Abstract
The genetically encoded biosensors, which could transform the input of specific metabolic concentrations into output of gene expression levels, have been developed by hacking the sensing and regulatory systems of the cell such as allosteric transcription factors (aTFs) and riboswitches. In this chapter, we first introduce the classification and functional mechanism of genetically encoded biosensor. Furthermore, the applications of biosensor in the development of microbial cell factories including high-throughput screening and dynamic metabolic engineering are reviewed. Finally, the future perspectives on biosensors and their applications are discussed.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Biosensor
- Allosteric transcription factors
- Riboswitch
- Synthetic biology
- Microbial cell factory
- High-throughput screening
- Dynamic metabolic engineering
More and more microbial cell factories have been constructed for the production of valuable products such as biofuels, chemicals, materials, and nutraceuticals using renewable biomass sources (Cordova and Alper 2016; Liu et al. 2017a, b; Luo et al. 2019; Zhou et al. 2018), and this process has been further facilitated by the development of synthetic biology (Ng et al. 2015). Microorganisms have the ability to sense the change of a wide range of metabolites and modulate related pathways accordingly. This process is achieved by their sensing and regulatory systems such as allosteric transcription factors (aTFs) and riboswitches. With the aid of synthetic biology, the genetically encoded biosensors, which were designed and built by engineering the native sensing and regulatory systems of cells, have been widely applied in the high-throughput screening and metabolic regulation of the microbial strains (Koch et al. 2019; Michener et al. 2012). In this chapter, we focused on the constructions and applications of biosensors derived from allosteric transcription factors (aTFs) and riboswitches, and divided them into two categories, namely the protein-based biosensors and the RNA-based biosensors. Other types of biosensors, including the Förster resonance energy transfer (FRET)-based and two-component regulatory system (TCRS)-based biosensors that have not been used widespread in the development of microbial cell factories, will not be discussed here (refer to reviews (Greenwald et al. 2018; Ravikumar et al. 2017)).
4.1 The Classification of Genetically Encoded Biosensors
4.1.1 Protein-Based Biosensors
4.1.1.1 The Functional Mechanism of Protein-Based Biosensors
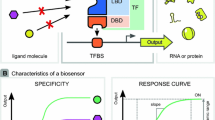
The protein-based biosensors were usually constructed by engineering aTFs, which could interact with specific small ligand molecules and change the activity of corresponding promoters (Fig. 4.1a) (Table 4.1) (De Paepe et al. 2017). The aTF typically consists of two function domains, namely the N-terminal ligand-binding domain (LBD) and the C-terminal DNA-binding domain (DBD). The binding of aTF on the transcription factor binding site (TFBS) of the promoter will increase or decrease the affinity of RNA polymerase (RNAP) to it, and the conformation changes of aTF induced by specific ligand will affect its binding to the promoter thus building a relationship between ligand concentration and promoter activity (Wan et al. 2019). Among the four general patterns of the aTF-mediated transcriptional regulation, patterns 3 and 4 were most employed due to the positive correlation between the input and output (Fig. 4.1b) (Mannan et al. 2017).
The two fundamental characteristics, namely responsive curve and specificity, were often used for the evaluation of the protein-based biosensor (Fig. 4.1c) (De Paepe et al. 2017). The responsive curve represents the relation between the input of ligand concentration and the output of promoter strength, which can be obtained by fitting the input and output into the Hill function as shown below:
where y is relative expression activity of the promoter (ymin and ymax are the minimum/maximum activities), x is the ligand concentration, K is the threshold, and n is the cooperativity (Meyer et al. 2019). And many important parameters of the biosensor could be acquired from the curve including basal, maximum, operational range, dynamic range, threshold, and sensitivity (Fig. 4.1c). Specificity determines the responsive of the biosensor to different ligand molecules.
4.1.1.2 Designing and Tuning Protein-Based Biosensors
In order to build a protein-based biosensor with favorable responsive curve in a host, specific aTF should be expressed properly, and applicable synthetic promoter needs to be designed and constructed. Sometimes, molecular modification on the aTF may be implemented to improve or change the specificity of biosensor (De Paepe et al. 2017). That is to say, the tuning of protein-based biosensor mainly focuses on aTF level and promoter level.
4.1.1.2.1 Tuning at aTF Level
To construct a protein-based biosensor responsive to a specific molecule, corresponding aTF must be chosen by consulting literatures or retrieving the databases such as RegulonDB (Gama-Castro et al. 2011), BRENDA (Placzek et al. 2017), and RegPrecise (Rodionov et al. 2013). Besides, transcriptome sequencing and analysis can also be used to identify specific aTF (Li et al. 2019). However, there may not be aTF in nature which responds to certain molecules. So the engineered aTFs responded to new non-natural ligands must be constructed, which could be achieved by the combination of rational design and directed evolution (Koch et al. 2019; Libis et al. 2016). For example, five amino acid positions located in the effector binding pocket (P8, T24, H80, Y82, and H93) of the L-arabinose-responsive aTF AraC were selected for simultaneous saturation mutagenesis, and the mutants that responded to mevalonate, triacetic acid lactone, and ectoine, respectively, were obtained by fluorescence-activated cell sorting (FACS)-mediated negative–positive dual screening (Chen et al. 2015; Tang et al. 2013; Tang and Cirino 2011). The computational design method is often used to reduce the design space. As an example, the Rosetta software was used in combination with single-residue saturation mutagenesis and error-prone PCR (epPCR)-based random mutagenesis for the construction of LacI mutants responding to fucose, gentiobiose, lactitol, and sucralose, respectively (Taylor et al. 2016). In addition, chimeric aTFs have also been built by fusing DBD and LBD from different proteins, and it is worth mentioning that the LBD could come from proteins other than aTF as long as it has demonstrable binding affinity to the ligand. For instance, benzoate-responsive aTFs were constructed by connecting benzoate LBDs to different DBDs with optimized linkers (Juárez et al. 2018).
The fundamental characteristics of the protein-based biosensors can also be optimized by introducing molecular modification into or tuning the expression level of the aTF. For example, the specificity of aTFMphR (that is derepressed by several naturally produced and semisynthetic macrolide antibiotics including erythromycin (ErA), josamycin, oleandomycin, narbomycin, methymycin, and pikromycin) to erythromycin was enhanced through epPCR and FACS; and its sensitivity was improved by introducing random mutagenesis to ribosome binding site (RBS) fortuning its expression level (Kasey et al. 2018).
