Abstract
Chalcones, the central precursor of flavonoids, are synthesized exclusively in plants from tyrosine and phenylalanine via the sequential reaction of phenylalanine ammonia-lyase (PAL), cinnamate-4-hydroxylase (C4H), 4-coumarate:coenzyme A ligase (4CL) and chalcone synthase (CHS). Chalcones are converted into the corresponding flavanones by the action of chalcone isomerase (CHI), or non-enzymatically under alkaline conditions. PAL from the yeast Rhodotorula rubra, 4CL from an actinomycete Streptomyces coelicolor A3(2), and CHS from a licorice plant Glycyrrhiza echinata, assembled as artificial gene clusters in different organizations, were used for fermentation production of flavanones in Escherichia coli. Because the bacterial 4CL enzyme attaches CoA to both cinnamic acid and 4-coumaric acid, the designed biosynthetic pathway bypassed the C4H step. E. coli carrying one of the designed gene clusters produced about 450 μg naringenin/l from tyrosine and 750 μg pinocembrin/l from phenylalanine. The successful production of plant-specific flavanones in bacteria demonstrates the usefulness of combinatorial biosynthesis approaches not only for the production of various compounds of plant and animal origin but also for the construction of libraries of "unnatural" natural compounds.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Flavonoid-derived natural products in plants are well known floral pigments [13, 46, 53, 56] that also function as pollen fertility factors [11, 19, 45, 62, 64], signal molecules for beneficial plant-microbe symbiosis in the rhizosphere [52, 63], and antimicrobial defense compounds [24, 28, 37, 65, 68]. Following reports of flavonoid compounds having cancer chemopreventive, antioxidant and antiasthmatic activities, there has been an explosion of interest in their use as health-promoting components of the human diet [1, 7, 10, 12, 15, 18, 21, 23, 29, 40, 42–44, 58, 59, 66]. These phenylpropanoid and flavonoid biosynthetic enzymes are therefore attractive metabolic engineering targets of processes to enhance or initiate the production of economically desirable traits or compounds. The phenylpropanoid and flavonoid biosynthetic pathways and their regulation have been the subjects of many studies [16, 20, 26, 55, 67]. Recent advances in the regulation of these pathways and the biochemistry of their specific enzymes and enzyme complexes have opened up strategies to increase flavonoid biosynthesis by genetic engineering [8, 9, 25].
In the plant phenylpropanoid pathway (Fig. 1), phenylalanine ammonia-lyase (PAL), which deaminates phenylalanine to yield cinnamic acid, is the first enzyme in the general phenylpropanoid pathway. Cinnamic acid is hydroxylated by cinnamate-4-hydroxylase (C4H) to 4-coumaric acid, which is then activated to 4-coumaroyl-CoA by the action of 4-coumarate:coenzyme A ligase (4CL). Chalcone synthase (CHS) catalyzes the stepwise condensation of three acetate units from malonyl-CoA with 4-coumaroyl-CoA to yield naringenin chalcone, the precursor of a large number of flavonoids. Naringenin chalcone is converted to naringenin by chalcone isomerase (CHI) or non-enzymatically in vitro under alkaline conditions [47].
Flavanone biosynthetic pathway in plants. The dotted arrows represent the expected bypass pathway for combinatorial biosynthesis of flavanones in Escherichia coli. PAL Phenylalanine ammonia-lyase, TAL tyrosine ammonia-lyase, C4H cinnamate-4-hydroxylase, 4CL 4-coumarate:coenzyme A ligase, ScCCL 4-coumarate/cinnamate:coenzyme A ligase from Streptomyces coelicolor A3(2), CHS chalcone synthase, CHI chalcone isomerase
The production of flavonoids by genetically engineered bacteria has not yet been reported, although the heterologous expression of phenylpropanoid biosynthetic enzymes in bacteria was described previously [2, 31, 41, 69]. One of the barriers to the production of these compounds is the difficulty in expressing active C4H, which could not be efficiently expressed in bacteria due to its instability and the lack of a specific cytochrome P-450 reductase [51]. We recently discovered a 4CL in the gram-positive, filamentous bacterium Streptomyces coelicolor A3(2) that can activate cinnamic acid to cinnamoyl-CoA, in addition to 4-coumaric acid to 4-coumaroyl-CoA [35]. Using the 4CL enzyme would bypass the C4H step for the production of pinocembrin chalcone from phenylalanine via the phenylpropanoid pathway. In this review article, we briefly describe the phenylpropanoid pathway in plants and the successful production of flavanones in Escherichia coli by inserting the bacterial 4-coumarate/cinnamate:CoA ligase gene within an artificial gene cluster for combinatorial biosynthesis. Successful heterologous production of plant-specific flavanones will allow a variety of flavonoids to be produced in bacteria by further genetic engineering, similar to the excellent results reported for the biosynthesis of polyketides [49, 50, 70].
