Abstract
Microbial cells have extensively been utilized to produce value-added bioactive compounds. Based on advancement in protein engineering, DNA recombinant technology, genome engineering, and metabolic remodeling, the microbes can be re-engineered to produce industrially and medicinally important platform chemicals. The emergence of co-culture system which reduces the metabolic burden and allows parallel optimization of the engineered pathway in a modular fashion restricting the formation of undesired byproducts has become an alternative way to synthesize and produce bioactive compounds. In this study, we present genetically engineered E. coli-based co-culture system to the de novo synthesis of apigetrin (APG), an apigenin-7-O-β-d-glucopyranoside of apigenin. The culture system consists of an upstream module including 4-coumarate: CoA ligase (4CL), chalcone synthase, chalcone flavanone isomerase (CHS, CHI), and flavone synthase I (FNSI) to synthesize apigenin (API) from p-coumaric acid (PCA). Whereas, the downstream system contains a metabolizing module to enhance the production of UDP-glucose and expression of glycosyltransferase (PaGT3) to convert API into APG. To accomplish this improvement in titer, the initial inoculum ratio of strains for making the co-culture system, temperature, and media component was optimized. Following large-scale production, a yield of 38.5 µM (16.6 mg/L) of APG was achieved. In overall, this study provided an efficient tool to synthesize bioactive compounds in microbial cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Metabolic engineering by remodeling metabolic system of the cultured organisms has become a critically important tool to synthesize bioactive secondary metabolites. There are indeed more substantial numbers of biofuels, drugs, and industrially essential chemicals being produced by metabolomics-modified organisms [2, 4, 25, 29, 41, 51]. Monoculture systems have generated most of these bioactive compounds under optimal conditions. Although monoculture system is an efficient way to produce platform chemicals, it suffers from several disadvantages including lack of an optimal environment for the functioning of all pathway-specific enzymes, the increment in metabolic burden due to reconstruction or heterologous expression of complex pathways, and the formation of undesired byproducts [56, 58].
Several secondary metabolites with high industrial and medicinal values are produced by microorganisms and plants [53, 54]. Among them, flavonoids are a class of ubiquitous metabolites found in different plant species with profound bioactivity. Most of them exist in their glycosylated form with more stability, bioactivity, and solubility. However, aglycones are also present in large quantities [21]. Among various flavonoid reported, apigenin (API) and its derivative have been proven to be a well-known anti-aging, anti-fungal, anti-tumor, and anti-inflammatory agents [27, 28, 44]. Moreover, its derivative apigenin-7-O-β-d-glucopyranoside, also known as apigetrin (APG), has shown additional anti-proliferative and anti-oxidant activity against reactive oxygen species (ROS) [36, 44]. Besides glycosylation, hydroxylation, malonylation, methylation, and sulfation are common modifications that diversify flavonoids. Therefore, there is an increasing demand for these molecules in clinics and food-processing practices. Since lower solubility in water, short retention time in the intestine, and lower absorption rate restrict the consumption of flavonoids, the development of water-soluble flavonoids has been implicated for the treatment of many medical problems [15]. Glycosylation is one of the ways to enhance water solubility, the stability, and pharmacokinetic properties of flavonoids [21, 22, 46]. The biological activities of most natural products are attributed to the sugar moieties attached to them [11]. While chemical synthesis is used to synthesize such derivatives, the use of site-specific glycosyltransferases (GTs) that catalyzes the formation of the glycosidic bond are often used for the in vitro and in vivo synthesis of glycosides [5, 6, 8, 14, 47, 49]. Due to the development of chemical and molecular biology techniques, APG has been synthesized by chemical, semi-, and biosynthetic approaches [26, 32, 38, 50]. Among them, most biological approaches for APG production were carried out using plant-originated genes including TAL, 4CL, CHS, CHI, FNSI (or FNSII), and GTs via reconstruction in recombinant plasmids and hosts such as E. coli, Bacillus, and Saccharomyces cerevisiae to construct a monoculture system [13, 19, 23, 35].
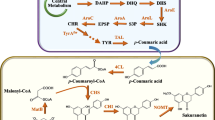
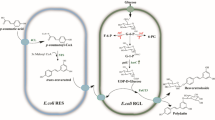
Recently, the co-culture system simultaneously culturing two distinct metabolically engineered strains for production of desired metabolites has become an exciting and alternative tool. Numerous studies have reported the use of E. coli, Bacillus, and S. cerevisiae as a co-culture system to produce plenty of known bioactive compounds such as flavonoids, alkaloids, terpenoids, etc. [7, 10, 18, 30, 55,56,57]. Therefore, in the present study, we described the use of a co-culture system by genetically engineered E. coli for the production of APG. To achieve this endeavor, the culture system was compartmentalized into two parts: upstream strain containing biosynthetic pathway for production of API and downstream strain for overproduction of UDP-glucose and heterologous expression of a glycosyltransferase (Fig. 1).
