Abstract
Genetic diversity is an important measure for the improvement of many crop species including Brassica. This study evaluated the genetic divergence among six Brassica species using simple sequence repeats (SSR). Ten SSR markers produced overall 21 alleles with an average of 2.1 alleles per primer. Out of 21, 18 alleles showed polymorphism (85.71%) and 3 alleles showed monomorphism (14.28%). A similarity matrix was constructed using Popgen32 software. Genetic identity ranged from 33.33 to 76.19%. B. nigra and B. campestris showed the highest identity, while the lowest identity was observed between B. campestris and B. oleracea. Sizes of amplified alleles were ranged from 70 to 290 bp. Polymorphic information content (PIC) varied from 0.37 to 0.71, with an average of 0.66 per primer. A dendrogram classified the genotypes into two main clusters. Cluster-A is further divided into cluster-C, which consists of B. carinata and B. oleracea. B. napus and B. juncea each form an independent cluster. Cluster-B consists of B. nigra and B. campestris, meaning that these two species are closely related to each other. The results indicated that these species can be isolated from each other at the molecular level by using molecular markers.
Zusammenfassung
Die genetische Diversität ist ein wichtiges Maß für die Verbesserung vieler Kulturpflanzen einschließlich Brassica. In der vorliegenden Studie wurde die genetische Divergenz zwischen 6 Brassica-Arten unter der Verwendung von Mikrosatelliten („simple sequence repeats“, SSR) untersucht. Dabei produzierten 10 SSR-Marker insgesamt 21 Allele mit einem Durchschnitt von 2,1 Allelen pro Primer. Von den 21 Allelen zeigten 18 Polymorphismus (85,71 %), und 3 Allele zeigten Monomorphismus (14,28 %). Mit der Popgen32-Software wurde eine Ähnlichkeitsmatrix erstellt. Die genetische Identität lag im Bereich von 33,33–76,19 %. Die höchste Identität wiesen B. nigra und B. campestris auf, während die niedrigste Identität zwischen B. campestris und B. oleracea beobachtet wurde. Die Größen der amplifizierten Allele lagen im Bereich von 70–290 bp. Der polymorphe Informationsgehalt (PIC) variierte von 0,37–0,71, mit einem Durchschnitt von 0,66 pro Primer. Ein Dendrogramm klassifizierte die Genotypen in 2 Hauptcluster. Cluster-A wird weiter unterteilt in Cluster-C, der aus B. carinata und B. oleracea besteht. Jeweils einen unabhängigen Cluster bilden B. napus und B. juncea. Cluster-B besteht aus B. nigra und B. campestris, was bedeutet, dass diese beiden Arten eng miteinander verwandt sind. Die Ergebnisse zeigten, dass diese Spezies auf molekularer Ebene unter Verwendung molekularer Marker voneinander isoliert werden können.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Introduction
The genus Brassica includes 37 varying species and has economic gains (Tsunoda et al. 1980). The family Brassicaceae is not only an important agricultural crop but also has commercial value. Controversy exists in numbering of basic chromosomes of Brassica, as their chromosome number X ranged from 7–11. Molecular research does not support Brassica species with chromosome number (X = 7) due to their multiple origins (Abbas et al. 2009). Brassica is commonly recognized as a flavoring crop, oilseed, and vegetable. In oil production worldwide, Brassica secures the second position, because its seeds have the potential of producing high quality oil. The raw materials have significant importance in production of biodiesel. In order to achieve future edible oil needs, Brassica breeding is essential to enhance the seed quality (Riaz et al. 2001). Different Brassica species, especially vegetables, play an important role in the human diet due to the presence of many compounds like vitamins, minerals, glucosinolates, and some phenolic compounds. Vegetables and some other plants of the Brassicaceae family are known as a major source of polyphenols (Harbaum et al. 2007). Molecular-based breeding of oilseed plants for superior quality and huge production are major sources of reduction in genomic variation in a few essential plants (Iqbal et al. 2015).