4.1.1.2.2 Tuning at Promoter Level
To build a protein-based biosensor in a host, synthetic responsive promoters need to be designed and constructed by inserting the TFBS into the promoter of this strain because the native promoter regulated by the aTF may lose its activity there. For example, FA/acyl-CoA-responsive promoters were built by inserting the TFBS of aTF FadR into a phage lambda promoter and a phage T7 promoter, respectively, and TFBS of LacI was added into the constructed synthetic promoters to eliminate leaky expression (Zhang et al. 2012). In addition, the fundamental characteristics could be modulated by changing the starting engineered promoter or the position and numbers of the TFBS. As an example, Siewers and coworkers have constructed several malonyl-CoA biosensors in Saccharomyces cerevisiae by inserting the TFBS of aTF FapR (FapO) into five native promoters, and improved the dynamic range and reduced the basal by adjusting the position and numbers of FapO (Dabirian et al. 2019b).
4.1.2 RNA-Based Biosensors
4.1.2.1 The Functional Mechanism of RNA-Based Biosensors
The RNA-based biosensors could be constructed by engineering the cis-acting metabolite-responsive riboswitches, which consist of ligand-binding (aptamer) domains that could bind with specific ligand when its abundance exceeds a threshold and expression platform that control the gene expression by interacting with various gene expression apparatus (Table 4.2) (Serganov and Patel 2007). In the natural world, riboswitches responsive to numerous small molecules including ion, purines, and their derivatives, amino acids, phosphorylated sugar, and so on have been found, and they could modulate gene expression by controlling transcription, translation, mRNA stability, and splicing (Fig. 4.2) (Serganov and Nudler 2013). Because the regulations on genes expression are achieved by modulating the secondary structure of mRNAs, RNA-based biosensors possess faster responses compared with the protein-based biosensors. In addition, they have a good transplantable character on account of the protein-free control process (Topp et al. 2010). For example, the glucosamine-6-phosphate riboswitch of B. subtilis was directly used for high-throughput screening of N-acetylglucosamine high-producing strain in S. cerevisiae (Lee and Oh 2015). The RNA-based biosensors also function in a dose-dependent manner, hence their fundamental characteristics for evaluation are the same as the protein-based biosensors mentioned above (Chang et al. 2012).
4.1.2.2 Designing and Tuning RNA-Based Biosensors
Duo to functional mechanism of riboswitches, the RNA-based biosensors are easily to be designed and built by adding the natural or engineered riboswitches into mRNAs (usually on 5′ untranslated region (UTR)), and the responsive characteristics could be tuned by modifying their sequences in aptamer region or expression platform and getting the mutants using high-throughput screening (Jang and Jung 2018; Page et al. 2018).
The dynamic range of the RNA-based biosensors can be improved by changing their promoter or RBS, and the anti-RBS sequence on the riboswitch also needs to be modified if it functions in an RBS sequestering manner. For example, RBS sequence on expression platform of the pyrimido[4,5-d]pyrimidine-2,4-diamine (PPDA) riboswitch was exchanged with the E. coli consensus RBS sequence (AGGAGG) for enhanced maximum of the biosensor firstly, and then high-throughput fluorescence-activated cell sorting (FACS)-based selection/counter selection methodology was used to identify anti-RBS sequences that give riboswitches with optimal OFF and ON states. Introducing these modifications improved the maximal expression and dynamic range of the biosensor by 8.2-folds and 80-folds, respectively (Kent and Dixon 2019). As an another example, Jiang et al. improved the dynamic range of a L-tryptophan riboswitch-based biosensor by changing its promoter and copy number (Jang and Jung 2018). To modulate the operational range of the riboswitch-based biosensor, the aptamer region can be modified to change the affinity between ligand and riboswitch. For instance, the dose–response curve of a l-tryptophan riboswitch-based biosensor was shifted toward higher ligand concentrations by exchanging a low affinity aptamer (Jang and Jung 2018).
The ligand specificity of the RNA-based biosensors may be enhanced or changed by modifying the aptamer regions (Robinson et al. 2014). For instance, the specially responsive ligand of the natural adenine riboswitch was turned to ammeline or azacytosine by introducing site-directed mutagenesis at U47 and U51 sites on the aptamer region that are responsible for the interaction with the ligand molecule (Dixon et al. 2010). In addition, “non-natural” synthetic riboswitches could be designed and constructed using corresponding aptamers found in the natural world or built artificially (Darmostuk et al. 2014; Kinghorn et al. 2017; Sun and Zu 2015). For example, an l-tryptophan riboswitch was built by selecting the N10 sequences connecting l-tryptophan aptamer region that had been reported previously with RBS and dual selection module (tetA-sgfp) in vivo (Yang et al. 2013). In addition, a statistical thermodynamic model has been proposed for the aptamer-based artificial riboswitch design (Espah Borujeni et al. 2016). In another example, self-cleaving ribozyme-based artificial riboswitches have been built by linking the thiM aptamer domain from E. coli into stem III of a fast-cleaving hammerhead ribozyme (HHR) (Wieland et al. 2009).
It is worth mentioning that new artificial RNA aptamers that bind to specific ligands could be easily constructed using the technology called systematic evolution of ligands by exponential enrichment (SELEX) in vitro (Darmostuk et al. 2014), and then the new aptamers will be used for the building of corresponding riboswitches (Jang et al. 2017). Alternatively, riboswitches responsive to new ligand can be also constructed by directly introducing a random-sequence library into the aptamer domain of a native or ready-made riboswitch and then conducting multiple rounds of dual genetic selection and FACS screening in vivo. Using this method, theophylline riboswitch that possesses a 2.3-fold dynamic range was obtained from the native ThiM#2 riboswitch (Page et al. 2018).
4.2 The Application of Genetically Encoded Biosensors
4.2.1 The Application in High-Throughput Screening
Because the metabolic networks and their regulations are very complex in the cell, high-throughput screening (HTS) is often used to obtain the best producer from the mutant libraries of enzymes or pathways (Lim et al. 2018). The genetically encoded biosensors could couple the target products’ concentrations with expression levels of the reporters, and then the best producer can be obtained by adaptive evolution or FACS.
4.2.1.1 Screening by Adaptive Evolution
To carry out adaptive evolution, appropriate reporter needs to be chosen to link cell growth with product concentration. For the biosensors whose expression levels are positive correlation to the concentrations of ligands, resistance maker could be used. For example, a tetracycline resistance protein TetA was used as the reporter of the aTF-based biosensor for directed evolution of a heterologous biosynthetic pathway of 1-butanol in E. coli (Dietrich et al. 2013). As the biosensors whose expression levels possess negative correlation with the ligand concentrations, negative selection marker needs to be used. For example, cytosine deaminase that has a cytotoxicity was used as the reporter of the GlcN6P riboswitch-based biosensor for the screening of the best mutant of the key pathway enzyme GFA1 for N-acetylglucosamine (GlcNAc) synthesis in S. cerevisiae (Lee and Oh 2015).
