Abstract
Mature sakacin A (SakA, encoded by sapA) and its cognate immunity protein (SakI, encoded by sapiA), and two SakA-derived chimeras mimicking the N-terminal end of mature enterocin P (EntP/SakA) and mature enterocin A (EntA/SakA) together with SakI, were fused to different signal peptides (SP) and cloned into the protein expression vectors pNZ8048 and pMG36c for evaluation of their production and functional expression by different lactic acid bacteria. The amount, antimicrobial activity, and specific antimicrobial activity of SakA and its chimeras produced by Lactococcus lactis subsp. cremoris NZ9000 depended on the SP and the expression vector. Only L. lactis NZ9000 (pNUPS), producing EntP/SakA, showed higher bacteriocin production and antimicrobial activity than the natural SakA-producer Lactobacillus sakei Lb706. The lower antimicrobial activity of the SakA-producer L. lactis NZ9000 (pNUS) and that of the EntA/SakA-producer L. lactis NZ9000 (pNUAS) could be ascribed to secretion of truncated bacteriocins. On the other hand, of the Lb. sakei Lb706 cultures transformed with the pMG36c-derived vectors only Lb. sakei Lb706 (pGUS) overproducing SakA showed a higher antimicrobial activity than Lb. sakei Lb706. Finally, cloning of SakA and EntP/SakA into pPICZαA and pKLAC2 permitted the production of SakA and EntP/SakA by recombinant Pichia pastoris X-33 and Kluyveromyces lactis GG799 derivatives although their antimicrobial activity was lower than expected from their production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lactic acid bacteria (LAB) secrete small, cationic, heat-stable antimicrobial peptides, commonly known as bacteriocins and many are more active than conventional antibiotics against pathogenic and drug-resistant Gram-positive bacteria, yet display no toxicity towards eukaryotic cells [16, 35]. Accordingly, bacteriocins produced by LAB may find use as natural antimicrobial peptides in food, medical, veterinary, and animal production applications, whereas bacteriocin-producing LAB could be evaluated for potential use as probiotics [8, 9, 23, 41]. However, the low bacteriocin yields obtained after their purification from natural producers and the high cost of synthetic bacteriocin synthesis drive the exploration of recombinant microbial systems for the biotechnological production of bacteriocins.
Bacteriocins produced by LAB may be classified into the class I lantibiotics, containing post-translationally modified amino acids, and the class II nonlantibiotics, containing non-modified amino acids. Class II bacteriocins may be further subdivided into the (1) pediocin-like (class IIa) bacteriocins, (2) two-peptide (class IIb) bacteriocins, (3) cyclic (class IIc) bacteriocins, and (4) nonpediocin one-peptide linear (class IId) bacteriocins [16, 43]. All pediocin-like bacteriocins have similar amino acid sequences, especially in their rather hydrophilic N-terminal half, which contains a disulfide bridge and a common YGNGVxC “pediocin box” motif, followed by a hinge and a somewhat more hydrophobic and diverse C-terminal half [31]. The target receptor for class IIa bacteriocins on sensitive cells has been identified as proteins of the sugar transporter mannose phosphotransferase system (Man-PTS) [21]. It has been suggested that the conserved N-terminal half of these bacteriocins initially interacts with the receptor to enable the C-terminal half to engage in more profound and active interactions with the receptor rendering the permeases open as pores, thereby causing leakage of solutes, disruption of membrane integrity, and eventually cell death [35].
Most class IIa bacteriocins are synthesized as biologically inactive precursors or prepeptides containing an N-terminal extension of the so-called double-glycine type (leader sequence) that is cleaved concomitantly with export across the cytoplasmic membrane by dedicated ATP-binding cassette transporters (ABC transporters) and their accessory proteins [32]. However, many secreted prokaryotic proteins and a few bacteriocins such as enterocin P [14] and hiracin JM79 [44] contain N-terminal extensions of the so-called Sec-type (signal peptide) which are proteolytically cleaved concomitantly with peptide externalization by the general secretory pathway (GSP) or Sec-dependent pathway. Secretory proteins are equipped with an N-terminal signal peptide (SP) that functions as a target and recognition signal for signal peptidases that remove the SP from the translocated protein, resulting in the extracellular release of the mature protein or peptide [24, 42]. Accordingly, the SP of secretory proteins may drive the access of fused mature proteins or peptides to the GSP for their secretion by the heterologous producer cells.
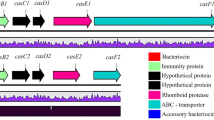
Sakacin A (SakA) is a class IIa, pediocin-like, antilisterial bacteriocin produced by Lactobacillus sakei Lb706 [33]. The sap locus responsible for SakA production in Lb. sakei Lb706 is located in a 60-kb pLSA60 plasmid harboring the sapAIPhKRTE operon. The genes sapA and saiA encode the 59 amino acid SakA precursor, consisting of a 18 amino acid leader sequence (LS sapA ) and the 41 amino acid mature bacteriocin (SakA), and the 90 amino acid immunity protein (SakI), respectively. The synthesis of SakA in Lb. sakei Lb706 is a temperature-sensitive process regulated by the peptide pheromone through a three-component regulatory system. In this respect, sap-Ph, sapK, and sapR encode the pheromone induction factor, the histidine protein kinase, and the response regulator, respectively. Finally, sapT and sapE encode an ABC transporter and its cognate accessory protein [20].
