Abstract
Hyaluronan (HA) plays a crucial role in the lubricating and buffering properties of synovial fluid. The purpose of this study was to examine the effects of interleukin (IL)-1β on HA degradation in cultured synovial membrane cells. The rabbit synovial membrane cell line HIG-82 was cultured with and without IL-1β. The amounts of HA of varying molecular weights in the medium were analyzed using high-performance liquid chromatography, the mRNA levels of HA synthase (HAS) and hyaluronidase (HYAL) were analyzed by means of real-time PCR, and HYAL activity was analyzed by HA zymography. The amounts of HA with a molecular weight lower than 300 kDa, and between 300 and 1900 kDa, in the culture medium of HIG-82 cells were significantly higher in the presence of IL-1β. However, the amount of HA with a molecular weight greater than 1900 kDa was significantly lower in the presence of IL-1β. Both HAS2 and HAS3 mRNA levels were upregulated by treatment with IL-1β. So, too, were the levels of HYAL1 and HYAL2 mRNA, which resulted in enhanced HYAL activity. However, HYAL activity was inhibited by transfection of HYAL2-siRNA. Our results suggest that IL-1β is a crucial factor in the fragmentation of HA in inflammatory joints.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hyaluronan (HA) is a glycosaminoglycan with an unmodified polymer of repeated disaccharides of d-glucuronic acid and N-acetyl-d-glucosamine. It is present in all vertebrates and certain bacterial pathogens. HA typically has a high molecular weight (800–1900 kDa) in its native state, and has various physiological and biological functions.11 In synovial fluid, high molecular weight-HA (HMW-HA) is essential for joint function, due to its physiological properties under physiological conditions. However, an increase in the level of low molecular weight-HA (LMW-HA) under inflammatory or pathologic conditions, such as osteoarthritis (OA), reduces the viscoelasticity of synovial fluid, leading to deterioration of joint lubrication.1 In addition, LMW-HA has effects on immune and inflammatory processes.16
In synovial fluid, the amount of HA and its molecular weight distribution is determined by a balance between the synthesis and degradation of HA. HA is synthesized on the cytoplasmic space of plasma membranes by three kinds of HA synthase (HAS1, HAS2, and HAS3).8 HAS1 and HAS2 polymerize HA chains of similar lengths (up to 2 × 103 kDa), whereas HAS3 produces shorter chains (200–300 kDa).8
The accumulation of LMW-HA is thus believed to be due to depolymerization with enzymatic cleavage17 and to nonenzymatic reactions, such as those triggered by reactive oxygen species.13 It has been suggested that the degradation of HA by hyaluronidase (HYAL) plays a crucial role in the development of OA.29 HYAL is a family of β-endoglucosidases that degrades HA into small fragments by cleaving internal β(1,4) linkages.10 In human, the HYAL genes HYAL1, HYAL2, HYAL3, HYAL4, PH-20, and HYALP1 have been identified.2 Among these, HYAL1 and HYAL2 are widely distributed in mammalian tissues,2,25 and are the principal mediators of HA catabolism.23 HYAL1 and 2 are mainly expressed in human synovial fluid. The expression level of HYAL2 in the synovial fluid of the knee joint was found to be significantly higher in patients with OA or rheumatoid arthritis (RA) than in healthy controls.29 HYAL1 was originally purified from human plasma, 4 and is highly localized in the major parenchymal organs.2 In contrast, HYAL2 is present in all tissues except adult brain.25 HYAL2 degrades HMW-HA into small HA fragments that have a molecular weight of approximately 20 kDa.12 However, the mechanisms responsible for modulation of HYAL1 and HYAL2 expression and activity in synovial membrane cells remain unclear.