4.2.1.2 Screening by Fluorescence-Activated Cell Sorting
The genetically encoded biosensors could also be applied for FACS by using fluorescence protein as the reporter. For instance, the yellow fluorescence protein (YFP) was acted as the reporter of a lysine biosensor in C. glutamicum, and then FACS was conducted for screening of pyruvate carboxylase variants created by error-prone PCR that enable improved l-lysine production from glucose (Kortmann et al. 2019).
4.2.2 The Application in Dynamic Metabolic Engineering
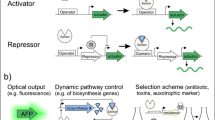
The genetically encoded biosensors also have wide applications in dynamic metabolic engineering, which is capable of dynamically coordinating the metabolic flux in a feedback manner and can avoid the adverse effects on cells caused by metabolic modification such as metabolic imbalance and accumulation of intermediate products (Lalwani et al. 2018; Shen et al. 2019; Xu 2018). Here, we divide these applications into three categories according to the regulation processes, namely dynamic pathway activation, dynamic pathway repression, and dynamic dual control (simultaneous activation and repression).
4.2.2.1 Dynamic Pathway Activation
Dynamic pathway activation can be used to redirect the flux from the native metabolism toward the target product by introducing a biosensor responsive to prevalent intermediate at the key branch points in the metabolic networks of the cell. For example, amalonyl-CoA biosensor was employed to alter the metabolic flux from central carbon metabolism into a heterologous 3-hydroxypropionic acid (3-HP) synthetic pathway by controlling the expression of the malonyl-CoA reductase derived from Chloroflexus aurantiacus, which enabled the dynamic switching between growth phase and production (David et al. 2016).
4.2.2.2 Dynamic Pathway Repression
The competitive pathways of target product were often knocked-outed to force more metabolic flux into the pathway of interest, while sometimes these competitive pathways may be necessary for the cell growth. In this situation, dynamic repression can be employed to redirect the flux toward target product. For instance, the lysine-OFF riboswitch was used to control the expression of citrate synthase (gltA), which is the key metabolic point of tricarboxylic acid (TCA) cycle, in a l-lysine-producing C. glutamicum strain, thus dynamically channel flux from central carbon metabolism into l-lysine synthesis (Zhou and Zeng 2015a). Similarly, a GlcN6P-OFF riboswitch was set as an intermediate metabolite biosensor that dynamically repressed the competitive pathways, namely peptidoglycan synthesis pathway and glycolysis pathway, in a GlcNAc-producing B. subtilis strain (Niu et al. 2018).
4.2.2.3 Dynamic Dual Control
To achieve the better and more precise control of the metabolic networks in a microbial cell factory, dynamic activation and repression on multiple targets simultaneously, which is widespread in the natural world, may be needed. This process can be realized by designing and building biosensors that possess opposite regulation effects. Xu et al. have constructed malonyl-CoA activating and repressing biosensors regulated by the aTF FapR, and controlled the malonyl-CoA source pathway (ACC) and the malonyl-CoA sink pathway (FAS) by the malonyl-CoA activating and repressing biosensors, respectively, which avoided the accumulation of intermediate product malonyl-CoA and balanced metabolism between cell growth and target product fatty acids formation (Xu et al. 2014). In another example, lysine-ON riboswitches were built by engineering a native lysine-OFF riboswitch from E. coli, and lysine-ON and lysine-OFF riboswitches were applied for the control of lysine transport protein and the key competitive pathway, namely TCA cycle, respectively (Zhou and Zeng 2015a).
Except for the double sensor mediated dynamically dual control, single biosensor-based dual control, which could be achieved by coupling the biosensor with some regulation tools acted as NOT gates, have also been reported. Yan and coworkers have presented a bifunctional dynamic control system based on biosensor and antisense RNA (as RNA), which can be used to upregulate and downregulate multiple genes simultaneously, and applied this system to achieve the dynamic flux distribution between native metabolism and the muconic acid biosynthetic pathway (Yang et al. 2018). In addition, the CRISPRi based NOT gate was also coupled with a biosensor to achieve the autonomous dual-control of metabolic flux in Bacillus subtilis (Wu et al. 2020).
4.3 Conclusions and Perspectives
Genetically encoded biosensors have been widely applied in the construction of efficient microbial cell factories. However, the building process of novel biosensors responsive to specific macular, which is the premise of all subsequent operations, is still time-consuming. Hence the computer-aided methods need to be further explored for accelerating biosensor design in the future. In addition, the biosensors-mediated feedback and dynamic regulation of the metabolic networks can be combined with the rising co-culture engineering strategy, which has been proved to be more advantageous in the synthesis of many products (Jones and Wang 2018), to achieve the coordination control of population dynamics. Furthermore, biosensors may also be used in the regulation of engineering spatial organization of metabolic enzymes, which can enhance flux into interested pathway and reduce their interactions with cellular background metabolism (Lee et al. 2012), for the reconstruction of the cell metabolism in space and time dimensions simultaneously.