Phenylpropanoid pathway in plants
Phenylalanine ammonia-lyase
As the first step in the phenylpropanoid pathway, PAL (EC 4.3.1.5) catalyzes the elimination of ammonia from phenylalanine to yield cinnamic acid (Fig. 1). PAL activity has been found in some fungi, bacteria, and in all higher plants but not in animals. In plants, PAL has been extensively studied for its roles in the formation of lignins, isoflavonoids, and other secondary metabolites. In addition, tyrosine ammonia-lyase (TAL) catalyzes the formation of 4-coumaric acid from tyrosine. Because some PALs have TAL activity [39, 54, 57], both phenylalanine and tyrosine are expected to be precursors in the combinatorial biosynthesis of flavonoids in bacteria.
Cinnamate-4-hydroxylase
Cinnamic acid is hydroxylated by the action of C4H (EC 1.14.13.11). This enzyme is a membrane-bound cytochrome P-450 hydroxylase that requires molecular oxygen and a reducing equivalent from NADPH delivered via cytochrome P-450 reductase [31, 51]. Therefore, efficient expression of a C4H gene in bacteria requires simultaneous expression of the specific P-450 reductase gene.
4-Coumarate:coenzyme A ligase
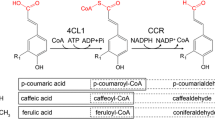
4-coumarate:CoA ligase (EC 6.2.1.12) catalyzes the conversion of 4-coumarate (4-hydroxycinnamate) and other substituted cinnamates, such as caffeate (3,4-dihydroxycinnamate) and ferulate (3-methoxy-4-hydroxycinnamate), into the corresponding CoA thiol esters, which are used for the biosynthesis of numerous phenylpropanoid-derived compounds, such as lignins, lignans, suberins, flavonoids, isoflavonoids, and various small phenolic compounds. 4CL-catalyzed CoA ester formation takes place via a two-step reaction. During the first step, 4-coumarate and ATP form a coumaroyl-adenylate intermediate with the simultaneous release of pyrophosphate. In the second step, the coumaroyl group is transferred to the sulfhydryl group of CoA, and AMP is released [3, 36]. The mechanism of formation of an adenylate intermediate is common among a number of enzymes with divergent functions, including luciferases, fatty acyl-CoA ligases, acetyl-CoA ligases, and specialized domains within peptide synthetase multi-enzymes. Despite their low overall amino acid sequence identity, the similar reaction mechanisms of these enzymes and the presence of conserved peptide motifs were used as criteria to classify them in a superfamily of adenylate-forming enzymes [22]. The relationship of 4CL to other adenylate-forming enzymes was substantiated recently by functional analysis of key 4CL amino acid residues that are conserved in other adenylate-forming enzymes [60].
Chalcone synthase
Chalcone synthase (EC 2.3.1.74) is a plant-specific polyketide synthase that uses a starter CoA-ester, typically 4-coumaroyl-CoA, derived from the phenylpropanoid pathway. It catalyzes three condensation reactions with malonyl-CoA and folds the resulting tetraketide intermediate into a new aromatic ring system [17, 33, 38]. After initial capture of the 4-coumaroyl moiety, each subsequent condensation step begins with decarboxylation of malonyl-CoA at the CHS active site, and the resulting acetyl-CoA carboanion then serves as the nucleophile for chain elongation. Ultimately, these reactions generate a tetraketide intermediate that cyclizes by a Claisen condensation into a hydroxylated aromatic ring system [17]. CHS supplies 4,2′,4′,6′-tetrahydroxychalcone to downstream enzymes that synthesize a wide variety of flavonoids, such as phytoalexins and anthocyanin pigments.