Materials and methods
Culture media and chemicals
LB (Luria–Bertani) and M9 minimal medium (per liter, 6 g Na2HPO4, 3 g K2HPO4, 0.5 g NaCl, 0.1% NH4CL, 1% glucose, 1 mM MgSO4·7H2O, 100 µM CaCl2, 1.46 mg biotin, 1 mg thiamine-HCl) containing 2% glucose were used for seed culture and substrate fermentation, respectively. Antibiotics including ampicillin, kanamycin, and chloramphenicol (Biobasics, Canada) were used in the final concentration of 100, 50 and 30 µg/mL, respectively. Ethyl acetate, methanol, acetonitrile, and dimethyl sulfoxide (DMSO) were purchased from Merck (Germany) or Sigma (USA). Nanodrop 2000 UV–Vis Spectrophotometer (Thermo, USA) and HPLC–PDA (Agilent, USA) were used for chromatographic analysis.
DNA manipulation, bacterial transformation, recombinant plasmid reconstruction
DNA manipulation such as DNA plasmid extraction, purification, digestion, and ligation followed standard protocols as described by [43]. pACYC-Deut-1 containing chalcone synthase (CHS) and chalcone flavanone isomerase(CHI) from Petunia hybrida was previously described [24, 31]. 4CL-2 [4-coumarate: CoA ligase (4CL2) from Nicotiana tabacum cv. Samsun] gene was digested from the pAC-4CL-STS plasmid by NcoI/NotI restriction enzyme and then transferred to the multicloning site 1 (MCS1) of pCDFDuet-1 bearing streptomycin resistance gene (Str) [3]. Similarly, flavone synthase I (FNSI) from parsley Petroselinum crispum cv was digested with NdeI/XhoI from pET15b [33] and cloned into MCS2 of this duet vector. Those vectors were then introduced into E. coli BL21(DE3) to construct the recombinant E. coli API to produce API. In another way, E. coli MA3 strain, previously constructed for enhancing production of UDP-glucose [40, 48], was transformed with the recombinant pQE-30 vector containing glycosyltransferase-encoded gene (PaGT3) [37, 39] to generate E. coli GLP. All strains and DNA plasmids were listed in Table 1.
Use of E. coli GLP strain for the synthesis of apigetrin
Escherichia coli GLP strain was cultured in 3 mL of LB medium, 220 rpm, and 37 °C for overnight. Then, 500 µL of this seed culture was transferred to 250 mL flask containing 50 mL minimal M9 medium plus 2% glucose and maintained at 37 °C, 220 rpm until its optical density (OD) at 600 nm achieved 0.6. Total 1 mM IPTG was then added and continuously cultured under the same condition for 3–4 h for induction of protein. Total 100 µM purified API was supplemented to the induced broth and continuously maintained until 60 h. Bioconversion of substrate was checked at 12 h of interval time.
Synthesis of apigetrin and apigenin from p-coumaric acid
Escherichia coli API was used to produce API by feeding various concentration of p-coumaric acid (PCA). Similarly, minimal M9 plus 2% glucose was used for culture with 37 °C and 220 rpm. To increase the bioconversion of substrate, PCA was supplemented gradually into the induced E. coli API culture broth to avoid its growth inhibition by excessive substrate, i.e., 20, 20, 30, and 30 µM of PCA. Moreover, the sample was taken out at 12-h intervals for production testing.
In another way, the E. coli MA2 strain [40, 48] was transformed with various recombinant vectors (pAC-4CL-FNSI, pAC-CHS_CHI, and pQE30-PaGT3) to generate E. coli APG strain and used for the synthesis of APG. PCA was also gradually supplemented to culture to study the dependence of substrate bioconversion upon a time. Mainly 20, 20, 30, and 30 µM PCA was gradually added at 12, 24, 36 and 48 h, respectively, and the broth culture was maintained until 60 h.
