Abstract
In vivo biosensors can recognize and respond to specific cellular stimuli. In recent years, biosensors have been increasingly used in metabolic engineering and synthetic biology, because they can be implemented in synthetic circuits to control the expression of reporter genes in response to specific cellular stimuli, such as a certain metabolite or a change in pH. There are many types of natural sensing devices, which can be generally divided into two main categories: protein-based and nucleic acid-based. Both can be obtained either by directly mining from natural genetic components or by engineering the existing genetic components for novel specificity or improved characteristics. A wide range of new technologies have enabled rapid engineering and discovery of new biosensors, which are paving the way for a new era of biotechnological progress. Here, we review recent advances in the design, optimization, and applications of in vivo biosensors in the field of metabolic engineering and synthetic biology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A biosensor is a biological entity that can be used to report the concentration of a desired compound. The first biosensor was invented to measure glucose in a biological sample using an immobilized glucose oxidase electrode [21]. Since then, numerous biosensors have been constructed in many forms, ranging from electrochemical, fluorescence tagged, nanomaterials, silica or quartz, and genetic constructs with reporter genes [52, 167]. Among them, in vivo biosensors are designed to enable continuous and real-time monitoring of target metabolites in biological systems, which provide communications into and out of the cell. In vivo biosensors have a number of advantages over other detection methods, which have been summarized in several reviews [27, 52, 167]. Currently, a broad spectrum of metabolites can be detected by in vivo biosensors non-invasively [137, 198]. The emerging in vivo biosensors has begun to be applied in many fields of biotechnology, such as diagnostics, biomedicine, food safety and processing, environmental monitoring, and security. Here, we focus on their applications in the field of metabolic engineering and synthetic biology.
Metabolic engineering enables microbes to produce various valuable products, and many attempts are required before an efficient cell factory is constructed [17]. However, evaluation of microbial phenotypes by the traditional analytical methods such as chromatography and mass spectrometry is slow and cumbersome, which becomes a major rate-limiting step in metabolic engineering. For example, random mutagenesis or genome-scale modifications generate large pools of mutant strains, which are difficult to analyse using the traditional analytical methods. Genetically encoded in vivo biosensors enable rapid and multiplexed phenotypic evaluation, offering the potential for high-throughput screening of novel or improved biocatalysts [32, 125, 137, 151, 198].
Metabolic engineering has greatly benefited from adaptive evolution approaches, which have been used as powerful tools to adapt microorganisms to stress conditions [13] or to improve product formation [5, 183]. Usually, an adaptive evolution approach selects mutants with improved growth and survival. Thus, mutants with improved stress tolerance can be selected [13], and mutants with improved product formation can also be selected if it is a growth-coupled product [5, 41, 183]. When there is no relationship between the desired product and growth, it is hard to use adaptive evolution. In vivo biosensors can sense and respond to desired metabolites, and couple the sensing with a fitness advantage by expressing a gene necessary for survival under selective conditions, expanding adaptive evolution to include phenotypes which are not linked to growth [20, 76, 99].
Another challenge in metabolic engineering is the inefficient use of cellular resources and imbalanced metabolism. For example, excessive expression of a certain pathway gene will burden the host cell and waste cellular resources to produce unnecessary mRNAs and proteins, while uncontrolled or deregulated metabolic pathways may lead to metabolic imbalance, and enzymes or intermediates can accumulate to toxic levels [196]. Furthermore, the balanced distribution of metabolic flux and concentration of key metabolites in a cell are constantly changing depending on the fermentation phases. Synthetic biology is able to design and build novel biomolecular components, networks, and pathways, and to use these constructs to control and regulate biological systems dynamically [152]. The application of synthetic biology in metabolic engineering can dynamically tune gene expression and intermediate accumulation to proper levels, improving the balance and productivity in microbes [22, 188, 196], but implementation of such control and regulation requires the development of intracellular metabolite sensors.
An exciting application of synthetic biology is focused on imaging or observation of chemicals [8, 23, 74, 135], which is also inspired by the availability of in vivo biosensors. The live concentration of metabolites enables metabolic engineers to determine the optimal production condition [135], environmentalists to measure pollutants or toxic compounds [8], or doctors to diagnose disease [23, 74]. Within these applications, new biological systems that do not exist in nature can be designed and built by the assembly of basic genetic parts and modules. Among these genetic parts, in vivo biosensors represent a significant contribution [167].
To date, only a few in vivo biosensors have been well characterized, which significantly limits their applications. Most in vivo biosensors have been designed based on a small number of naturally occurring regulatory systems such as transcription factors [32, 83, 101, 149, 174] and riboswitches [64, 96, 201, 203]. Novel in vivo biosensors have also been developed using rational design or random mutagenesis approaches to alter the effector specificities of existing genetic components [16, 19, 25, 65, 134, 159, 160, 185], and we will review their design and optimization strategies in detail in the following sections. Due to limited knowledge, a biosensor for a target product is difficult to generate if an associated transcription factor or riboswitch has not been identified. Advances in systems biology have facilitated the identification of responsive cellular machineries that respond to particular chemical or environmental stresses [22, 93, 123, 147, 158, 192], thus providing an alternative method for biosensor development without the need for isolated transcription factors or riboswitches.
The development of appropriate in vivo biosensors mostly relies on several key considerations, such as working range, sensitivity, specificity, and response time. The working range describes the maximum and minimum concentrations of a chemical that the sensor can detect. This range results in a meaningful and accurate output for the sensing system. The desired operational concentration varies by application. In the case of measuring pollutants or toxic compounds, the working range at nanomolar would be ideal, but the working range at millimolar would be more useful in optimizing a pathway to produce a target compound at grams. Several strategies for modulating biosensor operational range have been reported [19, 129]. The sensitivity can be quantified as the ratio of the incremental change in the sensor’s output to the incremental change of the measured input. An ideal sensor should have a large and preferably constant sensitivity in its operating range. Multiple approaches have been reported to increase the sensitivity [101, 150]. The specificity is the extent to which a biosensor is specific for a particular condition or metabolite. A low specificity may cause false and inaccurate readings when applying biosensors. Thus, the need for specific detection and response has to be addressed. Most protein-based biosensors involve the binding of the target molecule to its active site. Such interactions can be extremely specific due to the carefully evolved shape of the active site through natural selection. Another consideration of the biosensor is the response time to a stimulus. In vivo biosensors based on protein or nucleic acid act at the transcriptional level or translational level, which, therefore, results in a response time varied from seconds to hours behind actual metabolite changes. In most applications of biosensor, a shorter response time offers substantial advantages such as dynamic regulation [22, 188, 196] and real-time imaging of chemicals [3, 8, 11, 135, 136]. A shorter response time could be obtained using Spinach RNA or a low-molecular-weight protein as the reporter [62, 71, 88].
While biosensors have been reviewed previously [27, 137, 167, 198], this review focuses on the recent advances in the design and engineering of in vivo biosensors with increasing numbers of metabolite targets, mechanisms of action, and applications in the past 5 years. Here, two major categories of in vivo biosensors are classified based on their diverse mechanisms of sensing, i.e., protein-based and nucleic acid based in vivo biosensors. Finally, we discuss future directions to develop and apply in vivo biosensors.
Protein-based in vivo biosensors
Transcription factor (TF)-based in vivo biosensors
In nature, TFs have evolved to regulate gene expression in response to specific metabolites and have been widely used to construct in vivo biosensors [98]. These biosensors can offer high sensitivity and dynamic range, where small changes in ligand concentration are amplified through gene expression into large changes in protein abundance. As shown in Fig. 1a, when activated by the target effector molecule, the conformational changes in the activating TF enable it bind to specific DNA sequences in a promoter region and in turn to enhance expression of a reporter gene (e.g., fluorescent reporters, regulatory switches, or selection markers), while the conformational changes in the repressing TF upon effector binding release it from the promoter and allow the expression of the reporter gene. For biosensor design, overexpression of the relevant TF is usually necessary even if it is endogenous to the sensor cells [105, 149]. In general, TF-based biosensors offer a higher potential in their diversity and specificity due to the versatility and modularity of the functional groups in TFs. Besides, studies have shown that some prokaryotic TFs can also function in eukaryotes [83, 151, 174], further expanding their versatility. TF-based biosensors may have poor orthogonality and high background noise due to potential uncharacterized interactions between candidate TFs and operator sites [7, 80]. Thus, a successful reconstruction of a TF-based biosensor requires a clear understanding of transcriptional regulation network and protein expression system. Comparative genomics approaches enable in silico identification of TF-binding sites and regulon reconstruction, such as the LacI family [130] and the ROK (repressor, open reading frame, kinase) family [70].
To date, TFs have been reported to respond to sugars, vitamins, antibiotics, amino acids, secondary metabolites, metal ions, and lipid-related metabolites [98], serving as a large resource of biological components to directly design in vivo biosensors for various metabolites. Bioinformatic approaches and databases have also contributed to the collection, visualization, and analysis of TFs [118], and a Web-based tool has been recently developed to design biosensors by screening for multi-step enzymatic transformations converting non-detectable compounds into detectable ones [28]. Examples for different TF-based biosensors are provided in Table 1. For instance, TFs such as XylR and CebR were known to affect xylose and cello oligosaccharides catabolism, respectively, and a panel of biosensors was generated subsequently to sense xylose [57, 161, 174] and cellobiose [164]. Similarly, NADPH sensors were, respectively, identified in Escherichia coli using its native redox sensitive TF SoxR [149] and Saccharomyces cerevisiae using its native redox sensitive TF Yap1p [199]. According to this strategy, a variety of metabolite-actuated biosensors have been created, including genetically encoded sensors that respond to l-phenylalanine [94], l-tryptophan [39, 92], lactate [51], acrylate [136], glucarate [136], erythromycin [136], naringenin [102, 136], macrolide [68], cis,cis-muconic acid (CCM) [151], 3-hydroxypropionic acid [135, 202], 3,4-dihydroxy benzoate [66], butanol [32], and ferulic acid [97]. Furthermore, several biosensors have been constructed to detect lipid-related metabolites, such as alkane [180], fatty acid/fatty acyl-CoA [162], and malonyl-CoA [83, 186, 188] by re-assembling the natural alkane-responsive TF AlkR, bacterial regulator of fatty acid FadR, and bacterial regulator of fatty acid FapR, respectively. More recent efforts have further engineered natural TFs that can respond to arsenic (e.g., ArsR) [105], zinc (e.g., ZntR) [177], iron–sulfur (e.g., IscR) [50], copper (e.g., MarR) [56], ammonium (e.g., GlnR) [182], and choline (e.g., BetI) [139] into biosensors to measure environmentally and medically relevant concentrations of ions.
The number of known TFs is far lower than the number of metabolites of interest, limiting the application of TF-based biosensors. Hence, expanding the repertoire of compounds detectable by TFs is highly desirable. The alteration of TF effector specificities by various protein engineering approaches to recognize non-natural compounds enhances the availability of sensors with appropriate characteristics. It has already been demonstrated that computational approaches can be used for engineering specificity of TFs relying on a range of strategies that include structure-based predictions [160], metabolite analog searching [197], and computer-aided design [85]. In synergy with ongoing efforts of biosensor development, computational strategies offer immediate expansion for effectors of a particular TF.
Another potentially generalizable strategy is to create a synthetic TF by fusing a known metabolite-binding protein with a distinct functional domain, whereby the binding of the metabolite of interest brings about a change of activity of the functional domain [20, 43, 165, 194]. In one study, an S-adenosylmethionine biosensor was created by fusing the MetJ repressor to the transcriptional activation domain B42 [165]. The DNA-binding domains of the synthetic TF can be made more programmable using modules with known target DNA sequences such as the zinc finger (ZF) domains. This strategy was applied to generate a novel biosensor conferring maltose-regulated gene expression by fusing a programmable DNA-binding motif (zinc finger module) with a model ligand-binding protein (maltose-binding protein) [194].
Furthermore, random and saturation mutagenesis have shown to be promising in changing the specificity of TF-based biosensors. For example, the approach has been demonstrated in the engineering of the E. coli regulatory protein AraC for l-arabinose to activate gene expression in response to triacetic acid lactone [46, 159], or ectoine [16]. In another example, the homology modelling of protein, pobR, which shows high specificity for 4-hydroxy benzoate, led to the identification of key residues for creating 3,4-dihydroxy benzoate response [65]. Through the use of a targeted mutagenesis library, a final mutant was identified with significant induction by 3,4-dihydroxy benzoate. Similarly, variants of qacR were generated by computational protein design and screened for responsiveness to a new targeted effector vanillin [25]. The screen yielded two qacR mutants that respond to vanillin both in vitro and in vivo.
Förster resonance energy transfer (FRET)-based biosensors
Förster resonance energy transfer (FRET)-based genetically encoded biosensors are also used to sense in vivo metabolites. Usually, FRET sensors comprise an internal metabolite-binding protein (MBP) flanked by a pair of donor and acceptor fluorescent proteins (FPs) (Fig. 1b). When MBP binds the metabolite of interest, there is a conformational change to alter the intramolecular distance of the donor and acceptor FPs, where the emission from the donor FP excites the acceptor FP, giving a measurable radiometric index of a metabolite concentration. Cyan fluorescent protein (CFP) and yellow fluorescent protein (YFP) are often used as the pair of donor and acceptor FPs. Compared to TF-based biosensors, FRET sensors have a faster signal response by avoiding the time lag of TF-based biosensors between the induction of gene expression and the signal. Due to this ability to monitor molecules in real time, as shown in Table 1, FRET sensors have been widely applied in the studies of monitoring intracellular metabolite dynamics, or protein interaction and signal transduction [112]. However, FRET sensors are not used in screening due to their relatively low absolute signals, quantified as emission ratios of FRET acceptor over donor.
Over the years, FRET sensors have been developed to sense a broad range of metabolites, such as sugars [124], amino acids [3, 73, 110, 111, 154], pyruvate [140], lactate [103], and ions [26, 58], and stresses in the environment, such as redox status [120]. The key to constructing FRET sensors is the identification of appropriate MBPs that bind the target products. The reported vast number of metabolite-binding protein scaffolds in the presence and absence of the targets, such as TFs, periplasmic-binding proteins (PBPs), membrane proteins, and other types of ligand-sensing domain, can serve as important resources. For example, bacterial transcriptional proteins can be used as MBPs. A pyruvate sensor named Pyronic was constructed by flanking transcriptional regulator PdhR from E. coli with the FPs mTFP and Venus, at the N-terminus and C-terminus, respectively [140]. Exposure of Pyronic to increasing concentrations of pyruvate showed that it can respond to pyruvate between 10 μM to 1 mM. This enabled high temporal resolution of pyruvate at single cell level in live mammalian cells. Similarly, the lactate-binding TF LldR fused between mTFP and Venus at N- and C-termini, named as Laconic, was capable of quantifying lactate levels from 1 to 10 mM [103]. Beyond TFs, periplasmic-binding proteins have also been used to engineer FRET biosensors [53]. LivK, a bacterial periplasmic-binding protein, was sandwiched by CFP and YFP and expressed in E. coli and yeast, which allowed the quantification of leucine between 8 μM and 1 mM [110]. In a similar way, a lysine sensor was recently designed and developed using the lysine-binding periplasmic protein (LAO) from Salmonellaenterica serovar typhimurium LT2 strain as the binding element for the FRET sensor [3]. The developed sensor was found to be specific to lysine and can work in bacterial as well as yeast cell. Furthermore, membrane proteins also contributed the construction of FRET sensors. For example, ammonium transceptors, which are membrane proteins that possess both solute transport and receptor-like signaling activities, were applied to ammonium sensing [26]. Addition of ammonium to yeast cells expressing the sensors triggered concentration-dependent fluorescence intensity (FI) changes.
