Abstract
The weevil, Curculio elephas (Coleoptera, Curculionidae), is one of the most abundant and widespread chestnut feeding pests in Greece. Together with the tortricid moths Cydia splendana and C. fagiglandana (Lepidoptera, Tortricidae), C. elephas greatly affect chestnut production. This weevil occurs in a continuous range throughout Greece; however, its population structure is unknown. Information concerning its population structure could help in the future control of this pest. A 790 bp portion of the mitochondrial Cytochrome Oxidase I gene was sequenced for 160 C. elephas larvae from ten populations. Compared to other Curculio species haplotype diversity with 31 haplotypes and nucleotide diversity with six nucleotide substitutions was low. The Bayesian inference of phylogeography in concert with nested clade analysis revealed a loose geographic distribution of genetic diversity that was likely the joint effect of isolation by distance and gene flow. In addition, population indices and Bayesian-based calculations showed that the populations of C. elephas in Greece are at the phase of expansion. Dating the phylogeny suggests about 300,000 years before present as the start of divergence among the populations of C. elephas in Greece.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sweet chestnut, Castanea sativa Mill., is one of the most common trees of the Mediterranean flora and first acknowledged in human history between 10,600 to 8,900 years before present (Yasuda et al. 2000; Marinova et al. 2002). Since then, the interest in chestnut cultivation fluctuated with evidence of periods of intense use of chestnuts followed by periods of little use (Pite 1986; Santos et al. 2000; Conedera and Krebs 2008). Before the end of the last century, interest in chestnut cultivation increased. This is an economically promising industry and chestnut cultivation exhibits a potential that is not fully realized (Conedera and Krebs 2008; Diamandis 2008). The revitalization in chestnut cultivation is reflected in the increase of studies concerning C. sativa, which address the benefits of its wood and seeds as well as to its diseases and pests (Anagnostakis 1988; Paparati and Speranza 1999; Perlerou and Diamandis 2006). The biology, ethology and distribution of chestnut pests in Europe, and particularly those that feed in the nut have been extensively studied because they can greatly decrease chestnut production (Menu 1993; Menu and Debouzie 1995; Csóka 1997; Desouhant 1998; Speranza 1999; Hirka 2003; Avtzis 2012; Avtzis et al. 2012). The important weevil pest, Curculio elephas, was found to infest 35 % of chestnuts in Greece in 2012 (Avtzis et al. 2012).
Curculio elephas occurs in a continuous range throughout Greece including its islands (Avtzis et al. 2012). The interspecific relationships and movement of the weevil among locations is unknown. Phylogeographic results, including demographic, historical and population parameters, can be used to estimate the current status and future potential of a pest (Avise 1998). This information can be integrated into a pest management strategy. These results and conclusions rely on a robust estimate phylogeny which considers taxon sampling and phylogenetic methodology. The chestnut weevil has been included in a phylogenetic study that investigated the interspecific relationships among Curculio species, however, conclusions concerning their population genetics were not (Hughes and Vogler 2004).
This study endeavors to resolve the population structure of C. elephas in Greece, and the factors that contributed to the contemporary distribution of genetic diversity. We use Bayesian phylogenetic inference and other phylogenetic software to show that past geographic isolation and contemporary geneflow has influenced the population genetic structure of C. elephas in Greece.
Materials and methods
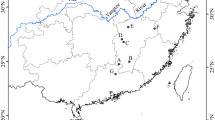
Chestnuts were manually opened and all living larvae were placed in individual vials with 100 % ethanol. We collected 160 C. elephas larvae from ten populations for this study (Table 1; Fig. 1). DNA from each larva was extracted using GenElute Kit (Sigma) and processed following the manufacturer’s protocol. Amplification of an ~800 bp locus from the 3′ end of mitochondrial Cytochrome Oxidase I (COI) gene was carried out with the primers Jerry and Pat (Simon et al. 1994) in 25 μl reactions containing 0.6 μl (3 u) of MyTaq™ (BioLine, GmBH, Germany), 5 μl of the provided 5× MyTaq™ red reaction buffer (BioLine, GmBH, Germany), 0.75 μl of each primer (20 μΜ), 8 μl of DNA extract and ddH2O to the volume of 25 μl. PCR conditions included an initial denaturation step at 94 °C for 3 min, followed by 40 cycles at 94 for 30 s, 45 for 30 s, and 72 °C for 1 min with a final extension step that lasted 5 min at 72 °C. PCR products were then purified with PureLink™ PCR Purification Kit (Invitrogen) and sequenced with an ABI 3730XL at CEMIA SA (Larissa, Greece) using both primers.