Several cytokines and growth factors, including interleukin-1 beta (IL-1β) and TNF-α have been detected in the synovial fluid of patients with inflammatory disease of the joints.20 IL-1β is highly expressed in synovial fluid in OA joints, and a 10-fold higher level of IL-1β was detected in the synovial membrane of patients with OA, as compared to individuals with normal synovial membrane.26 IL-1β is a principal proinflammatory cytokine. It enhances the catabolic action of several cells, such as fibroblasts, and strongly interacts with receptors on the surface of synovial membrane.26 Because IL-1β is abundant in synovial fluid from joints of individuals with OA and RA, it has been implicated in the progression of these conditions.21,26 A previous study demonstrated the effect of IL-1β on HA synthesis in synovial membrane cells.14 However, the mechanism underlying the LMW-HA accumulation caused by IL-1β has not been clarified.
We hypothesized that IL-1β plays a crucial role in HA fragmentation in synovial fluid. Therefore, we used the synovial membrane cell line HIG-82 to elucidate the effects of IL-1β on the synthesis and degradation of HA by HYALs.
Materials and Methods
Cell Culture
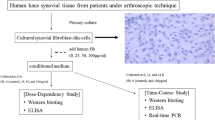
Rabbit knee synovial membrane cell line HIG-82 was purchased from American Type Culture Collection (Manassas, VA). The cells were cultured on 100-mm culture dishes (Corning, New York, NY), and the cultures were maintained in 10 mL α-minimum essential medium (α-MEM, Sigma–Aldrich, St. Louis, MO), supplemented with 292 μg/mL l-glutamine (Katayama Chemical, Osaka, Japan), 60 mg/mL kanamycin sulfate (Meiji Seika, Tokyo, Japan), 50 U/mL penicillin G (Sigma–Aldrich), and 10% fetal bovine serum (FBS, Mitsubishikasei, Tokyo, Japan) under an atmosphere of 5% CO2 in a humidified incubator. The medium was changed every second day.
Stimulation of HIG-82 Cells by IL-1β
The medium was replaced with another containing 0.5% FBS 12 h before the experiments. Then, HIG-82 cells were incubated in medium without FBS in the presence or absence of recombinant human IL-1β (Sigma–Aldrich; 0.1, 1.0, and 10 ng/mL) for 1–72 h.
Quantification of HA Concentration in the Culture Medium of HIG-82 Cells
The amount and size of HA in media were analyzed according to previously described method.28 Briefly, the medium was moved to extraction columns (Bond Elut SCX and BondElut SAX; GL Sciences, Tokyo, Japan). After washing, the solvents were eluted by 3 mL of 50 mM MeOH/HCl. The samples were then dried and dissolved in 500 μL of 0.1 M NaCl.
High-performance liquid chromatography (HPLC; Waters 600E, Waters, Tokyo, Japan) with gel filtration columns (Ohpak KB-804 for the fraction under 1,000 kDa, Ohpak KB-806 for the fraction of 1000–20,000 kDa; Shodex, Tokyo, Japan) was used. Elution was carried out with 0.1 mM NaCl, at a flow rate of 1.0 mL/min. The column effluent was monitored by a differential refractometer (RI DETECTOR 504R; GL Sciences, Tokyo, Japan). HAS1 and HAS2 synthesize HA with up to 2 × 103 kDa, usually defined as high molecular weight-HA, whereas HAS3 polymerases HA with 200–300 kDa, can be defined as low molecular weight-HA.8 Therefore, the accumulation levels of HA of various molecular weights were quantified based on the calibration curves for pure HAs with molecular weights of 300 and 1900 kDa (Denki Kagaku Kogyo, Tokyo, Japan).
Quantitative Real-time PCR Analysis
The mRNA levels of HAS2, HAS3, HYAL1, and HYAL2 were examined by quantitative real-time PCR analysis using a LightCycler® system (Roche Diagnosics, Tokyo, Japan) and QuantiTectTM SYBR® Green PCR Master Mix (QIAGEN, Tokyo, Japan). Total RNA was extracted from the cell cultures using Trizol® Reagent (Gibco BRL, Gaithersburg, MD). First-strand cDNA was synthesized from 1 μg total RNA using Rever Tra Ace-α (Toyobo, Osaka, Japan). Table 1 shows the sequences of the primers for HAS2, HAS3, HYAL1, HYAL2, and glyceraldehyde 3-phosphate dehydrogenase (GAPDH). The signals of HASs and HYALs were evaluated in a qualitative manner, relative to the GAPDH signals. Normalized Ct values were expressed relative to the controls.