Experimental co-culture for production of apigetrin
Effect of initial inoculum ratio on the production of apigetrin
Escherichia coli API and GLP were separately cultured by 3 mL of LB broth medium containing appropriate antibiotics at 37 °C and 220 rpm for overnight. Five hundred µL (unless stated) of each strain was subsequently diluted into 50 mL of M9 minimal media supplementing necessary antibiotics in 250 mL flasks; then the flasks were incubated at 37 °C and 220 rpm till the OD600nm of 0.6. The culture broths were centrifuged, and the cell pellets were resuspended in 50 mL of M9 media plus 2% glucose. The two populations were mixed in different ratios by OD, i.e., various ratios of API/GLP = 1: 1; 2:1; 4:1; 6:1; 8:1 and 10:1 (v/v) to generate a co-culture system. The broth culture was added with 1 mM IPGT and continuously cultured for 3–4 h at 37 °C, and 220 rpm to induce protein expression. Subsequently, the system was supplemented by total 100 µM PCA, i.e., 20, 20, 30, 30 µM at 12, 24, 36, and 48 h. Formation of NRN, API, and APG was investigated at interval time of 12 h until 60 h of culture. Similarly, inverse various ratios (in which API was kept in constant, but GLP was varied) of API/GLP, i.e. 1:2, 1:4, 1:6, 1:8, and 1:10 were also used for evaluating the production of APG under the same conditions and substrate feeding as described above.
Effect of culture temperature
Various range of culture temperature from 25 to 37 °C, i.e., 25, 27, 30, 32, and 37 °C was used to investigate the effect on the production of API and its derivatives. The total PCA concentration of 100 µM was supplemented to the culture under 220 rpm. The converted naringenin, API, and APG concentration were checked after interval time of 12 h until 60 h of culture.
Analysis and quantification
HPLC–DAD analysis was performed by injecting 20 µL of the samples on an Agilent 1260 HPLC system equipped with a photodiode array detector (DAD), degasser, an autosampler. An Agilent Zorbax SB C18 column (250 mm × 4.6 mm i.d., 5 μm; Agilent, Santa Clara, CA, USA) was used. The mobile phase of 0.1% trifluoroacetic acid (TFA) aqueous solution (solvent A) and acetonitrile (solvent B) were used with 1 mL/min flow rate. The concentration of acetonitrile during the binary gradient condition was as: 0–10 min, 0–60%; 10–20 min, 75%; 20–35 min, 75–90%. Peak detection was carried out at UV absorbance at 295 nm whereas.
Naringenin, API, and APG were purified using an MPLC instrument equipped the silica gel RP-packed column (YMC gel ODS-A, AA12SA5, Japan). The corresponding peaks were collected and then lyophilized by freeze drier (FDU-1200, Eyela, Japan). All chemicals were dissolved in DMSO-d6 and subjected to NMR analysis. A series of concentrations ranging from 10 to 100 mg/L of the product were prepared to construct a calibration curve. The molecular mass of the compounds was determined in LC–ESI–MS using Phenomenex Synergi Polar-RP column (150 × 4.6 mm, 4 µm), positive-ion mode.
Statistical analysis
The student’s t test was performed on the biological replicates to determine the statistical significance of the difference between control and experiment samples at each time point. Differences with a P value < 0.05 were considered statistically significant.
Results
Production of apigenin from p-coumaric acid using E. coli API strain
Since glucose increases the intracellular acetyl-CoA concentration via glycolysis and acts as the primary carbon source for flavonoid production [45], instead of LB, M9 minimal media with 2% glucose was used to restrict the rapid degradation of PCA [31]. The recombinant E. coli harboring biosynthetic gene cluster for API production was constructed using 4-coumarate: CoA ligase (4CL2) gene from Nicotiana tabacum cv. Samsun [3], chalcone synthase (CHS) and chalcone flavanone isomerase(CHI) from Petunia hybrida and Medicago sativa, respectively, as described early, and flavone synthase I (FNSI) from parsley Petroselinum crispum cv. [33]. When cultured at 37 °C, 220 rpm, and fed with 20 µM of PCA, the formation of naringenin (NRN) and API were observed by HPLC analysis (Fig. 2b). Further analysis by LC–ESI–MS resulted in the mass – charge ratio (m/z) of [M + H]+ of 272.9 and 270.8 for NRN and API in positive mode, respectively (Fig. 3a, b). Also, the time and concentration-dependent study showed highest concentration of API (31.8 µM) at 60 h, when 100 µM PCA was fed entirely (Fig. 4). Moreover, the yield of conversion was not improved when a higher concentration of PCA was supplied into the broth culture.