Chalcone isomerase
In the last stages of the biosynthesis of flavanone, chalcone isomerase (EC 5.5.1.6) catalyzes the intramolecular cyclization of 4,2′,4′,6′-tetrahydroxychalcone (chalcone) and 6-deoxychalcone (4,2,4-trihydroxychalcone), both derived from the upstream enzyme CHS, into (2S)-naringenin (5,7,4-trihydroxyflavanone) and (2S)-5-deoxyflavanone (7, 4-dihydroxyflavanone), respectively [27, 48]. Although chalcones spontaneously cyclize in alkaline solution to produce an enantiomeric mixture of flavanones, CHI directs formation of biologically active (2S)-flavanones [4, 5, 34].
4-Coumarate/cinnamate:coenzyme A ligase from S. coelicolor A3(2)
The filamentous, soil-living, gram-positive bacterial genus Streptomyces is characterized by the ability to produce a wide variety of secondary metabolites, including antibiotics, and by complex morphological differentiation culminating in sporulation [30]. S. coelicolor A3(2) has been most extensively and intensively characterized among Streptomyces by the research group of D.A. Hopwood, and recently the whole genome has been sequenced (http://www.sanger.ac.uk/Projects/S_coelicolor) [6]. In the database, a gene (SCD10.15) encoding a 522-amino-acid protein has been annotated as a 4CL gene. The protein has higher sequence similarity to plant 4CLs than to bacterial acyl-CoA ligases; it shows 44% identity and 58% similarity to Arabidopsis At4CL2. This is the first bacterial protein that shows end-to-end sequence similarity to plant 4CLs over 40% identity. The recombinant protein, expressed in E. coli, had distinct 4CL activity, but its substrate specificity was unique; the enzyme efficiently converted cinnamate, which is a very poor substrate for plant 4CLs [35]. The enzyme was therefore named ScCCL, for S. coelicolor A3(2) cinnamate:CoA ligase (Fig. 2). The 4-coumarate/cinnamate:CoA ligase activity of the enzyme is useful for the production of flavanones by combinatorial biosynthesis in bacteria in that it can bypass the C4H step, which is apparently difficult to express in bacteria, as described above.
Flavanone fermentation in bacteria
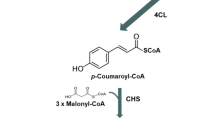
For the purpose of production of flavanones in E. coli, an artificial gene cluster was constructed that contained three genes of heterologous origins: PAL from a yeast Rhodotorula rubra, 4CL from an actinomycete S. coelicolor A3(2), and CHS from a licorice plant Glycyrrhiza echinata (Fig. 3) [32]. The chalcones were expected to be converted in vitro to the corresponding flavanones by raising the pH of the culture broth to 9. Because PAL uses phenylalanine and tyrosine as substrates, 4CL attaches CoA to both cinnamate and 4-coumarate, and CHS forms chalcones from cinnamate-CoA and 4-coumarate-CoA, the respective artificial gene cluster was expected to direct the synthesis of pinocembrin from phenylalanine and naringenin from tyrosine (Fig. 1). We first generated a gene cluster in which PAL, 4CL and CHS genes were placed in this order under the control of the T7 promoter (PT7) and the ribosome-binding sequence in the pET vector. The translational initiation codons of 4CL and CHS overlapped with the termination codons of the preceding genes. This type of gene organization is often found in bacterial gene clusters, such as those for antibiotic and xenobiotic biosynthesis. Plasmid pET26b-3GS carrying the gene cluster, however, caused E. coli BL21 (DE3) to produce only a very small amount of pinocembrin and naringenin, even when the amino acid precursors phenylalanine and tyrosine were added; the bacteria produced 0.2 μg pinocembrin/l and 0.57 μg naringenin/l , when 2 mM of each of the amino acid precursors was supplied. By contrast, 4-coumaric acid and cinnamic acid accumulated in large amounts, about 9 mg/l and 12 mg/l, respectively.