Substrate Gel Electrophoresis (Zymography)
Zymography was performed according to a previously described technique, with some modifications.6 Polyacrylamide gel containing human umbilical cord HA (ICN Biomedical, Costa Mesa, CA) was used for gel electrophoresis. Nine milliliters of culture medium was concentrated 30 times, and a sample of 2 μL was diluted with 0.0625 M Tris/HCl, pH 6.8, containing 10% glycerol and 0.5% bromphenol blue. Gel electrophoresis was performed by use of a running buffer containing 0.25 M Tris and 0.192 M glycine at pH 9.0. The gels were incubated at 37 °C in the buffer containing 0.25 M NaCl and 0.1 M sodium formate, pH 3.7, for 14 h. After fixation with 7% acetic acid for 1 h, the gels were stained with 0.5% Alcian blue in 3% acetic acid. Destaining was performed with 50% methanol and 10% acetic acid for 30 min. Gels were then washed twice with 5% methanol and 7% acetic acid for 1 h. The experiments were repeated in triplicate. The gels were dried, and scanned by means of an image analyzer (VersaDoc 5000MP, Bio-Rad, Hercules, CA). Gel analysis software (Quantity One, Bio-Rad) was used for quantitative analysis of the detected bands. The signal intensity was evaluated relative to the control signals.
Transfection of HIG-82 Cells with HYAL2-Small Interfering RNA
Three different HYAL2-small interfering (si) RNA duplexes targeting the firefly luciferase gene were designed using a rabbit HYAL2 cDNA sequence (GenBank; AY603960) and then chemically synthesized (B-Bridge, Osaka, Japan; Table 2). GL3 Luciferase Duplex (B-Bridge) was used as a negative control.
The HIG-82 cells were treated with HYAL2-siRNA or 1 μM GL3 Luciferase Duplex in antibiotic-free α-MEM with 2% siFECTOR (B-Bridge) and 20% FBS, and incubated for 40 h at 37 °C. The optimum concentration of FBS for the transfection was defined as 20% in a preliminary study.
Statistical Analysis
The experiments were repeated in triplicate. Means and standard deviations were calculated from the data obtained and then subjected to Student’s t-test or one-way ANOVA followed by Scheffe’s multiple comparisons test using Graphpad Prism 4.0a software (Graphpad Software, San Diego, CA) to examine mean differences at the 1% and 5% levels of significance.
Results
Concentration and Size of HA in Culture Medium of HIG-82 Cells
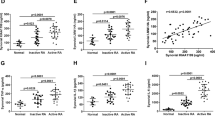
In the culture medium of HIG-82 cells, the concentration of HA with a molecular weight lower than 300 kDa (LMW-HA) was significantly higher (p < 0.01) after 72 h of IL-1β treatment, as compared with control (Fig. 1a). After treatment with IL-1β, the concentration of HA with a molecular weight between 300 and 1,900 kDa (medium molecular weight-HA; MMW-HA) was significantly higher (p < 0.05), whereas the concentration of HA with a molecular weight higher than 1900 kDa (HMW-HA) was significantly lower (p < 0.05) (Figs. 1b and 1c).
Concentration of HA, by size, in the medium of HIG-82 cell culture HIG-82 cells were treated with 1 ng/mL IL-1β for 0–72 h. The concentration and size of HA in the medium were analyzed by means of HPLC with gel filtration columns and differential refractometer. n = 3, **p < 0.01, *p < 0.05. (a) HA with a molecular weight lower than 300 kDa. (b) HA with a molecular weight from 300 to 1900 kDa, and (c) HA with a molecular weight greater than 1900 kDa
Gene Expressions of HAS2 and HAS3 in Cultured HIG-82 Cells
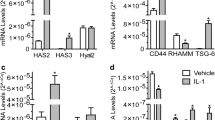
HAS2 mRNA levels were upregulated significantly (p < 0.01, after 3 h; p < 0.05, after 1 and 6 h) by treatment with 1 ng/mL IL-1β, as compared with untreated controls (Fig. 2a).