Two-component system (TCS) biosensor
TCSs allow microbes to sense environmental changes or target metabolites. The previous studies have demonstrated that TCS could be designed and engineered as in vivo biosensors for different applications [131]. Typically, a TCS system includes a histidine kinase (HK) and a response regulator (RR) (Fig. 1c). The presence of environmental changes or target metabolites induces autophosphorylation of HK, followed by phosphoryl transfer to the RR. The phosphorylated RR further regulates the expression of a reporter gene. HK is a transmembrane protein. The C-terminal autokinase domain of HK is usually in the cytosol, and its N-terminal domain for stimulus sensing could be localized in the extracytoplasmic compartment (periplasm, inner or outer membrane, or even extracellular space). The localization of the N-terminal sensor domain enables the TCS sensor to sense metabolites in different cytoplasm or extracellular space, which is not possible for many other sensors if the metabolite is membrane-impermeable. Examples are provided in Table 1.
The HK domains of natural TCS, which have a variety of signal recognition capabilities, could be adopted to construct a broad range of TCS sensors. In E. coli, an HydHG (also known as ZraSR) TCS was found to have a response to zinc and lead [81]. The HydHG TCS was subsequently engineered to respond to different zinc concentrations, with the threshold level reduced via the introduction of a self-activation loop and elevated by genomic integration [126]. Likewise, the same TCS also contributed to the development of a lead sensing and removal system [67].
It is reported that the chimeric HK strategy can be employed for the development of powerful TCS sensors, in which the sensory domain (N-terminal) and catalytic domain (C-terminal) from different HKs are combined. Because the EnvZ–ompR complex in E. coli is a well-studied TCS that is widely distributed in bacteria, the sensory and catalytic domains of EnvZ are commonly used in domain swapping strategies. For example, E. coli, which does not have the methanol sensing apparatus, was engineered to sense methanol by employing chimeric HK strategy [145]. The TCS biosensor was constructed by fusing the methanol sensing domain of Paracoccus denitrificans FlhS with the catalytic domain of E. coli EnvZ. The constructed chimeric FlhS/EnvZ TCS enabled the cell to sense methanol by the expression of ompC and GFP (green fluorescent protein) gene regulated by ompC promoter. Similarly, a chimeric DcuS/EnvZ (DcuSZ) TCS was constructed by fusing the sensor HK of DcuS with the cytoplasmic catalytic domain of EnvZ [48]. The output signals produced by the chimeric DcuSZ TCS were enough to detect fumarate concentration quantitatively. Another demonstration of the chimeric strategy is the development of malate TCS sensor based on chimeric MalK/EnvZ [49], which was created by fusing the sensor domain of MalK (Bacillus subtilis) to the catalytic domain of EnvZ. In the same manner, a chimeric AauS/EnvZ TCS was built by integration of the sensing domain of AauS (Pseudomonas putida) with the catalytic domain of EnvZ to control the expression of the ompC gene in response to acidic amino acids [132].
G-protein-coupled receptors (GPCRs)-based biosensors
The GPCRs’ system (also known as seven transmembrane receptors, 7TMRs) is a class of transmembrane receptors that tranduces various extracellular signals into intracellular responses, making itself an attractive candidate for engineering and development of in vivo biosensors [109]. As shown in Fig. 1d, when GPCRs are activated by extracellular signals, the GPCR would couple to a G-protein composed of α-, β-, and γ-subunits, dissociating the heterodimer of β-, and γ-subunits from the α-subunit and activating downstream effectors. Similar to other protein-based biosensors, the versatile nature of GPCRs makes them broadly useful for metabolite sensing (Table 1). However, GPCR-based biosensors are usually limited to extracellular sensing.
A recent study was reported to construct medium-chain fatty acid biosensors using GPCRs known to bind fatty acids in mammalian (OR1G1 and GPR40) [113]. One of the sensors detected even-chain C8–C12 fatty acids with dynamic range up to 17-fold increase after activation. Introduction of a synthetic response unit gave a further improvement in the response to decanoic acid using the GPCR OR1G1. Later, the same group swapped OR1G1 with GPCRs known to bind serotonin, and kept the rest of the sensor cell intact, which resulted in a serotonin sensor [38]. Among the six known human serotonin GPCRs, 5-HT4-based sensor performed better, and the sensor was also demonstrated to work in the 96-well plate format with reliable statistical parameters. In another example, using the odorant receptors specific for the volatile fragrance compounds, a series of GPCR-based sensors were constructed to sense the volatile fragrance compounds eugenol, coumarin, dihydrojasmone, and acetophenone [114], upon which the authors designed a synthetic multi-layered gaseous-fragrance-programmable analog-to-digital converter (ADC). Similarly, a GPCR-based pH sensor was created to sense pH value with the help of the human proton-activated cell-surface receptor TDAG8 [6]. Besides organic acids, this sensor could also be adjusted by CO2 that shifts the CO2-bicarbonate balance toward hydrogen ions.
For GPCRs, the structure-based drug design of receptors received limited success until recently due to the low progress in determination of GPCR crystal structures [69]. Currently, the high-resolution crystal structures of GPCRs enabled the ligand discovery for several pharmaceutically relevant receptors [75, 79]. However, the structure-based approach has not been applied in the construction of GPCR biosensors.
Enzyme-coupled biosensors
Another kind of protein-based in vivo biosensor relies on enzyme-coupled reactions. This biosensor takes advantage of enzyme-catalysed reactions that can convert a target metabolite into detectable chromogenic and fluorogenic substances (Fig. 1e). The development of this sensor requires a priori knowledge of relevant reactions. Examples of these biosensors can be found in Table 1.
In some cases, the required reactions can be accomplished in one-step. For example, an enzyme-coupled biosensor for l-3,4-dihydroxyphenylalanine (l-DOPA) was developed by converting l-DOPA into the yellow, fluorescent pigment betaxanthin with the help of DOPA dioxygenase [29]. Using this sensor, a tyrosine hydroxylase with 2.8-fold improved activity was identified, which was able to further improve the yields of l-DOPA. In another study, a biosensor for free CoA-SH was developed through a coupled reaction by monitoring NADH production with α-ketoglutarate dehydrogenase (α-KGDH), which can be quantified by absorbance at 340 nm [86].
In other cases, a cascade of reactions was required to form a detectable molecule. As an example, xylose or lactate can be detected via enzyme-coupled biosensors. It was shown that pyranose oxidase and lactate oxidase would oxidize xylose and lactate, respectively, with concomitant production of byproduct, hydrogen peroxide (H2O2). The H2O2 byproduct was subsequently used to oxidize Amplex UltraRed by horseradish peroxidase, leading to the generation of the detectable, fluorescent compound resorufin [171]. In principle, the assay described in this work allowed for use of any oxidase enzyme, which could be considered as a generalized method to construct enzyme-coupled biosensors for other products of interest. In a similar example, an enzyme-coupled biosensor for succinic acid was established using fumarate reductase and peroxidase [157]. In this reaction, fumarate reductase can catalyse succinic acid to fumarate and produce H2O2, which can be further detected by the mentioned peroxidase-dependent fluorescent assay.
To create enzyme-coupled biosensors, there are also several cases combining the use of enzymatic and chemical reactions. Recently, an enzymatic assay for cytidine was developed by combining cytidine deaminase (CDA) and the indophenol method [33]. CDA catalysed the cleavage of cytidine to uridine and NH3, the latter of which can be accurately determined using the indophenol method. The sensor enabled a sensitive and convenient assay for cytidine detection, which was used to screen cytidine producing strains. The strategy was also adopted by the same group to construct a biosensor for adenosine [34].
Nucleic acid-based biosensors
RNA-based biosensors
Nucleic acid-based sensors work as an alternative to protein-based biosensing systems. Compared to protein-based sensors, nucleic acid-based in vivo biosensors are usually specific to a limited number of metabolites, but show advantages such as faster response time and decreased metabolic burden on cells [89]. Furthermore, nucleic acid-based biosensors can, in principle, be combined as tandem copies, leading to a tighter regulation of gene expression.
There are two popular kinds of nucleic acid-based biosensors, which are based on RNA or DNA. In general, RNA riboswitch is a genetic control element of a messenger RNA molecule that binds a small molecule, resulting in a change in production of the protein encoded by the mRNA. RNA riboswitch can selectively bind to various metabolites (via RNA aptamer domains) and change its own structure, which thereby modulates the transcription of a measurable readout such as fluorescence, enzyme activity, and cell growth (Fig. 2a). Aptazyme is a variation of the RNA riboswitch, in which the output domain is a hammerhead ribozyme whose activity depends on aptamer binding status where ligand binding induces the RNA to cleave itself (Fig. 2a). RNA riboswitches have been engineered as in vivo biosensors responsive to target molecules of interest [137, 198]. Currently, as shown in Table 1, most of the RNA-based biosensors have been reported in bacterial systems, and few could be found in yeast, due to the fact that the majority of riboswitches discovered in bacteria are non-functional in yeast.
Schematic of nucleic acid-based sensors. a RNA-based biosensors. Left: binding ligand results in changes in mRNA secondary structure and renders the ribosome binding site (RBS) inaccessible to ribosome; Middle: in the absence of ligand, the intrinsic terminator stops transcription, while binding ligand eliminates the terminator structure; Right: binding ligand activates the hammerhead ribozyme and triggers self-cleavage, which subsequently leads to mRNA degradation. b DNA-based biosensors. Binding of target metabolites to the promoter enables the expression of reporter gene
Aptamers are the sensing elements of the riboswitch construct. Metabolite-binding RNA aptamers targeting small molecules of interest can be mined from natural metabolite-regulation systems. For example, the lysine riboswitch is one natural riboswitch. Based on it, genetic RNA biosensors were constructed to monitor changes in the intracellular concentration of lysine [173, 190, 200, 201]. Furthermore, the construction of theophylline sensors using the well-characterized theophylline aptamer as the sensor element has also been demonstrated [37, 96, 119, 169]. Ribozymes have been developed as in vivo biosensors for N-acetylneuraminate [191] and glucosamine 6-phosphate [77], while the Spinach RNA aptamer, which was described to emit green fluorescence on binding with 3,5-difluoro-4-hydroxybenzylidene imidazolinone (DFHBI) [122], was applied in the development of thiamine 5′-pyrophosphate, [193] and S-adenosyl-l-homocysteine [155] biosensors. These examples will be further elaborated in the application section. On the other hand, it was reported that riboswitches were found for specific heavy metal ions using comparative sequence analysis to search bacterial genomes and metagenomic sequences [47]. This study finally demonstrated and validated the discovery of riboswitch RNAs to sense Ni2+ and Co2+. In yeast, an RNA sensor was established in response to neomycin by attaching a synthetic neomycin aptamer to the catalytic core of the type III hammerhead ribozyme (HHR) from Schistosoma mansoni [72]. These innovations enabled the development of neomycin-dependent RNA modules that switch gene expression up to 25-fold. It can also be used as a general strategy to create novel yeast synthetic riboswitches by combining an in vitro selected synthetic aptamer to the catalytic core of ribozyme such as HHR.
Metabolite-binding RNA aptamers can also be generated towards a ligand of choice by screening a pool of synthetic aptamer libraries. In particular, in the past 20 years, the systematic evolution of ligands by exponential enrichment (SELEX) and its enhancements, such as High-Fidelity (Hi-Fi) SELEX, semi-automated SELEX, capillary electrophoresis (CE) SELEX, capture-SELEX, and branched SELEX, have yielded new aptamers that respond to a plethora of compounds [55]. The traditional SELEX can select aptamers using iterative cycles of binding, partition, and amplification. In a recent example, a single-stranded RNA was isolated that can respond to naringenin from combinatorially prepared nucleic acid libraries using SELEX [64, 185]. The isolated riboswitch contributed to the construction of a naringenin sensor, which displayed an increased fluorescent signal generation after activation upon binding with naringenin. In another study, a new platform, Hi-Fi SELEX, was reported that introduced novel fixed-region blocking elements to improve the functional diversity of the starting library [121]. Then, the improvement of Hi-Fi SELEX to aptamer selections was demonstrated using human α-thrombin as the target. Besides, to shorten the time of selection in SELEX, capillary electrophoresis (CE) was reported to be more efficient for the separation of bound and unbound DNA/RNA. CE–SELEX has been reported to select DNA aptamers for large-molecule targets, such as cancer biomarker HE4 protein [36], and small-molecule targets, such as N-methyl mesoporphyrin IX (NMM) [189]. These studies demonstrated that CE-SELEX reduced the selection time and the number of cycles. Later, a fraction collection approach was developed in capillary electrophoresis SELEX (FCE-SELEX) for the partition of a bound DNA-target complex [95]. By integrating fraction collection with a facile oil seal method, in a single round of selection, a streptavidin-binding aptamer (SBA) was generated. There are also other emerged solutions to detect the binding of a target molecule to an RNA. In two recent studies, it was demonstrated that select riboswitch readout domains from the existing riboswitches, called expression platforms, can be engineered using simple design rules to host natural or synthetic aptamers to create novel chimeric RNAs that regulate transcription both in vitro and in vivo [14, 15]. In another example, the aptamer domain of a known switch that regulates expression of the marker TetA linked to GFP was replaced with 40 random bases [204], and then, the library was transformed into E. coli. The dualistic nature of TetA allowed the application of dual genetic selection to identify riboswitches that expressed TetA-GFP only in response to bisphenol A. Furthermore, structure information could be used to help the screening. For example, structure-guided chemical genetic screening of the natural adenine-binding add A aptamer was performed, and two additional ligands [pyrimido(4,5-d)pyrimidine-2,4-diamine and 2-aminopyrimido(4,5-d)pyrimidin-4(3H)-one] were identified that bound to the aptamer with high affinity, exhibiting improved in vivo gene-induction properties and reduced cellular toxicity [134]. The same group also proposed targeted mutagenesis of native riboswitches to change their specificity toward rationally designed synthetic ligand analogs [133]. In parallel, a novel bioinformatics-guided selection and screening strategy was used to rapidly isolate aptamers against two different targets: the human interleukin (IL)-10 and human 4-1BB receptors [82].