References
Binder S, Schendzielorz G, Stäbler N, Krumbach K, Hoffmann K, Bott M, Eggeling L (2012) A high-throughput approach to identify genomic variants of bacterial metabolite producers at the single-cell level. Genome Biol 13:R40. https://doi.org/10.1186/gb-2012-13-5-r40
Chang AL, Wolf JJ, Smolke CD (2012) Synthetic RNA switches as a tool for temporal and spatial control over gene expression. Curr Opin Biotechnol 23:679–688. https://doi.org/10.1016/j.copbio.2012.01.005
Chen W, Zhang S, Jiang P, Yao J, He Y, Chen L, Gui X, Dong Z, Tang SY (2015) Design of an ectoine-responsive AraC mutant and its application in metabolic engineering of ectoine biosynthesis. Metab Eng 30:149–155. https://doi.org/10.1016/j.ymben.2015.05.004
Chen X-F, Xia X-X, Lee SY, Qian Z-G (2017) Engineering tunable biosensors for monitoring putrescine in Escherichia coli. Biotechnol Bioeng 115:1014. https://doi.org/10.1002/bit.26521
Choi SL, Rha E, Lee SJ, Kim H, Kwon K, Jeong YS, Rhee YH, Song JJ, Kim HS, Lee SG (2014) Toward a generalized and high-throughput enzyme screening system based on artificial genetic circuits. ACS Synth Biol 3:163–171. https://doi.org/10.1021/sb400112u
Chou HH, Keasling JD (2013) Programming adaptive control to evolve increased metabolite production. Nat Commun 4:1–8. https://doi.org/10.1038/ncomms3595
Cordova LT, Alper HS (2016) Central metabolic nodes for diverse biochemical production. Curr Opin Chem Biol 35:37–42. https://doi.org/10.1016/j.cbpa.2016.08.025
Dabirian Y, Gonçalves Teixeira P, Nielsen J, Siewers V, David F (2019a) FadR-based biosensor-assisted screening for genes enhancing fatty acyl-CoA pools in Saccharomyces cerevisiae. ACS Synth Biol 8(8):1788–1800. https://doi.org/10.1021/acssynbio.9b00118
Dabirian Y, Li X, Chen Y, David F, Nielsen J, Siewers V (2019b) Expanding the dynamic range of a transcription factor-based biosensor in Saccharomyces cerevisiae. ACS Synth Biol 8(9):1968–1975. https://doi.org/10.1021/acssynbio.9b00144
Darmostuk M, Rimpelova S, Gbelcova H, Ruml T (2014) Current approaches in SELEX: an update to aptamer selection technology. Biotechnol Adv 33:1141–1161. https://doi.org/10.1016/j.biotechadv.2015.02.008
David F, Nielsen J, Siewers V (2016) Flux control at the Malonyl-CoA node through hierarchical dynamic pathway regulation in Saccharomyces cerevisiae. ACS Synth Biol 5:224–233. https://doi.org/10.1021/acssynbio.5b00161
De Los Santos ELC, Meyerowitz JT, Mayo SL, Murray RM (2016) Engineering transcriptional regulator effector specificity using computational design and in vitro rapid prototyping: developing a vanillin sensor. ACS Synth Biol 5:287–295. https://doi.org/10.1021/acssynbio.5b00090
De Paepe B, Peters G, Coussement P, Maertens J, De Mey M (2017) Tailor-made transcriptional biosensors for optimizing microbial cell factories. J Ind Microbiol Biotechnol 44:623–645. https://doi.org/10.1007/s10295-016-1862-3
De Paepe B, Maertens J, Vanholme B, De Mey M (2019) Chimeric LysR-type transcriptional biosensors for customizing ligand specificity profiles toward flavonoids. ACS Synth Biol 8:318–331. https://doi.org/10.1021/acssynbio.8b00326
Dietrich J a, Shis DL, Alikhani A, Keasling JD (2013) Transcription factor-based screens and synthetic selections for microbial small-molecule biosynthesis. ACS Synth Biol 2:47–58. https://doi.org/10.1021/sb300091d
Dixon N, Duncan JN, Geerlings T, Dunstan MS, McCarthy JEG, Leys D, Micklefield J (2010) Reengineering orthogonally selective riboswitches. Proc Natl Acad Sci 107:2830–2835. https://doi.org/10.1073/pnas.0911209107
Doong SJ, Gupta A, Prather KLJ (2018) Layered dynamic regulation for improving metabolic pathway productivity in Escherichia coli. Proc Natl Acad Sci 115:2964–2969. https://doi.org/10.1073/pnas.1716920115
Eckdahl TT, Campbell AM, Heyer LJ, Poet JL, Blauch DN, Snyder NL, Atchley DT, Baker EJ, Brown M, Brunner EC, Callen SA, Campbell JS, Carr CJ, Carr DR, Chadinha SA, Chester GI, Chester J, Clarkson BR, Cochran KE, Doherty SE, Doyle C, Dwyer S, Edlin LM, Evans RA, Fluharty T, Frederick J, Galeota-Sprung J, Gammon BL, Grieshaber B, Gronniger J, Gutteridge K, Henningsen J, Isom B, Itell HL, Keffeler EC, Lantz AJ, Lim JN, McGuire EP, Moore AK, Morton J, Nakano M, Pearson SA, Perkins V, Parrish P, Pierson CE, Polpityaarachchige S, Quaney MJ, Slattery A, Smith KE, Spell J, Spencer M, Taye T, Trueblood K, Vrana CJ, Whitesides ET (2015) Programmed evolution for optimization of orthogonal metabolic output in bacteria. PLoS One 10:1–27. https://doi.org/10.1371/journal.pone.0118322
Endoh T, Sugimoto N (2015) Rational design and tuning of functional RNA switch to control an allosteric intermolecular interaction. Anal Chem 87:7628–7635. https://doi.org/10.1021/acs.analchem.5b00765
Espah Borujeni A, Mishler DM, Wang J, Huso W, Salis HM (2016) Automated physics-based design of synthetic riboswitches from diverse RNA aptamers. Nucleic Acids Res 44:1–13. https://doi.org/10.1093/nar/gkv1289
Furukawa K, Ramesh A, Zhou Z, Weinberg Z, Vallery T, Winkler WC, Breaker RR (2015) Bacterial riboswitches cooperatively bind Ni2+ or Co2+ ions and control expression of heavy metal transporters. Mol Cell 57:1088–1098. https://doi.org/10.1016/j.molcel.2015.02.009
Gama-Castro S, Salgado H, Peralta-Gil M, Santos-Zavaleta A, Muniz-Rascado L, Solano-Lira H, Jimenez-Jacinto V, Weiss V, García-Sotelo JS, López-Fuentes A, Porrón-Sotelo L, Alquicira-Hernández S, Medina-Rivera A, Martínez-Flores I, Alquicira-Hernández K, Martínez-Adame R, Bonavides-Martínez C, Miranda-Ríos J, Huerta AM, Mendoza-Vargas A, Collado-Torres L, Taboada B, Vega-Alvarado L, Olvera M, Olvera L, Grande R, Morett E, Collado-Vides J (2011) RegulonDB version 7.0: Transcriptional regulation of Escherichia coli K-12 integrated within genetic sensory response units (Gensor Units). Nucleic Acids Res 39:98–105. https://doi.org/10.1093/nar/gkq1110