HAS3 mRNA levels were also upregulated significantly (p < 0.01, after 1 and 3 h; p < 0.05, after 6 h) by treatment with 1 ng/mL IL-1β (Fig. 3b).
Gene Expressions of HYAL1 and HYAL2 in Cultured HIG-82 Cells
HYAL1 mRNA level was immediately downregulated (p < 0.01) after 1-h stimulation with 1 ng/mL IL-1β, as compared with untreated control, and continued to decrease until 12-h stimulation (Fig. 3a).
In contrast, HYAL2 mRNA levels were upregulated significantly (p < 0.01, after 3 h; p < 0.05, after 1, 6, and 12 h) by treatment with 1 ng/mL IL-1β (Fig. 3b).
HYAL Activity in Culture Medium of HIG-82 Cells
Because of the enzymatic activity of HAYL occurred in the culture medium, and only a small amount remained in the cell layer,22 the culture medium was subjected to HA zymography. A band was detected at 57 kDa, which corresponds to the molecular weight of HYAL2, on the gel at pH 3.7 (Fig. 4a). HAYL activity was not detected clearly in the untreated control. HYAL activity was enhanced significantly by IL-1β treatment, in a dose-dependent manner (0–10 ng/mL), after 14 h (Fig. 4b). No HYAL activity was detected in the experimental or control groups at pH 7.0 (data not shown).
HYAL activity in HIG-82 cells, as detected by HA zymography. HIG-82 cells were treated with 0, 0.1, 1.0, and 10 ng/mL IL-1β for 14 h. (a) Nine milliliters of the culture medium of synovial membrane cells was collected, and concentrated 30 times to obtain detectable amounts of HYAL activity. Gel electrophoresis was performed using polyacrylamide gel containing human umbilical cord HA. The gels were incubated at 37 °C, pH 3.7, for 14 h, and stained with 0.5% Alcian blue. HYAL activity was visualized as a band at 57 kDa (arrow). (b) The gels were scanned by means of the image analyzer. The signal intensity of bands was evaluated relative to the control signals using the gel analysis software. n = 3, **p ≤ 0.01, *p ≤ 0.05
Effect of HYAL2-siRNA on HYAL2 Activity in Cultured HIG-82 Cells
HYAL2-siRNA-transfected and the control (treated with GL3 Luciferase Duplex) HIG-82 cells were incubated with and without 1 ng/mL IL-1β. After 14-h incubation, the HYAL activity was enhanced significantly by the treatment with IL-1β in the control cells (Figs. 5a and 5b). However, the HYAL activity became much lower in the HYAL2-siRNA-transfected cells than that in the control cells after the treatment with IL-1β.
HYAL activity in HIG-82 cell culture with and without HYAL2-siRNA transfection HIG-82 cells at 80% confluence were treated with HYAL2-siRNA. GL3 luciferase duplex was used as a negative control. (a) Forty hours after transfection, the HIG-82 cells were incubated with and without 1 ng/mL IL-1β for 12 h. The HYAL activity in the culture medium was quantified by HA zymography. HYAL activity was visualized as a band at 57 kDa (arrow). (b) The gels were scanned by means of the image analyzer. The signal intensity of bands was evaluated relative to the control signals using the gel analysis software. n = 3, *p ≤ 0.05
Discussion
In the present study, levels of HAS2 and HAS3 mRNA were upregulated by IL-1β treatment in HIG-82 cells. In our previous study, expression of HAS2 and HAS3 mRNAs, but not HAS1 mRNA, was detected in the synovial membrane of rabbit knee joints in both in vivo and in vitro experiments.27 HAS1 mRNA was not detected in the present study (data not shown). Concentrations of LMW-HA and MMW-HA in culture medium were higher, whereas that of HMW-HA was lower, in the presence of IL-1β.