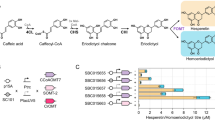
Organization of the artificial gene clusters used for production of flavanones in E. coli. A In pET26b-3GS, PAL from a yeast, ScCCL from an actinomycete, and CHS from a licorice plant were placed under the control of the T7 promoter and ribosome-binding sequence in pET26b. The initiation codons of ScCCL and CHS overlap the termination codon of the preceding genes. In pET26b-rbs-3GS, the three genes (all with the ribosome-binding sequence at appropriate positions) are co-transcribed from the T7 promoter in front of PAL. In pET26b-PT7-3GS, all three genes contain their own T7 promoter and ribosome-binding sequences. B Accumulation of naringenin and pinocembrin in the culture broth of E. coli harboring the artificial gene clusters
Accumulation of 4-coumaric acid and cinnamic acid suggested that 4CL and CHS were not expressed efficiently. We therefore constructed pET26b-rbs-3GS, in which the three genes were transcribed by a single PT7 in front of PAL, and each of the three contained rbs at appropriate positions and pET26b-PT7-3GS, in which all three genes contained both PT7 and a ribosome-binding sequence (Fig. 3). Placement of a ribosome-binding sequence in front of each gene (plasmid pET26b-rbs-3GS) enhanced the yields of pinocembrin and naringenin by about 45-fold and 85-fold, respectively. Consistent with this, SDS-PAGE of a cell-lysate prepared from E. coli harboring pET26b-rbs-3GS revealed the presence of large amounts of soluble PAL, 4CL and CHS. Furthermore, large amounts of flavanones (about 750 μg pinocembrin/l and 450 μg naringenin/l) were produced. These findings show the importance of efficient transcription from the T7 promoter and efficient translation from the ribosome-binding sequence.
The yields of flavanones were still low. We expect that an increase in the amount of a precursor, malonyl-CoA, which is present at 4–90 μM (0.01–0.23 nmol/mg dry weight) in E. coli under normal cultural conditions [6], by overexpression of the acetyl-CoA carboxylase gene [14] would lead to enhancement of the yields. A fermentation condition to remove the ammonia produced by the action of PAL would also increase the yields. The use of a mutant strain of Corynebacterium glutamicum, e.g., a phenylalanine or tyrosine fermenter, would release us from the need to provide amino acids to the culture. Concerning the host cells, yeast and fungi, such as Saccharomyces cerevisiae and Aspergillus oryzae, may be useful for functional expression of P-450 genes that are required for the further conversion of flavanones.
Concluding remarks
The genomes of Streptomyces are useful sources of genes coding for a variety of functions, since these bacteria produce a variety of unique secondary metabolites. Genome sequencing of additional Streptomyces species could very well lead to the discovery of enzymes that are applicable to the bioconversion and combinatorial biosynthesis of useful compounds. The production of plant-specific flavanones in E. coli serves as an example of the usefulness of this type of metabolic engineering. We believe that it can be applied to the production of a variety of compounds of plant and animal origin in bacteria. In addition, the this approach together with combinatorial biosynthesis will allow the construction of libraries of "unnatural" natural products.
References
Alekel DL, St Germain A, Pererson CT, Hanson KB, Stewart JW, Toda T (2000) Isoflavone-rich soy protein attenuates bone loss in the lumber spine of perimenopausal women. Am J Clin Nutr 72:844–852
Baedeker M, Schulz GE (1999) Overexpression of a designed 2.2 kb gene of eukaryotic phenylalanine ammonia-lyase in E. coli. FEBS Lett 457:57–60
Becker-André M, Schulze-Lefert P, Hahlbrock K(1991) Structural comparison, modes of expression, and putative cis-acting elements of the two 4-coumarate:CoA ligase genes in potato. J Biol Chem 266:8551–8559