Molecular markers have developed a huge range of applications in the field of molecular biology that may include phylogenetic study, evolution, ecology, population genetics, and study of complicated genomic traits in both plants and animals. These different applications are achievable due to rapid advances in biological and molecular techniques (Morin et al. 2004). DNA-based markers contribute towards the utilization of useful genes in plant genetic engineering (Agrama and Tuinstra 2003). Different markers can be used to estimate genetic diversity, like amplified fragment length polymorphisms (AFLP), restriction fragment length polymorphisms (RFLP), single nucleotide polymorphisms (SNP), simple sequence repeats (SSR), and random amplified polymorphic DNA (RAPD; Pejic et al. 1998).
Different molecular markers have been used for the desired plant selection and to help plant breeders in selecting the plant having the desired traits directly or indirectly. DNA markers are recognized as useful markers for application in plant molecular breeding as they are usable, abundant, phenotypically unbiased, and time and stage independent (Hoshino et al. 2012). Microsatellites or SSR markers are the most popularly used PCR-based markers. These markers are successfully utilized across plant species for practical approaches as they are co-dominant, locus specific, hypervariable, multi-allelic, and robust. The genome-wide distribution of SSR markers makes them suitable markers for gene mapping and high-ranked genotyping, and helpful in marker-assisted breeding and gene mapping (Kong et al. 2000; Edwards et al. 2007; Murray et al. 2009). The objective of the current research was to estimate the genetic diversity and polymorphic information contents (PIC) among Brassica species using PCR-based SSR markers.
Materials and Methods
Experimental Site and Location
A screen house study entitled “Genetic diversity analysis of Brassica species using PCR based SSR markers” was conducted in the Centre of Agricultural Biochemistry and Biotechnology (CABB), University of Agriculture, Faisalabad, Pakistan. Faisalabad is located at a latitude of 31.421045 and longitude of 73.089294, having a semiarid environment.
Plant Material
The Brassica genotypes used in this research were provided by the University of Agriculture, Faisalabad (UAF). Six genotypes (Table 1) were sown in compost and sand (1:3) in the growth room of CABB, University of Agriculture, Faisalabad, Pakistan. Healthy and fresh leaves were taken and stored at −20 °C for good-quality DNA extraction.
DNA Extraction and Quantification
DNA was extracted from each collected sample using standard 2xCTAB extraction buffer as described by Ramos-Gómez et al. (2014). To confirm the DNA extraction, samples were run on 1% agarose gel and DNA quantification was measured by using Nanodrop ND-1000 (Thermo Scientific, Wilmington, DE, USA). DNA dilutions (25 ng/µl) were prepared for PCR reaction.
PCR-Based SSR Analysis and Agarose Gel Electrophoresis
SSR analysis was carried out using 10 primer pairs. A few important properties of these primers are displayed in Table 2. The PCR (Bio-Rad, California, USA) protocol was conducted in 25 μl of mixture using standard protocols reported by Tommasini et al. (2003). The PCR cycles consist of beginning denaturation at 94 °C for 5 min, followed by 35 cycles of denaturation at 94 °C for 1 min, annealing at 55 °C for 30 sec (based on primer), extension at 72 °C for 1 min, and a final extension at 72 °C for 10 min. SSR (PCR) amplification bands were analyzed by 3% (w/v) agarose gel electrophoresis at 120 voltage, and amplified bands were viewed on a gel documentation system (Bio-Rad GDS, California, USA).
Statistical Data Analysis
Bands scoring were done by using (1) for presence and (0) for absence of bands in order to obtain the genetic similarity matrix. Genetic similarities were estimated through Popgene32 software version 1.44 (Department of Renewable Resources, University of Alberta, Canada) (Yeh et al. 2002). The dendrogram was built based on the unweighted pair group method with arithmetic average (UPGMA) displaying genetic relationships between six genotypes. Polymorphism information content (PIC) for every marker was estimated using Power-marker software (Liu and Muse 2005).
