie Van Leeuwenhoek 89, 55–69 (2006). https://doi.org/10.1007/s10482-005-9009-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-005-9009-y