Bednar RA (1990) Reactivity and pH dependence of thiol conjugation to N-ethylmaleimide: detection of a conformational change in chalcone isomerase. Biochemistry 29:3684–3690
Bednar RA, Fried WB, Lock YW, Pramanik B (1989) Chemical modification of chalcone isomerase by mercurials and tetrathionate. J Biol Chem 264:14272–14276
Bentley SD, Chater KF, Cerdeno-Tarraga AM, Challis GL, Thomson NR, James KD, Harris DE, Quail MA, Kieser H, Harper D, Bateman A, Brown S, Chandra G, Chen CW, Collins M, Cronin A, Fraser A, Goble A, Hidalgo J, Hornsby T, et al (2002) Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417:141–147
Bingham SA, Atkinson C, Liggins J, Bluck L, Coward A (1998) Phyto-estrogen: where are we now? Brit J Nutr 79:393–406
Bradley JM, Davis KM, Deroles SC, Bloor SJ, Lewis DH (1998) The maize Lc regulatory gene up-regulates the flavonoid biosynthetic pathway of Petunia. Plant J 13:381–392
Bradley JM, Deroles SC, Boase, Bloor S, Swinny E, Davis KM (1999) Variation in the ability of the maize Lc regulatory gene to upregulate flavonoid biosynthesis in heterologous systems. Plant Sci 140:31–39
Bravo L (1998) Polyphenols: chemistry, dietary sources, metabolism, and nutritional significance. Nutr Rev 56:317–333
Burbulis I, Iacobucci M, B Shirley (1996) A null mutation in the first enzyme of flavonoid biosynthesis does not affect male fertility in Arabidopsis. Plant Cell 8:1013–1025
Cassidy A, Bingham S (1995) Biological effects of isoflavones in young women: importance of the chemical composition of soybean products. Brit J Nutr 74:587–601
Davies K, Bloor S, Spiller G (1998) Production of yellow colour in flowers: redirection of flavonoid biosynthesis in Petunia. Plant J 13:259–266
Davis MS, Solbiati J, Cronan, Jr JE (2000) Overproduction of acetyl-CoA carboxylase activity increases the rate of fatty acid biosynthesis in Escherichia coli. J Biol Chem 275:28593–28598
Davis SR, Dalais FS, Simpson ER, Murkias AL (1999) Phytoestrogens in health and disease. Recent Prog Horn Res 54:185–210
Dixon RA, Steele CL (1999) Flavonoids and isoflavonoids – a gold mine for metabolic engineering. Trends Plant Sci 4:394–400
Ferrer JL, Jez JM, Bowman ME, Dixon RA, Noel JP (1999) Structure of chalcone synthase and the molecular basis of plant polyketide biosynthesis. Nat Struct Biol 6:775–784
File SE, Jarrett N, Fluck E, Duffy R, Casey K, Wiseman H (2001) Eating soya improves human memory. Psychopharmacology 157:430–436
Fischer R, Budde I, Hain R (1997) Stilbene synthase gene expression causes changes in flower colour and male sterility in tobacco. Plant J 11:489–498
Forkmann G, Martens S (2001) Metabolic engineering and applications of flavonoids. Curr Opin Biotechnol 12:155–160
Frits WA, Coward L, Wang J, CA Lamartiniere (1998) Dietary genistein: prenatal mammary cancer prevention, bioavailability and toxicity testing in rat. Carcinogenesis 19:2152–2158
Fulda M, Heinz E, FP Wolter. (1994) The fadD gene of Escherichia coli K-12 is located close to rnd at 39.6 min of the chromosomal map and is a new member of the AMP-binding protein family. Mol Gen Genet 242:241–249
Galati G, Teng S, Moridani MY, Chan TS, O'Brien PJ (2000) Cancer chemoprevention and apoptosis mechanisms induced by dietary polyphenolics. Drug Methabol Drug Interact 17:311–349
Grayer RJ, Harborne JB (1994) A survey of antifungal compounds from higher plants, 1982–1993. Phytochemistry 37:19–42
Grotewold E, Chamberlin M, Snook M, Siame B, Butler L, Swenson J, Maddock S, St Clair G, Bowen B (1998) Engineering secondary metabolism in maize cells by ectopic expression of transcription factors. Plant Cell 10:721–740
Hahlbrock K, Grisebach H (1979) Enzymatic controls in the biosynthesis of lignin and flavonoids. Annu Rev Plant Physiol 30:105–130