Results
Level of Polymorphism
A set of ten SSR primers was used to evaluate the level of polymorphism among six Brassica species. Primers displayed a different level of polymorphism among all six genotypes. Overall, 21 alleles were generated by SSR markers with a mean of 2.1 alleles per primer. Out of ten, only three primers (BRMS-019, Na10-G10 and Na12-B09) gave both polymorphic and monomorphic bands while remaining seven showed only polymorphic band pattern. The highest number of alleles was produced by the two primers BRMS-019 and Na10-G10, while a lower number of alleles was produced by primer BRMS-027 (Table 3).
Polymorphic Information
For all SSR primers, the quantity of amplified alleles (QAA), quantity of polymorphic alleles (QPA), quantity of monomorphic alleles (QMA), percentage of polymorphic alleles (PPA), and polymorphism information content (PIC) were calculated. Out of the total, eighteen alleles showed polymorphism (85.71%) and three alleles showed monomorphism (14.28%). The percentage of polymorphic alleles was 88.33%. All primers produced specific, effective, and measurable alleles. The amplified alleles ranged from 70 to 290 bp. All primers displayed different levels of polymorphism among six Brassica species. Distinction capacities for every single genotype were determined using PIC values. PIC varied from 0.37 to 0.71, with a mean of 0.66 per marker. Nine SSR primers indicated PIC numbers more than 0.5, which means there is a huge genetic variation present among the six Brassica species (Table 3). PIC values indicated that primers were equally distributed in populations. Polymorphic information observed by SSR primers can be used for the production of hybrid genotypes.
Similarity Matrix and Cluster Analysis
Genetic similarity among and distance between the six Brassica species were estimated using Popgene32 software based on Nei’s UPGMA (Yeh et al. 2002). Genetic similarity ranged from 33.33 to 76.19%. The similarity matrix clearly showed that B. nigra and B. campestris were closely related species (76.19%), and that B. campestris and B. oleracea were the most dissimilar species (33.33%). Likewise, B. oleracea and B. carinata were 71.43% similar, and thus the second most similar species. Results of genetic similarity matrix and distance displayed that all six Brassica species are associated with each other (Table 4).
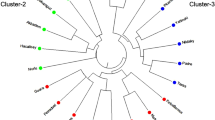
The dendrogram was constructed using Popgene32 software based on Nei’s UPGMA based on SSR data. The dendrogram clustered the genotypes into two major clusters, A and B.
Cluster-A is further divided into cluster-C which consists of B. carinata and B. oleracea. B. napus and B. juncea each form an independent cluster. Cluster-B consists of B. nigra and B. campestris, meaning that these two species are closely related to each other (Fig. 1).
Discussion
In the present study, ten SSR primers were used to estimate the genetic diversity between six Brassica species. Overall, 21 alleles were generated by SSR markers, with a mean of 2.1 alleles/primer. Another study investigated a mean of 2 alleles per primer using 7 SSR markers for 31 B. napus cultivars (Uzunova and Ecke 1999); 3.9 alleles per primer were reported by Varghese et al. (2000) for 31 rapeseed winter genotypes using 6 SSR markers; and Tonguç and Griffiths (2004) noticed 4.44 alleles per primer among 54 Brassica oleracea genotypes with the help of 13 SSR primer pairs. In other research, 4 loci per primer were measured for Brassica napus cultivars using morphological markers noticed by Yu et al. (2005). However, 8.7 to 9.8 bands were observed in Brassica carinata by Teklewold and Becker (2006). These variations in number of bands might be due to the procedure or material used in those studies. In the current research, the mean number of alleles per primer was 2.1, which is closely related to the average number of alleles reported by Hvarleva et al. (2007). In 2003, Tang and Knapp (2003) performed an experiment for 47 accession of native American land races and wild genotypes of cultivated sunflower using 122 SSR markers. These markers produced 1341 alleles between the 47 genotypes with an average of 12 alleles per primer, the average number of alleles between wild genotypes was three time higher (9.7) then native American land races (3.5). In another latest research, Raza et al. (2018) observed 227 bands by 20 RAPD markers with a mean of 11.35 bands per marker and showed 86.34% polymorphism among ten cultivated sunflower genotypes.