The production of SakA and immunity to the bacteriocin has been obtained in the recombinant non-bacteriocin-producing host Lb. sakei Lb790 but not in other LAB, probably because signal-transducing systems were not functioning properly [1]. Accordingly, as a further attempt to improve the production and functional expression of this tightly regulated bacteriocin by other microbial hosts we have evaluated the fusion of mature sakacin A (SakA, encoded by sapA) and its cognate immunity protein (SakI, encoded by sapiA), and of two SakA derived chimeras mimicking the N-terminal end of mature enterocin P (EntP/SakA) and mature enterocin A (EntA/SakA) to different signal peptides and their expression by Lactococcus lactis subsp. cremoris NZ9000 and Lb. sakei Lb706. Moreover, the cloning of SakA and the EntP/SakA-derived chimera in different yeast protein expression vectors has also permitted the evaluation of their functional expression by the yeasts Pichia pastoris X-33 and Kluyveromyces lactis GG799.
Materials and methods
Microbial strains, plasmids, and growth conditions
The microbial strains and plasmids used in this study are listed in Table 1. Lactobacillus sakei Lb706 was used as the source of sapA (SakA) and saiA (SakI), whereas L. lactis MG1363, Enterococcus faecium P13, and E. hirae DCH5 were used as the source of the signal peptides Usp45 (SP usp45 ), EntP (SP entP ), and HirJM79 (SP hirJM79 ), respectively. The primary amino acid and nucleotide sequence of the SP usp45 , SP entP , and SP hirJM79 are given by the GenBank accession numbers M60178, AF005278, and DQ664500, respectively. The GenBank accession number for the structural sakacin A (sapA) and immunity (saiA) genes of Lb. sakei Lb706 is Z46867. The lactococcal strains were propagated at 32 °C in M17 broth (Oxoid Ltd., Basingstoke, UK) supplemented with 0.5 % (w/v) glucose (GM17). The enterococcal strains and the lactobacilli were grown in MRS broth (Oxoid) at 32 °C. Pichia pastoris X-33 (Invitrogen Life Technologies, Barcelona, Spain) and K. lactis GG799 (New England Biolabs, Ipswich, MA, USA) were cultured in YPD medium (Sigma-Aldrich Inc., St. Louis, MO, USA) at 30 °C with shaking. Escherichia coli JM109 (Invitrogen) was grown in LB broth (Sigma) at 37 °C with shaking. Listeria spp. strains were cultured in BHI broth (Oxoid) at 32 °C. Agar plates were made by addition of 1.5 % (w/v) agar (Oxoid) to the liquid media. When necessary, chloramphenicol (Sigma), Zeocin (Invitrogen), and ampicillin (Sigma) were used at a concentration of 5, 25, and 50 μg/ml, respectively. Cell dry weights of late exponential phase bacterial cultures, expressed as cell dry mass, were determined gravimetrically.
Basic genetic techniques and enzymes
Total genomic DNA from Lb. sakei Lb706, L. lactis MG1363, E. faecium P13, and E. hirae DCH5 was isolated using the Wizard® DNA Purification Kit (Promega, Madison, WI, USA). Plasmid DNA isolation was carried out using the QIAprep Spin Miniprep Kit (QIAGEN, Hilden, Germany), as suggested by the manufacturer, but cells were suspended with lysozyme (40 mg ml−1) and mutanolysin (500 U ml−1) and incubated at 37 °C for 10 min to weaken the cell wall before following the kit instructions. DNA restriction enzymes were supplied by New England Biolabs. Ligations were performed with the T4 DNA ligase (Roche Molecular Biochemicals, Mannheim, Germany). E. coli JM109 and K. lactis GG799 competent cells were transformed as described by the supplier (Invitrogen and New England Biolabs, respectively). Competent P. pastoris X-33 cells were obtained as recommended by the supplier (Invitrogen) and transformed by electroporation with a Gene Pulser™ and Pulse Controller apparatus (Bio-Rad Laboratories, Hercules, CA, USA), as previously described [29].
PCR amplification and nucleotide sequencing
Oligonucleotide primers were purchased from Sigma-Genosys Ltd. (Cambridge, UK). Conditions for PCR amplifications were performed as previously described [8]. The PCR-generated fragments were purified by a NucleoSpin® Extract II Kit (Macherey-Nagel GmbH & Co. KG, Düren, Germany) for cloning and nucleotide sequencing. Nucleotide sequencing of purified PCR products was done using the ABI PRISM® BigDye™ Terminator cycle sequence reaction kit and the automatic DNA sequencer ABI PRISM, model 377 (Applied Biosystems, Foster City, CA, USA), at the Unidad de Genómica (Facultad de Ciencias Biológicas, Universidad Complutense de Madrid, Spain).