Greenwald EC, Mehta S, Zhang J (2018) Genetically encoded fluorescent biosensors illuminate the spatiotemporal regulation of signaling networks. Chem Rev 118:11707–11794. https://doi.org/10.1021/acs.chemrev.8b00333
Hanko EKR, Minton NP, Malys N (2018) A transcription factor-based biosensor for detection of Itaconic acid. ACS Synth Biol 7:1436–1446. https://doi.org/10.1021/acssynbio.8b00057
Hao Z, Lou H, Zhu R, Zhu J, Zhang D, Zhao BS, Zeng S, Chen X, Chan J, He C, Chen PR (2014) The multiple antibiotic resistance regulator MarR is a copper sensor in Escherichia coli. Nat Chem Biol 10:21–28. https://doi.org/10.1038/nchembio.1380
Ho JCH, Pawar SV, Hallam SJ, Yadav VG (2018) An improved whole-cell biosensor for the discovery of lignin-transforming enzymes in functional metagenomic screens. ACS Synth Biol 7:392–398. https://doi.org/10.1021/acssynbio.7b00412
Jang S, Jung GY (2018) Systematic optimization of L-tryptophan riboswitches for efficient monitoring of the metabolite in Escherichia coli. Biotechnol Bioeng 115:266–271. https://doi.org/10.1002/bit.26448
Jang S, Jang S, Xiu Y, Kang TJ, Lee S-H, Koffas MAG, Jung GY (2017) Development of artificial riboswitches for monitoring of naringenin in vivo. ACS Synth Biol 6:2077–2085. https://doi.org/10.1021/acssynbio.7b00128
Jha RK, Kern TL, Fox DT, Strauss CEM (2014) Engineering an Acinetobacter regulon for biosensing and high-throughput enzyme screening in E. coli via flow cytometry. Nucleic Acids Res 42:8150–8160. https://doi.org/10.1093/nar/gku444
Jones JA, Wang X (2018) Use of bacterial co-cultures for the efficient production of chemicals. Curr Opin Biotechnol 53:33–38. https://doi.org/10.1016/j.copbio.2017.11.012
Juárez JF, Lecube-Azpeitia B, Brown SL, Johnston CD, Church GM (2018) Biosensor libraries harness large classes of binding domains for construction of allosteric transcriptional regulators. Nat Commun 9:3101. https://doi.org/10.1038/s41467-018-05525-6
Kasey CM, Zerrad M, Li Y, Cropp TA, Williams GJ (2018) Development of transcription factor-based designer macrolide biosensors for metabolic engineering and synthetic biology. ACS Synth Biol 7:227–239. https://doi.org/10.1021/acssynbio.7b00287
Kent R, Dixon N (2019) Systematic evaluation of genetic and environmental factors affecting performance of translational riboswitches. ACS Synth Biol 8:884–901. https://doi.org/10.1021/acssynbio.9b00017
Ketterer S, Gladis L, Kozica A, Meier M (2016) Engineering and characterization of fluorogenic glycine riboswitches. Nucleic Acids Res 44:5983–5992. https://doi.org/10.1093/nar/gkw465
Kinghorn AB, Fraser LA, Lang S, Shiu SCC, Tanner JA (2017) Aptamer bioinformatics. Int J Mol Sci 18(12):2516. https://doi.org/10.3390/ijms18122516
Klauser B, Atanasov J, Siewert LK, Hartig JS (2015) Ribozyme-based aminoglycoside switches of gene expression engineered by genetic selection in S. cerevisiae. ACS Synth Biol 4:516–525. https://doi.org/10.1021/sb500062p
Knudsen JD, Carlquist M, Gorwa-Grauslund M (2014) NADH-dependent biosensor in Saccharomyces cerevisiae: principle and validation at the single cell level. AMB Express 4:1–12. https://doi.org/10.1186/s13568-014-0081-4
Koch M, Pandi A, Borkowski O, Cardoso Batista A, Faulon J-L (2019) Custom-made transcriptional biosensors for metabolic engineering. Curr Opin Biotechnol 59:78–84. https://doi.org/10.1016/j.copbio.2019.02.016
Kortmann M, Mack C, Baumgart M, Bott M (2019) Pyruvate carboxylase variants enabling improved lysine production from glucose identified by biosensor-based high-throughput fluorescence-activated cell sorting screening. ACS Synth Biol 8:274–281. https://doi.org/10.1021/acssynbio.8b00510
Kwon KK, Yeom SJ, Lee DH, Jeong KJ, Lee SG (2018) Development of a novel cellulase biosensor that detects crystalline cellulose hydrolysis using a transcriptional regulator. Biochem Biophys Res Commun 495:1328–1334. https://doi.org/10.1016/j.bbrc.2017.11.157
Lalwani MA, Zhao EM, Avalos JL (2018) Current and future modalities of dynamic control in metabolic engineering. Curr Opin Biotechnol 52:56–65. https://doi.org/10.1016/j.copbio.2018.02.007
Leavitt JM, Wagner JM, Tu CC, Tong A, Liu Y, Alper HS (2017) Biosensor-enabled directed evolution to improve muconic acid production in Saccharomyces cerevisiae. Biotechnol J 12:1–9. https://doi.org/10.1002/biot.201600687
Lee SW, Oh MK (2015) A synthetic suicide riboswitch for the high-throughput screening of metabolite production in Saccharomyces cerevisiae. Metab Eng 28:143–150. https://doi.org/10.1016/j.ymben.2015.01.004
Lee H, DeLoache WC, Dueber JE (2012) Spatial organization of enzymes for metabolic engineering. Metab Eng 14:242–251. https://doi.org/10.1016/j.ymben.2011.09.003
Li L, Tu R, Song G, Cheng J, Chen W, Li L, Wang L, Wang Q (2019) Development of a synthetic 3-dehydroshikimate biosensor in Escherichia coli for metabolite monitoring and genetic screening. ACS Synth Biol 8(2):297–306. https://doi.org/10.1021/acssynbio.8b00317
Libis V, Delépine B, Faulon JL (2016) Sensing new chemicals with bacterial transcription factors. Curr Opin Microbiol 33:105–112. https://doi.org/10.1016/j.mib.2016.07.006
Lim HG, Jang S, Jang S, Seo SW, Jung GY (2018) Design and optimization of genetically encoded biosensors for high-throughput screening of chemicals. Curr Opin Biotechnol 54:18–25. https://doi.org/10.1016/j.copbio.2018.01.011
Liu L, Guan N, Li J, Shin H, Du G, Chen J (2017a) Development of GRAS strains for nutraceutical production using systems and synthetic biology approaches: advances and prospects. Crit Rev Biotechnol 37:139–150. https://doi.org/10.3109/07388551.2015.1121461
Liu Y, Zhuang Y, Ding D, Xu Y, Sun J, Zhang D (2017b) Biosensor-based evolution and elucidation of a biosynthetic pathway in Escherichia coli. ACS Synth Biol 6:837–848. https://doi.org/10.1021/acssynbio.6b00328