Currently, only a limited number of RNA aptamers have transitioned into effective and widely used intracellular biosensors, and many do not reliably function in a cellular context. To solve this problem, it was proposed that recurrent structural folds found in natural RNA aptamers and small nucleolytic ribozymes can be reprogrammed to host a broad spectrum of small molecule-binding sites while preserving the robust folding and highly stable architectural properties of the parent [127]. Using scaffolds derived from two different riboswitch aptamer domains and a ribozyme, a diverse set of aptamers were obtained to recognize 5-hydroxytryptophan, serotonin, and dopamine. Recently, computational tools were developed to rationally design riboswitches using the existing aptamers [55, 169]. A statistical thermodynamic model was developed to predict the sequence–structure–function relationship for translation-regulating riboswitches [4]. Using the model, automated computational design of 62 synthetic riboswitches was carried out using six different RNA aptamers to sense diverse chemicals (theophylline, tetramethylrosamine, fluoride, dopamine, thyroxine, and 2,4-dinitrotoluene). Moreover, the development of fluorescent biosensors that exhibited a response to (2′-5′,3′-5′) cyclic guanosine monophosphate-adenosine monophosphate (2′,3′-cGAMP) was reported [10]. The biosensor was designed by making rational mutations to natural riboswitch aptamers that recognize the related molecule 3′,3′-cyclic di-GMP. In another example, a fluorescent biosensor for c-di-GMP was first generated by fusing a natural GEMM-I (genes for the environment, membranes, and motility) riboswitch aptamer to Spinach [71]. Then, rational mutations to the ligand-binding pocket guided by structure were exploited to alter specificity of the RNA-based biosensor to c-AMP-GMP. Later, the same group constructed a series of improved biosensors that were up to 450% brighter and 13 times faster, which together enabled detection of c-di-GMP from picomolar to micromolar concentrations [175].
DNA-based biosensors
DNA can also serve to detect molecules in vivo. In nature, cells are evolved with responsive promoters to sense stress or metabolites. Like other sensors, DNA-based sensors are also comprised of different genetic elements: a sensing element (metabolite-responsive promoters) and a reporter gene (Fig. 2b). Identification of the appropriate promoter is usually considered as the most crucial step. Examples for different DNA-based sensors are provided in Table 1.
Currently, many inducible promoters have been reported for the inducer molecule, such as isopropyl-beta-d-thiogalactopyranoside (IPTG) [1], arabinose [116], lactose [87], and tetracycline (Tc)-regulated gene expression system [107]. Meanwhile, many promoters that respond to target molecules could be mined from the published literature. The devices lysGE, pN, and pA from C. glutamicum ATCC13032 were reported to be capable of responding to lysine [9]. When these devices were transferred to E. coli, the promoter pA responded to the change of extracellular lysine concentration [176]. Metallothioneins (MTs) are metal-binding proteins involved in cellular protection from metal toxicity. After investigating promoters that drove the expression of MTs, the CdMT promoter was found to have a good response to cadmium ions [2]. In another example, by mining literature, sucrose responsive promoters were identified from selected candidates which were repressed during growth on glucose, and upregulated during growth on sucrose [179].
When intermediate metabolites accumulate to toxic levels, one of the native stress responses is the change of promoter activity, which allowed the identification of novel metabolite-responsive promoters. For example, promoters that responded to farnesyl pyrophosphate (FPP) were identified by whole-genome transcriptome analysis in E. coli [22]. In Aspergillus niger, a low pH-inducible promoter, Pgas, was also identified by transcriptional analysis [192]. The strength of Pgas was independent of acid type and acid ion concentration, showing dependence on pH only. Similarly, application of transcriptome analysis was also applied to identify responsive promoters to 1-butanol [147] and 1,4-butanediol (1,4-BDO) [158]. Moreover, novel threonine promoters were discovered upon the proteomic analysis of E. coli in response to extracellular threonine [93]. In parallel, a phenylalanine-responsive promoter was screened from a promoter library [100].
Rational design and engineering of responsive promoters were also reported. Recently, a set of promoters that were inducible under acidic conditions were selected from literature searches and data mining [128]. By engineered the upstream activating sequence, the YGP1 promoter was successfully engineered for improved performance at low pH. The engineering strategy outlined for the YGP1 promoter was subsequently used to build low pH induction into native promoters lacking this response.
Applications of in vivo biosensors in synthetic biology and metabolic engineering
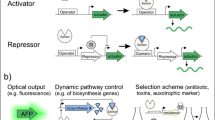
In addition to the examples described above, there are many advanced applications of the in vivo biosensors, such as dynamic regulation of pathways, optimizing the product titers, enhancing substrate utilization and precursor availability, isolation and improvement of desired enzymes, as well as in vivo monitoring of the target compounds (Table 1). We will discuss these applications in detail below. Applications in each category will be broadly grouped according to biosensor types.
Dynamic regulation
Balanced utilization of cellular resources, in particular the control of metabolic intermediate fluxes and biosynthetic pathway genes, is important for the production of value-added chemicals in microorganisms. Despite recent advances in synthetic biology, most control systems for heterologous pathways still rely on constitutive or inducible promoters, for which the outputs are optimized for a particular condition and are static in nature, unable to respond to growth and environmental changes [22]. Moreover, these static control systems may not be suitable when piecing together a complicated pathway with biosynthetic modules with mismatched input/output levels, or when there is a need to minimize the accumulation of potentially toxic intermediates [60]. In contrast, dynamic regulation enables microorganisms to adapt to changes in their internal or external conditions. A dynamic regulation system consists of a sensing component, which can detect the metabolite of interest or physiological state, and a regulator component, which converts the sensor signal to into a transcriptional signal, often resulting in the upregulation or downregulation of a key pathway gene [196]. The ability to autonomously regulate gene expression and, consequently, metabolic flux through the biosynthetic pathways in response to its environment enables cells to utilize resources efficiently and improve productivity, and also minimize the need for chemical inducers and human intervention of fermentations. Dynamic regulation has been hailed as a new frontier in metabolic engineering [12, 104, 166]. Since the one of the first demonstrations of the dynamic flux control to improve yield and productivity of a heterologous pathways by Farmer and Liao in 2000 [40], the number of dynamic regulation systems has grown, particularly in the past 5 years. Here, we present recent applications of riboswitch-, transcription factor-, and promoter-based biosensors.
Lysine-responsive riboswitches from E. coli and B. subtilis were used to control competing but essential metabolic by-pathways of lysine biosynthesis in Corynebacterium glutamicum [201]. Citrate synthetase (gltA) gene encoding the first enzyme of the tricarboxylic acid (TCA) cycle was placed under the control of the lysine-OFF riboswitches, which repress translation initiation in the presence of lysine, thus repressing TCA cycle activity. This allowed metabolic fluxes to be directed towards lysine biosynthesis instead of cell growth, and enhanced lysine yield by up to 63% in the resulting LPECRS strain. To expand the C. glutamicum engineering toolbox and explore the potential synergistic effects of a combined ON–OFF control, a lysine-ON riboswitch was developed by screening a riboswitch library with randomized sequences between the lysine aptamer and the ribosomal-binding site using the tetA-based dual selection system [200]. A lysine-ON riboswitch, which upregulated the expression of the downstream gene by more than 18-fold in the presence of 0.1 mM lysine, was identified from the screening. The lysine-ON riboswitch was then introduced into the LPECRS strain [201] to control the expression of lysE, a gene encoding a lysine transporter protein. The lysine production yield of the resulting strain with the dual lysine-OFF and -ON riboswitches was enhanced by 21% over the parent LPECRS strain.
Chou et al. developed the feedback-regulated evolution of phenotype (FREP) system that regulated the mutation rate of the cell population inversely to the target metabolite concentration [20]. Metabolite sensors were assembled from natural or synthetic metabolite-responsive TF to gauge the concentration of the target metabolite and regulate the mutator mutD5 expression. Under low metabolite concentration, mutation rate was increased to generate greater diversity in the cell population. Conversely, mutation rate was decreased in the presence of high metabolite concentration, thereby fixing the genotype of the cell population to the high production phenotype. As proof-of-concept, FREP was used to evolve E. coli strains for increased production of tyrosine and isoprenoid.
Malonyl-CoA is an important building block for a variety of biofuels, fine chemicals, and pharmaceuticals, and is the rate-limiting precursor in the chain elongation reaction during the synthesis of these compounds [42, 45, 187]. Hence, optimizing metabolic flux control through the malonyl-CoA node has also been the subject of numerous microbial cell factory engineering efforts. Central to the development of malonyl-CoA-responsive biosensors for regulating the malonyl-CoA pathways is the transcription factor, FapR, from B. subtilis [144]. FapR binds to its DNA operator sequence in the absence of malonyl-CoA to repress the downstream gene, and upon binding of malonyl-CoA, changes its conformation and releases from the operator sequence to allow transcription of the downstream gene [143]. FapR and its cognate operator sequence were transplanted into E. coli to build a malonyl-CoA sensor [188], and the sensor was applied in a feedback regulatory circuit to regulate fatty acid synthetic pathway in E. coli, improving the productivity and titer by 33 and 34%, respectively [90]. Discovery of the PGAP promoter in E. coli, which is positively regulated by FapR in the absence of malonyl-CoA, enabled the creation of a feed-forward regulatory circuit that activates fatty acid synthesis in the presence of high malonyl-CoA concentrations [186]. When implemented together with the feedback circuit, the malonyl-CoA regulatory system improved fatty acid synthesis by more than twofold compared to the unregulated E. coli strain. In a hierarchical dynamic control system developed by David et al. for improved production of 3-hydroxy propionic acid (3-HP) in S. cerevisiae [24], malonyl-CoA was channeled to fatty acid synthesis in the growth phase by subjecting the fatty acid synthase gene to the control of the glucose-inducible PHXT1 promoter. In the fermentation stage, the fatty acid synthesis will be downregulated due to glucose limitation, and the malonyl-CoA accumulated will be directed towards 3-HP production by FapR which upregulates the expression of the malonyl-CoA reductase gene.
A dynamic sensor-regulator system (DSRS) based on the fatty acid/acyl-CoA-responsive TF, FadR, was developed to regulate the expression of genes involved in the production of fatty acid-based products [196]. The FadR sensor was used to regulate the pyruvate decarboxylase and alcohol dehydrogenase genes for ethanol production, as well as the acyl-CoA synthase and wax–ester synthase genes in response to fatty acid accumulation, achieving 1.5 g/L of fatty acid ethyl ester titer and a threefold improvement in theoretical yield in E. coli over the non-dynamically regulated strain, as well as increased strain stability.
The competition for cellular resources between native and heterologous pathways in a production host often results in poor cell growth and productivity [78]. Other than using glucose-responsive promoters such as PHXT1 as sensors to initiate the redirection of metabolic flux after the growth phase, quorum sensing (QS) systems, which enable bacterial cells to detect local cell population density via extracellular signaling molecules and regulate gene expression in a coordinated manner in response [108], have emerged as an attractive method of separating growth from production phases in industrial microorganisms. In recent years, QS circuits have been applied in the dynamic redirection of metabolic flux from the central metabolic pathways in the growth phase to heterologous pathways for the production of isopropanol and 1,4-butanediol in E. coli [91, 153], and para-hydroxybenzoic acid in S. cerevisiae [178]. A tunable pathway-independent genetic control module based on the Esa QS system from Pantoea stewartii was engineered to regulate target genes in E. coli. In this system, the trigger to redirect glycolytic flux into heterologous pathways can be optimized for individual systems by controlling the strength of the promoter–ribosomal-binding site driving the expression of 3-oxohexanoylhomoserine lactone synthase, EsaI [54]. This module was applied to improve myo-inositol titers by 5.5-fold, and increase titers of glucaric acid and shikimate from undetectable to more than 0.8 g/L and 100 mg/L, respectively.
Farnesyl diphosphate (FPP) is a common precursor for mevalonate pathway-based isoprenoids, which includes high-value compounds such as drugs (artemisin and taxol), flavors and fragrances, and advanced biofuels. However, FPP accumulation is toxic to common production hosts such as E. coli and S. cerevisiae. Hence, maintaining a high metabolic flux through the FPP node while keeping its intracellular concentration low is important to developing industrial isoprenoid production strains. Creating a dynamic control system to achieve this goal would require a biosensor that can detect and regulate the pathway flux through the FPP node. To address the lack of known natural sensors for FPP, Dahl et al. applied whole-genome transcript arrays to identify promoters that respond to the accumulation of toxic FPP, and applied the promoters to regulate the production of FPP in the isoprenoid biosynthetic pathway in E. coli [22]. Consequently, amorphadiene production in the dynamically regulated E. coli strain was increased twofold over production strains with inducible or constitutive promoters. This strategy circumvents the need for known sensors for metabolites of interest, of which there are very limited numbers, and it also eliminated the need for expensive inducers. In another study targeting the branch point at the FPP node, a set of ergosterol-repressed promoters from the yeast native ergosterol biosynthetic pathway were used to downregulate the expression of ERG9 when ergosterol was in excess [195]. This feedback regulation of ERG9 enhanced the metabolic flux towards the production of a non-native isoprenoid, amorpha-4,11-diene, and improved its titer by up to fivefold in yeast. Using glucose-responsive promoters as sensors, Xie et al. developed a sequential control strategy to dynamically control the downstream, upstream, and competitive pathways from the FPP node of the carotenoid production yeast strain [184]. In this system, a glucose-repressed-modified GAL regulation system was used to regulate tHMG1, the rate-limiting gene of the upstream mevalonate (MVA) pathway, and the beta-carotene biosynthetic pathway, while the ERG9 gene, encoding squalene synthase which commits FPP to the competing ergosterol biosynthetic pathway was placed under the control of a glucose-inducible promoter, PHXT1. In the growth stage, the ergosterol synthetic pathway, which is essential for cell viability, was expressed under high glucose concentration to sustain cell growth. In the second stage, under glucose limitation, tHMG and the beta-carotene pathway genes were induced, while the squalene pathway was downregulated, maximizing carotenoid production. Subsequently, a titre of 1165 mg/L carotenoids was achieved using this inducer-free strategy in a fed-batch fermentation.
In other promoter-based dynamic regulation applications, a low pH-inducible promoter, Pgas, was identified using transcriptome analysis for the dynamic control of Aspergillus niger for the itaconic acid production [192]. Placing the cis-aconitate decarboxylase (CAD) gene under the control of the low pH-inducible promoter, Pgas, enhanced itaconic acid titer in A. niger by more than fivefold compared to the strain with CAD placed under a strong constitutive promoter, PgpdA.