In another study, researchers used 12 SSR primer pairs to evaluate the diversity in 90 genotypes of interspecific crosses among B. napus and Brassica campestris. Total 33 allels were produced by 12 SSR primers with a mean of 2.66 alleles per primer. Out of these, 32 alleles showing 95.83% polymorphism were reported by Kanwal and Nawaz (2014).
A total of 38 SSR primers were used to estimate the genetic diversity among 28 sunflower cultivars and 102 alleles with a mean of 2.32 alleles/primer were observed. Genetic similarities varied from 0.25 to 0.90, while this study noticed similarities ranged from 33.33 to 76.19. PIC ranged from 0.09 to 0.62 with a mean of 0.41 as reported by Darvishzadeh et al. (2010).
However, a few researchers have described that genotypes cannot be characterized based on their origin. Some outcomes have been noticed in Ethiopian bananas by Birmeta et al. (2002), Ethiopian and Eritrean sorghum (Ayana et al. 2000), green mustard (Rabbani et al. 1998), Chinese vegetables Brassica (Ren et al. 1995), and rapeseed (Mailer et al. 1994). Important heredity differences were observed among Chinese and European cultivars of Brassica napus because of variation in genetic material as indicated by Shengwu et al. (2003).
In this research, the observed allele size ranged from 70 to 290 bp, and this outcome is quite close to the band size measured by Cartea et al. (2003) and Teklewold and Becker (2006). Out of the total (21), 18 alleles were polymorphic and showed 85.71% polymorphism among amplified alleles, and this high value of polymorphism indicates the huge genetic diversity among genotypes. However, primers with highly polymorphic alleles are more useful to evaluate the genetic distances and polymorphisms among genotypes investigated by Pradhan et al. (2004) and Sobotka et al. (2004). The present research observed 85.71% polymorphism, while Yu et al. (2005) observed 56% polymorphism in 11 genotypes of oilseed rape, and Teklewold and Becker (2006) observed 20–83% polymorphism among 43 Ethiopian mustard verities by using fifty RAPD markers.
PIC is the important feature for molecular research and the PIC value is a function of allele number and frequency. 9 out of 10 primers display PIC values higher than 0.5, which indicates the uniqueness of the SSR fingerprint. These values were comparable with the results assessed by Tonguç and Griffiths (2004), who used 13 SSR markers and obtained the same PIC values for 54 Brassica genotypes. In another study, PIC varied from 0.04 to 0.80 with a mean of 0.50 in Brassica napus, as observed by Plieske and Struss (2001). The PIC value in this study varied from 0.375 to 0.719, with a mean of 0.66. These results are very similar to Kanwal and Nawaz (2014), who noticed PIC varying from 0.395 to 0.726 with an average of 0.584 for each primer.
Conclusion
The conducted research outlines that genetic diversity and polymorphism exist among the six studied Brassica species. The allele number is directly and positively related to the genetic diversity and PIC values. The difference in PIC values with the reported values is due to the different genotypes and sample sizes. Results could be used to recognize different species, which could be utilized for future breeding programs aimed at creating genetic diversity in the local Brassica germplasm.