HPLC trace analysis of the biotransformation products of recombinant E. coli recombinant strains. a authentic PCA and API standard compounds, b E. coli API supplemented with the substrate PCA for production of API, c synthesis of APG by E. coli APG strain utilizing PCA as initial substrate, d E. coli GLP used for bioconversion of API to APG, and e co-culture of API and GLP (ratio of 1:2) for production of APG. triangle, PCA peak; star, APG peak; diamond, API peak; and 4 pointed star shape, naringenin peak
Synthesis of apigetrin using E. coli APG strain using PCA as initial substrate
To avoid growth inhibition of E. coli by extra-cellular addition of PCA [51], 10 µM PCA was initially supplemented to the induce broth culture. Post substrate addition after 5 h, the production of NRN, API, and APG were analyzed. As evident from Fig. 3c, we observed the presence of APG peak with retention time of 20.50 min in HPLC chromatogram (Fig. 2c) with m/z of [APG + H] + = 433.0. Further analysis of bioconversion at 12 h interval showed 77.7% PCA to be consumed, while the maximum yield of APG, NRN, and API reached 15.5, 32.8, and 27.4 µM after 60-h culture, respectively (Fig. 5).
Biotransformation of API to APG by E. coli GLP
To compare the yield of APG production, 10 µM API was initially supplemented to induced culture broth, which gradually increased up to 100 µM in total at 48 h maintaining till 60 h. As evident from HPLC chromatogram monitored at the 12-h interval, the maximum accumulation of APG reached to 45.8 µM (19.8 mg/L) after 60 h (Figs. 2d, 6), where the rate of biotransformation of API to APG was higher than a synthesis of APG from PCA.
De novo co-culture for production of APG
Optimization of inoculum ratio in co-culture
Co-culture for the production of natural products requires testing proper ratio of initial inoculum because each strain was designed to carry specific work. Mainly, E. coli API and GLP were stepwise reconstructed to synthesis API and APG.
By changing the ratio of API/GLP from 1:1 to 10:1, the production of APG concentration was achieved to maximum, yielding of 32.5 µM (14.1 mg/L) or 2.1-fold compared to the use of mono-culture E. coli APG (Figs. 2e, 7). Hence, E. coli GLP expressed the strong bioactivity of PaGT3 again as proved by biotransformation of API to APG too. Moreover, the higher the ratio of E. coli API the more production of APG. Concurrently, the more production of API the more consumption of initial substrate, PCA.
In contrast, the ratio of API/GLP was changed from 1/2 to 1/10 the maximum concentration of APG has achieved the highest yield of 23.4 µM at a ratio of 1/10 and gradually decreased to 15.6 µM at the ratio of 1/2. Therefore, it was observed that higher APG concentration in accompanies with a higher ratio of GLP/API. However, the ratio of E. coli API was constant then the maximum yield of APG just achieved 72% (23.4/32.5 × 100) compared to the previous case. Based on above result analysis the ratio of API/GLP was selected between 8 and 10 for further experiments.
Effect of temperature on co-culture for production of APG
Temperature is one of the most important factors that affect bacterial growth and heterologous expression of enzymes. It has been proven that 35–37 °C is a suitable range for E. coli culture. However, glycosyltransferases are optimally expressed at a lower temperature. Thereby, various range of temperature was tested for co-culture to obtain higher production of APG. As a result, API was strongly synthesized, PCA concentration was fast consumed resulting in the highest yield of APG of 38.5 µM (16.6 mg/L) (Fig. 8) when the range of temperature maintained between 25 and 32 °C. However, the production of API was not increased when the temperature reached 37 °C with a maximal yield of APG was 32.5 µM.
Structural confirmation of NRN, API, and APG
The structures of API, APG, and NRN were confirmed by 1D and 2D NMR. We observed two meta-coupled [δH 6.19 (H-6) and 6.48 (H-8), each 1H, d, J = 2.0 Hz] and four ortho-coupled [δH 7.92 (H-2′ and H-6′) and 6.92 (H-3′ and H-5′), each 2H, d, J = 9.0 Hz] aromatic protons of API in 1H NMR spectrum. Also, the signals of one olefinic [δH 6.76 (1H, s, H-2)] and one chelated hydroxyl [δH 12.95 (1H, s, 5-OH)] protons were observed. The 13C NMR spectrum of API exhibited signals of 15 carbons for a flavone skeleton (Table 2). Comparison of the 1H and 13C NMR data of API with the reported values [1, 49] as well as detailed analysis of its HMBC correlations (Fig. 9) elucidated its structure as 5,7,4′-trihydroxyflavone. Similarly, the other compounds were characterized as APG (apigenin-7-O-β-d-glucoside) [12, 49] and 2S-5,7,4′-trihydroxyflavanone (naringenin) [52], which is in good agreement with their 1H and 13C NMR chromatograms, earlier report, and combination with their HMBC evidence (Figs. S2, S3, and S4, supporting information).