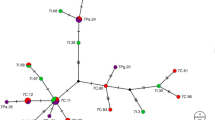
a Distribution of collected C. elephas populations in Greece. Circles indicate the phylogenetic clades and the remaining haplotype as shown in (b). b 50 % majority rule consensus tree derived from the Bayesian analysis of C. elephas mitochondrial haplotypes. Numbers above branches indicate Bayesian posterior probabilities (>0.85). c Nested Clade Analysis of C. elephas mitochondrial DNA haplotypes. Empty circles indicate missing haplotypes whereas the size of each parallelogram is proportional to the individuals it represents. Color of clade is common in all three figures. (Color figure online)
Data analysis
Sequences were visualized using Chromas v. 1.45 and then aligned using Clustal X (Thompson et al. 1997) with the default settings. After haplotypes were identified, and to exclude cases of base misincorporation as a result of PCR error (Kobayashi et al. 1999), those represented by only a single individual were verified by additional sequencing of an independent amplicon. The sequences of all C. elephas and C. glandium (outgroup) haplotypes were deposited in GenBank (Accession Nos: KC663584–KC663614), and referenced by haplotype designations provided below. Bayesian-based inference was performed with MrBayes version 3.1.1 (Ronquist and Huelsenbeck 2003) using the nucleotide substitution model F81 (Felsenstein 1981) as suggested by MrModeltest v2.1 (Nylander et al. 2004). However, prior fixed values for rates and nucleotide frequencies were not used as recommended (Ronquist et al. 2011). The sequence of C. glandium was used as outgroup in the above-mentioned analysis. The number of generations was initially set to 5,000,000 with a sampling frequency of 100 generations in a dual run each with four chains. After 1,500,000 generations, stationarity was achieved with the average standard deviation of split frequencies ranging between 0.00298 and 0.00191. Therefore, the last 8,500 trees of each run were used to compute the 50 % majority rule consensus tree and posterior probabilities. The frequency and spatial distribution of the mitochondrial DNA haplotypes were used to test the null hypothesis of random association of maternal lineages with geographic locations. A statistical parsimony network (Templeton et al. 1992) was created using TCS (Clement et al. 2000) and no ambiguous loops were detected. Haplotypes were then clustered into a hierarchical series of groups (Templeton et al. 1992) and the resulting nested network was statistically evaluated with GeoDis v. 2.0 (Posada et al. 2000).
Patterns of molecular diversity based on the mitochondrial DNA sequences between and within populations were assessed by estimating haplotype and nucleotide diversity (π) (Nei 1987) and the average number of nucleotide differences (Tajima 1983) using the software DNAsp version 5 (Rozas and Rozas 1999). Isolation by distance was evaluated by Mantel test (Mantel 1967), as this is inferred in the program Mantel non-parametric calculator version 2.0 (Liedloff 1999). Natural logarithms of geographical linear distances (km) between localities were correlated with the respective Tamura and Nei (1993) genetic distances, calculated with MEGA v.5 (Tamura et al. 2011), with 1,000 random iterations to obtain statistical inferences at α = 1 %. Sequence data were also examined for evidence of demographic expansion using two approaches: (i) mismatch distribution and (ii) Tajima’s D statistics (Tajima 1989). When at equilibrium, the distribution of pairwise differences is expected to yield a multimodal distribution, whereas sudden demographic expansion is commonly associated with unimodal distributions (Slatkin and Hudson 1991; Rogers and Harpending 1992). In addition, Tajima’s D statistics were used as a complementary approach to investigate the demographic history of the species. Both mismatch distribution among haplotypes (with 1,000 randomizations) and Tajima’s D were calculated using DNAsp version 5 (Rozas and Rozas 1999). In addition, the demographic history of C. elephas in Greece was assessed through a lineage through time (LTT) plot, as this is implemented in the Bayesian phylogenetic software Beast (Drummond and Rambaut 2007). Finally, an analysis of molecular variance (AMOVA) was used to examine the amount of genetic variability partitioned within and among populations as well as among groups of populations with Arlequin ver.3.0 (Excoffier et al. 2005). To elucidate the contribution of different factors, populations were clustered according to geographic distribution using a non-parametric approach (Excoffier et al. 1992). In addition, a spatial analysis of molecular variance was used to define the groups of populations using SAMOVA 1.0 (Dupanloup et al. 2002). This software seeks the composition of a user-defined number (K) of groups of geographic adjacent populations that maximize Φ CT. The program was run for 10,000 iterations for K = 2 to 9 from each of 100 random initial conditions.