Recombinant plasmids derived from pNZ8048
The primers and inserts used for the construction of the recombinant plasmids are listed in Table 2. Plasmid derivatives were constructed as follows: the primer pairs USPBS-F/USPSKA-R, EPBS-F/EPSA-R, and HNZ-F/SAPSH-R were used for PCR amplification from total genomic DNA of L. lactis MG1363, E. faecium P13, and E. hirae DCH5 of the NU, NP, and NH fragments encoding the SP usp45 , SP entP , and SP hirJM79 , respectively, with a tail complementary to the N-terminal sequence of SakA. Primers USPBS-F/USEA-R were used for PCR amplification from total genomic DNA of L. lactis MG1363 of a 137-bp BspHI fragment (NUA) containing the SP usp45 and the N-terminal nucleotide sequence ACCACTCATAGTGGTAAATAT::sapA mature (entA/sapA), encoding the first 7 amino acids (TTHSGKY) from EntA fused to mature sapA. Primers USPBS-F/USEP-R were used for PCR amplification from L. lactis MG1363 of a 117-bp BspHI fragment (NUP) containing the SP usp45 and the N-terminal nucleotide sequence of the GCTACG::sapA mature (entP/sapA) encoding the first 2 amino acids (AT) from EntP fused to mature sapA. Primers SKA-F/SKAJEI-R were used for PCR amplification from total genomic DNA of Lb. sakei Lb706 of a 428-bp HindIII fragment (A) containing mature sapA and saiA. Primers SEA-F/SKAJEI-R were used for PCR amplification from Lb. sakei Lb706 of a 440-bp HindIII fragment (B) containing the entA/sapA chimera and saiA. Primers SEP-F/SKAJEI-R were used for PCR amplification from Lb. sakei Lb706 of a 431-bp HindIII fragment (C) containing the entP/sapA chimera and saiA. Mixtures of fragments NU and A, of fragments NP and A, and of fragments NH and A were used as templates to amplify the PCR products NUS, NPS, and NHS encoding mature sapA fused to the SP usp45 , SP entP , and SP hirJM79 , respectively, and saiA. Similarly, mixtures of fragments NUA and B, and NUP and C were used as templates to amplify the 532-bp BspHI-HindIII fragment NUAS, and the 523-pb BspHI-HindIII fragment NUPS, encoding the EntA/SakA and the EntP/SakA chimeras, respectively, fused to the SP usp45 , and SakI. Fragments NUS, NPS, NHS, NUAS, and NUPS were digested with the corresponding restriction enzymes and inserted into pNZ8048, with transcription of the cloned bacteriocins under control of the inducible P nisA promoter, which was also digested with NcoI/HindIII. The ligation mixtures were used to transform L. lactis NZ9000 competent cells. The resulting plasmid derivatives pNUS, pNPS, pNHS, pNUAS, and pNUPS, respectively, were confirmed by bacteriocinogenicity tests, PCR, and sequencing of the inserts.
Recombinant plasmids derived from pMG36c
To construct recombinant plasmids derived from pMG36c the purified pNUS, pNUAS, and pNUPS plasmids were used as templates by using the primer pair USP-F/SKAJEI-R for generation of SacI-HindIII 538-bp (GUS), 550-bp (GUAS), and 541-bp (GUPS) fragments, respectively, containing the P 32 promoter and ribosome binding site (RBS) of pMG36c fused in frame with the genes of interest. The purified inserts were digested with the corresponding restriction enzymes and inserted into plasmid pMG36c digested with the same enzymes. The ligation mixtures were used to transform L. lactis NZ9000 and Lb. sakei Lb706 competent cells. The plasmid derivatives pGUS, pGUAS, and pGUPS were checked by bacteriogenicity tests, PCR, and sequencing of the inserts.
Antimicrobial activity of the recombinant bacterial strains
The antimicrobial activity of individual LAB colonies was examined by the stab-on-agar test (SOAT), as previously described [14]. Recombinant cultures of L. lactis NZ9000 were induced for production of SakA or the EntA/SakA and EntP/SakA chimeras when they reached an optical density at 600 nm (OD600) of 0.5, using as inducer crude nisin A (NisA) from the supernatant of L. lactis BB24 (NisA producer), at a final concentration of 10 ng ml−1. Cell-free culture supernatants from all transformants were obtained by centrifugation of cultures at 12,000g at 4 °C for 10 min, adjusted to pH 6.2 with 1 M NaOH, filtered through 0.2-μm-pore-size filters (Whatman Int. Ltd., Maidstone, UK), and stored at −20 °C until use. The antimicrobial activity of the supernatants was examined by a microtiter plate assay (MPA) as described (Gutiérrez et al. [30]), using E. faecium T136 and Pediococcus damnosus CECT4797 as indicator microorganisms. In the MPA, growth inhibition of the sensitive culture was measured spectrophotometrically at 620 nm with a microtitre Labsystems iEMS plate reader (Labsystems, Helsinki, Finland). One bacteriocin unit (BU) was defined as the reciprocal of the highest dilution of the bacteriocin causing 50 % growth inhibition (50 % of the turbidity of the control culture without bacteriocin). The antimicrobial activity of most recombinant LAB was also tested against Listeria spp. obtained from the CECT (Colección Española de Cultivos Tipo, Valencia, Spain), using the MPA.