Discussion
It is observed that flavonoids are stable in the form of glycosides (O-, C- or N-glycosidic linked derivatives) in vascular plants. Till today, synthesis of flavonoid glycosides was carried out by a chemical approach with several disadvantages such as protection of un-reacted moiety, generation of various derivatives under severe conditions [50]. However, glycosyltransferase acts as a biocatalyst to catalyze the attachment of glycosyl moiety to various flavonoid acceptor in vitro and in vivo. Microbial synthesis of glycosides by mono-culture requires the construction of recombinant plasmids with numbers of genes, inhibition of bacterial chemo-physiological process, and challenge to achieve optimal culture conditions. For example, biological synthesis of flavanones (naringenin, pinocembrin) by recombinant mono-culture E. coli has been carried out by [16, 19, 24, 35]. Also, flavones (API, chrysin) have been synthesized by FNSI [34] including its derivative, genkwanin [23]. Therefore, as an alternative approach, co-culture system was designed in this study to fix the bottlenecks of monoculture. For example, resveratrol, afzelechin, muconic acid, and 3-aminobenzoic acid have been successfully produced by co-culture approach [9, 18, 55, 57]. Here, we have experimentally biosynthesized API and APG from PCA by monoculture system, where the maximal yield of each metabolite was 31.8 (8.6 mg/L) and 15.5 µM (6.7 mg/L), respectively. Besides, API substrate was also used for direct biotransformation by E. coli GLP and resulted in 45.8 µM of APG (19.8 mg/L). Furthermore, co-culture based on the modular system including upstream (E. coli API) and downstream (E. coli GLP) was optimized by the ratio of inoculum and temperature condition. By changing the various ratio of inoculum API/GLP, it resulted in the maximal yield of APG of 32.5 µM. Furthermore, the best yield of APG was achieved (38.5 µM or 16.6 mg/L) under optimal temperature (32 °C). Thereby, it meant increment in 2.5-fold of APG compared to mono-culture E. coli APG (16.6 mg/L), however, still lower than direct biotransformation of API by E. coli GLP (19.8 mg/L). For reference, the yield of API was mentioned in different records such as 30 mg/L [23] or 13 mg/L [34] via mono-culture. In comparison to other literature, the titer of afzelechin and resveratrol was achieved 40.7 ± 0.1, and 22.6 mg/L by E. coli co-culture; our yield of APG was in comparative level.
To further increase the titer of APG, several works need to be continuously carried out including strain improvement of E. coli API such as over-expression of malonyl-CoA [25, 34] or biosynthesis pathway of naringenin from the direct amino acid by the introduction of tyrosine ammonium lyase (TAL) or phenylalanine ammonia-lyase (PAL) [34, 35]. Similarly, improvement of E. coli GLP can be achieved by strengthening of PaGT3 via site-directed mutagenesis [20, 42]. Finally, it is necessary to optimize the culture conditions for co-culture system [17, 18].
Conclusions
Bioactive compounds are becoming essential in clinical practices, production of cosmetics, adjuvant, antibiotics as well as functional foods. Therefore, biosynthesis of bioactive compounds has evolved from simple to sophisticated approaches. Combination of genetic engineering, recombinant protein, metabolic engineering, and fermentation technology creates a novel ways for biosynthesis of these valuables including co-culture system. Our observations showed co-culture as an applicable and reproductive approach, which can be used to solve the bottlenecks of monoculture, yielding higher production of APG under simple conditions.
References
Alwahsh MA, Khairuddean M, Chong WK (2015) Chemical constituents and antioxidant activity of Teucrium barbeyanum Aschers. Rec Nat Prod 9:159–163
Atsumi S, Cann AF, Connor MR et al (2008) Metabolic engineering of Escherichia coli for 1-butanol production. Metab Eng 10:305–311. https://doi.org/10.1016/j.ymben.2007.08.003
Beekwilder J, Wolswinkel R, Jonker H et al (2006) Production of resveratrol in recombinant microorganisms. Appl Environ Microbiol 72:5670–5672. https://doi.org/10.1128/AEM.00609-06