Liu C, Zhang B, Liu YM, Yang KQ, Liu SJ (2018) New intracellular shikimic acid biosensor for monitoring shikimate synthesis in Corynebacterium glutamicum. ACS Synth Biol 7:591–601. https://doi.org/10.1021/acssynbio.7b00339
Luo X, Reiter MA, D’Espaux L, Wong J, Denby CM, Lechner A, Zhang Y, Grzybowski AT, Harth S, Lin W, Lee H, Yu C, Shin J, Deng K, Benites VT, Wang G, Baidoo EEK, Chen Y, Dev I, Petzold CJ, Keasling JD (2019) Complete biosynthesis of cannabinoids and their unnatural analogues in yeast. Nature 567:123–126. https://doi.org/10.1038/s41586-019-0978-9
Lynch SA, Gallivan JP (2009) A flow cytometry-based screen for synthetic riboswitches. Nucleic Acids Res 37:184–192. https://doi.org/10.1093/nar/gkn924
Lynch SA, Desai SK, Sajja HK, Gallivan JP (2007) A high-throughput screen for synthetic riboswitches reveals mechanistic insights into their function. Chem Biol 14:173–184. https://doi.org/10.1016/j.chembiol.2006.12.008
Machado LFM, Dixon N (2016) Development and substrate specificity screening of an: in vivo biosensor for the detection of biomass derived aromatic chemical building blocks. Chem Commun 52:11402–11405. https://doi.org/10.1039/c6cc04559f
Mannan AA, Liu D, Zhang F, Oyarzún DA (2017) Fundamental design principles for transcription-factor-based metabolite biosensors. ACS Synth Biol 6:1851–1859. https://doi.org/10.1021/acssynbio.7b00172
Merulla D, Van Der Meer JR (2016) Regulatable and modulable background expression control in prokaryotic synthetic circuits by auxiliary repressor binding sites. ACS Synth Biol 5:36–45. https://doi.org/10.1021/acssynbio.5b00111
Meyer A, Pellaux R, Potot S, Becker K, Hohmann HP, Panke S, Held M (2015) Optimization of a whole-cell biocatalyst by employing genetically encoded product sensors inside nanolitre reactors. Nat Chem 7:673–678. https://doi.org/10.1038/nchem.2301
Meyer AJ, Segall-Shapiro TH, Glassey E, Zhang J, Voigt CA (2019) Escherichia coli “marionette” strains with 12 highly optimized small-molecule sensors. Nat Chem Biol 15:196–204. https://doi.org/10.1038/s41589-018-0168-3
Michener JK, Thodey K, Liang JC, Smolke CD (2012) Applications of genetically-encoded biosensors for the construction and control of biosynthetic pathways. Metab Eng 14:212–222. https://doi.org/10.1016/j.ymben.2011.09.004
Muranaka N, Sharma V, Nomura Y, Yokobayashi Y (2009) Efficient design strategy for whole-cell and cell-free biosensors based on engineered riboswitches. Anal Lett 42:108–122. https://doi.org/10.1080/00032710802568556
Mustafi N, Grünberger A, Kohlheyer D, Bott M, Frunzke J (2012) The development and application of a single-cell biosensor for the detection of l-methionine and branched-chain amino acids. Metab Eng 14:449–457. https://doi.org/10.1016/j.ymben.2012.02.002
Ng CY, Khodayari A, Chowdhury A, Maranas CD (2015) Advances in de novo strain design using integrated systems and synthetic biology tools. Curr Opin Chem Biol 28:105–114. https://doi.org/10.1016/j.cbpa.2015.06.026
Niu T, Liu Y, Li J, Koffas M, Du G, Alper HS, Liu L (2018) Engineering a glucosamine-6-phosphate responsive glmS ribozyme switch enables dynamic control of metabolic flux in Bacillus subtilis for overproduction of N-acetylglucosamine. ACS Synth Biol 7:2423–2435. https://doi.org/10.1021/acssynbio.8b00196
Nomura Y, Yokobayashi Y (2007) Reengineering a natural riboswitch by dual genetic selection. J Am Chem Soc 129:13814–13815. https://doi.org/10.1021/ja076298b
Page K, Shaffer J, Lin S, Zhang M, Liu JM (2018) Engineering riboswitches in vivo using dual genetic selection and fluorescence-activated cell sorting. ACS Synth Biol 7(9):2000–2006. https://doi.org/10.1021/acssynbio.8b00099
Peters G, De Paepe B, De Wannemaeker L, Duchi D, Maertens J, Lammertyn J, De Mey M (2018) Development of N-acetylneuraminic acid responsive biosensors based on the transcriptional regulator NanR. Biotechnol Bioeng 115:1855–1865. https://doi.org/10.1002/bit.26586
Placzek S, Schomburg I, Chang A, Jeske L, Ulbrich M, Tillack J, Schomburg D (2017) BRENDA in 2017: new perspectives and new tools in BRENDA. Nucleic Acids Res 45:D380–D388. https://doi.org/10.1093/nar/gkw952
Porter EB, Polaski JT, Morck MM, Batey RT (2017) Recurrent RNA motifs as scaffolds for genetically encodable small-molecule biosensors. Nat Chem Biol 13:295–301. https://doi.org/10.1038/nchembio.2278
Ravikumar S, Baylon MG, Park SJ, Choi J i (2017) Engineered microbial biosensors based on bacterial two-component systems as synthetic biotechnology platforms in bioremediation and biorefinery. Microb Cell Factories 16:1–10. https://doi.org/10.1186/s12934-017-0675-z
Rebets Y, Schmelz S, Gromyko O, Tistechok S, Petzke L, Scrima A, Luzhetskyy A (2018) Design, development and application of whole-cell based antibiotic-specific biosensor. Metab Eng 47:263. https://doi.org/10.1016/j.ymben.2018.03.019
Robinson CJ, Vincent HA, Wu MC, Lowe PT, Dunstan MS, Leys D, Micklefield J (2014) Modular riboswitch toolsets for synthetic genetic control in diverse bacterial species. J Am Chem Soc 136:10615–10624. https://doi.org/10.1021/ja502873j
Rode AB, Endoh T, Sugimoto N (2015) Tuning riboswitch-mediated gene regulation by rational control of aptamer ligand binding properties. Angew Chem Int Ed Engl 54:905–909. https://doi.org/10.1002/anie.201407385
Rodionov DA, Kazanov MD, Kazakov AE, Sutormin RA, Leyn SA, Ravcheev DA, Kovaleva GY, Arkin AP, Dubchak I, Riehl W, Novichkov PS (2013) RegPrecise 3.0—a resource for genome-scale exploration of transcriptional regulation in bacteria. BMC Genomics 14:745. https://doi.org/10.1186/1471-2164-14-745
Rogers JK, Church GM (2016) Genetically encoded sensors enable real-time observation of metabolite production. Proc Natl Acad Sci 113:2388–2393. https://doi.org/10.1073/pnas.1600375113
Rogers JK, Guzman CD, Taylor ND, Raman S, Anderson K, Church GM (2015) Synthetic biosensors for precise gene control and real-time monitoring of metabolites. Nucleic Acids Res 43:7648–7660. https://doi.org/10.1093/nar/gkv616