Cloning of the sakacin A structural gene (sapA) and its EntP/SakA (GCTACG::sapA) chimera in P. pastoris X-33 and K. lactis GG799, and determination of their antimicrobial activity
The primers and inserts used for the construction of the recombinant plasmids are listed in Table 2. Derivatives of plasmids pPICZαA and pKLAC2 were constructed as follows: primers JJSA-F/JJSA-R and JJPSA-F/JJSA-R were used for PCR amplification from total genomic DNA of Lb. sakei Lb706 DNA of the 158-bp and 161-bp XhoI–NotI fragments KR-SA and KR-PSA, respectively, carrying the α-factor Kex2 signal-protease cleavage site without the Glu-Ala spacer fused to mature SakA and to the EntP/SakA chimera. The KR-SA and KR-PSA fragments were digested with the restriction enzymes mentioned above and the resulting 138-bp and 141-bp fragments were ligated into the pPICZαA and pKLAC2 vectors at appropriate sites to generate plasmids pPSA and pKSA, and pPPSA and pKPSA, respectively. Competent E. coli JM109 cells were transformed with these plasmids and the resulting transformants were confirmed by DNA sequencing. Subsequently, SacI-linearized pPSA and pPPSA and SacII-linearized pKSA and pKPSA were used to transform P. pastoris X-33 and K. lactis GG799, respectively. The P. pastoris X-33SA and P. pastoris X-33PSA transformants were selected from YPD agar supplemented with Zeocin (100 and 1,000 μg ml−1) and sorbitol (1 M). The K. lactis GG799SA and K. lactis GG799PSA transformants were selected on YCB (New England Biolabs) agar supplemented with Tris–HCl Buffer (30 mM) and acetamide (5 mM). Plates were incubated at 30 °C for 3–5 days. The presence of the integrated pPSA, pKSA, pPPSA, and pKPSA linearized plasmids in the genome of the transformed yeasts was confirmed by bacteriocinogenicity tests and by PCR and sequencing of the inserts.
The antimicrobial activity of individual P. pastoris X33SA, P. pastoris X33PSA, K. lactis GG799SA, and K. lactis GG799PSA transformants was screened by a streak-on-agar test (STOAT). Briefly, the P. pastoris X33SA and P. pastoris X33PSA transformants were streaked onto BMMY buffered methanol complex medium [1 % yeast extract, 2 % peptone, 100 mM potassium phosphate (pH 6), 1.34 % yeast nitrogen base (YNB) without amino acids, 4 × 10−5 % biotin, and 0.5 % methanol] agar and grown at 30 °C to induce production of the bacteriocins. During the incubation period, methanol was added daily to the plates at 0.5 % (v/v) final concentration to maintain the induction. The K. lactis GG799SA and K. lactis GG799PSA transformants were streaked onto YPGal (1 % yeast extract, 2 % peptone and 2 % galactose) agar. After incubation of the plates at 30 °C for 48 h, 40 ml of MRS soft agar containing about 1 × 105 cfu ml−1 of the indicator microorganism P. damnosus CECT4797 was added, and the plates were further incubated at 30 °C overnight for development of inhibition zones.
To determine the growth of the recombinant yeasts and the antimicrobial activity of their supernatants, the P. pastoris X33SA and P. pastoris X33PSA bacteriocin producers were grown in the buffered glycerol complex medium BMGY [1 % yeast extract, 2 % peptone, 100 mM potassium phosphate (pH 6), 1.34 % YNB without amino acids, 4 × 10−5 % biotin, and 1 % glycerol] at 30 °C until an OD600 of approximately 2–6 was reached. Cells were then harvested by centrifugation (5,000×g at 4 °C for 10 min), and resuspended to an OD600 of 1 in BMMY medium. Similarly, the K. lactis GG799SA and K. lactis GG799PSA cultures were grown in the YPD medium at 30 °C until an OD600 of approximately 2–6 was reached. Cells were then harvested by centrifugation (5,000×g at 4 °C for 10 min), washed with YPGal, and resuspended to an OD600 of 1 in fresh YPGal medium. The P. pastoris and K. lactis cultures were incubated at 30 °C for 36 h with shaking (250 rpm). During growth, samples were collected periodically for determination of their OD600, bacteriocin production, and the antimicrobial activity of their supernatants by the MPA.
Production of specific anti-SakA polyclonal antibodies and ELISA
The peptide fragment SAJJ (NH2-CSGWASGLAGM-COOH), deduced from the C-terminal amino acid sequence of SakA, was selected as the antigen for the generation of antibodies of predetermined specificity against SakA. The synthetic peptide SAJJ was synthesized by Invitrogen Ltd. (Paisley, Scotland, UK), with a peptide purity of greater than 95 %. The peptide SAJJ was conjugated to the keyhole limpet hemocyanin (KLH) carrier protein as a SAJJ-KLH conjugate, 1:2 (w/w), using the components of the inject maleimide-activated mariculture KLH kit (Perbio Science, Rockford, IL, USA) for use as the immunogen. Rabbits (New Zealand White Females) were immunized with SAJJ-KLH, as described previously [9]. Serum was obtained from blood samples incubated overnight at 4 °C, centrifuged at 1,000×g at room temperature for 15 min, and stored at −20 °C until use. The enzyme-linked immunosorbent assay (ELISA) procedures for determination of antiserum specificity and sensitivity were performed as described previously [9]. A noncompetitive indirect ELISA (NCI-ELISA) was developed to detect and quantify SakA, EntP/SakA, and EntA/SakA in the supernatants of the recombinant bacterial and yeast producer cells. Briefly, wells of flat-bottomed polystyrene microtitre plates (Maxisorp, Nunc, Roskilde, Denmark) were coated overnight (4 °C) with supernatants from Lb. sakei Lb706 or the recombinant LAB and yeast hosts. After addition of the anti-JJSA-KLH antibodies and the goat anti-rabbit immunoglobulin G peroxidase conjugate (Cappel Laboratories, West Chester, PA, USA), bound peroxidase was determined with ABTS (2,2′-azino-bis[3-ethylbenzthiazoline-6-sulfonic acid]) (Sigma) as the substrate by measuring the absorbance of the wells at 405 nm with a Labsystems iEMS reader (Labsystems) with a built-in software package for data analysis.