Estimations of divergence times on the COI locus among the major phylogenetic clades of our analyses were carried out with the Bayesian inference method implemented in MDIV software (Nielsen and Wakeley 2001). Using a Markov Chain Monte Carlo simulation of the coalescence process in concert with the HKY model of sequence evolution to correct for multiple substitutions, we generated maximum likelihood estimators for the population parameter theta θ (θ = 2Ν e μ), the migration rate between populations since divergence M (M = 2Ν e m) as well as the time since divergence in mutational time T (T = t div/2Ν e ), with Ν e being the female haploid effective size, μ the per locus mutation rate, m the migration rate and t the number of years since divergence. Initially the method was tested for different values of M (1, 3, 5, 10 and 20) and T (5, 10, 15 and 20); only then did we set our prior values for M = 10 and T = 20, as they produced consistent posterior distributions. MDIV analyses were run for 5 × 106 generations with a burn-in period of 5 × 105, and repeated three times to insure convergence. The coalescent-scaled parameter T was converted to number of years since divergence t div according to the formula t div = T(θ)/2μ. For the calculation of μ, it was assumed one generation of C. elephas per year and a neutral rate of 2 % per million years (Papadopoulou et al. 2010), and thus for the 790 bp long locus μ was 7.9 × 10−6 substitutions/site/year.
Results
A total of 31 mitochondrial DNA COI haplotypes were found among 160 C. elephas individuals (Table 1; Fig. 1a). Haplotype diversity was 0.7041 (±0.0405), whereas nucleotide diversity (π) among all haplotypes was 0.001668 (±0.001157) (Table 2). Mean number of pairwise nucleotide differences among all individuals was 3.114 (±1.2), with a maximum number of six nucleotide differences. Phylogenetic analysis of these 31 haplotypes supported the monophyly of C. elephas when C. glandium was used as an outgroup (data not shown here). The majority of haplotypes occurred as an unresolved cluster and the remaining haplotypes were distributed among three major clades with substantial structure. These major clades (clade 1, 2 and 3) were supported with high (>0.85) posterior probability values (Fig. 1b). Even though clade 1 consisted of haplotypes that were found only in population “Melivia” (4) the other two clades showed substantial structure. Clade 2, contained two subclades (2-1 and 2-2) with geographically isolated haplotypes (haplotypes of subclade 2-1 were found only in population “Aghiasos” (5) and the ones of subclade 2-2 only in population “Halkida”(6)), whereas the other two haplotypes (Ele 19 and Ele 28) could be found in populations “Arnea” (10), “Halkida” (6), and “Western Crete” (7). Clade 3 had only one statistically supported deeper subclade (3-1) that was present in the populations “Paiko” (9), “Kastri” (8), and “Aghio Oros”(1) and one more haplotype (Ele 16) that was spread in populations “Aghio Oros” (1), “Western Crete” (7), and “Eastern Crete” (2).
The parsimony network did not contain any reticulations, indicating the absence of homoplasy among haplotypes. Evaluation of the total network (Fig. 1c) as inferred from GeoDis revealed that the current pattern of differentiation resulted from “restricted gene flow with isolation by distance.”. The pattern of isolation by distance was also tested using a Mantel test that rejected the null hypothesis of no association between genetic (Tamura–Nei) and geographic distances (g = 3.2357, p = 0.01). Analysis of molecular variance (AMOVA), however, did not show a strong genetic structure when populations were grouped according to various geographic combinations (data not shown) whereas SAMOVA showed that the value of ΦCT was larger when K = 2 indicating that partitioning into two groups was more appropriate. Several different approaches, including mismatch analysis (Fig. 2b), LTT (Fig. 2a) and Tajima’s D statistic (D = −2.25371 and p < 0.01) (Table 2), indicated a consistent population expansion of C. elephas.