Improving product titer and productivity
Naturally occurring organisms are seldom optimized to produce industrially relevant molecules, and extensive modifications to the cells are often required to achieve economically viable production. To this end, directed genome evolution approaches, such as multiplex automated genome engineering (MAGE) [172], trackable multiplex recombineering (TRMR) [141], yeast oligo-mediated genome engineering (YOGE) [31], the heritable recombination system [138], and RNAi-assisted genome evolution (RAGE) [148], are highly efficient for strain development, while combinatorial pathway optimization methods such as customized optimization of metabolic pathways by combinatorial transcriptional engineering (COMPACTER) [35] have been shown to optimize the expression of pathway genes. However, our ability to create the genetic diversity using these methods far outstrips the throughput of current analytical methods. Biosensors provide an alternative to bridge this gap by converting hard-to-quantify phenotypes, for example accumulation of non-colorimetric small molecules, to easily measurable parameters such as fluorescence and growth, thereby providing a quantifiable link between genotype and phenotype amendable for high-throughput assays. Here, we present some of the recent applications of the various forms of biosensors to address the high-throughput screening bottleneck.
Driven by an l-valine responsive sensor based on the Lrp transcription regulator from C. glutamicum, the l-valine production in C. glutamicum was improved by 25%, and byproduct formation was reduced by more than threefold through adaptive laboratory evolution [99]. The alcohol-responsive BmoR transcriptional activator from Pseudomonas butanova was engineered to a 1-butanol biosensor regulating tetA expression, and enabled the isolation of an E. coli strain with 35% improvement in 1-butanol-specific productivity through optimization of pathway enzyme expression [32].
Using a multi-parametric screening approach, a functional design for biosensors based on LysR-type transcriptional regulators (LTTRs) in S. cerevisiae was developed [151]. Notably, this approach enabled the direct transplantation of small molecule-binding prokaryotic transcriptional activators and their cognate DNA operators into S. cerevisiae to create metabolite biosensors without the need for further engineering. As proof-of-principle, the CCM and naringenin biosensors developed by this approach were applied for in vivo screening of cells, and successfully identified strains with improved product titers.
Liang et al. developed a strategy termed sensor-assisted transcriptional regulator engineering (SATRE) based on the mevalonate-responsive biosensor to improve acetyl-CoA flux from methanol to mevalonate in the methylotrophic cell factory, Methylobacterium extorquens AM1 [84]. Two million variants from a QscR transcriptional regulator library were screened in one round by flow cytometry using the mevalonate biosensor-based system, yielding a mutant strain (Q49) with 60% increase in mevalonate concentration, and capable of producing 2.67 g/L of mevalonate in a fed-batch fermentation.
To eliminate the amplification of unproductive cheater populations during selection process, a toggled selection scheme was developed in E. coli based on the tolC selector gene regulated by a naringenin-responsive TF, TtgR [129]. The selection scheme switches between positive selection using sodium docedyl-sulfate resistance conferred to the host by the activation of tolC expression in a naringenin overproducer and negative selection using colicin E1 to eliminate any evolutionary escapees with constitutive tolC expression. In four rounds of evolution targeting the expression of key pathway genes using MAGE [172], naringenin production from glucose in E. coli was increased by 36-fold to reach 61 mg/L.
In addition to screening pathway optimization libraries, biosensors can also be applied to maintain high productivity in fermentation. By putting an auxotrophic selection marker under the control free fatty acid (FFA)-responsive transcription factor, FadR, an isogenic E. coli population is continuously enriched for non-genetic variants with enhanced FFA production when grown under selection pressure [181]. When this strategy, termed in vivo population quality control (PopQC), was applied to a fed-batch fermentation of an FFA overproducing strain, a titer of 21.5 g/L, which was fourfold enhanced over the non-selective fermentation condition, was achieved.
A naringenin-responsive riboswitch based on a novel synthetic aptamer was developed to activate GFP expression upon naringenin binding [185]. This riboswitch biosensor successfully identified naringenin overproducing E. coli strains in co-culture through flow cytometry-based high-throughput screening. The theophylline riboswitch [30] regulating the tetracycline resistance gene, tetA, was used to facilitate the selection of a combinatorial library consisting of differing promoters, RBS, origins of replication, and chaperone proteins influencing the expression of the caffeine demethylase enzyme, which converts caffeine to theophylline, for improved theophylline production in E. coli [37]. The negative Ni2+-selection system based on a self-cleaving NeuAc (N-acetylneuraminic acid)-responsive ribozyme regulating the tetA gene was used to select for improved NeuAc overproducing E. coli strains from combinatorial RBS and randomly mutagenized NeuAc synthase gene libraries [191]. This selection system improved the titer of the NeuAc biosynthetic pathway, which was previously recalcitrant to rational metabolic engineering, by up to 34%, with the final strain achieving a titer of 8.31 g/L NeuAc. Using a similar Ni2+-selection, a lysine riboswitch which represses the expression of the downstream tetA gene upon binding lysine was used to screen the promoter libraries of the phosphoenolpyruvate carboxylase for improved lysine producing E. coli strains in the selection media, yielding a variant with significantly improved lysine titer (0.6 g/L) [190]. Based on a putative E. coli FMN riboswitch, Meyer et al. engineered an FMN-responsive ribozyme, which self-cleaves in the presence of FMN to activate the expression of GFP, to screen for B. subtilis variants with improved vitamin B12 production [106]. In this workflow, engineered B. subtilis cells, which convert cellobiose to vitamin B2, were co-confined with E. coli sensor cells harboring the FMN ribozyme sensors in nanolitre reactors. Vitamin B2 produced by B. subtilis will be taken up by E. coli sensor cells and converted to FMN, which then activates GFP fluorescence. Using E. coli sensor cells and fluorescence-activated cell sorting (FACS), B. subtilis variants with elevated cellobiose-to-vitamin B12 conversion efficiency were effectively isolated from a genome shuffling library.
Aptamers are often applied as biosensors to regulate gene expression in prokaryotes by exposing the RBS of the regulated gene in the presence of the target ligand, hence upregulating expression [30, 117, 156, 163, 168, 191]. However, it is difficult to replicate this strategy in S. cerevisiae because of the difference in the eukaryotic translation initiation mechanism. To address this issue, a self-cleaving, glucosamine 6-phosphate (GlcN6P)-responsive ribozyme glmS was integrated into the 3′-untranslated region of the cytosine deaminase (fcy1) gene in S. cerevisiae to create a suicide riboswitch for evolutionary engineering [77]. The presence of GlcN6P activates glmS self-cleavage and destabilizes the fcy1 mRNA, making the S. cerevisiae host less susceptible to the toxic conversion of fluorocytosine to fluorouracil by FCY1. Using this selection system, improved mutants of the glutamine–fructose-6-phosphate transaminase and haloacid dehalogenase-like phosphatase enzymes in the N-acetyl glucosamine (GlcNAc) production pathway were identified, and an improved GlcNAc producer strain capable of producing more than 3 mg GlcNAc/g dry cell weight was isolated.
A biosensor composed of a lysine-responsive promoter regulating the expression of GFP was used to select for randomly mutagenized lysine producing E. coli strains using FACS, yielding a mutant strain with 21% titer improvement [176]. Liu et al. engineered a selection circuit to optimize l-tryptophan production [92]. In this circuit, the L-Trp biosensor, based on the nascent leader peptide, tnaC, of the tnaCAB operon, regulates the expression of the maltase (malQ) gene, and sustains the growth of E. coli strains which overproduce L-Trp when maltose is introduced as the sole carbon source. This method was successfully used to enrich for L-Trp overproducers from site-specific saturation mutagenesis libraries to genome-wide random mutagenesis libraries.
Improving substrate utilization and production of key precursors
In addition to facilitating direct enhancement of product titer, biosensors have also been applied to improve the substrate utilization and concentration of key precursors in biosynthetic pathways. To improve xylose utilization in S. cerevisiae, a xylose-responsive gene circuit was engineered in S. cerevisiae based on the xylose binding repressor, XylR, from Staphylococcus xylosus [174]. Using FACS-based screening, a sugar transporter HXT14 mutant with 6.5-fold improvement in xylose transportation was identified from a random mutagenesis library. Using the FerC repressor from Sphingobium sp. SYK-6, a biosensor that detects substituted cinnamic acid CoA-esters was developed in E. coli [97]. The biosensor can detect 13 different substituted cinnamic acid-based compounds, which are characteristic products from the enzymatic degradation of lignin, and was applied to screen for efficient feruloyl esterases against different biomass sources.
A biosensor based on the FapR TF enabled the identification of two gene targets from a genome-wide cDNA library that improved intracellular malonyl-CoA concentration in S. cerevisiae and subsequently enhanced 3-HP titer by 120% [83]. L-Trp is the key intermediate in the biosynthesis deoxyviolacein, a useful natural product with a variety of bioactivities. Fang et al. applied the TnaC-based L-Trp biosensor to sequentially improve the upstream module for the production of the L-Trp in E. coli through combinatorial optimization of pathway RBS, and then the downstream module for the conversion of L-Trp to deoxyviolacein through the random mutagenesis of VioB, the rate-limiting enzyme [39]. Using this strategy, termed intermediate-sensor-assisted push–pull strategy, the L-Trp titer from the upstream module was first enhanced by about 100-fold. When coupled with the engineered downstream module, the resultant strain showed a 4.4-fold improvement in deoxyviolacein production with a titer of 1.92 g/L, representing the highest deoxyviolacein production reported.
Isolation and improvement of desired enzymes
As biosensors can be coupled to the expression of fluorescent or selection markers, they have been employed as high-throughput screening or selection assays to identify key genes or directed evolution of single enzymes. The synthetic gene circuit based on the S-adenosylmethionine (SAM)-responsive MetJ transcriptional repressor of E. coli enabled the screening of a genomic library to identify the GAL11 gene as a novel multicopy enhancer of SAM levels in S. cerevisiae [165]. Based on a phenol-responsive TF, DmpR, a genetic circuit capable of detecting a phenol compound was developed to facilitate the rapid discovery of novel enzymes from metagenomic libraries that can act on synthetic phenyl substrates to produce phenol [18].
Isolation of improved N-acetyl-l-glutamate kinase (NAGK) mutants with more than 20-fold less sensitivity toward l-arginine inhibition, which in turn improved the C. glutamicum production of l-arginine, was achieved using the pSenLys-Spc biosensor, based on the LysG TF [142], while an NADPH-responsive biosensor based on the transcriptional regulator SoxR was engineered to facilitate the high-throughput screening of the NADPH-dependent Lactobacillus brevi alcohol dehydrogenase (ADH) variant library for improved activity on the 4-methyl-2-pentanone substrate [149]. In one round of FACS screening, 106 ADH variants from error-prone PCR (epPCR) and active site saturation mutagenesis libraries were screened to yield a variant with 36% improved activity. A single cell sorting system based on the 3,4-dihydroxy benzoate (34DHB)-responsive pcaU transcription factor from Acinetobacter sp. ADP1 was used to screen for dehydroshikimate dehydratase enzymes with improved activity on dehydroshikimate to produce 34DHB [66]. A chimeric aspartate kinase was evolved through random mutagenesis followed by Ni2+-selection using a high-throughput synthetic RNA device which comprises of a lysine-sensing riboswitch regulating the tetA selection module [173].
In vivo monitoring
The ability to observe the real-time formation of metabolites in production strains is desirable as it allows engineers to rapidly evaluate strain phenotypes as well as effects of process parameters [135]. To this end, various biosensors have been developed to monitor the production of value-added chemicals. Using a diacid-responsive transcriptional activator, CdaR, a biosensor for the real-time monitoring of glucarate biosynthesis was developed, and this biosensor was subsequently applied to identify superior enzyme variants [136], while an Lrp-based biosensor was developed for the live cell imaging of l-valine production to monitor single cell productivity and study the phenotypic pattern of microbial production strains [115]. As no natural TF for 3-HP exists, Rogers and Church developed a strategy to quantify 3-HP concentration by intra-cellularly converting 3-HP to acrylate, which can be measured by the acuR transcription factor-based biosensor [135]. Using this method, process parameters leading to 23-fold higher production of 3HP than previously reported was identified. Using a cobalamin ribosensor engineered from a putative riboswitch in Propionibacterium freudenreichii spp. shermanii DSM 20270, a sensitive detection method for vitamin B12 content in fermented foods was developed [203].
A genetically encoded sensor, termed Spinach riboswitch, based on the fusion of natural riboswitches with the Spinach aptamer which binds the DFHBI fluorophore enabled the live imaging of dynamic changes in intracellular TPP concentrations in individual E. coli cells [193]. Using an inverted fusion design strategy, an S-adenosyl-L-homocysteine (SAH) riboswitch was coupled with a modified Spinach2 aptamer to generate an RNA-based fluorescent SAH biosensor capable of monitoring in vivo methylthioadenosine nucleosidase activity, which has potential applications in evaluation of inhibitor drug efficacies [155]. Exploiting the naturally occurring AdoCbl-responsive riboswitch from E. coli, intracellular sensors of coenzyme B12 were engineered to examine the synthesis and the import of coenzyme B12 in E. coli, generating data pertinent to improving the understanding of vitamin B12 biology [44].
FRET-based biosensors are non-invasive and have short response time, making them good candidates for real-time monitoring of cellular processes. An FRET-based biosensor for monitoring trehalose-6-phosphate (T6P) was developed from the fusion of the trehalose repressor (TreR) from E. coli with eCFP and Venus [124]. This sensor enabled the in vivo monitoring of T6P dynamics in E. coli and S. cerevisiae, which is virtually impossible for conventional metabolite analytical methods due to their invasiveness and low sensitivity, and it can potentially be applied to evaluate trehalose pathway-targeting drugs against pathogenic nematodes. In another example, an FRET-based methionine biosensor based on methionine binding protein (MetN) from E. coli between CFP and YFP was developed for the in vivo imaging of methionine flux [111].
In other applications, using the CysR transcription factor from C. glutamicum, which is activated by O-acetyl serine (OAS) and O-acetyl homoserine (OAH)—intermediates of sulfur containing amino acids l-cysteine and l-methionine synthesis, a biosensor was developed to allow the study of consequences of sulfur limitation in single cells such as in pathogens under the conditions that may exist in their hosts [59]. To develop a low cost and equipment-free monitoring method for micronutrient zinc in blood, Watstein et al. developed a whole-cell biosensor based on a pair of zinc-responsive TFs, Zur, a transcriptional repressor, and ZntR, a transcriptional activator [177] in response to different extracellular zinc concentrations, the whole-cell biosensor can produce different colored pigments (violacein, lycopene, and β-carotene) as qualitative indication zinc levels. A QS-based amplifier based on metal ion-induced promoters was developed to improve the performance of environmental metal ion biosensing [61].
Perspectives and concluding remarks
Advances in genome sequencing and synthetic biology have greatly enhanced our ability to identify, build, modify, and apply in vivo biosensors. Over the last few years, applications of in vivo biosensors have moved from simple metabolite sensing and reporting to more complex genetic systems, such as dynamic and multi-layered genetic circuits, offering more advanced tools for engineering biosynthetic pathways and whole cells.