Purification of SakA, EntP/SakA, and EntA/SakA, mass spectrometry analysis, and N-terminal amino acid sequencing
SakA, EntP/SakA, and EntA/SakA were purified from Lb. sakei Lb706 and L. lactis NZ9000 (pNUS), L. lactis NZ9000 (pNUPS), and L. lactis NZ9000 (pNUAS), respectively. SakA and EntP/SakA were also purified from K. lactis GG799SA and P. pastoris X-33PSA and K. lactis GG799PSA, respectively, as previously described [8, 45]. Briefly, supernatants from early stationary phase cultures of recombinant LAB strains and 0.5 l of the recombinant yeasts were precipitated with ammonium sulfate, desalted by gel filtration, and subjected to cationic-exchange and hydrophobic-interaction chromatography, followed by reversed-phase chromatography in a fast-protein liquid chromatography system (RP-FPLC) (GE Healthcare, Barcelona, Spain). Purified fractions were subjected to matrix-assisted laser desorption/ionization time-of-flight (MALDI–TOF) mass spectrometry, as previously described [30]. For N-terminal sequencing, the purified bacteriocins were subjected to automatic Edman degradation and sequence on polyvinylidene difluoride membranes (PVDF) in a Procise 494 HT Sequencing System (Applied Biosystems Inc., Foster City, CA, USA), according to the manufacturer’s standard methods, at the Centro de Investigaciones Biológicas (CIB, Madrid, Spain).
Results
Heterologous production and functional expression of SakA and two SakA-derived chimeras, EntP/SakA and EntA/SakA, by different LAB strains
The mature bacteriocin SakA and two SakA-derived chimeras, EntP/SakA and EntA/SakA, fused to different signal peptides (SP usp45 , SP entP , and SP hirJM79 ) and cloned into the protein expression vectors pNZ8048, under control of the inducible P nisA promoter, and into pMG36c, carrying the P 32 constitutive promoter, have been evaluated for their production and functional expression by L. lactis subsp. cremoris NZ9000 and Lb. sakei Lb706. Cloning of SP usp45 , SP entP , and SP hirJM79 fused to mature sapA (SakA) and saiA (SakI) into pNZ8048 afforded the NisA-induced derived vectors pNUS, pNPS, and pNHS, respectively, whereas cloning of SP usp45 fused to mature sapA (SakA) and saiA (SakI) into pMG36c afforded the constitutive pGUS. Similarly, cloning of the SP usp45 fused to entP/sapA (EntP/SakA) and saiA (SakI) and fused to entA/sapA (EntA/SakA) and saiA (SakI) into pNZ8048 and pMG36c afforded the NisA-induced derived vectors pNUPS and pNUAS, and the constitutive pGUPS and PGUAS, respectively. Transformation of competent L. lactis subsp. cremoris NZ9000 and Lb. sakei Lb706 with the pNZ8048- and pMG36c-derived vectors yielded recombinant strains which were further checked for bacteriocin production.
The production and functional expression of the SakA, EntP/SakA, and EntA/SakA in the supernatants of the recombinant LAB strains were quantified by specific anti-SakA antibodies in an NCI-ELISA and by a microtitre plate assay (MPA), respectively. None of the control strains transformed with the empty vectors, except the natural Lb. sakei Lb706 SakA producer, showed production of SakA. The production of SakA by L. lactis NZ9000 transformed with the pNZ8048 derivatives was 1.6-fold higher for L. lactis NZ9000 (pNUS) and identical for L. lactis NZ9000 (pNHS), as compared to production of SakA by Lb. sakei Lb706, whereas it was non-detectable for L. lactis NZ9000 (pNPS). The production of EntP/SakA by L. lactis NZ9000 (pNUPS) and of EntA/SakA by L. lactis NZ9000 (pNUAS) was 1.5- and 1.3-fold higher, respectively, than that of SakA by Lb. sakei Lb706 (Table 3). The production of SakA and its chimeras by L. lactis NZ9000, transformed with the pMG36c-derivatives, was from 0.9-fold lower to 2.3-fold higher than production of SakA by Lb. sakei Lb706 (Table 3). Finally, the production of SakA, EntP/SakA, and EntA/SakA by Lb. sakei Lb706 transformed with the pMG36c-derivatives pGUS, pGUPS, and pGUAS, respectively, was from 1.1- to 2.2-fold higher than production of SakA by Lb. sakei Lb706 (Table 3).