It was demonstrated that IL-1β increased glucosamine incorporation into glycosaminoglycan in cultured synovial membrane cells,14 suggesting the enhancement of HA synthesis activity by IL-1β. In addition, IL-1β induced accumulation of LMW-HA in cultured synovial membrane cells that were derived from patients with OA and RA.9 In human lung fibroblasts, LMW-HA was reported to accumulate in the presence of IL-1β.19 These findings are consistent with the results of the present study.
Our results suggest that the increase in LMW-HA in synovial cell culture may be partly due to the upregulation of HAS3 induced by IL-1β. However, increased MMW-HA and decreased HMW-HA in culture decrease upregulation of HAS2 in synovial cells, which suggests that degradation of HMW-HA is enhanced.
The activity of HYALs can be regulated by cytokines and growth factors.3 In the present study, HYAL activity was enhanced by treatment with IL-1β on HA zymography, suggesting that HAYL plays a role in the accumulation of LMW-HA and MMW-HA, and the decrease in HMW-HA, after IL-1β treatment of HIG-82 cells.
In addition, the gene expressions of HYAL1 and HYAL2 differed: HYAL1 mRNA expression was significantly downregulated, whereas HYAL2 mRNA level was significantly upregulated, by treatment with IL-1β. A previous study reported a high level of HYAL2 expression in the synovial membrane of patients with OA, suggesting a crucial role for HYAL2 in the degradation of HA in the joints of OA patients.29
In monolayer culture, most HYAL activity is quickly secreted into the culture medium and is not retained by the cell layer.22 In addition, HYAL activity can be detected only at an acidic pH of 4.5–pH 3.7.3 Therefore, in the present study, the culture medium was subjected to HA zymography at pH 3.7. HYAL activity was detected only at pH 3.7; none was detected at a neutral pH (data not shown). This result is consistent with that of a previous study, which found that HYAL activity in synovial fluid can be detected with HA zymography only at the optimal pH of 4.0, whereas no activity was detectable at pH 5.0 to 7.0.15
HYAL2 has been detected in lysosomes12 and is linked by a glycosylphosphatidylinositol (GPI) anchor to the outer cell membrane,18 thus contributing to the initial cleaving of HMW-HA.12 This membrane-associated HYAL2 is believed to be active in acid-neutral conditions (pH 6.0–7.0) in association with CD44, the predominant receptor for HA.7 However, HYAL2 released from cell membrane has been reported to be active in acidic conditions,12 implying that the free-form HYAL2 released from synovial membrane cells may be inactive at physiological pH in synovial fluid, and active when HYAL2 is present in lysosomal compartments.
HYAL1 is a lysosomal enzyme, and is active only at an acidic pH (optimum pH, 3.8–4.3).5 HYAL1 degrades HA of any molecular size, a process that generates mainly tetrasaccharides or hexasaccharides.4,5
HYAL2 degrades HMW-HA into small fragments of HA with a molecular weight of approximately 20 kDa12; however, the expression and localization of HYAL1 differ from those of HYAL2, and HYAL1 digests small fragments of HA produced by HYAL2 activity.2 However, the role of HYAL1 in HA degradation has not been fully elucidated, and further study will be undertaken in the near future.
HYAL1 and HYAL2 may have mutually exclusive roles in the catabolism of HA in synovial membrane cells. Although it has been suggested that HYAL1 and HYAL2 act either in sequence in the degradation of HA, or have distinct functions with respect to the size of HA,24 their roles in synovial membrane cells remain to be resolved.
To examine the role of HYAL2 in HA degradation induced by IL-1β in HIG-82 cells, we attempted to inhibit expression of HYAL2 mRNA by HYAL2-siRNA transfection. HYAL activity was indeed inhibited, suggesting that HYAL2 plays a crucial role in the enhancement of HA degradation by IL-1β.
In conclusion, we showed that IL-1β induces fragmentation of HA by HYAL activity, which indicates that, under inflammatory conditions, IL-1β is a crucial factor in the degradation of HA in synovial fluid.
References
Bjelle, A., T. Andersson, and K. Granath. Molecular weight distribution of hyaluronic acid of human synovial fluid in rheumatic diseases. Scand. J. Rheumatol. 12(2):133–138, 1983.