To accelerate the development of biosensors, it is critical to expand the repertoire of detectable metabolites and products. Currently, there are only a limited number of biosensors compared to the vast number of small molecules of interest. Genomic and transcriptomic analysis can help researchers to uncover more TF- or promoter-based in vivo biosensors, while advances in protein engineering as well as computational design can help researchers to better tailor the specificity of TFs. In addition, methods to study genome-wide RNA secondary structures [170] can help researchers to identify novel RNA-based sensors.
With the increasing complexity of biochemical production strategies enabled by synthetic biology, there is a greater need for standardized and streamlined engineering methods, as demonstrated by Gupta et al. [54]. Another implication of more complex genetic designs is that biosynthetic pathways will become increasingly modularized to enable parallel optimization, which presents opportunities for the application of biosensors to dynamically regulate flux between modules [60]. Hence, better control over the dynamic ranges of in vivo biosensors is needed to match the output and input of adjacent modules. This can be achieved with the development of more standardized genetic parts as well as protein engineering.
The challenges presented here are non-trivial, but offer great opportunities to advance the field of in vivo biosensors. Going forward, we anticipate increasing adoption of in vivo biosensors in metabolic engineering and synthetic biology for the optimization of strain development, and the dynamic regulation of biosynthetic pathways.
Abbreviations
- ADC:
-
Analog-to-digital converter
- ADH:
-
Alcohol dehydrogenase
- 1,4-BDO:
-
1,4-Butanediol
- CAD :
-
Cis-aconitate decarboxylase gene
- CCM:
-
Cis,cis-muconic acid
- CDA:
-
Cytidine deaminase
- 2′,3′-cGAMP:
-
(2′-5′,3′-5′) Cyclic guanosine monophosphate-adenosine monophosphate
- CE:
-
Capillary electrophoresis
- CFP:
-
Cyan fluorescent protein
- COMPACTER:
-
Customized optimization of metabolic pathways by combinatorial transcriptional engineering
- DFHBI:
-
3,5-Difluoro-4-hydroxybenzylidene imidazolinone
- 34DHB:
-
3,4-Dihydroxy benzoate
- l-DOPA:
-
l-3,4-Dihydroxyphenylalanine
- DSRS:
-
Dynamic sensor-regulator system
- epPCR:
-
Error-prone PCR
- FACS:
-
Fluorescence activated cell sorting
- fcy1 :
-
Cytosine deaminase gene
- FFA:
-
Free fatty acid
- FI:
-
Fluorescence intensity
- FP:
-
Fluorescent proteins
- FPP:
-
Farnesyl pyrophosphate
- FREP:
-
Feedback-regulated evolution of phenotype
- FRET:
-
Förster resonance energy transfer
- GEMM:
-
Genes for the environment, membranes, and motility
- GFP:
-
Green fluorescent protein
- GlcN6P:
-
Glucosamine 6-phosphate
- GlcNAc:
-
N-acetyl glucosamine
- gltA :
-
Citrate synthetase gene
- GPCR:
-
G-protein-coupled receptors
- HHR:
-
Hammerhead ribozyme
- Hi-Fi:
-
High-fidelity
- HK:
-
Histidine kinase
- 3-HP:
-
3-Hydroxy propionic acid
- IL:
-
Interleukin
- IPTG:
-
Isopropyl-beta-d-thiogalactopyranoside
- α-KGDH:
-
α-Ketoglutarate dehydrogenase
- LAO:
-
Lysine-binding periplasmic protein
- LTTR:
-
LysR-type transcriptional regulator
- MAGE:
-
Multiplex automated genome engineering
- malQ :
-
Maltase gene
- MBP:
-
Metabolite-binding protein
- MetN:
-
Methionine-binding protein
- MT:
-
Metallothionein
- MVA:
-
Mevalonate
- NAGK:
-
N-acetyl-l-glutamate kinase
- NeuAC:
-
N-acetylneuramine acid
- NMM:
-
N-methyl mesoporphyrin IX
- OAH:
-
O-acetyl homoserine
- OAS:
-
O-acetyl serine
- PBP:
-
Periplasmic-binding proteins
- PopQC:
-
Population quality control
- QS:
-
Quorum sensing
- RAGE:
-
RNAi-assisted genome evolution
- RBS:
-
Ribosome binding site
- ROK:
-
Repressor, open reading frame, kinase
- RR:
-
Response regulator
- SAM:
-
S-adenosylmethionine
- SAH:
-
S-adenosyl-l-homocysteine
- SATRE:
-
Sensor-assisted transcriptional regulator engineering
- SBA:
-
Streptavidin-binding aptamer
- SELEX:
-
Systematic evolution of ligands by exponential enrichment
- T6P:
-
Trehalose-6-phosphate
- Tc:
-
Tetracycline
- TCA:
-
Tricarboxylic acid
- TCS:
-
Two-component system
- TF:
-
Transcriptional factor
- TreR:
-
Trehalose repressor
- TRMR:
-
Trackable multiplex recombineering
- YFP:
-
Yellow fluorescent protein
- YOGE:
-
Yeast oligo-mediated genome engineering
- ZF:
-
Zinc finger
References
Amann E, Brosius J, Ptashne M (1983) Vectors bearing a hybrid trp-lac promoter useful for regulated expression of cloned genes in Escherichia coli. Gene 25:167–178. https://doi.org/10.1016/0378-1119(83)90222-6
Amaro F, Turkewitz AP, Martíngonzález A, Gutiérrez JC (2014) Functional GFP-metallothionein fusion protein from Tetrahymena thermophila: a potential whole-cell biosensor for monitoring heavy metal pollution and a cell model to study metallothionein overproduction effects. Biometals 27:195. https://doi.org/10.1007/s10534-014-9704-0
Ameen S, Ahmad M, Mohsin M, Qureshi MI, Ibrahim MM, Abdin MZ, Ahmad A (2016) Designing, construction and characterization of genetically encoded FRET-based nanosensor for real time monitoring of lysine flux in living cells. J Nanobiotechnol 14:49. https://doi.org/10.1186/s12951-016-0204-y
Amin EB, Mishler DM, Wang J, Walker H, Salis HM (2016) Automated physics-based design of synthetic riboswitches from diverse RNA aptamers. Nucleic Acids Res 44:1–13. https://doi.org/10.1093/nar/gkv1289
Atsumi S, Liao JC (2008) Directed evolution of Methanococcus jannaschii citramalate synthase for biosynthesis of 1-propanol and 1-butanol by Escherichia coli. Appl Environ Microbiol 74:7802–7808. https://doi.org/10.1128/aem.02046-08
Ausländer D, Ausländer S, Charpin-El Hamri G, Sedlmayer F, Müller M, Frey O, Hierlemann A, Stelling J, Fussenegger M (2014) A synthetic multifunctional mammalian pH sensor and CO2 transgene-control device. Mol Cell 55:397–408. https://doi.org/10.1016/j.molcel.2014.06.007
Becker K, Beer C, Freitag M, Kück U (2015) Genome-wide identification of target genes of a mating-type α-domain transcription factor reveals functions beyond sexual development. Mol Microbiol 96:1002–1022. https://doi.org/10.1111/mmi.12987
Bereza-Malcolm LT, Mann G, Franks AE (2015) Environmental sensing of heavy metals through whole cell microbial biosensors: a synthetic biology approach. ACS Synth Biol 4:535–546. https://doi.org/10.1021/sb500286r
Binder S, Schendzielorz G, Stäbler N, Krumbach K, Hoffmann K, Bott M, Eggeling L (2012) A high-throughput approach to identify genomic variants of bacterial metabolite producers at the single-cell level. Genome Biol 13:R40. https://doi.org/10.1186/gb-2012-13-5-r40
Bose D, Su Y, Marcus A, Raulet DH, Hammond MC (2016) An RNA-based fluorescent biosensor for high-throughput analysis of the cGAS-cGAMP-STING pathway. Cell Chem Biol 23:1539–1549. https://doi.org/10.1016/j.chembiol.2016.10.014
Brink DP, Borgström C, Tueros FG, Gorwa-Grauslund MF (2016) Real-time monitoring of the sugar sensing in Saccharomyces cerevisiae indicates endogenous mechanisms for xylose signaling. Microb Cell Fact 15:183. https://doi.org/10.1186/s12934-016-0580-x
Brockman IM, Prather KLJ (2015) Dynamic metabolic engineering: new strategies for developing responsive cell factories. Biotechnol J 10:1360–1369. https://doi.org/10.1002/biot.201400422
Caspeta L, Chen Y, Ghiaci P, Feizi A, Buskov S, Hallström BM, Petranovic D, Nielsen J (2014) Altered sterol composition renders yeast thermotolerant. Science 346:75–78. https://doi.org/10.1126/science.1258137
Ceres P, Garst AD, Marcano-Velázquez JG, Batey RT (2013) Modularity of select riboswitch expression platforms enables facile engineering of novel genetic regulatory devices. ACS Synth Biol 2:463–472. https://doi.org/10.1021/sb4000096
Ceres P, Trausch JJ, Batey RT (2013) Engineering modular ‘ON’RNA switches using biological components. Nucleic Acids Res 41:10449–10461. https://doi.org/10.1093/nar/gkt787
Chen W, Zhang S, Jiang P, Yao J, He Y, Chen L, Gui X, Dong Z, Tang SY (2015) Design of an ectoine-responsive AraC mutant and its application in metabolic engineering of ectoine biosynthesis. Metab Eng 30:149–155. https://doi.org/10.1016/j.ymben.2015.05.004
Chen X, Gao C, Guo L, Hu G, Luo Q, Liu J, Nielsen J, Chen J, Liu L (2017) DCEO biotechnology: tools to design, construct, evaluate, and optimize the metabolic pathway for biosynthesis of chemicals. Chem Rev. https://doi.org/10.1021/acs.chemrev.6b00804 (In press)
Choi S-L, Rha E, Lee SJ, Kim H, Kwon K, Jeong Y-S, Rhee YH, Song JJ, Kim H-S, Lee S-G (2013) Toward a generalized and high-throughput enzyme screening system based on artificial genetic circuits. ACS Synth Biol 3:163–171. https://doi.org/10.1021/sb400112u
Chong H, Ching CB (2016) Development of colorimetric-based whole-cell biosensor for organophosphorus compounds by engineering transcription regulator DmpR. ACS Synth Biol 5:1290–1298. https://doi.org/10.1021/acssynbio.6b00061
Chou HH, Keasling JD (2013) Programming adaptive control to evolve increased metabolite production. Nat Commun 4:2595. https://doi.org/10.1038/ncomms3595
Clark LC, Lyons C (1962) Electrode systems for continuous monitoring in cardiovascular surgery. Ann N Y Acad Sci 102:29–45. https://doi.org/10.1111/j.1749-6632.1962.tb13623.x
Dahl RH, Zhang F, Alonso-Gutierrez J, Baidoo E, Batth TS, Redding-Johanson AM, Petzold CJ, Mukhopadhyay A, Lee TS, Adams PD, Keasling JD (2013) Engineering dynamic pathway regulation using stress-response promoters. Nat Biotechnol 13:1039–1046. https://doi.org/10.1038/nbt.2689
Danino T, Prindle A, Kwong GA, Skalak M, Li H, Allen K, Hasty J, Bhatia SN (2015) Programmable probiotics for detection of cancer in urine. Sci Transl Med 7:289ra284. https://doi.org/10.1126/scitranslmed.aaa3519
David F, Nielsen J, Siewers V (2016) Flux control at the malonyl-CoA node through hierarchical dynamic pathway regulation in Saccharomyces cerevisiae. ACS Synth Biol 5:224. https://doi.org/10.1021/acssynbio.5b00161
de los Santos ELC, Meyerowitz JT, Mayo SL, Murray RM (2016) Engineering transcriptional regulator effector specificity using computational design and in vitro rapid prototyping: developing a vanillin sensor. ACS Synth Biol 5:287–295. https://doi.org/10.1021/acssynbio.5b00090
De Michele R, Ast C, Loqué D, Ho C-H, Andrade SL, Lanquar V, Grossmann G, Gehne S, Kumke MU, Frommer WB (2013) Fluorescent sensors reporting the activity of ammonium transceptors in live cells. eLife 2:e00800. https://doi.org/10.7554/elife.00800
Dekker L, Polizzi KM (2017) Sense and sensitivity in bioprocessing—detecting cellular metabolites with biosensors. Curr Opin Chem Biol 40:31–36. https://doi.org/10.1016/j.cbpa.2017.05.014
Delépine B, Libis V, Carbonell P, Faulon J-L (2016) SensiPath: computer-aided design of sensing-enabling metabolic pathways. Nucleic Acids Res 44:226–231. https://doi.org/10.1093/nar/gkw305
DeLoache WC, Russ ZN, Narcross L, Gonzales AM, Martin VJJ, Dueber JE (2015) An enzyme-coupled biosensor enables (S)-reticuline production in yeast from glucose. Nat Chem Biol 11:465–471. https://doi.org/10.1038/nchembio.1816
Desai SK, Gallivan JP (2004) Genetic screens and selections for small molecules based on a synthetic riboswitch that activates protein translation. J Am Chem Soc 126:13247–13254. https://doi.org/10.1021/ja048634j
DiCarlo JE, Conley AJ, Penttilä M, Jäntti J, Wang HH, Church GM (2013) Yeast oligo-mediated genome engineering (YOGE). ACS Synth Biol 2:741. https://doi.org/10.1021/sb400117c
Dietrich JA, Shis DL, Alikhani A, Keasling JD (2013) Transcription factor-based screens and synthetic selections for microbial small-molecule biosynthesis. ACS Synth Biol 2:47–58. https://doi.org/10.1021/sb300091d
Dong H, Liu Y, Zu X, Li N, Li F, Zhang D (2015) An enzymatic assay for high-throughput screening of cytidine-producing microbial strains. PLoS One 10:e0121612. https://doi.org/10.1371/journal.pone.0121612
Dong H, Zu X, Zheng P, Zhang D (2015) A rapid enzymatic assay for high-throughput screening of adenosine-producing strains. Microb Biotechnol 8:230–238. https://doi.org/10.1111/1751-7915.12189
Du J, Yuan Y, Si T, Lian J, Zhao H (2012) Customized optimization of metabolic pathways by combinatorial transcriptional engineering. Nucleic Acids Res 40:e142. https://doi.org/10.1093/nar/gks549
Eaton RM, Shallcross JA, Mael LE, Mears KS, Minkoff L, Scoville DJ, Whelan RJ (2015) Selection of DNA aptamers for ovarian cancer biomarker HE4 using CE-SELEX and high-throughput sequencing. Anal Bioanal Chem 407:6965–6973. https://doi.org/10.1007/s00216-015-8665-7