References
Abbas SJ, Farhatullah MK, Khan I, Munir I (2009) Molecular analysis of genetic diversity in Brassica species. Pak J Bot 41:167–176
Agrama H, Tuinstra M (2003) Phylogenetic diversity and relationships among sorghum accessions using SSRs and RAPDs. Afr J Biotechnol 2:334–340
Ayana A, Bryngelsson T, Bekele E (2000) Genetic variation of Ethiopian and Eritrean sorghum (Sorghum bicolor (L.) Moench) germplasm assessed by random amplified polymorphic DNA (RAPD). Genet Resour Crop Evol 47:471–482
Birmeta G, Nybom H, Bekele E (2002) RAPD analysis of genetic diversity among clones of the Ethiopian crop plant Ensete ventricosum. Euphytica 124:315–325
Cartea ME, Picoaga A, Soengas P, Ordás A (2003) Morphological characterization of kale populations from northwestern Spain. Euphytica 129:25–32
Darvishzadeh R, Azizi M, Hatami-Maleki H, Bernousi I, Mandoulakani BA, Jafari M, Sarrafi A (2010) Molecular characterization and similarity relationships among sunflower (Helianthus annuus L.) inbred lines using some mapped simple sequence repeats. Afr J Biotechnol 9:7280–7288
Edwards D, Forster JW, Chagné D, Batley J (2007) What Are SNPs? Assoc Mapp Plants. https://doi.org/10.1007/978-0-387-36011-9_3
Harbaum B, Hubbermann EM, Wolff C, Herges R, Zhu Z, Schwarz K (2007) Identification of Flavonoids and Hydroxycinnamic acids in Pak Choi varieties (brassica campestris L. ssp. chinensis var. communis) by HPLC–ESI-MS n and NMR and their Quantification by HPLC–DAD. J Agric Food Chem 55:8251–8260
Hoshino AA, Bravo JP, Morelli KA, Nobile PM (2012) Microsatellites as tools for genetic diversity analysis. In: Caliskan M (ed) Genetic diversity in microorganisms. Intech Open, London
Hvarleva T, Bakalova A, Chepinski I, Hristova-Cherbadji M, Hristov M, Atanasov A (2007) Characterization of Bulgarian sunflower cultivars and inbred lines with microsatellite markers. Biotechnol Biotechnol Equip 21:408–412
Iqbal S, Hamim I, Haque S, Nath UK (2015) Genetic diversity analysis of mustard (Brassica spp.) germplasm using molecular marker for selection of short duration genotypes. Afr J Biotechnol 14:1439–1448
Kanwal M, Nawaz I (2014) Genetic diversity analysis of Brassica napus/Brassica campestris progenies using microsatellite markers. Pak J Bot 46:779–787
Kong L, Dong J, Hart G (2000) Characteristics, linkage-map positions, and allelic differentiation of Sorghum bicolor (L.) Moench DNA simple-sequence repeats (SSRs). Theor Appl Genet 101:438–448
Liu K, Muse SV (2005) PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Mailer R, Scarth R, Fristensky B (1994) Discrimination among cultivars of rapeseed (Brassica napus L.) using DNA polymorphisms amplified from arbitrary primers. Theor Appl Genet 87:697–704
Morin PA, Luikart G, Wayne RK (2004) SNPs in ecology, evolution and conservation. Trends Ecol Evol (Amst) 19(4):208–216. https://doi.org/10.1016/j.tree.2004.01.009
Murray SC, Rooney WL, Hamblin MT, Mitchell SE, Kresovich S (2009) Sweet sorghum genetic diversity and association mapping for brix and height. Plant Genome 2:48–62
Pejic I, Ajmone-Marsan P, Morgante M, Kozumplick V, Castiglioni P, Taramino G, Motto M (1998) Comparative analysis of genetic similarity among maize inbred lines detected by RFLPs, RAPDs, SSRs, and AFLPs. Theor Appl Genet 97:1248–1255
Plieske J, Struss D (2001) Microsatellite markers for genome analysis in Brassica. I. Development in Brassica napus and abundance in Brassicaceae species. Theor Appl Genet 102:689–694
Pradhan A, Yan G, Plummer J (2004) Development of DNA fingerprinting keys for the identification of radish cultivars. Aust J Exp Agric 44:95–102