Hahlbrock K, Zilg H, Grisebach H (1970) Stereochemistry of the enzymatic cyclisation of 4,2′,4′-trihydroxychalcone to 7,4′-dihydroxyflavanone by isomerases from mung bean seedlings. Eur J Biochem 15:13–18
Harborne JB (1999) The comparative biochemistry of phytoalexin induction in plants. Biochem Syst Ecol 27:335–368
Hollman PC, Katan MB (1997) Absorption, metabolism and health effects of dietary flavonoids in man. Biomed Pharmacother 51:305–310
Horinouchi S,Beppu T (1994) A-factor as a microbial hormone that controls cellular differentiation and secondary metabolism in Streptomyces griseus. Mol Microbiol 12:859–864
Hotze M, Schröder G, Schröder J (1995) Cinnamate 4-hydroxylase from Cantharanthus roseus, and a strategy for functional expression of plant cytochrome P450 proteins as translational fusions with P450 reductase in E. coli. FEBS Lett 374:345–350
Hwang EI, Kaneko M, Ohnishi Y, Horinouchi S (2003) Production of plant-specific flavanones by Escherichia coli containing an artificial gene cluster. Appl Environ Microbiol 69:2699–2706
Jez JM, Austin MB, Ferrer JL, Bowman ME, Schröder J, Noel JP (2000) Structural control of polyketide formation in plant-specific polyketide synthase. Chem Biol 7:919–930
Jez JM, Noel JP (2002) Reaction mechanism of chalcone isomerase. J Biol Chem 277:1361–1369
Kaneko M, Ohnishi Y, Horinouchi S (2003) Cinnamate:coenzyme A ligase from the filamentous bacterium Streptomyces coelicolor A3(2). J Bacteriol 185:20–27
Knobloh KH, Hahlbrock K (1975) Isoenzymes of p-coumarate:CoA ligase from cell suspension of Glycine max. Eur J Biochem 52:311–320
Knott J, Martens S, Forkmann G (2000) Induction of resistance mechanism against fungal infection of cultivars of Rosa. Polyphenols Commun 2:627–628
Kreuzaler F, Hahlbrock K (1975) Enzymic synthesis of an aromatic ring from acetate units. Eur J Biochem 56:205–213
Kyndt JA, Meyer TE, Cusanovich MA, Van Beeumen JJ (2002) Characterization of a bacterial tyrosine ammonia lyase, a biosynthetic enzyme for the photoactive yellow protein. FEBS Lett 512:240–244
Lamartiniere CA (2000) Protection against breast cancer with genistein: a component of soy. Am J Clin Nutr 71:1705S-1707S
Lee D, Douglas CJ (1996) Two divergent members of a tobacco 4-coumarate:coenzyme A ligase (4CL) gene family. Plant Physiol 112:193–205
Le Marchand L (2002) Cancer preventive effects of flavonoids – a review. Biomed Pharmacother 56:296–301
MerzDemlow BE, Duncan AM, Wangen KE, Xu X, Carr TP, Phipps WR, Kurzer MS (2000) Soy isoflavones improve plasma lipids in normocholesterolemic, premenopausal women. Am J Clin Nutr 71:1462–1469
Messina MJ (1999) Legumes and soybeans: overview of their nutritional profiles and health effects. Am J Clin Nutr 70:439S-450S
Mo Y, Nagel C, Taylor L (1992) Biochemical complementation for chalcone synthase mutants defines a role for flavonols in functional pollen. Plant Biol 89:7213–7217
Mol J, Grotewold E, Koes R (1998) How genes paint flowers and seeds. Trends Plant Sci 3:212–217
Mol JNM, Robbins MP, Dixon RA, Veltkamp E (1985) Spontaneous and enzymic rearrangement of naringenin chalcone to flavanone. Phytochemistry 24:2267–2269
Moustafa E, Wong E (1967) Purification and properties of chalcone-flavanone isomerase from soya bean seed. Phytochemistry 6:625–632
Pfeifer BA, Admiraal SJ, Gramajo H, Cane DE, Khosla C (2001) Biosynthesis of complex polyketides in a metabolically engineered strain of E. coli. Science 291:1790–1792
Pfeifer BA, Khosla C (2001) Biosynthesis of polyketides in heterologous hosts. Microbiol Mol Biol Rev 65:106–118
Pompon D, Louerat B, Bronine A, Urban P (1996) Yeast expression of animal and plant P450s in optimized redox environments. Methods Enzymol 272:51–64
Pueppke JL (1996) The genetics and biochemical basis for nodulation of legumes by rhizobia. Crit Rev Biotechnol 16:1-51
Quattrocchio F, Wing J, van der Woude K, Souer E, de Vetten N, Mol J, Koes R (1999) Molecular analysis of the anthocyanin 2 gene of petunia and its role in the evolution of flower color. Plant Cell 11:1433–1444