Eckdahl TT, Campbell AM, Heyer LJ, Poet JL, Blauch DN, Snyder NL, Atchley DT, Baker EJ, Brown M, Brunner EC (2015) Programmed evolution for optimization of orthogonal metabolic output in bacteria. PLoS One 10:e0118322. https://doi.org/10.1371/journal.pone.0118322
Ehrenworth AM, Claiborne T, Peralta-Yahya P (2017) Medium-throughput screen of microbially produced serotonin via a G-protein-coupled receptor-based sensor. Biochemistry. https://doi.org/10.1021/acs.biochem.7b00605 (In press)
Fang M, Wang T, Zhang C, Bai J, Zheng X, Zhao X, Lou C, Xing X-H (2015) Intermediate-sensor assisted push-pull strategy and its application in heterologous deoxyviolacein production in Escherichia coli. Metab Eng 33:41–51. https://doi.org/10.1016/j.ymben.2015.10.006
Farmer WR, Liao JC (2000) Improving lycopene production in Escherichia coli by engineering metabolic control. Nat Biotechnol 18:533–537. https://doi.org/10.1038/75398
Feist AM, Zielinski DC, Orth JD, Schellenberger J, Herrgard MJ, Palsson BØ (2010) Model-driven evaluation of the production potential for growth-coupled products of Escherichia coli. Metab Eng 12:173–186. https://doi.org/10.1016/j.ymben.2009.10.003
Felnagle EA, Chaubey A, Noey EL, Houk KN, Liao JC (2012) Engineering synthetic recursive pathways to generate non-natural small molecules. Nat Chem Biol 8:518–526. https://doi.org/10.1038/nchembio.959
Feng J, Jester BW, Tinberg CE, Mandell DJ, Antunes MS, Chari R, Morey KJ, Rios X, Medford JI, Church GM, Fields S, Baker D (2015) A general strategy to construct small molecule biosensors in eukaryotes. eLife 4:e10606. https://doi.org/10.7554/elife.10606
Fowler CC, Brown ED, Li Y (2010) Using a riboswitch sensor to examine coenzyme B(12) metabolism and transport in E. coli. Chem Biol 17:756. https://doi.org/10.1016/j.chembiol.2010.05.025
Fowler ZL, Gikandi WW, Koffas MA (2009) Increased malonyl coenzyme A biosynthesis by tuning the Escherichia coli metabolic network and its application to flavanone production. Appl Environ Microbiol 75:5831–5839. https://doi.org/10.1128/AEM.00270-09
Frei CS, Wang Z, Qian S, Deutsch S, Sutter M, Cirino PC (2016) Analysis of amino acid substitutions in AraC variants that respond to triacetic acid lactone. Protein Sci 25:804. https://doi.org/10.1002/pro.2873
Furukawa K, Ramesh A, Zhou Z, Weinberg Z, Vallery T, Winkler Wade C, Breaker Ronald R (2015) Bacterial riboswitches cooperatively bind Ni2+ or Co2+ ions and control expression of heavy metal transporters. Mol Cell 57:1088–1098. https://doi.org/10.1016/j.molcel.2015.02.009
Ganesh I, Ravikumar S, Lee SH, Park SJ, Hong SH (2013) Engineered fumarate sensing Escherichia coli based on novel chimeric two-component system. J Biotechnol 168:560–566. https://doi.org/10.1016/j.jbiotec.2013.09.003
Ganesh I, Ravikumar S, Yoo IK, Hong SH (2015) Construction of malate-sensing Escherichia coli by introduction of a novel chimeric two-component system. Bioprocess Biosyst Eng 38:1–8. https://doi.org/10.1007/s00449-014-1321-3
Giel JL, Nesbit AD, Mettert EL, Fleischhacker AS, Wanta BT, Kiley PJ (2013) Regulation of iron–sulphur cluster homeostasis through transcriptional control of the Isc pathway by [2Fe–2S]–IscR in Escherichia coli. Mol Microbiol 87:478–492. https://doi.org/10.1111/mmi.12052
Goers L, Ainsworth C, Goey CH, Kontoravdi C, Freemont PS, Polizzi KM (2017) Whole-cell Escherichia coli lactate biosensor for monitoring mammalian cell cultures during biopharmaceutical production. Biotechnol Bioeng 114:1290–1300. https://doi.org/10.1002/bit.26254
Goers L, Kylilis N, Tomazou M, Wen KY, Freemont P, Polizzi K (2013) Engineering microbial biosensors. Microbial synthetic biology. Elsevier, Amsterdam, pp 119–156
Grünewald FS (2013) Periplasmic binding proteins in biosensing applications. Advances in chemical bioanalysis. Springer, Berlin, pp 205–235
Gupta A, Brockman Reizman IM, Reisch CR, Prather KLJ (2017) Dynamic regulation of metabolic flux in engineered bacteria using a pathway-independent quorum-sensing circuit. Nat Biotechnol 35:273–279. https://doi.org/10.1038/nbt.3796
Hallberg ZF, Su Y, Kitto RZ, Hammond MC (2017) Engineering and in vivo applications of riboswitches. Annu Rev Biochem 86:1611–1625. https://doi.org/10.1146/annurev-biochem-060815-014628
Hao Z, Lou H, Zhu R, Zhu J, Zhang D, Zhao BS, Zeng S, Chen X, Chan J, He C, Chen PR (2014) The multiple antibiotic resistance regulator MarR is a copper sensor in Escherichia coli. Nat Chem Biol 10:21–28. https://doi.org/10.1038/nchembio.1380
Hector RE, Mertens JA (2017) A synthetic hybrid promoter for xylose-regulated control of gene expression in Saccharomyces yeasts. Mol Biotechnol 59:24–33. https://doi.org/10.1007/s12033-016-9991-5
Hessels AM, Chabosseau P, Bakker MH, Engelen W, Rutter GA, Taylor KM, Merkx M (2015) eZinCh-2: a versatile, genetically encoded FRET sensor for cytosolic and intraorganelle Zn2+ imaging. ACS Chem Biol 10:2126–2134. https://doi.org/10.1021/acschembio.5b00211
Hoffmann K, Grünberger A, Lausberg F, Bott M, Eggeling L (2013) Visualization of imbalances in sulfur assimilation and synthesis of sulfur-containing amino acids at the single-cell level. Appl Environ Microbiol 79:6730–6736. https://doi.org/10.1128/AEM.01804-13
Holtz WJ, Keasling JD (2010) Engineering static and dynamic control of synthetic pathways. Cell 140:19–23. https://doi.org/10.1016/j.cell.2009.12.029
Hsu CY, Chen BK, Hu RH, Chen BS (2016) Systematic design of a quorum sensing-based biosensor for enhanced detection of metal ion in Escherichia coli. IEEE Trans Biomed Circuits Syst 10:593–601. https://doi.org/10.1109/TBCAS.2015.2495151
Imperial College London iGEM project (2010) Parasite detection with a rapid response. http://2010.igem.org/Team:Imperial_College_London. Accessed 24 Jan 2018
Immethun CM, Ng KM, DeLorenzo DM, Waldron-Feinstein B, Lee Y-C, Moon TS (2016) Oxygen-responsive genetic circuits constructed in Synechocystis sp. PCC 6803. Biotechnol Bioeng 113:433–442. https://doi.org/10.1002/bit.25722
Jang S, Jang S, Xiu Y, Kang TJ, Lee S-H, Koffas MA, Jung GY (2017) Development of artificial riboswitches for monitoring of naringenin in vivo. ACS Synth Biol. https://doi.org/10.1021/acssynbio.7b00128 (In press)
Jha RK, Chakraborti S, Kern TL, Fox DT, Strauss CEM (2015) Rosetta comparative modeling for library design: engineering alternative inducer specificity in a transcription factor. Proteins Struct Funct Bioinf 83:1327–1340. https://doi.org/10.1002/prot.24828
Jha RK, Kern TL, Fox DT, Strauss CEM (2014) Engineering an Acinetobacter regulon for biosensing and high-throughput enzyme screening in E. coli via flow cytometry. Nucleic Acids Res 42:8150–8160. https://doi.org/10.1093/nar/gku444
Kannan Maruthamuthu M, Ganesh I, Ravikumar S, Hong SH (2015) Evaluation of zraP gene expression characteristics and construction of a lead (Pb) sensing and removal system in a recombinant Escherichia coli. Biotechnol Lett 37:659–664. https://doi.org/10.1007/s10529-014-1732-x
Kasey C, Zerrad M, Li Y, Cropp TA, Williams GJ (2017) Development of transcription factor-based designer macrolide biosensors for metabolic engineering and synthetic biology. ACS Synth Biol. https://doi.org/10.1021/acssynbio.7b00287 (In press)
Katritch V, Cherezov V, Stevens RC (2013) Structure-function of the G protein-coupled receptor superfamily. Annu Rev Pharmacol Toxicol 53:531–556. https://doi.org/10.1146/annurev-pharmtox-032112-135923
Kazanov MD, Li X, Gelfand MS, Osterman AL, Rodionov DA (2013) Functional diversification of ROK-family transcriptional regulators of sugar catabolism in the Thermotogae phylum. Nucleic Acids Res 41:790–803. https://doi.org/10.1093/nar/gks1184
Kellenberger CA, Wilson SC, Saleslee J, Hammond MC (2013) RNA-based fluorescent biosensors for live cell imaging of second messengers cyclic di-GMP and cyclic AMP-GMP. J Am Chem Soc 135:4906. https://doi.org/10.1021/ja311960g
Klauser B, Atanasov J, Siewert LK, Hartig JS (2015) Ribozyme-based aminoglycoside switches of gene expression engineered by genetic selection in S. cerevisiae. ACS Synth Biol 4:516–525. https://doi.org/10.1021/sb500062p
Ko W, Kim S, Lee HS (2017) Engineering a periplasmic binding protein for amino acid sensors with improved binding properties. Org Biomol Chem. https://doi.org/10.1039/C7OB01847A (In press)
Kotula JW, Kerns SJ, Shaket LA, Siraj L, Collins JJ, Way JC, Silver PA (2014) Programmable bacteria detect and record an environmental signal in the mammalian gut. Proc Natl Acad Sci USA 111:4838–4843. https://doi.org/10.1073/pnas.1321321111
Kumari P, Ghosh E, Shukla AK (2015) Emerging approaches to GPCR ligand screening for drug discovery. Trends Mol Med 21:687–701. https://doi.org/10.1016/j.molmed.2015.09.002
Leavitt JM, Wagner JM, Tu CC, Tong A, Liu Y, Alper HS (2017) Biosensor-enabled directed evolution to improve muconic acid production in Saccharomyces cerevisiae. Biotechnol J. https://doi.org/10.1002/biot.201600687 (In press)
Lee S-W, Oh M-K (2015) A synthetic suicide riboswitch for the high-throughput screening of metabolite production in Saccharomyces cerevisiae. Metab Eng 28:143–150. https://doi.org/10.1016/j.ymben.2015.01.004
Lee SK, Chou H, Ham TS, Lee TS, Keasling JD (2008) Metabolic engineering of microorganisms for biofuels production: from bugs to synthetic biology to fuels. Curr Opin Biotechnol 19:556–563. https://doi.org/10.1016/j.copbio.2008.10.014
Lee Y, Basith S, Choi S (2017) Recent advances in structure-based drug design targeting class a G protein-coupled receptors utilizing crystal structures and computational simulations. J Med Chem. https://doi.org/10.1021/acs.jmedchem.6b01453 (In press)
Lefrançois P, Euskirchen GM, Auerbach RK, Rozowsky J, Gibson T, Yellman CM, Gerstein M, Snyder M (2009) Efficient yeast ChIP-Seq using multiplex short-read DNA sequencing. BMC Genom 10:37. https://doi.org/10.1186/1471-2164-10-37
Leonhartsberger S, Huber A, Lottspeich F, Böck A (2001) The hydH/G genes from Escherichia coli code for a zinc and lead responsive two-component regulatory system. J Mol Biol 307:93–105. https://doi.org/10.1006/jmbi.2000.4451
Levay A, Brenneman R, Hoinka J, Sant D, Cardone M, Trinchieri G, Przytycka TM, Berezhnoy A (2015) Identifying high-affinity aptamer ligands with defined cross-reactivity using high-throughput guided systematic evolution of ligands by exponential enrichment. Nucleic Acids Res 43:e82. https://doi.org/10.1093/nar/gkv534
Li S, Si T, Wang M, Zhao H (2015) Development of a synthetic malonyl-CoA sensor in Saccharomyces cerevisiae for intracellular metabolite monitoring and genetic screening. ACS Synth Biol 4:1308–1315
Liang W-F, Cui L-Y, Cui J-Y, Yu K-W, Yang S, Zhang C, Xing X-H (2017) Biosensor-assisted transcriptional regulator engineering for Methylobacterium extorquens AM1 to improve mevalonate synthesis by increasing the acetyl-CoA supply. Metab Eng 39:159–168. https://doi.org/10.1016/j.ymben.2016.11.010
Libis V, Delépine B, Faulon J-L (2016) Expanding biosensing abilities through computer-aided design of metabolic pathways. ACS Synth Biol 5:1076–1085. https://doi.org/10.1021/acssynbio.5b00225
Lin JL, Zhu J, Wheeldon I (2016) Rapid ester biosynthesis screening reveals a high activity alcohol-O-acyltransferase (AATase) from tomato fruit. Biotechnol J 11:700–707. https://doi.org/10.1002/biot.201500406
Lin LL, Hsu WH (1997) Lactose-induced expression of Bacillus sp. TS-23 amylase gene in Escherichia coli regulated by a T7 promoter. Lett Appl Microbiol 24:365. https://doi.org/10.1046/j.1472-765X.1997.00146.x
Litke JL, You M, Jaffrey SR (2016) Developing fluorogenic riboswitches for imaging metabolite concentration dynamics in bacterial cells. Methods Enzymol 572:315–333. https://doi.org/10.1016/bs.mie.2016.03.021
Liu CC, Arkin AP (2010) The case for RNA. Science 330:1185–1186. https://doi.org/10.1126/science.1199495
Liu D, Xiao Y, Evans B, Zhang F (2015) Negative feedback regulation of fatty acid production based on a malonyl-CoA sensor-actuator. ACS Synth Biol 4:132–140. https://doi.org/10.1021/sb400158w
Liu H, Lu T (2015) Autonomous production of 1,4-butanediol via a de novo biosynthesis pathway in engineered Escherichia coli. Metab Eng 29:135–141. https://doi.org/10.1016/j.ymben.2015.03.009
Liu S-D, Wu Y-N, Wang T-M, Zhang C, Xing X-H (2017) Maltose utilization as a novel selection strategy for continuous evolution of microbes with enhanced metabolite production. ACS Synth Biol. https://doi.org/10.1021/acssynbio.7b00247 (In press)
Liu Y, Li Q, Zheng P, Zhang Z, Liu Y, Sun C, Cao G, Zhou W, Wang X, Zhang D (2015) Developing a high-throughput screening method for threonine overproduction based on an artificial promoter. Microb Cell Fact 14:121. https://doi.org/10.1186/s12934-015-0311-8