Saeki K, Tominaga M, Kawai-Noma S, Saito K, Umeno D (2016) Rapid diversification of beti-based transcriptional switches for the control of biosynthetic pathways and genetic circuits. ACS Synth Biol 5:1201–1210. https://doi.org/10.1021/acssynbio.5b00230
Schendzielorz G, Dippong M, Grünberger A, Kohlheyer D, Yoshida A, Binder S, Nishiyama C, Nishiyama M, Bott M, Eggeling L (2014) Taking control over control: use of product sensing in single cells to remove flux control at key enzymes in biosynthesis pathways. ACS Synth Biol 3:21–29. https://doi.org/10.1021/sb400059y
Serganov A, Nudler E (2013) A decade of riboswitches. Cell 152:17–24. https://doi.org/10.1016/j.cell.2012.12.024
Serganov A, Patel DJ (2007) Ribozymes, riboswitches and beyond: regulation of gene expression without proteins. Nat Rev Genet 8:776–790. https://doi.org/10.1038/nrg2172
Shen X, Wang J, Li C, Yuan Q, Yan Y (2019) Dynamic gene expression engineering as a tool in pathway engineering. Curr Opin Biotechnol 59:122–129. https://doi.org/10.1016/j.copbio.2019.03.019
Siedler S, Schendzielorz G, Binder S, Eggeling L, Bringer S, Bott M (2014a) SoxR as a single-cell biosensor for NADPH-consuming enzymes in Escherichia coli. ACS Synth Biol 3:41–47. https://doi.org/10.1021/sb400110j
Siedler S, Stahlhut SG, Malla S, Maury JÔ, Neves AR (2014b) Novel biosensors based on flavonoid-responsive transcriptional regulators introduced into Escherichia coli. Metab Eng 21:2–8. https://doi.org/10.1016/j.ymben.2013.10.011
Siedler S, Khatri NK, Zsohár A, Kjærbølling I, Vogt M, Hammar P, Nielsen CF, Marienhagen J, Sommer MOA, Joensson HN (2017) Development of a bacterial biosensor for rapid screening of yeast p-coumaric acid production. ACS Synth Biol 6:1860–1869. https://doi.org/10.1021/acssynbio.7b00009
Sinha J, Reyes SJ, Gallivan JP (2010) Reprogramming bacteria to seek and destroy an herbicide. Nat Chem Biol 6:464–470. https://doi.org/10.1038/nchembio.369
Skjoedt ML, Snoek T, Kildegaard KR, Arsovska D, Eichenberger M, Goedecke TJ, Rajkumar AS, Zhang J, Kristensen M, Lehka BJ, Siedler S, Borodina I, Jensen MK, Keasling JD (2016) Engineering prokaryotic transcriptional activators as metabolite biosensors in yeast. Nat Chem Biol 12:951–958. https://doi.org/10.1038/nchembio.2177
Stoddard CD, Widmann J, Trausch JJ, Marcano-Velázquez JG, Knight R, Batey RT (2013) Nucleotides adjacent to the ligand-binding pocket are linked to activity tuning in the purine riboswitch. J Mol Biol 425:1596–1611. https://doi.org/10.1016/j.jmb.2013.02.023
Su Y, Hickey SF, Keyser SGL, Hammond MC (2016) In vitro and in vivo enzyme activity screening via RNA-based fluorescent biosensors for S-Adenosyl- l-homocysteine (SAH). J Am Chem Soc 138:7040–7047. https://doi.org/10.1021/jacs.6b01621
Sun H, Zu Y (2015) A highlight of recent advances in aptamer technology and its application. Molecules 20:11959–11980. https://doi.org/10.3390/molecules200711959
Tang SY, Cirino PC (2011) Design and application of a mevalonate-responsive regulatory protein. Angew Chem Int Ed Engl 50:1084–1086. https://doi.org/10.1002/anie.201006083
Tang SY, Qian S, Akinterinwa O, Frei CS, Gredell J a, Cirino PC (2013) Screening for enhanced triacetic acid lactone production by recombinant Escherichia coli expressing a designed triacetic acid lactone reporter. J Am Chem Soc 135:10099–10103. https://doi.org/10.1021/ja402654z
Taylor ND, Garruss AS, Moretti R, Chan S, Arbing MA, Cascio D, Rogers JK, Isaacs FJ, Kosuri S, Baker D, Fields S, Church GM, Raman S (2015) Engineering an allosteric transcription factor to respond to new ligands. Nat Methods 13:177–183. https://doi.org/10.1038/nmeth.3696
Taylor ND, Garruss AS, Moretti R, Chan S, Arbing MA, Cascio D, Rogers JK, Isaacs FJ, Kosuri S, Baker D, Fields S, Church GM, Raman S (2016) Engineering an allosteric transcription factor to respond to new ligands. Nat Methods 13:177–183. https://doi.org/10.1038/nmeth.3696
Topp S, Reynoso CMK, Seeliger JC, Goldlust IS, Desai SK, Murat D, Shen A, Puri AW, Komeili A, Bertozzi CR (2010) Synthetic riboswitches that induce gene expression in diverse bacterial species. Appl Environ Microbiol 76:7881–7884
Trabelsi H, Koch M, Faulon JL (2018) Building a minimal and generalizable model of transcription factor-based biosensors: showcasing flavonoids. Biotechnol Bioeng 115(9):2292–2304. https://doi.org/10.1002/bit.26726
Uchiyama T, Watanabe K (2008) Substrate-induced gene expression (SIGEX) screening of metagenome libraries. Nat Protoc 3:1202–1212. https://doi.org/10.1038/nprot.2008.96
Umeyama T, Okada S, Ito T (2013) Synthetic gene circuit-mediated monitoring of endogenous metabolites: identification of GAL11 as a novel multicopy enhancer of S-adenosylmethionine level in yeast. ACS Synth Biol 2:425–430. https://doi.org/10.1021/sb300115n
van Sint Fiet S, van Beilen JB, Witholt B (2006) Selection of biocatalysts for chemical synthesis. Proc Natl Acad Sci 103:1693–1698. https://doi.org/10.1073/pnas.0504733102
Wachsmuth M, Findeiß S, Weissheimer N, Stadler PF, Mörl M (2013) De novo design of a synthetic riboswitch that regulates transcription termination. Nucleic Acids Res 41:2541–2551. https://doi.org/10.1093/nar/gks1330
Wan X, Marsafari M, Xu P (2019) Engineering metabolite-responsive transcriptional factors to sense small molecules in eukaryotes: current state and perspectives. Microb Cell Factories 18:61. https://doi.org/10.1186/s12934-019-1111-3
Wang J, Gao D, Yu X, Li W, Qi Q (2015) Evolution of a chimeric aspartate kinase for L-lysine production using a synthetic RNA device. Appl Microbiol Biotechnol 99:8527–8536. https://doi.org/10.1007/s00253-015-6615-0
Wang M, Li S, Zhao H (2016) Design and engineering of intracellular-metabolite-sensing/regulation gene circuits in Saccharomyces cerevisiae. Biotechnol Bioeng 113:206–215. https://doi.org/10.1002/bit.25676