The evaluation of the antimicrobial activity of the recombinant LAB revealed that most of the L. lactis NZ9000 transformed hosts showed a much lower (from no activity to 0.14-fold lower) antimicrobial activity than that of Lb. sakei Lb706, except for L. lactis NZ9000 (pNUPS), with a 1.9- and a 6.4-fold higher antimicrobial activity, depending on the indicator strain, than that of Lb. sakei Lb706. On the other hand, supernatants of Lb. sakei Lb706 (pGUS) showed a 2.3- and a 2.9-fold higher antimicrobial activity, whereas those of Lb. sakei Lb706 (pGUPS) and Lb. sakei Lb706 (pGUAS) showed a lower antimicrobial activity (0.36- to 0.62-fold lower) than Lb. sakei Lb706. It should be noted that, although most of the recombinant L. lactis NZ9000 and Lb. sakei Lb706 hosts produced a higher extracellular amount of SakA, EntP/SakA, and EntA/SakA than Lb. sakei Lb706, only L. lactis NZ9000 (pNUPS) and Lb. sakei Lb706 (pGUS) showed a 1.3- and 4.5-fold, and a 1.0- and 1.3-fold higher, respectively, specific antimicrobial activity than the SakA produced by Lb. sakei Lb706 (Table 3).
Supernatants of the recombinant LAB strains also showed antagonistic activity against Listeria spp. by the MPA (Table 4). The pattern of the antagonistic activity of their supernatants was similar to that observed against the indicator bacteria E. faecium T136 and P. damnosus CECT4797 (Table 3). Supernatants of L. lactis NZ9000 (pNUS) showed a 0.04- to 0.63-fold lower antagonistic activity, those of L. lactis NZ9000 (pNUPS) a 1.0- to 5.4-fold higher antagonistic activity, and supernatants of L. lactis NZ9000 (pNUAS) from no activity to a 0.25-fold lower antagonistic activity, as compared to that of Lb. sakei Lb706 against the evaluated Listeria strains. A low antilisterial activity was also observed for L. lactis NZ9000 transformed with plasmids pGUS, pGUPS, and pGUAS (from no activity to a 1.0-fold antimicrobial activity as compared to that of Lb. sakei Lb706). However, Lb. sakei Lb706 (pGUS) showed a 1.3- to 5.0-fold higher antilisterial activity than Lb. sakei Lb706, whereas Lb. sakei Lb706 (pGUPS) and Lb. sakei Lb706 (pGUAS) showed a lower antilisterial activity than Lb. sakei Lb706 against Listeria spp. (Table 4).
Purification of SakA, EntP/SakA, and EntA/SakA, mass spectrometry analysis, and N-terminal amino acid sequencing
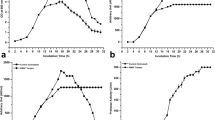
The SakA produced by Lb. sakei Lb706 and L. lactis (pNUS), the EntP/SakA produced by L. lactis (pNUPS), and the EntA/SakA produced by L. lactis (pNUAS) were purified to homogeneity following a previously used chromatographic procedure (results not shown). MALDI–TOF MS analysis of the purified SakA from Lb. sakei Lb706 showed a major peak of 4,306.9 Da close to its predicted molecular mass of 4,308.8 Da (Fig. 1a), whereas the molecular mass of the SakA produced by L. lactis (pNUS) showed a much lower molecular mass of 3,805.3 Da (Fig. 1b). Similarly, the molecular mass of the EntP/SakA produced by L. lactis (pNUPS) was 4,410.5 Da (Fig. 1c), whereas that of the EntA/SakA produced by L. lactis (pNUAS) showed a molecular mass of 3,805.8 Da (Fig. 1d). Given that the obtained molecular masses of SakA and EntA/SakA produced by L. lactis NZ9000 are almost identical to each other but lower than their predicted theoretical molecular masses of 4,308.8 and 4,769.3 Da, respectively, it could be that both occur via abnormal processing. Accordingly, determination of the N-terminal amino acid sequence of both bacteriocins by Edman degradation demonstrated that they started with the amino acid sequence NGVY-, meaning that both have lost their first five (ARSYG-) and nine (TTHSGKYYG-) N-terminal amino acids, respectively, during processing and secretion out of the L. lactis NZ9000 producer hosts.