Rösler J, Krekel F, Amrhein N, Schmid J (1997) Maize phenylalanine ammonia-lyase has tyrosine ammonia-lyase activity. Plant Physiol 113:175–179
Schröder J (1997) A family of plant-specific polyketide synthases: facts and predictions. Trends Plant Sci 2:373–378
Schwinn K, Markham K, Given N (1994) Floral flavonoids and the potential for pelargonidin biosynthesis in commercial Chrysanthemum cultivars. Phytochemistry 35:145–150
Scott DA, Hammond PM, Brearly GM, Price CP (1992) Identification by high-performance liquid chromatography of tyrosine ammonia-lyase activity in purified fraction of Phaseolus vulgaris phenylalanine ammonia-lyase. J Chromatog B 573:309–312
Setchell KD (1998) Phytoestrogens: the biochemistry, physiology, and implications for human health of soy isoflavones. Am J Clin Nutr 68:1333S-1346S
Setchell KD, Cassidy A (1999) Dietary isoflavones: biological effects and relevance to human health. J Nutr 129:758S-767S
Stuible HP, Büttner D, Ehlting J, Hahlbrock K, Kombrink E (2000) Mutational analysis of 4-coumarate:CoA ligase identifies functionally important amino acids and verifies its relationship to other adenylate-forming enzymes. FEBS Lett 467:117–122
Takamura Y, Nomura. G (1988) Changes in the intracellular concentration of acetyl-CoA and malonyl-CoA in relation to the carbon and energy metabolism of Escherichia coli K12. J Gen Microbiol 134:2249–2253
Taylor LP, Jorgensen R (1992) Conditional male fertility in chalcone synthase-deficient Petunia. J Hered 83:11–17
Van Rhijn R, Vanderleyden J (1995) The Rhizobium-plant symbiosis. Microbiol Rev 59:124–142
van der Meer I, Stam M, van Tunen A, Mol J, Stuitje R (1992) Antisense inhibition of flavonoid biosynthesis in Petunia anthers results in male sterility. Plant Cell 4:253–262
Volpin H, Elkind Y, Okon Y, Kapulnik Y (1994) A vesicular arbuscular mycorrhizal fungus (Glomus intraradix) induces a defense response in alfalfa roots. Plant Physiol 104:683–689
Wang HK, Xia Y, Yang ZY, Natschke SL, Lee KH (1998) Recent advances in the discovery and development of flavonoids and their analogues as antitumor and anti-HIV agents. Adv Exp Med Biol 439:191–225
Weisshaar B, Jenkins GI (1998) Phenylpropanoid biosynthesis and its regulation. Curr Opin Plant Biol 1:251–257
Welle R, Schröder G, Schilts E, Grisbach H, Schröder J(1991) Induced plant responses to pathogen attack: analysis and heterologous expression of the key enzyme in the biosynthesis of phytoalexins in soybean (Glycine max L. Merr. cv. Harosoy 63). Eur J Biochem 196:423–430
Yamaguchi T, Kurosaki F, Suh DY, Sankawa U, Nishioka M, Akiyama T, Shibuya M, Ebizuka Y (1999) Cross-reaction of chalcone synthase and stilbene synthase overexpressed in E. coli. FEBS Lett 460:457–461
Zawada RJX, Khosla C (1999) Heterologous expression, purification, reconstitution and kinetic analysis of an extended type II polyketide synthase. Chem Biol 6:607–615
Acknowledgements
The work from this laboratory was supported by a Research Grant of the Noda Institute for Scientific Research, by the Bio Design Program of the Ministry of Agriculture, Forestry, and Fisheries of Japan (BDP-03-VI-2–2), and by a grant from the Industrial Technology Research Grant Program 2000 of the New Energy and Industrial Technology Development Organization of Japan (00A03004).
Author information
Authors and Affiliations
Corresponding author
Additional information
Dedicated to Professor Sir David Hopwood.
Rights and permissions
About this article
Cite this article
Kaneko, M., Hwang, E.I., Ohnishi, Y. et al. Heterologous production of flavanones in Escherichia coli: potential for combinatorial biosynthesis of flavonoids in bacteria. J IND MICROBIOL BIOTECHNOL 30, 456–461 (2003). https://doi.org/10.1007/s10295-003-0061-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-003-0061-1