Watstein DM, McNerney MP, Styczynski MP (2015) Precise metabolic engineering of carotenoid biosynthesis in Escherichia coli towards a low-cost biosensor. Metab Eng 31:171–180. https://doi.org/10.1016/j.ymben.2015.06.007
Weigand JE, Sanchez M, Gunnesch EB, Zeiher S, Schroeder R, Suess B (2008) Screening for engineered neomycin riboswitches that control translation initiation. RNA 14:89–97. https://doi.org/10.1261/rna.772408
Wieland M, Hartig JS (2008) Improved aptazyme design and in vivo screening enable riboswitching in bacteria. Angew Chem Int Ed Engl 47:2604–2607. https://doi.org/10.1002/anie.200703700
Wieland M, Benz A, Klauser B, Hartig JS (2009) Artificial ribozyme switches containing natural riboswiteh aptamer domains. Angew Chem Int Ed Engl 48:2715–2718. https://doi.org/10.1002/anie.200805311
Win MN, Smolke CD (2007) A modular and extensible RNA-based gene-regulatory platform for engineering cellular function. Proc Natl Acad Sci 104:14283–14288. https://doi.org/10.1073/pnas.0703961104
Woolston BM, Roth T, Kohale I, Liu DR, Stephanopoulos G (2018) Development of a formaldehyde biosensor with application to synthetic methylotrophy. Biotechnol Bioeng 115:206–215. https://doi.org/10.1002/bit.26455
Wu Y, Chen T, Liu Y, Tian R, Lv X, Li J, Du G, Chen J, Ledesma-Amaro R, Liu L (2020) Design of a programmable biosensor-CRISPRi genetic circuits for dynamic and autonomous dual-control of metabolic flux in Bacillus subtilis. Nucleic Acids Res 48(2):996–1009
Xiao Y, Bowen CH, Liu D, Zhang F (2016) Exploiting nongenetic cell-to-cell variation for enhanced biosynthesis. Nat Chem Biol 12:339–344. https://doi.org/10.1038/nchembio.2046
Xiao Y, Jiang W, Zhang F (2017) Developing a genetically encoded, cross-species biosensor for detecting ammonium and regulating biosynthesis of cyanophycin. ACS Synth Biol 6:1807–1815. https://doi.org/10.1021/acssynbio.7b00069
Xiong D, Lu S, Wu J, Liang C, Wang W, Wang W, Jin JM, Tang SY (2017) Improving key enzyme activity in phenylpropanoid pathway with a designed biosensor. Metab Eng 40:115–123. https://doi.org/10.1016/j.ymben.2017.01.006
Xiu Y, Jang S, Jones JA, Zill NA, Linhardt RJ, Yuan Q, Jung GY, Koffas MAG (2017) Naringenin-responsive riboswitch-based fluorescent biosensor module for Escherichia coli co-cultures. Biotechnol Bioeng 114:2235–2244. https://doi.org/10.1002/bit.26340
Xu P (2018) Production of chemicals using dynamic control of metabolic fluxes. Curr Opin Biotechnol 53:12–19. https://doi.org/10.1016/j.copbio.2017.10.009
Xu P, Li L, Zhang F, Stephanopoulos G, Koffas M (2014) Improving fatty acids production by engineering dynamic pathway regulation and metabolic control. Proc Natl Acad Sci 111:11299–11304. https://doi.org/10.1073/pnas.1406401111
Yang J, Seo SW, Jang S, Shin S-I, Lim CH, Roh T-Y, Jung GY (2013) Synthetic RNA devices to expedite the evolution of metabolite-producing microbes. Nat Commun 4:1413. https://doi.org/10.1038/ncomms2404
Yang P, Wang J, Pang Q, Zhang F, Wang J, Wang Q, Qi Q (2017) Pathway optimization and key enzyme evolution of N-acetylneuraminate biosynthesis using an in vivo aptazyme-based biosensor. Metab Eng 43:21–28. https://doi.org/10.1016/j.ymben.2017.08.001
Yang Y, Lin Y, Wang J, Wu Y, Zhang R, Cheng M, Shen X, Wang J, Chen Z, Li C, Yuan Q, Yan Y (2018) Sensor-regulator and RNAi based bifunctional dynamic control network for engineered microbial synthesis. Nat Commun 9:1–10. https://doi.org/10.1038/s41467-018-05466-0
You M, Litke JL, Jaffrey SR (2015) Imaging metabolite dynamics in living cells using a spinach-based riboswitch. Proc Natl Acad Sci U S A 112:E2756–E2765. https://doi.org/10.1073/pnas.1504354112
Zhang F, Carothers JM, Keasling JD (2012) Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat Biotechnol 30:354–359. https://doi.org/10.1038/nbt.2149
Zhang J, Barajas JF, Burdu M, Ruegg TL, Dias B, Keasling JD (2017) Development of a transcription factor-based lactam biosensor. ACS Synth Biol 6:439–445. https://doi.org/10.1021/acssynbio.6b00136
Zheng S, Hou J, Zhou Y, Fang H, Wang TT, Liu F, Wang FS, Sheng JZ (2018) One-pot two-strain system based on glucaric acid biosensor for rapid screening of myo-inositol oxygenase mutations and glucaric acid production in recombinant cells. Metab Eng 49:212–219. https://doi.org/10.1016/j.ymben.2018.08.005
Zhou LB, Zeng AP (2015a) Exploring lysine riboswitch for metabolic flux control and improvement of L-lysine synthesis in Corynebacterium glutamicum. ACS Synth Biol 4:729–734. https://doi.org/10.1021/sb500332c
Zhou LB, Zeng AP (2015b) Engineering a lysine-ON riboswitch for metabolic control of lysine production in Corynebacterium glutamicum. ACS Synth Biol 4:1335–1340. https://doi.org/10.1021/acssynbio.5b00075
Zhou S, Ainala SK, Seol E, Nguyen TT, Park S (2015) Inducible gene expression system by 3-hydroxypropionic acid. Biotechnol Biofuels 8:1–8. https://doi.org/10.1186/s13068-015-0353-5
Zhou YJ, Kerkhoven EJ, Nielsen J (2018) Barriers and opportunities in bio-based production of hydrocarbons. Nat Energy 3:925–935. https://doi.org/10.1038/s41560-018-0197-x
Zhu X, Wang X, Zhang C, Wang X, Gu Q (2015) A riboswitch sensor to determine vitamin B12 in fermented foods. Food Chem 175:523–528. https://doi.org/10.1016/j.foodchem.2014.11.163
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Wu, Y., Du, G., Chen, J., Liu, L. (2020). Genetically Encoded Biosensors and Their Applications in the Development of Microbial Cell Factories. In: Singh, V., Singh, A., Bhargava, P., Joshi, M., Joshi, C. (eds) Engineering of Microbial Biosynthetic Pathways. Springer, Singapore. https://doi.org/10.1007/978-981-15-2604-6_4
Download citation
DOI: https://doi.org/10.1007/978-981-15-2604-6_4
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-2603-9
Online ISBN: 978-981-15-2604-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)