Liu Y, Zhuang Y, Ding D, Xu Y, Sun J, Zhang D (2017) Biosensor-based evolution and elucidation of a biosynthetic pathway in Escherichia coli. ACS Synth Biol 6:837–848. https://doi.org/10.1021/acssynbio.6b00328
Luo Z, Zhou H, Jiang H, Ou H, Li X, Zhang L (2015) Development of a fraction collection approach in capillary electrophoresis SELEX for aptamer selection. Analyst 140:2664–2670. https://doi.org/10.1039/C5AN00183H
Ma AT, Schmidt CM, Golden JW (2014) Regulation of gene expression in diverse cyanobacterial species by using theophylline-responsive riboswitches. Appl Environ Microbiol 80:6704–6713. https://doi.org/10.1128/aem.01697-14
Machado LFM, Dixon N (2016) Development and substrate specificity screening of an in vivo biosensor for the detection of biomass derived aromatic chemical building blocks. Chem Commun 52:11402–11405. https://doi.org/10.1039/C6CC04559F
Mahr R, Frunzke J (2015) Transcription factor-based biosensors in biotechnology: current state and future prospects. Appl Microbiol Biotechnol 100:79–90. https://doi.org/10.1007/s00253-015-7090-3
Mahr R, Gätgens C, Gätgens J, Polen T, Kalinowski J, Frunzke J (2015) Biosensor-driven adaptive laboratory evolution of l-valine production in Corynebacterium glutamicum. Metab Eng 32:184–194. https://doi.org/10.1016/j.ymben.2015.09.017
Mahr R, von Boeselager RF, Wiechert J, Frunzke J (2016) Screening of an Escherichia coli promoter library for a phenylalanine biosensor. Appl Microbiol Biotechnol 100:6739–6753. https://doi.org/10.1007/s00253-016-7575-8
Mannan AA, Liu D, Zhang F, Oyarzún DA (2017) Fundamental design principles for transcription-factor-based metabolite biosensors. ACS Synth Biol. https://doi.org/10.1021/acssynbio.7b00172 (In press)
Marin AM, Souza EM, Pedrosa FO, Souza LM, Sassaki GL, Baura VA, Yates MG, Wassem R, Monteiro RA (2013) Naringenin degradation by the endophytic diazotroph Herbaspirillum seropedicae SmR1. Microbiology 159:167–175. https://doi.org/10.1099/mic.0.061135-0
Martín AS, Ceballo S, Ruminot I, Lerchundi R, Frommer WB, Barros LF (2013) A genetically encoded FRET lactate sensor and its use to detect the warburg effect in single cancer cells. PLoS One 8:e57712. https://doi.org/10.1371/journal.pone.0057712
McNerney MP, Watstein DM, Styczynski MP (2015) Precision metabolic engineering: the design of responsive, selective, and controllable metabolic systems. Metab Eng 31:123–131. https://doi.org/10.1016/j.ymben.2015.06.011
Merulla D, van der Meer JR (2016) Regulatable and modulable background expression control in prokaryotic synthetic circuits by auxiliary repressor binding sites. ACS Synth Biol 5:36–45. https://doi.org/10.1021/acssynbio.5b00111
Meyer A, Pellaux R, Potot S, Becker K, Hohmann HP, Panke S, Held M (2015) Optimization of a whole-cell biocatalyst by employing genetically encoded product sensors inside nanolitre reactors. Nat Chem 7:673. https://doi.org/10.1038/NCHEM.2301
Meyer V, Wanka F, Van GJ, Arentshorst M, Ca VDH, Ram AF (2011) Fungal gene expression on demand: an inducible, tunable, and metabolism-independent expression system for Aspergillus niger. Appl Environ Microb 77:2975–2983. https://doi.org/10.1128/AEM.02740-10
Miller MB, Bassler BL (2001) Quorum sensing in bacteria. Annu Rev Microbiol 55:165–199. https://doi.org/10.1146/annurev.micro.55.1.165
Miyano K, Sudo Y, Yokoyama A, Hisaoka-Nakashima K, Morioka N, Takebayashi M, Nakata Y, Higami Y, Uezono Y (2014) History of the G protein-coupled receptor (GPCR) assays from traditional to a state-of-the-art biosensor assay. J Pharmacol Sci 126:302–309. https://doi.org/10.1254/jphs.14R13CP
Mohsin M, Abdin MZ, Nischal L, Kardam H, Ahmad A (2013) Genetically encoded FRET-based nanosensor for in vivo measurement of leucine. Biosens Bioelectron 50:72–77. https://doi.org/10.1016/j.bios.2013.06.028
Mohsin M, Ahmad A (2014) Genetically-encoded nanosensor for quantitative monitoring of methionine in bacterial and yeast cells. Biosens Bioelectron 59:358–364. https://doi.org/10.1016/j.bios.2014.03.066
Mohsin M, Ahmad A, Iqbal M (2015) FRET-based genetically-encoded sensors for quantitative monitoring of metabolites. Biotechnol Lett 37:1919–1928. https://doi.org/10.1007/s10529-015-1873-6
Mukherjee K, Bhattacharyya S, Peralta-Yahya P (2015) GPCR-based chemical biosensors for medium-chain fatty acids. ACS Synth Biol 4:1261–1269. https://doi.org/10.1021/sb500365m
Müller M, Ausländer S, Spinnler A, Ausländer D, Sikorski J, Folcher M, Fussenegger M (2017) Designed cell consortia as fragrance-programmable analog-to-digital converters. Nat Chem Biol 13:309–316. https://doi.org/10.1038/nchembio.2281
Mustafi N, Grünberger A, Mahr R, Helfrich S, Nöh K, Blombach B, Kohlheyer D, Frunzke J (2014) Application of a genetically encoded biosensor for live cell imaging of l-valine production in pyruvate dehydrogenase complex-deficient Corynebacterium glutamicum strains. PLoS One 9:e85731. https://doi.org/10.1371/journal.pone.0085731
Newman JR, Fuqua C (1999) Broad-host-range expression vectors that carry the L-arabinose-inducible Escherichia coli araBAD promoter and the araC regulator. Gene 227:197–203. https://doi.org/10.1016/S0378-1119(98)00601-5
Nomura Y, Yokobayashi Y (2007) Reengineering a natural riboswitch by dual genetic selection. J Am Chem Soc 129:13814–13815. https://doi.org/10.1021/ja076298b
Novichkov PS, Kazakov AE, Ravcheev DA, Leyn SA, Kovaleva GY, Sutormin RA, Kazanov MD, Riehl W, Arkin AP, Dubchak I (2013) RegPrecise 3.0—a resource for genome-scale exploration of transcriptional regulation in bacteria. BMC Genom 14:745. https://doi.org/10.1186/1471-2164-14-745
Ohbayashi R, Akai H, Yoshikawa H, Hess WR, Watanabe S (2016) A tightly inducible riboswitch system in Synechocystis sp. PCC 6803. J Gen Appl Microbiol 62:154–159. https://doi.org/10.2323/jgam.2016.02.002
Oku M, Hoseki J, Ichiki Y, Sakai Y (2013) A fluorescence resonance energy transfer (FRET)-based redox sensor reveals physiological role of thioredoxin in the yeast Saccharomyces cerevisiae. FEBS Lett 587:793–798. https://doi.org/10.1016/j.febslet.2013.02.003
Ouellet E, Foley JH, Conway EM, Haynes C (2015) Hi-Fi SELEX: a high-fidelity digital-PCR based therapeutic aptamer discovery platform. Biotechnol Bioeng 112:1506–1522. https://doi.org/10.1002/bit.25581
Paige JS, Wu KY, Jaffrey SR (2011) RNA mimics of green fluorescent protein. Science 333:642–646. https://doi.org/10.1126/science.1207339
Park J-N, Sohn MJ, Oh D-B, Kwon O, Rhee SK, Hur C-G, Lee SY, Gellissen G, Kang HA (2007) Identification of the cadmium-inducible Hansenula polymorpha SEO1 gene promoter by transcriptome analysis and its application to whole-cell heavy-metal detection systems. Appl Environ Microbiol 73:5990–6000. https://doi.org/10.1128/AEM.00863-07
Peroza EA, Ewald JC, Parakkal G, Skotheim JM, Zamboni N (2015) A genetically encoded Förster resonance energy transfer sensor for monitoring in vivo trehalose-6-phosphate dynamics. Anal Biochem 474:1–7. https://doi.org/10.1016/j.ab.2014.12.019
Pfleger BF, Pitera DJ, Newman JD, Martin VJ, Keasling JD (2007) Microbial sensors for small molecules: development of a mevalonate biosensor. Metab Eng 9:30–38. https://doi.org/10.1016/j.ymben.2006.08.002
Pham VD, Ravikumar S, Lee SH, Hong SH, Yoo I-K (2013) Modification of response behavior of zinc sensing HydHG two-component system using a self-activation loop and genomic integration. Bioprocess Biosyst Eng 36:1185–1190. https://doi.org/10.1007/s00449-012-0845-7
Porter EB, Polaski JT, Morck MM, Batey RT (2017) Recurrent RNA motifs as scaffolds for genetically encodable small-molecule biosensors. Nat Chem Biol 13:295–301. https://doi.org/10.1038/nchembio.2278
Rajkumar AS, Liu G, Bergenholm D, Arsovska D, Kristensen M, Nielsen J, Jensen MK, Keasling JD (2016) Engineering of synthetic, stress-responsive yeast promoters. Nucleic Acids Res 44:e136. https://doi.org/10.1093/nar/gkw553
Raman S, Rogers JK, Taylor ND, Church GM (2014) Evolution-guided optimization of biosynthetic pathways. Proc Natl Acad Sci USA 111:17803–17808. https://doi.org/10.1073/pnas.1409523111
Ravcheev DA, Khoroshkin MS, Laikova ON, Tsoy OV, Sernova NV, Petrova SA, Rakhmaninova AB, Novichkov PS, Gelfand MS, Rodionov DA (2014) Comparative genomics and evolution of regulons of the LacI-family transcription factors. Front Microbiol 5:294. https://doi.org/10.3389/fmicb.2014.00294
Ravikumar S, Baylon MG, Park SJ, J-i Choi (2017) Engineered microbial biosensors based on bacterial two-component systems as synthetic biotechnology platforms in bioremediation and biorefinery. Microb Cell Fact 16:62. https://doi.org/10.1186/s12934-017-0675-z
Ravikumar S, Ganesh I, Maruthamuthu MK, Hong SH (2015) Engineering Escherichia coli to sense acidic amino acids by introduction of a chimeric two-component system. Korean J Chem Eng 32:2073–2077. https://doi.org/10.1007/s11814-015-0024-z
Robinson CJ, Medina-Stacey D, Wu MC, Vincent HA, Micklefield J (2016) Rewiring riboswitches to create new genetic circuits in bacteria. Methods Enzymol 575:319–348. https://doi.org/10.1016/bs.mie.2016.02.022
Robinson CJ, Vincent HA, Wu M-C, Lowe PT, Dunstan MS, Leys D, Micklefield J (2014) Modular riboswitch toolsets for synthetic genetic control in diverse bacterial species. J Am Chem Soc 136:10615–10624. https://doi.org/10.1021/ja502873j
Rogers JK, Church GM (2016) Genetically encoded sensors enable real-time observation of metabolite production. Proc Natl Acad Sci USA 113:2388–2393. https://doi.org/10.1073/pnas.1600375113
Rogers JK, Guzman CD, Taylor ND, Raman S, Anderson K, Church GM (2015) Synthetic biosensors for precise gene control and real-time monitoring of metabolites. Nucleic Acids Res 43:7648–7660. https://doi.org/10.1093/nar/gkv616
Rogers JK, Taylor ND, Church GM (2016) Biosensor-based engineering of biosynthetic pathways. Curr Opin Biotechnol 42:84–91. https://doi.org/10.1016/j.copbio.2016.03.005
Romanini DW, Peralta-Yahya P, Mondol V, Cornish VW (2012) A heritable recombination system for synthetic Darwinian evolution in yeast. ACS Synth Biol 1:602. https://doi.org/10.1021/sb3000904
Saeki K, Tominaga M, Kawai-Noma S, Saito K, Umeno D (2016) Rapid diversification of BetI-based transcriptional switches for the control of biosynthetic pathways and genetic circuits. ACS Synth Biol 5:1201–1210. https://doi.org/10.1021/acssynbio.5b00230
San Martín A, Ceballo S, Baeza-Lehnert F, Lerchundi R, Valdebenito R, Contreras-Baeza Y, Alegría K, Barros LF (2014) Imaging mitochondrial flux in single cells with a FRET sensor for pyruvate. PLoS One 9:e85780. https://doi.org/10.1371/journal.pone.0085780
Sandoval NR, Kim JY, Glebes TY, Reeder PJ, Aucoin HR, Warner JR, Gill RT (2012) Strategy for directing combinatorial genome engineering in Escherichia coli. Proc Natl Acad Sci USA 109:10540–10545. https://doi.org/10.1073/pnas.1206299109
Schendzielorz G, Dippong M, Grünberger A, Kohlheyer D, Yoshida A, Binder S, Nishiyama C, Nishiyama M, Bott M, Eggeling L (2014) Taking control over control: use of product sensing in single cells to remove flux control at key enzymes in biosynthesis pathways. ACS Synth Biol 3:21. https://doi.org/10.1021/sb400059y
Schujman GE, Guerin M, Buschiazzo A, Schaeffer F, Llarrull LI, Reh G, Vila AJ, Alzari PM, De Mendoza D (2006) Structural basis of lipid biosynthesis regulation in Gram-positive bacteria. EMBO J 25:4074–4083. https://doi.org/10.1038/sj.emboj.7601284
Schujman GE, Paoletti L, Grossman AD, de Mendoza D (2003) FapR, a bacterial transcription factor involved in global regulation of membrane lipid biosynthesis. Dev Cell 4:663–672. https://doi.org/10.1016/S1534-5807(03)00123-0
Selvamani V, Ganesh I, Mk Maruthamuthu, Eom GT, Hong SH (2017) Engineering chimeric two-component system into Escherichia coli from Paracoccus denitrificans to sense methanol. Biotechnol Bioprocess Eng 22:225–230. https://doi.org/10.1007/s12257-016-0484-y
Seo SW, Kim D, Latif H, O’Brien EJ, Szubin R, Palsson BO (2014) Deciphering Fur transcriptional regulatory network highlights its complex role beyond iron metabolism in Escherichia coli. Nat Commun 5:4910. https://doi.org/10.1038/ncomms5910
Shi S, Choi YW, Zhao H, Tan MH, Ang EL (2017) Discovery and engineering of a 1-butanol biosensor in Saccharomyces cerevisiae. Bioresour Technol. https://doi.org/10.1016/j.biortech.2017.06.114 (In press)
Si T, Luo Y, Bao Z, Zhao H (2014) RNAi-assisted genome evolution in Saccharomyces cerevisiae for complex phenotype engineering. ACS Synth Biol 4:283–291. https://doi.org/10.1021/sb500074a