Dating the divergence times between the major phylogenetic clades, suggested that clades began to diverge after the “Mindel” ice age that ended about 300.000 before present (BP) in the Alpine regions. By the start of the following ice age (Riss), the major clades of C. elephas in Greece began formed (Table 3). The average migration rate estimate was M = 0.173, suggesting that on average one female was exchanged between the clades about every six generations. However, this rate was significantly lower (M = 0.14) among the well-defined clades compared to the rates estimated between the smaller clades and the cluster of remaining haplotypes (M = 0.18–0.22) (Table 3).
Discussion
Our results suggest that divergence among the clades of C. elephas in Greece occurred during the Pleistocene approximately 200,000 years before present (Table 3). The broad distribution pattern of C. elephas in Greece, in concert with their recent phylogenetic divergence (only six nucleotides differed among the 790 mitochondrial DNA nucleotides) suggest that isolation by distance contributed to the occurrence of unique haplotypes/clades in populations peripheral to “Hortiatis” (3), “Melivia” (4), and “Arnea” (10). The outcome of the Nested Clade analysis and the Mantel tests support this observation and suggest that restricted gene flow with isolation by distance shaped the current pattern of intraspecific diversity of C. elephas.
Pleistocene environment likely influenced the genetic diversity of C. elephas in Greece. During Pleistocene, Greece experienced repeated periods of extreme cold with a roughly 100,000 years cycle interrupted by short warm interglacial periods (Messerli 1967), similar to the ones known in the Alps and northern Europe. The glacial sequences, Skamnellian, Vlasian, and Tymphian stages, in Greece (Sibrava et al. 1986; Tzedakis et al. 1997) have the equivalent names Mindel, Riss, and Würm respectively, which apply to the classic Alpine morphostratigraphical periods (Penck and Brückner 1909). Our results suggest that the main divergence events began during the Skamnellian stage and emerged in the subsequent interglacial stage (300000–190,000 years before present), before the Vlasian stage began. The origin of divergence in the Skamnellian stage seems plausible as this glacial stage gave rise to the most extensive glacial deposits (Hughes et al. 2003; Woodward et al. 2004) which separated the populations of C. elephas and initiated lineage divergence. The glaciers formed during these ice ages, and particularly during the Vlasian and Tymphian stages were of a lesser magnitude (Hughes et al. 2003) and likely allowed for movement of weevils among regions and subsequent gene flow.
Contemporary factors may have also influenced the observed distribution of haplotype diversity. A rough estimation of migration rates among the different clades indicates an average migration of one female every six generations. In the case of chestnut weevil, gene flow may have been increased due to human activities, due to the historical cultivation of chestnut (Conedera et al. 2004). Chestnuts have been commonly transported all over Greece and consequently, weevils were also transported to distant regions. The human-mediated transport of chestnut weevils has quite likely blurred the phylogeographic signal of C. elephas in Greece resulting in the current distribution of genetic diversity. A comparison between the haplotype diversity for C. elephas with that of other Curculio species supports further the action of a homogenizing factor among pest’s populations in Greece. Aoki et al. (2008) found 114 haplotypes among 204 C. hilgendorfi samples while Aoki et al. (2009) identified 41 haplotypes among the 115 C. sikkimensis individuals analyzed. Only Toju and Sota (2006) found comparable results, with 23 haplotypes among the 246 individuals of C. camelliae analyzed. Gene flow was suggested as a plausible explanation for the observed diversity (Toju and Sota 2006). Gene flow should have affected the observed distribution of haplotypes of C. elephas in Greece.
Curculio elephas populations are expanding (Table 2). This pest exhibits the potential to proliferate rapidly which underlines the urgent need for a strategy to restrict its movement. Chemical insecticides are commonly used to control C. elephas in Greece. The increased use of these chemicals throughout the potential range of C. elephas is not recommended because of their environmental impact. Species-specific biological agents for the control of C. elephas would minimize environmental impact. Entomopathogenic fungus and nematodes are potentially efficient control agents of chestnut weevil (Paparati and Speranza 1999; Kepenekci et al. 2004; Karagoz et al. 2009). The effectiveness of these entomopathogens was deduced from laboratory tests and limited field trials thus generalization of their effectiveness throughout the potential range of C. elephas is limited. Field experiments with entomopathogenic nematodes, fungus and other biological control agents are needed to test the efficiency of these agents throughout the potential range of C. elephas.