Mass spectrometry analysis of purified sakacin A from Lb. sakei Lb706 (a), purified sakacin A from L. lactis NZ9000 (pNUS) (b), purified enterocin P/sakacin A from L. lactis NZ9000 (pNUPS) (c), and purified enterocin A/sakacin A from L. lactis NZ9000 (pNUAS) (d). Numbers indicate the molecular mass in daltons of most of the observed peptide fragments
Heterologous production and functional expression of SakA and EntP/SakA by P. pastoris and K. lactis
Cloning of the PCR fragments containing the α-factor Kex2 signal cleavage fused to mature sapA (SakA) or to mature entP/sapA (EntP/SakA) into the expression vectors pPICZαA and pKLAC2 afforded the P. pastoris X-33 and K. lactis GG799 derivatives, respectively, from which to determine the production and functional expression of SakA and EntP/SakA. Since the P. pastoris X-33SA, producer of SakA, and P. pastoris X-33PSA, producer of EntP/SakA, did not show a defined direct antimicrobial activity when streaked in selective agar plates with the SakA-sensitive indicator P. damnosus CECT4797, they were selected on the basis of their high Zeocin resistance (1,000 μg ml−1). However, K. lactis GG799SA, producer of SakA, and K. lactis GG799PSA, producer of EntP/SakA, showed a low but defined direct antimicrobial activity against P. damnosus CECT4797. Colonies of P. pastoris X-33 and K. lactis GG799, transformed with the linearized control plasmids, were used as bacteriocin-negative controls to discard the possibility that the antimicrobial activity exerted by the recombinant hosts was due to metabolites other than bacteriocins.
The heterologous production and functional expression of the SakA and EntP/SakA produced by the recombinant yeasts were determined by an NCI-ELISA and an MPA, respectively. The growth and production of SakA by P. pastoris X-33SA were lower than those of EntP/SakA by P. pastoris X-33PSA. The largest production of SakA and EntP/SakA by the recombinant P. pastoris producers was 1.8- and 24.2-fold higher, respectively, than production of SakA by Lb. sakei Lb706. However, neither the SakA nor the EntP/SakA produced by the P. pastoris producers showed a measurable antimicrobial activity (Table 5). On the other hand, the growth of K. lactis GG799SA and K. lactis GG799PSA was similar, whereas the production of SakA was 11.4-fold higher and that of the EntP/SakA 55.2-fold higher than production of SakA by Lb. sakei Lb706. However, the antimicrobial activity and the specific antimicrobial activity of either the SakA or the EntP/SakA produced by the K. lactis derivatives were much lower, ranging from 0.01- to 0.06-fold lower for both, than those of the SakA produced by Lb. sakei Lb706 (Table 5).
Purification and mass spectrometry analysis of the SakA and EntP/SakA produced by P. pastoris and K. lactis
Although not a measurable or a very low antimicrobial activity was observed in the supernatants of the recombinant yeasts, their supernatants were subjected to protein purification following the procedure used for the recombinant LAB producers. Interestingly, a high antimicrobial activity was observed after the gel filtration, cationic exchange, hydrophobic interaction, and reverse-phase chromatography steps of all evaluated supernatants, except in those from P. pastoris X-33SA (results not shown). The theoretical molecular mass of SakA is 4,308.8 Da and that of the EntP/SakA is 4,410.0 Da. However, MALDI–TOF MS analysis of the purified EntP/SakA produced by P. pastoris X-33PSA showed a minor peptide fragment of 4,439.9 Da and major fragments of higher molecular mass (5.4–5.8 kDa). Similarly, the purified SakA from K. lactis GG799SA showed a fragment of 4,351.6 Da as well as fragments of higher molecular mass (>4.4 kDa), whereas the EntP/SakA produced by K. lactis G799PSA also showed a minor peak of 4,285.3 Da and numerous fragments of higher molecular mass (5.5–6.1 kDa) (Fig. 2).
Discussion
If the use of bacteriocins as natural antimicrobial agents in food, veterinary, and medical applications is ever to meet the high expectations of the research community, a high level production of active bacteriocins in homologous and heterologous microbial hosts is essential. Production of bacteriocins by LAB is essentially based on the expression of native biosynthetic genes, by exchanging or replacing leader peptides and/or dedicated processing and secretion systems (ABC transporters) or by fusion of mature bacteriocins to signal peptides that act as secretion signals [8, 9, 30]. Accordingly, optimization of bacteriocin gene expression and protein production would help the development of LAB as microbial cell factories for production and delivery of bacteriocins of biotechnological interest.
For the heterologous production of SakA and two SakA-derived chimeras, EntP/SakA and EntA/SakA, by L. lactis subsp. cremoris NZ9000 and Lb. sakei Lb706, the mature SakA and its chimeras were fused to different SPs. The SP usp45 , SP entP , and SP hirJM79 encode the 27 amino acid SP of the major extracellular Usp45 protein from L. lactis MG1363 [48], the 27 amino acid SP of the bacteriocin EntP [14], and the 30 amino acid SP of the bacteriocin hiracin JM79 [44], respectively. SakA holds the N-terminal ARS amino acid sequence before the YGNGVxC consensus sequence of the class IIa bacteriocins. However, since positively charged amino acids immediately following the SP cleavage site may interfere with the secretion machinery or passage of the protein through the bacterial membrane [37], the EntP/SakA chimera was constructed by displacement of the arginine (R) residue from position +2 to position +3 of the mature bacteriocin, thus mimicking the N-terminal amino acid sequence of EntP (ATR), a bacteriocin homologous to SakA [14] which is known to be overproduced and adequately processed by recombinant L. lactis and P. pastoris [8, 29]. The EntA/SakA chimera was also constructed to hold the TTHSGKY amino acid sequence before the YGNGVxC consensus sequence of the class IIa bacteriocins, thus mimicking the N-terminal amino acid sequence of EntA [2], which is also overproduced and adequately processed by recombinant L. lactis [9] and the yeasts P. pastoris, K. lactis, Hansenula polymorpha, and Arxula adeninivorans [10].