Bhan N, Xu P, Koffas MAG (2013) Pathway and protein engineering approaches to produce novel and commodity small molecules. Curr Opin Biotechnol 24:1137–1143. https://doi.org/10.1016/j.copbio.2013.02.019
Brazier-Hicks M, Edwards R (2013) Metabolic engineering of the flavone-C-glycoside pathway using polyprotein technology. Metab Eng 16:11–20. https://doi.org/10.1016/j.ymben.2012.11.004
Brazier-Hicks M, Evans KM, Gershater MC et al (2009) The C-glycosylation of flavonoids in cereals. J Biol Chem 284:17926–17934. https://doi.org/10.1074/jbc.M109.009258
Brenner K, You L, Arnold FH (2008) Engineering microbial consortia: a new frontier in synthetic biology. Trends Biotechnol 26:483–489. https://doi.org/10.1016/j.tibtech.2008.05.004
Buettner FFR, Ashikov A, Tiemann B et al (2013) C. elegans DPY-19 is a C-mannosyltransferase glycosylating thrombospondin repeats. Mol Cell 50:295–302. https://doi.org/10.1016/j.molcel.2013.03.003
Camacho-Zaragoza JM, Hernández-Chávez G, Moreno-Avitia F et al (2016) Engineering of a microbial coculture of Escherichia coli strains for the biosynthesis of resveratrol. Microb Cell Fact 15:163. https://doi.org/10.1186/s12934-016-0562-z
Chanos P, Mygind T (2016) Co-culture-inducible bacteriocin production in lactic acid bacteria. Appl Microbiol Biotechnol 100:4297–4308. https://doi.org/10.1007/s00253-016-7486-8
Chaudhary AK, Dhakal D, Sohng JK (2013) An insight into the “-omics” based engineering of Streptomycetes for secondary metabolite overproduction. Biomed Res Int 2013:968518. https://doi.org/10.1155/2013/968518
Giang PM, Son PT (2004) Apigenin 7-O-β-d-glucoside from the leaves of Acanthus integrifolius T. Anders., Acanthaceae. J Chem 42:496–498
Gurung RB, Kim EH, Oh TJ, Sohng JK (2013) Enzymatic synthesis of apigenin glucosides by glucosyltransferase (YjiC) from Bacillus licheniformis DSM 13. Mol Cells 36:355–361. https://doi.org/10.1007/s10059-013-0164-0
Hamilton ML, Caulfield JC, Pickett JA, Hooper AM (2009) C-Glucosylflavonoid biosynthesis from 2-hydroxynaringenin by Desmodium uncinatum (Jacq.) (Fabaceae). Tetrahedron Lett 50:5656–5659. https://doi.org/10.1016/j.tetlet.2009.07.118
Havsteen BH (2002) The biochemistry and medical significance of the flavonoids. Pharmacol Ther 96(2–3):67–202
Hwang EI, Kaneko M, Ohnishi Y, Horinouchi S (2003) Production of plant-specific flavanones by Escherichia coli containing an artificial gene cluster. Appl Environ Microbiol 69:2699–2706. https://doi.org/10.1128/aem.69.5.2699
Jones JA, Koffas MAG (2016) Optimizing metabolic pathways for the improved production of natural products. In: O’Connor SE (ed) Synthetic biology and metabolic engineering in plants and microbes part a: metabolism in microbes, 1st edn. Elsevier Inc, pp 179–193. https://doi.org/10.1016/bs.mie.2016.02.010
Jones JA, Vernacchio VR, Sinkoe AL et al (2016) Experimental and computational optimization of an Escherichia coli co-culture for the efficient production of flavonoids. Metab Eng 35:55–63. https://doi.org/10.1016/j.ymben.2016.01.006
Kaneko M, Il Hwang E, Ohnishi Y, Horinouchi S (2003) Heterologous production of flavanones in Escherichia coli: potential for combinatorial biosynthesis of flavonoids in bacteria. J Ind Microbiol Biotechnol 30:456–461. https://doi.org/10.1007/s10295-003-0061-1
Kim HL, Kim AH, Park MB et al (2013) Altered sugar donor specificity and catalytic activity of pteridine glycosyltransferases by domain swapping or site-directed mutagenesis. BMB reports 46:37–40
Kren V, Martínková L (2001) Glycosides in medicine: “The role of glycosidic residue in biological activity”. Curr Med Chem 8:1303–1328. https://doi.org/10.2174/0929867013372193
Langenhan JM, Peters NR, Guzei IA et al (2005) Enhancing the anticancer properties of cardiac glycosides by neoglycorandomization. Proc Natl Acad Sci USA 102:12305–12310. https://doi.org/10.1073/pnas.0503270102
Lee H, Kim BG, Kim M, Ahn JH (2015) Biosynthesis of two flavones, apigenin and genkwanin, in Escherichia coli. J Microbiol Biotechnol 25:1442–1448. https://doi.org/10.4014/jmb.1503.03011