Csoka, A. B., G. I. Frost, and R. Stern. The six hyaluronidase-like genes in the human and mouse genomes. Matrix Biol. 20(8):499–508, 2001.
Flannery, C. R., C. B. Little, C. E. Hughes, and B. Caterson. Expression and activity of articular cartilage hyaluronidases. Biochem. Biophys. Res. Commun. 251(3):824–829, 1998.
Frost, G. I., T. B. Csoka, T. Wong, and R. Stern. Purification, cloning, and expression of human plasma hyaluronidase. Biochem. Biophys. Res. Commun. 236(1):10–15, 1997.
Frost, G. I., and R. Stern. A microtiter-based assay for hyaluronidase activity not requiring specialized reagents. Anal. Biochem. 251(2):263–269, 1997.
Guntenhoner, M. W., M. A. Pogrel, and R. Stern. A substrate-gel assay for hyaluronidase activity. Matrix 12(5):388–396, 1992.
Harada, H., and M. Takahashi. CD44-dependent intracellular and extracellular catabolism of hyaluronic acid by hyaluronidase-1 and -2. J. Biol. Chem. 282(8):5597–5607, 2007.
Itano, N., T. Sawai, M. Yoshida, P. Lenas, Y. Yamada, M. Imagawa, T. Shinomura, M. Hamaguchi, Y. Yoshida, Y. Ohnuki, et al. Three isoforms of mammalian hyaluronan synthases have distinct enzymatic properties. J. Immunol. 274(35):25085–25092, 1999.
Konttinen, Y. T., H. Saari, and D. C. Nordstrom. Effect of interleukin-1 on hyaluronate synthesis by synovial fibroblastic cells. Clin. Rheumatol. 10(2):151–154, 1991.
Kreil, G. Hyaluronidases—a group of neglected enzymes. Protein Sci. 4(9):1666–1669, 1995.
Laurent, T. C., and J. R. Fraser. Hyaluronan. Faseb J. 6(7):2397–2404, 1992.
Lepperdinger, G., B. Strobl, and G. Kreil. HYAL2, a human gene expressed in many cells, encodes a lysosomal hyaluronidase with a novel type of specificity. J. Biol. Chem. 273(35):22466–22470, 1998.
McNeil, J. D., O. W. Wiebkin, W. H. Betts, and L. G. Cleland. Depolymerisation products of hyaluronic acid after exposure to oxygen-derived free radicals. Ann. Rheum. Dis. 44(11):780–789, 1985.
Meyer, F. A., I. Yaron, and M. Yaron. Synergistic, additive, and antagonistic effects of interleukin-1 beta, tumor necrosis factor alpha, and gamma-interferon on prostaglandin E, hyaluronic acid, and collagenase production by cultured synovial fibroblasts. Arthritis Rheum. 33(10):1518–1525, 1990.
Nagaya, H., T. Ymagata, S. Ymagata, K. Iyoda, H. Ito, Y. Hasegawa, and H. Iwata. Examination of synovial fluid and serum hyaluronidase activity as a joint marker in rheumatoid arthritis and osteoarthritis patients (by zymography). Ann. Rheum. Dis. 58(3):186–188, 1999.
Oertli, B., B. Beck-Schimmer, X. Fan, and R. P. Wuthrich. Mechanisms of hyaluronan-induced up-regulation of ICAM-1 and VCAM-1 expression by murine kidney tubular epithelial cells: hyaluronan triggers cell adhesion molecule expression through a mechanism involving activation of nuclear factor-kappa B and activating protein-1. J. Immunol. 161(7):3431–3437, 1998.
Orkin, R. W., and B. P. Toole. Isolation and characterization of hyaluronidase from cultures of chick embryo skin- and muscle-derived fibroblasts. J. Biol. Chem. 255(3):1036–1042, 1980.