References
Anagnostakis SL (1988) Cryphonectria parasitica cause of chestnut blight. Adv Plant Pathol 6:123–136
Aoki K, Kato M, Murakami N (2008) Glacial bottleneck and postglacial recolonization of a seed parasitic weevil, Curculio hilgendorfi, inferred from mitochondrial DNA variation. Mol Ecol 17:3276–3289
Aoki K, Kato M, Murakami N (2009) Phylogeographical patterns of a generalist acorn weevil: insight into the biogeographical history of broadleaved deciduous and evergreen forests. BMC Evol Biol 9:103
Avise JC (1998) The history and purview of phylogeography: a personal reflection. Mol Ecol 7:371–379
Avtzis DN (2012) The distribution of Pammene fasciana L. (Lepidoptera, Tortricidae) in Greece: a present yet underestimated chestnut-feeding pest. Int J Pest Sci 58:115–119
Avtzis DN, Perlerou C, Diamandis S (2012) Geographic distribution of chestnut feeding insects in Greece. J Pest Sci. doi:10.1007/s10340-012-0451-0
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659
Conedera M, Krebs P (2008) History, present situation and perspective of chestnut cultivation in Europe. Acta Hortic 784:23–27
Conedera M, Manetti MC, Giudici F, Amorini E (2004) Distribution and economic potential of the sweet chestnut (Castanea sativa Mill.) in Europe. Ecol mediterr 30:179–193
Csóka G (1997) Increased insect damage in Hungarian forests under drought impact. Biologia 52:159–162
Desouhant E (1998) Selection of fruits for oviposition by chestnut weevil, Curculio elephas. Entomol Exp Appl 86:71–78
Diamandis S (2008) Sweet chestnut (Castanea sativa): a nut tree with great potential still to be exploited. Acta Hortic 784:37–42
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214
Dupanloup I, Schneider S, Excoffier L (2002) A simulated annealing approach to define the genetic structure of populations. Mol Ecol 12:2571–2581
Excoffier L, Smouse PE, Quattro M (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: applications to human mitochondrial DNA restriction data. Genetics 131:479–491
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Felsenstein J (1981) Evolution trees from gene frequencies and quantitative characters: finding maximum likelihood estimates. Evolution 35:1229–1242
Hirka A (2003) Investigations on carpophagous insects of oaks in Hungary. PhD Thesis, University of West Hungary, Sopron
Hughes J, Vogler AP (2004) The phylogeny of acorn weevils (genus Curculio) from mitochondrial and nuclear DNA sequences: the problem of incomplete data. Mol Phylogenet Evol 32:601–615
Hughes PD, Gibbard PL, Woodward JC (2003) Relict rock glaciers as indicators of Mediterranean palaeoclimate during the Last Glacial Maximum (Late Würmian) of northwest Greece. J Quat Sci 18:431–440
Karagoz M, Gulcu B, Hazir S, Kaya HK (2009) Laboratory evaluation of Turkish entomopathogenic nematodes for suppression of the chestnut pests, Curculio elephas (Coleoptera: Curculionidae) and Cydia splendana (Lepidoptera: Tortricidae). Biocontrol Sci Tech 19:755–768
Kepenekci I, Gokce A, Gaugler R (2004) Virulence of three species of entomopathogenic nematodes to chestnut weevil, Curculio elephas (Coleoptera: Curculionidae). Nematropica 34:199–204
Kobayashi N, Tamura K, Aotsuka T (1999) PCR error and molecular population genetics. Biochem Genet 37:317–321
Liedloff A (1999) MANTEL, version 2.0: the nonparametric test calculator. School of Natural Resource Sciences, QLD
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Marinova E, Tchakalova E, Stoyanova D, Grozeva S, Dotscheva E (2002) Ergebnisse archobotanischer Untersuchungen aus dem Neolithikum und Chalcolithikum in Südwestbulgarien. Archaeologia Bulgarica 6:1–11
Menu F (1993) Strategies of emergence in the chestnut weevil Curculio elephas (Coleoptera: Curculionidae). Oecologia 96:383–390
Menu F, Debouzie D (1995) Larval development variation and adult emergence in the chestnut weevil Curculio elephas Gyllenhal (Col., Curculionidae). J Appl Entomol 119:279–284
Messerli B (1967) Die eiszeitliche und die gegenwartige Vertgletscherung im Mittelemeeraum. Geograph Helv 22:105–228
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nielsen R, Wakeley J (2001) Distinguishing migration from isolation: a Markov Chain Monte Carlo approach. Genetics 158:885–896