Rabbani MA, Iwabuchi A, Murakami Y, Suzuki T, Takayanagi K (1998) Genetic diversity in mustard (Brassica juncea L.) germplasm from Pakistan as determined by RAPDs. Euphytica 103:235–242
Ramos-Gómez S, Busto MD, Perez-Mateos M, Ortega N (2014) Development of a method to recovery and amplification DNA by real-time PCR from commercial vegetable oils. Food Chem 158:374–383
Raza A, Shaukat H, Ali Q, Habib M (2018) Assessment of RAPD markers to analyse the genetic diversity among sunflower (Helianthus annuus L.) genotypes. Turk J Agric Food Sci Technol 6:107–111
Ren J, McFerson JR, Li R, Kresovich S, Lamboy WF (1995) Identities and relationships among Chinese vegetable Brassicas as determined by random amplified polymorphic DNA markers. J Am Soc Hortic Sci 120:548–555
Riaz A, Li G, Quresh Z, Swati M, Quiros C (2001) Genetic diversity of oilseed Brassica napus inbred lines based on sequence-related amplified polymorphism and its relation to hybrid performance. Plant Breed 120:411–415
Shengwu H, Ovesna J, Kucera L, Kucera V, Vyvadilová M (2003) Evaluation of genetic diversity of Brassica napus germplasm from China and Europe assessed by RAPD markers. Plant Soil Environ 49:106–113
Sobotka R, Dolanska L, Curn V, Ovesná J (2004) Fluorescence-based AFLPs occur as the most suitable marker system for oilseed rape cultivar identification. J Appl Genet 45:161–174
Tang S, Knapp SJ (2003) Microsatellites uncover extraordinary diversity in native American land races and wild populations of cultivated sunflower. Theor Appl Genet 106:990–1003
Teklewold A, Becker HC (2006) Geographic pattern of genetic diversity among 43 Ethiopian mustard (Brassica carinata A. Braun) accessions as revealed by RAPD analysis. Genet Resour Crop Evol 53(6):1173–1185. https://doi.org/10.1007/s10722-005-2011-4
Tommasini L et al (2003) The development of multiplex simple sequence repeat (SSR) markers to complement distinctness, uniformity and stability testing of rape (Brassica napus L.) varieties. Theor Appl Genet 106:1091–1101
Tonguç M, Griffiths PD (2004) Genetic relationships of Brassica vegetables determined using database derived simple sequence repeats. Euphytica 137:193–201
Tsunoda S, Hinata K, Gómez-Campo C (1980) Brassica crops and wild allies. Biology and breeding. Bull Torrey Bot Club 108(3):388. https://doi.org/10.2307/2484725
Uzunova M, Ecke W (1999) Abundance, polymorphism and genetic mapping of microsatellites in oilseed rape (Brassica napus L.). Plant Breed 118:323–326
Varghese J, Rudolph B, Uzunova M, Ecke W (2000) Use of 5′-anchored primers for the enhanced recovery of specific microsatellite markers in Brassica napus L. Theor Appl Genet 101:115–119
Yeh F, Yang R, Boyle T, Ye Z, Mao J (2002) Popgen 32, microsoftware windows based freeware for population genetic analysis. Molecular Biology and Biotechnology Center, Edmonton
Yu C, Hu S, Zhao H, Guo A, Sun G (2005) Genetic distances revealed by morphological characters, isozymes, proteins and RAPD markers and their relationships with hybrid performance in oilseed rape (Brassica napus L.). Theor Appl Genet 110:511–518
Acknowledgements
We are very thankful to the members of Centre of Agricultural Biochemistry and Biotechnology (CABB), University of Agriculture, Faisalabad, Pakistan for their support and encouragement to conduct this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
A. Raza, S.S. Mehmood, F. Ashraf, and R.S.A. Khan declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Raza, A., Mehmood, S.S., Ashraf, F. et al. Genetic Diversity Analysis of Brassica Species Using PCR-Based SSR Markers. Gesunde Pflanzen 71, 1–7 (2019). https://doi.org/10.1007/s10343-018-0435-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10343-018-0435-y