Rai, S. K., F. M. Duh, V. Vigdorovich, A. Danilkovitch-Miagkova, M. I. Lerman, and A. D. Miller. Candidate tumor suppressor HYAL2 is a glycosylphosphatidylinositol (GPI)-anchored cell-surface receptor for jaagsiekte sheep retrovirus, the envelope protein of which mediates oncogenic transformation. Proc. Natl. Acad Sci. USA 98(8):4443–4448, 2001.
Sampson, P. M., C. L. Rochester, B. Freundlich, and J. A. Elias. Cytokine regulation of human lung fibroblast hyaluronan (hyaluronic acid) production. Evidence for cytokine-regulated hyaluronan (hyaluronic acid) degradation and human lung fibroblast-derived hyaluronidase. J. Clin. Invest. 90(4):1492–1503, 1992.
Schlaak, J. F., I. Pfers, K. H. Meyer Zum Buschenfelde, and E. Marker-Hermann. Different cytokine profiles in the synovial fluid of patients with osteoarthritis, rheumatoid arthritis and seronegative spondylarthropathies. Clin. Exp. Rheumatol. 14(2):155–162, 1996.
Smith, J. B., M. H. Bocchieri, L. Sherbin-Allen, M. Borofsky, and J. L. Abruzzo. Occurrence of interleukin-1 in human synovial fluid: detection by RIA, bioassay and presence of bioassay-inhibiting factors. Rheumatol. Int. 9(2):53–58, 1989.
Stair-Nawy, S., A. B. Csoka, and R. Stern. Hyaluronidase expression in human skin fibroblasts. Biochem. Biophys. Res. Commun. 266(1):268–273, 1999.
Stern, R. Hyaluronan catabolism: a new metabolic pathway. Eur. J. Cell Biol. 83(7):317–325, 2004.
Stern, R. Hyaluronan metabolism: a major paradox in cancer biology. Pathol. Biol. (Paris) 53(7):372–382, 2005; (Epub 2005 Jan 19).
Strobl, B., C. Wechselberger, D. R. Beier, and G. Lepperdinger. Structural organization and chromosomal localization of Hyal2, a gene encoding a lysosomal hyaluronidase. Genomics 53(2):214–219, 1998.
Suzuki, T., N. Segami, M. Nishimura, and T. Nojima. Co-expression of interleukin-1beta and tumor necrosis factor alpha in synovial tissues and synovial fluids of temporomandibular joint with internal derangement: comparison with histological grading of synovial inflammation. J. Oral Pathol. Med. 31(9):549–557, 2002.
Tanimoto, K., S. Ohno, K. Fujimoto, K. Honda, C. Ijuin, N. Tanaka, T. Doi, M. Nakahara, and K. Tanne. Proinflammatory cytokines regulate the gene expression of hyaluronic acid synthetase in cultured rabbit synovial membrane cells. Connect. Tissue Res. 42(3):187–195, 2001.
Tanimoto, K., A. Suzuki, S. Ohno, K. Honda, N. Tanaka, T. Doi, K. Yoneno, M. Ohno-Nakahara, Y. Nakatani, M. Ueki, et al. Effects of TGF-beta on hyaluronan anabolism in fibroblasts derived from the synovial membrane of the rabbit temporomandibular joint. J. Dent. Res. 83(1):40–44, 2004.
Yoshida, M., S. Sai, K. Marumo, T. Tanaka, N. Itano, K. Kimata, and K. Fujii. Expression analysis of three isoforms of hyaluronan synthase and hyaluronidase in the synovium of knees in osteoarthritis and rheumatoid arthritis by quantitative real-time reverse transcriptase polymerase chain reaction. Arthritis Res. Ther. 6(6):514–520, 2004.
Acknowledgment
This research was supported by a Grant-in-Aid (No. 19659540) for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology, Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Associate Editor Ceng Dong oversaw the review of this article.
Rights and permissions
About this article
Cite this article
Tanimoto, K., Yanagida, T., Tanne, Y. et al. Modulation of Hyaluronan Fragmentation by Interleukin-1 Beta in Synovial Membrane Cells. Ann Biomed Eng 38, 1618–1625 (2010). https://doi.org/10.1007/s10439-010-9927-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10439-010-9927-3