Nylander J, Ronquist F, Huelsenbeck JP, Nieves-Aldrey JL (2004) Bayesian phylogenetic analysis of combined data. Syst Biol 53:47–67
Papadopoulou A, Anastasiou I, Vogler AP (2010) Revisiting the insect mitochondrial molecular clock: the mid-Aegean trench calibration. Mol Biol Evol 27:1659–1672
Paparati B, Speranza S (1999) Management of chestnut weevil (Curculio spp.) insect key-pests in central Italy. Acta Hortic 693:551–556
Penck A, and Brückner E (1909) Die Alpen im Eiszeitalter. Tauchnitz: Leipzig: 3, 1199
Perlerou C, Diamandis S (2006) Identification and geographic distribution of vc types of Cryphonectria parasitica and occurence of hypovirulence in Greece. Forest Pathol 36:413–421
Pite JR (1986) Terres de castanide. Homme et paysages du châtaignier de l’antiquité a nos jours, Paris
Posada D, Crandall KA, Templeton AR (2000) GeoDis: a program for the cladistic nested analysis of the geographical distribution of genetic haplotypes. Mol Ecol 9:487–488
Rogers AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol Evol 9:553–569
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Ronquist F, Huelsenbeck JP, Teslenko M (2011) Draft MrBayes version 3.2 Manual: Tutorials and Model Summaries. Distributed with the software from http://brahms.biology.rochester.edu/software.html
Rozas J, Rozas R (1999) DnaSP version 3: an integrated program for molecular population genetics and molecular evolution analysis. Bioinformatics 15:174–175
Santos L, Vidal RJR, Jalut G (2000) History of vegetation during the Holocene in the Courel and Queixa Sierras, Galicia, northwest Iberian Peninsula. J Quaternary Res 15:621–632
Sibrava V, Bowen DQ, Richmond GM (1986) Quaternary Glaciations in the Northern Hemisphere. Quaternary Science Reviews, 5
Simon C, Frati F, Beckenbach A, Crespi B, Liu H, Flook P (1994) Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved PCR primers. Ann Entomol Soc Am 87:651–701
Slatkin M, Hudson RR (1991) Pairwise comparisons of mitochondrial DNA sequences in stable and exponentially growing populations. Genetics 129:555–562
Speranza S (1999) Chestnut pests in central Italy. Acta Hortic 494:417–423
Tajima F (1983) Evolutionary relationship of DNA sequences in finite populations. Genetics 105:437–460
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Templeton AR, Crandall KA, Sing FC (1992) A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genetics 132:619–633
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Toju H, Sota T (2006) Phylogeography and the geographic cline in the armament of a seed-predatory weevil: effects of historical events vs. natural selection from the host plant. Mol Ecol 15:4161–4173
Tzedakis PC, Andrieu V, de Beaulieu JL, Crowhurst S, Follieri M, Hooghiemstra H, Magri D, Reille M, Sadori L, Shackleton NJ, Wijmstra TA (1997) Comparison of terrestrial and marine records of changing climate of the last 500,000 years. Earth Planet Sci Lett 150:171–176
Woodward JC, Macklin MG, Smith GR (2004) Pleistocene glaciation in the mountain of Greece. In: Ehlers J, Gibbard PL (eds) Quaternary glaciations—extent and chronology—part 1: Europe. Elsevier, Amsterdam, pp 155–173
Yasuda Y, Kitagawa H, Nakagawa T (2000) The earliest record of major anthropogenic deforestation in the Ghab Valley, northwest Syria: a palynological study. Quatern Int 73(74):127–136
Acknowledgments
The authors thank two anonymous reviewers as well as the subject editor for their comments that improved the quality of the manuscript. DNA also thanks G. Csóka and A. Hirka for their valuable support with the literature, S. Diamandis for setting the first ideas on this research plan, and N. Avtzis for his support and inspiration.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Stauffer.
Rights and permissions
About this article
Cite this article
Avtzis, D.N., Cognato, A.I. Genetic structure of Curculio elephas Gyll. (Coleoptera, Curculionidae) in Greece: an important pest of sweet chestnut. J Pest Sci 86, 491–497 (2013). https://doi.org/10.1007/s10340-013-0504-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10340-013-0504-z