Leonard E, Lim KH, Saw PN, Koffas MAG (2007) Engineering central metabolic pathways for high-level flavonoid production in Escherichia coli. Appl Environ Microbiol 73:3877–3886. https://doi.org/10.1128/AEM.00200-07
Leonard E, Yan Y, Fowler ZL et al (2008) Strain improvement of recombinant Escherichia coli for efficient production of plant flavonoids. Mol Pharm 5:257–265. https://doi.org/10.1021/mp7001472
Leonard E, Yan Y, Lim KH, Koffas MAG (2005) Investigation of two distinct flavone synthases for plant-specific flavone biosynthesis in Saccharomyces cerevisiae. Appl Environ Microbiol 71:8241–8248. https://doi.org/10.1128/AEM.71.12.8241-8248.2005
Lim HS, Kim OS, Kim BY, Jeong SJ (2016) Apigetrin from Scutellaria baicalensis Georgi inhibits neuroinflammation in BV-2 microglia and exerts neuroprotective effect in HT22 hippocampal cells. J Med Food 19:1032–1040. https://doi.org/10.1089/jmf.2016.0074
Lin XH, Pan JB, Zhang XJ (2017) Anti-inflammatory and anti-oxidant effects of apigetrin on LPS-induced acute lung injury by regulating Nrf2 and AMPK pathways. Biochem Biophys Res Commun. https://doi.org/10.1016/j.bbrc.2017.07.071
Liu T, Khosla C (2010) Genetic engineering of Escherichia coli for biofuel production. Annu Rev Genet 44:53–69. https://doi.org/10.1146/annurev-genet-102209-163440
Magdouli S, Brar SK, Blais JF (2016) Co-culture for lipid production: advances and challenges. Biomass Bioenerg 92:20–30. https://doi.org/10.1016/j.biombioe.2016.06.003
Malla S, Koffas MAG, Kazlauskas RJ, Kim BG (2012) Production of 7-O-methyl aromadendrin, a medicinally valuable flavonoid, in Escherichia coli. Appl Environ Microbiol 78:684–694
Marín L, Gutiérrez-Del-Río I, Yagüe P et al (2017) De novo biosynthesis of apigenin, luteolin, and eriodictyol in the actinomycete Streptomyces albus and production improvement by feeding and spore conditioning. Front Microbiol 8:921. https://doi.org/10.3389/fmicb.2017.00921
Martens S, Forkmann G, Matern U, Lukačin R (2001) Cloning of parsley flavone synthase I. Phytochemistry 58:43–46. https://doi.org/10.1016/S0031-9422(01)00191-1
Miyahisa I, Funa N, Ohnishi Y et al (2006) Combinatorial biosynthesis of flavones and flavonols in Escherichia coli. Appl Microbiol Biotechnol 71:53–58. https://doi.org/10.1007/s00253-005-0116-5
Miyahisa I, Kaneko M, Funa N et al (2005) Efficient production of (2S)-flavanones by Escherichia coli containing an artificial biosynthetic gene cluster. Appl Microbiol Biotechnol 68:498–504. https://doi.org/10.1007/s00253-005-1916-3
Nasr Bouzaiene N, Chaabane F, Sassi A et al (2016) Effect of apigenin-7-glucoside, genkwanin and naringenin on tyrosinase activity and melanin synthesis in B16F10 melanoma cells. Life Sci 144:80–85. https://doi.org/10.1016/j.lfs.2015.11.030
Noguchi A, Kunikane S, Homma H et al (2009) Identification of an inducible glucosyltransferase from Phytolacca americana L. cells that are capable of glucosylating capsaicin. Plant Biotechnol 26:285–292. https://doi.org/10.5511/plantbiotechnology.26.285
Oyama K, Kondo T (2004) Total synthesis of apigenin 7,4′-di-O-β-glucopyranoside, a component of blue flower pigment of Salvia patens, and seven chiral analogues. Tetrahedron 60:2025–2034. https://doi.org/10.1016/j.tet.2004.01.001
Ozaki S, Imai H, Iwakiri T et al (2012) Regioselective glucosidation of trans-resveratrol in Escherichia coli expressing glucosyltransferase from Phytolacca americana. Biotech Lett 34:475–481. https://doi.org/10.1007/s10529-011-0784-4
Pandey RP, Malla S, Simkhada D et al (2012) Production of 3-O-xylosyl quercetin in Escherichia coli. Appl Microbiol Biotechnol 97(5):1889–1901. https://doi.org/10.1007/s00253-012-4438-9
Park JH, Oh JE, Lee KH et al (2012) Rational design of Escherichia coli for l-isoleucine production. ACS Synth Biol 1:532–540. https://doi.org/10.1021/sb300071a
Ramos A, Olano C, Braña AF et al (2009) Modulation of deoxysugar transfer by the elloramycin glycosyltransferase ElmGT through site-directed mutagenesis. J Bacteriol 191:2871–2875. https://doi.org/10.1128/JB.01747-08
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, third. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Samet I, Villareal MO, Motojima H et al (2015) Olive leaf components apigenin 7-glucoside and luteolin 7-glucoside direct human hematopoietic stem cell differentiation towards erythroid lineage. Differentiation 89:146–155. https://doi.org/10.1016/j.diff.2015.07.001