Siedler S, Schendzielorz G, Binder S, Eggeling L, Bringer S, Bott M (2014) SoxR as a single-cell biosensor for NADPH-consuming enzymes in Escherichia coli. ACS Synth Biol 3:41. https://doi.org/10.1021/sb400110j
Silva-Rocha R, De LV (2012) Broadening the signal specificity of prokaryotic promoters by modifying cis-regulatory elements associated with a single transcription factor. Mol BioSyst 8:1950–1957. https://doi.org/10.1039/c2mb25030f
Skjoedt ML, Snoek T, Kildegaard KR, Arsovska D, Eichenberger M, Goedecke TJ, Rajkumar AS, Zhang J, Kristensen M, Lehka BJ (2016) Engineering prokaryotic transcriptional activators as metabolite biosensors in yeast. Nat Chem Biol 12:951–958. https://doi.org/10.1038/nchembio.2177
Smolke CD, Silver PA (2011) Informing biological design by integration of systems and synthetic biology. Cell 144:855–859. https://doi.org/10.1016/j.cell.2011.02.020
Soma Y, Hanai T (2015) Self-induced metabolic state switching by a tunable cell density sensor for microbial isopropanol production. Metab Eng 30:7–15. https://doi.org/10.1016/j.ymben.2015.04.005
Steffen V, Otten J, Engelmann S, Radek A, Limberg M, Koenig BW, Noack S, Wiechert W, Pohl M (2016) A toolbox of genetically encoded FRET-based biosensors for rapid l-lysine analysis. Sensors 16:1604. https://doi.org/10.3390/s16101604
Su Y, Hickey SF, Keyser SG, Hammond MC (2016) In vitro and in vivo enzyme activity screening via RNA-based fluorescent biosensors for s-adenosyl-l-homocysteine (SAH). J Am Chem Soc 138:7040–7047. https://doi.org/10.1021/jacs.6b01621
Suess B, Fink B, Berens C, Stentz R, Hillen W (2004) A theophylline responsive riboswitch based on helix slipping controls gene expression in vivo. Nucleic Acids Res 32:1610–1614. https://doi.org/10.1093/nar/gkh321
Sun L, Zhang H, Yuan H, Tu R, Wang Q, Ma Y (2013) A double-enzyme-coupled assay for high-throughput screening of succinic acid-producing strains. J Appl Microbiol 114:1696–1701. https://doi.org/10.1111/jam.12175
Szmidt-Middleton HL, Ouellet M, Adams PD, Keasling JD, Mukhopadhyay A (2013) Utilizing a highly responsive gene, yhjX, in E. coli based production of 1,4-butanediol. Chem Eng Sci 103:68–73. https://doi.org/10.1016/j.ces.2013.06.044
Tang S-Y, Qian S, Akinterinwa O, Frei CS, Gredell JA, Cirino PC (2013) Screening for enhanced triacetic acid lactone production by recombinant Escherichia coli expressing a designed triacetic acid lactone reporter. J Am Chem Soc 135:10099–10103. https://doi.org/10.1021/ja402654z
Taylor ND, Garruss AS, Moretti R, Chan S, Arbing MA, Cascio D, Rogers JK, Isaacs FJ, Kosuri S, Baker D, Fields S, Church GM, Raman S (2016) Engineering an allosteric transcription factor to respond to new ligands. Nat Meth 13:177–183. https://doi.org/10.1038/nmeth.3696
Teo WS, Chang MW (2015) Bacterial XylRs and synthetic promoters function as genetically encoded xylose biosensors in Saccharomyces cerevisiae. Biotechnol J 10:315–322. https://doi.org/10.1002/biot.201400159
Teo WS, Hee KS, Chang MW (2013) Bacterial FadR and synthetic promoters function as modular fatty acid sensor-regulators in Saccharomyces cerevisiae. Eng Life Sci 13:456–463. https://doi.org/10.1002/elsc.201200113
Topp S, Gallivan JP (2007) Guiding bacteria with small molecules and RNA. J Am Chem Soc 129:6807–6811. https://doi.org/10.1021/ja0692480
Toussaint M, Bontemps C, Besserer A, Hotel L, Gérardin P, Leblond P (2016) Whole-cell biosensor of cellobiose and application to wood decay detection. J Biotechnol 239:39–46. https://doi.org/10.1016/j.jbiotec.2016.10.003
Umeyama T, Okada S, Ito T (2013) Synthetic gene circuit-mediated monitoring of endogenous metabolites: identification of GAL11 as a novel multicopy enhancer of S-adenosylmethionine level in yeast. ACS Synth Biol 2:425–430. https://doi.org/10.1021/sb300115n
Venayak N, Anesiadis N, Cluett WR, Mahadevan R (2015) Engineering metabolism through dynamic control. Curr Opin Biotechnol 34:142–152. https://doi.org/10.1016/j.copbio.2014.12.022
Vigneshvar S, Sudhakumari CC, Senthilkumaran B, Prakash H (2016) Recent advances in biosensor technology for potential applications—an overview. Front Bioeng Biotechnol 4:11. https://doi.org/10.3389/fbioe.2016.00011
Vinkenborg JL, Karnowski N, Famulok M (2011) Aptamers for allosteric regulation. Nat Chem Biol 7:519–527. https://doi.org/10.1038/nchembio.609
Wachsmuth M, Findei㟠S, Weissheimer N, Stadler PF, MãRl M (2013) De novo design of a synthetic riboswitch that regulates transcription termination. Nucleic Acids Res 41:2541–2551. https://doi.org/10.1093/nar/gks1330
Wan Y, Qu K, Zhang QC, Flynn RA, Manor O, Ouyang Z, Zhang J, Spitale RC, Snyder MP, Segal E, Chang HY (2014) Landscape and variation of RNA secondary structure across the human transcriptome. Nature 505:706–709. https://doi.org/10.1038/nature12946
Wang BL, Ghaderi A, Zhou H, Agresti J, Weitz DA, Fink GR, Stephanopoulos G (2014) Microfluidic high-throughput culturing of single cells for selection based on extracellular metabolite production or consumption. Nat Biotechnol 32:473–478. https://doi.org/10.1038/nbt.2857
Wang HH, Isaacs FJ, Carr PA, Sun ZZ, Xu G, Forest CR, Church GM (2009) Programming cells by multiplex genome engineering and accelerated evolution. Nature 460:894–898. https://doi.org/10.1038/nature08187
Wang J, Gao D, Yu X, Li W, Qi Q (2015) Evolution of a chimeric aspartate kinase for l-lysine production using a synthetic RNA device. Appl Microbiol Biotechnol 99:8527–8536. https://doi.org/10.1007/s00253-015-6615-0
Wang M, Li S, Zhao H (2016) Design and engineering of intracellular-metabolite-sensing/regulation gene circuits in Saccharomyces cerevisiae. Biotechnol Bioeng 113:206–215. https://doi.org/10.1002/bit.25676
Wang XC, Wilson SC, Hammond MC (2016) Next-generation RNA-based fluorescent biosensors enable anaerobic detection of cyclic di-GMP. Nucleic Acids Res 44:e139. https://doi.org/10.1093/nar/gkw580
Wang Y, Li Q, Zheng P, Guo Y, Wang L, Zhang T, Sun J, Ma Y (2016) Evolving the l-lysine high-producing strain of Escherichia coli using a newly developed high-throughput screening method. J Ind Microbiol Biotechnol 43:1227–1235. https://doi.org/10.1007/s10295-016-1803-1
Watstein DM, McNerney MP, Styczynski MP (2015) Precise metabolic engineering of carotenoid biosynthesis in Escherichia coli towards a low-cost biosensor. Metab Eng 31:171–180. https://doi.org/10.1016/j.ymben.2015.06.007
Williams TC, Averesch NJH, Winter G, Plan MR, Vickers CE, Nielsen LK, Krömer JO (2015) Quorum-sensing linked RNA interference for dynamic metabolic pathway control in Saccharomyces cerevisiae. Metab Eng 29:124–134. https://doi.org/10.1016/j.ymben.2015.03.008
Williams TC, Espinosa MI, Nielsen LK, Vickers CE (2015) Dynamic regulation of gene expression using sucrose responsive promoters and RNA interference in Saccharomyces cerevisiae. Microb Cell Fact 14:43. https://doi.org/10.1186/s12934-015-0223-7
Wu W, Zhang L, Yao L, Tan X, Liu X, Lu X (2015) Genetically assembled fluorescent biosensor for in situ detection of bio-synthesized alkanes. Sci Rep 5:10907. https://doi.org/10.1038/srep10907
Xiao Y, Bowen CH, Liu D, Zhang F (2016) Exploiting nongenetic cell-to-cell variation for enhanced biosynthesis. Nat Chem Biol 12:339. https://doi.org/10.1038/nchembio.2046
Xiao Y, Jiang W, Zhang F (2017) Developing a genetically encoded, cross-species biosensor for detecting ammonium and regulating biosynthesis of cyanophycin. ACS Synth Biol. https://doi.org/10.1021/acssynbio.7b00069 (In press)
Xie W, Lv X, Ye L, Zhou P, Yu H (2015) Construction of lycopene-overproducing Saccharomyces cerevisiae by combining directed evolution and metabolic engineering. Metab Eng 30:69–78. https://doi.org/10.1016/j.ymben.2015.04.009
Xie W, Ye L, Lv X, Xu H, Yu H (2015) Sequential control of biosynthetic pathways for balanced utilization of metabolic intermediates in Saccharomyces cerevisiae. Metab Eng 28:8. https://doi.org/10.1016/j.ymben.2014.11.007
Xiu Y, Jang S, Jones JA, Zill NA, Linhardt RJ, Yuan Q, Jung GY, Koffas MAG (2017) Naringenin-responsive riboswitch-based fluorescent biosensor module for Escherichia coli co-cultures. Biotechnol Bioeng 114:2235–2244. https://doi.org/10.1002/bit.26340
Xu P, Li L, Zhang F, Stephanopoulos G, Koffas M (2014) Improving fatty acids production by engineering dynamic pathway regulation and metabolic control. Proc Natl Acad Sci USA 111:11299–11304. https://doi.org/10.1073/pnas.1406401111
Xu P, Ranganathan S, Fowler ZL, Maranas CD, Koffas MA (2011) Genome-scale metabolic network modeling results in minimal interventions that cooperatively force carbon flux towards malonyl-CoA. Metab Eng 13:578–587. https://doi.org/10.1016/j.ymben.2011.06.008
Xu P, Wang W, Li L, Bhan N, Zhang F, Koffas MAG (2014) Design and kinetic analysis of a hybrid promoter-regulator system for malonyl-CoA sensing in E. coli. ACS Chem Biol 9:451–458. https://doi.org/10.1021/cb400623m
Yang J, Bowser MT (2013) Capillary electrophoresis–SELEX selection of catalytic DNA aptamers for a small-molecule porphyrin target. Anal Chem 85:1525–1530. https://doi.org/10.1021/ac302721j
Yang J, Seo SW, Jang S, Shin SI, Lim CH, Roh TY, Jung GY (2013) Synthetic RNA devices to expedite the evolution of metabolite-producing microbes. Nat Commun 4:1413. https://doi.org/10.1038/ncomms2404
Yang P, Wang J, Pang Q, Zhang F, Wang J, Wang Q, Qi Q (2017) Pathway optimization and key enzyme evolution of N-acetylneuraminate biosynthesis using an in vivo aptazyme-based biosensor. Metab Eng 43:21–28. https://doi.org/10.1016/j.ymben.2017.08.001
Yin X, Shin H-D, Li J, Du G, Liu L, Chen J (2017) Pgas, a low-pH-induced promoter, as a tool for dynamic control of gene expression for metabolic engineering of Aspergillus niger. Appl Environ Microbiol 83:e03216–e03222. https://doi.org/10.1128/aem.03222-16
You M, Litke JL, Jaffrey SR (2015) Imaging metabolite dynamics in living cells using a Spinach-based riboswitch. Proc Natl Acad Sci USA 112:E2756–E2765. https://doi.org/10.1073/pnas.1504354112
Younger AK, Dalvie NC, Rottinghaus AG, Leonard JN (2016) Engineering modular biosensors to confer metabolite-responsive regulation of transcription. ACS Synth Biol 6:311–325. https://doi.org/10.1021/acssynbio.6b00184
Yuan J, Ching C (2015) Dynamic control of ERG9 expression for improved amorpha-4,11-diene production in Saccharomyces cerevisiae. Microb Cell Fact 14:38. https://doi.org/10.1186/s12934-015-0220-x
Zhang F, Carothers JM, Keasling JD (2012) Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat Biotechnol 30:354–359. https://doi.org/10.1038/nbt.2149
Zhang J, Barajas JF, Burdu M, Ruegg TL, Dias B, Keasling JD (2017) Development of a transcription factor-based lactam biosensor. ACS Synth Biol 6:439–445. https://doi.org/10.1021/acssynbio.6b00136
Zhang J, Jensen MK, Keasling JD (2015) Development of biosensors and their application in metabolic engineering. Curr Opin Chem Biol 28:1–8. https://doi.org/10.1016/j.cbpa.2015.05.013
Zhang J, Sonnenschein N, Pihl TPB, Pedersen KR, Jensen MK, Keasling JD (2016) Engineering an NADPH/NADP+ redox biosensor in yeast. ACS Synth Biol 5:1546–1556. https://doi.org/10.1021/acssynbio.6b00135
Zhou L, Zeng A (2015) Engineering a Lysine-ON riboswitch for metabolic control of lysine production in Corynebacterium glutamicum. ACS Synth Biol 4:1335–1340. https://doi.org/10.1021/acssynbio.5b00075
Zhou L, Zeng A (2015) Exploring lysine riboswitch for metabolic flux control and improvement of l-lysine synthesis in Corynebacterium glutamicum. ACS Synth Biol 4:729–734. https://doi.org/10.1021/sb500332c
Zhou S, Ainala SK, Seol E, Nguyen TT, Park S (2015) Inducible gene expression system by 3-hydroxypropionic acid. Biotechnol Biofuels 8:169. https://doi.org/10.1186/s13068-015-0353-5
Zhu X, Wang X, Zhang C, Wang X, Gu Q (2015) A riboswitch sensor to determine vitamin B12 in fermented foods. Food Chem 175:523–528. https://doi.org/10.1016/j.foodchem.2014.11.163
Zorawski M, Shaffer J, Velasquez E, Liu JM (2016) Creating a riboswitch-based whole-cell biosensor for bisphenol A. FASEB J 30(805):3
Acknowledgements
We acknowledge funding supports from the State Key Laboratory of Microbial Technology Open Projects Fund in China (Project no. M2017-02 to S.S.), the National Research Foundation Singapore (NRF2013-THE001-095 to E.L.A.), and the Visiting Investigator Programme of Agency for Science, Technology, and Research, Singapore and US Department of Energy (DE-SC0018260 to H.Z.).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Shi, S., Ang, E.L. & Zhao, H. In vivo biosensors: mechanisms, development, and applications. J Ind Microbiol Biotechnol 45, 491–516 (2018). https://doi.org/10.1007/s10295-018-2004-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-018-2004-x