Takamura Y, Nomura G (1988) Changes in the intracellular concentration of acetyl-CoA and Malonyl-CoA in relation to the carbon and energy metabolism of Escherichia coli K12. Microbiology 134:2249–2253. https://doi.org/10.1099/00221287-134-8-2249
Thorson JS, Vogt T (2005) Glycosylated natural products. In: Wong CH (ed) Carbohydrate-based drug discovery. Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, pp 685–711
Thuan NH, Pandey RP, Thuy TTT et al (2013) Improvement of regio-specific production of myricetin-3-O-α-l-rhamnoside in engineered Escherichia coli. Appl Biochem Biotechnol 171:1956–1967. https://doi.org/10.1007/s12010-013-0459-9
Thuan NH, Park JW, Sohng JK (2013) Toward the production of flavone-7-O-β-d-glucopyranosides using Arabidopsis glycosyltransferase in Escherichia coli. Process Biochem 48:1744–1748. https://doi.org/10.1016/j.procbio.2013.07.005
Thuan NH, Sohng JK (2013) Recent biotechnological progress in enzymatic synthesis of glycosides. J Ind Microbiol Biotechnol 40(12):1329–1356. https://doi.org/10.1007/s10295-013-1332-0
Wang J, Zhou R-G, Wu T et al (2012) Total synthesis of apigenin. J Chem Res 36:121–122. https://doi.org/10.3184/174751912X13285269293913
Watts K, Lee P, Schmidt-Dannert C (2006) Biosynthesis of plant-specific stilbene polyketides in metabolically engineered Escherichia coli. BMC Biotechnol 6:22. https://doi.org/10.1186/1472-6750-6-22
Xu S, Shang MY, Liu GX et al (2013) Chemical constituents from the Rhizomes of Smilax glabra and their antimicrobial activity. Molecules 18:5265–5287. https://doi.org/10.3390/molecules18055265
Yan Y, Li Z, Koffas MAG (2008) High-yield anthocyanin biosynthesis in engineered Escherichia coli. Biotechnol Bioeng 100:126–140. https://doi.org/10.1002/bit.21721
Yu O, Shi J, Hession AO et al (2003) Metabolic engineering to increase isoflavone biosynthesis in soybean seed. Phytochemistry 63:753–763. https://doi.org/10.1016/S0031-9422(03)00345-5
Zhang H, Li Z, Pereira B, Stephanopoulos G (2015) Engineering E. coli-E. coli cocultures for production of muconic acid from glycerol. Microb Cell Fact 14:134. https://doi.org/10.1186/s12934-015-0319-0
Zhang H, Pereira B, Li Z, Stephanopoulos G (2015) Engineering Escherichia coli coculture systems for the production of biochemical products. Proc Natl Acad Sci USA 112:8266–8271. https://doi.org/10.1073/pnas.1506781112
Zhang H, Stephanopoulos G (2016) Co-culture engineering for microbial biosynthesis of 3-amino-benzoic acid in Escherichia coli. Biotechnol J 11:981–987. https://doi.org/10.1002/biot.201600013
Zhou K, Qiao K, Edgar S, Stephanopoulos G (2015) Distributing a metabolic pathway among a microbial consortium enhances production of natural products. Nat Biotechnol 33:377–383. https://doi.org/10.1038/nbt.3095
Acknowledgements
This work was supported by the National Foundation for Science and Technology Development of Vietnam (NAFOSTED) (106-NN.02-2014.25). We are grateful to Dr. Jules Beekwilder (Plant Research International, The Netherlands) for kindly providing pAC-4CL-STS plasmid, Dr. Shin-ichi Ozaki (Yamaguchi University, Japan) for providing plasmid pQE3-PaGT3 plasmid, Dr. Stefan Marten (Biotecnologia dei Prodotti Naturali, San Michele all’Adige, Italy) for FNSI, Dr. Mattheos A. G. Koffas (Rensselaer Polytechnic Institute, Troy, New York 12180, United States) and Dr. Sailesh Malla (Technical University of Denmark) for CHS and CHI genes, respectively.
Author information
Authors and Affiliations
Contributions
NHT and ACK conceived the study, designed experiments, analyzed data, and wrote the manuscript. NHT, DVC, and NXC performed the experiments, and NXC performed NMR study and analyzed the NMR data. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Thuan, N.H., Chaudhary, A.K., Van Cuong, D. et al. Engineering co-culture system for production of apigetrin in Escherichia coli. J Ind Microbiol Biotechnol 45, 175–185 (2018). https://doi.org/10.1007/s10295-018-2012-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-018-2012-x