In this work, the development of specific anti-SakA antibodies and an NCI-ELISA has permitted the detection and quantification of SakA and its derived chimeras in the supernatants of the recombinant LAB strains, as well as the determination of their specific antimicrobial activity. Apparently, the production and antimicrobial activity of the SakA, EntP/SakA, and EntA/SakA produced by the recombinant LAB strains depend on the SP, the expression vector, and the host strain (Table 3). The production of SakA and its derivatives by the recombinant LAB strains may also rely on the expression of the SakI immunity (saiA) gene. It has been demonstrated that bacteriocin producers are protected from their own bacteriocins by the concomitant expression of a cognate immunity protein, that the expression of the immunity protein may increase bacteriocin production, and that the immunity proteins form a strong complex with the receptor proteins, thereby preventing producer cells from being killed [8, 35]. The production and secretion of SakA by the recombinant L. lactis NZ9000 derivatives was observed when sapA was fused to SP usp45 and SP hirJM79 . However, the SP entP was unable to drive the production of SakA by L. lactis NZ9000. This was an unexpected result because the SP entP fused to mature pediocin PA-1 (PedA-1) [38] and EntA [9, 39] permitted the production and secretion of both bacteriocins by different recombinant L. lactis hosts. In this context, secretion levels in Gram-positive bacteria may become not only affected by variations in the SPs but also by differences in the N-terminal part of the mature peptide or protein that may become only evident after fusion with some SP. It is possible that the N-terminal amino acid sequence of mature SakA may affect its secretion because positive charges at the N-terminus may affect export of the peptide by the Sec-dependent pathway [37]. Mature SakA may also remain N-terminally associated to the cell membrane via a Sec-type signal peptide that is not cleaved off during secretion [7]. It has been also hypothesized that variations in secretion capacities can be governed by post-transcriptional factors such as secondary structure of mRNA, codon usage, and translation efficiency [22]. The molecular folding of SakA inside L. lactis may also maintain the prepeptide in a secretion-incompetent conformation [40]. In any case, protein secretion is a preferred means of protein expression in the development of LAB as cell factories for production of biologically active bacteriocins, and SP usp45 fused to mature sapA and its chimeras adequately drive their production and secretion by the recombinant L. lactis and Lb. sakei hosts (Table 3).
The role of the expression vectors in terms of production and antimicrobial activity of the target bacteriocins seems to depend on the host strain, genes of interest, promoter, and vector copy number (Table 3). Plasmid pNZ8048 contains the high copy number heterogramic replicon of the lactococcal plasmid pSH71 with a unique NcoI cleavage site, downstream of the nisA ribosomal binding sequence (RBS), used for translational fusions inducible by NisA [19, 36]. The expression vector pMG36c contains the low copy replication origin of the lactococcal plasmid pWV01 and the strong P 32 promoter to drive the constitutive transcription of inserted genes into the multicloning site (MCS) of pUC18 [49]. In this work, production of SakA and its chimeras by recombinant L. lactis and Lb. sakei hosts is not closely associated with the protein expression vector used, and this observation is in contrast to results in which production of bacteriocins is higher by pNZ8048-derived LAB transformants than by the pMG36c-derived ones [8, 9, 30, 38, 39, 45]. For optimization of protein expression, inducible systems are often considered superior to constitutive expression systems, because the former enable achievement of sufficient biomass prior to initiation of target protein expression and consequent metabolic burden of the cell. However, other factors such as mRNA stability and secondary structure may steer protein production from the recombinant L. lactis and Lb. sakei hosts [22].
Of interest is also the observation that supernatants of most recombinant L. lactis NZ9000, transformed with the pNZ8048-derived expression vectors, show a lower antimicrobial activity than that expected from the production of SakA, EntP/SakA, and EntA/SakA, except for L. lactis NZ9000 (pNUPS), with an antimicrobial activity in the range of that deduced from its production of EntP/SakA. On the other hand, all recombinant L. lactis NZ9000 hosts transformed with pMG36c-derived vectors show non-measurable or very poor antimicrobial activity (Table 3). It may occur that the short induction time for bacteriocin production from nisin-inducible systems most probably prevents bacteriocins from attaching to cell walls, forming aggregates, and/or undergoing protease degradation [8, 30]. A high bacteriocin production does not always correspond to a high antimicrobial activity. The low antimicrobial activity of the SakA and its chimeras produced by the recombinant LAB hosts may depend on many factors which are difficult to determine. It is possible that regulatory responses to secretion stress activate quality control networks involving folding factors and housekeeping proteases [18]. Differences in the Sec-dependent translocation and Sec machinery in the different LAB strains, differences in protein folding, and conformational modifications of the bacteriocin to a less extracellular active form may also account for a low antagonistic activity of the secreted bacteriocins [46]. In this respect, the four cysteine residues present in SakA and presumably involved in the formation of two disulfide bonds (DSB) may also play a role in the folding, structural integrity, and antimicrobial activity of the produced bacteriocins [25]. Bacteriocin self-aggregation may also decrease the antagonistic activity of bacteriocins [9].
MALDI–TOF MS analysis of the bacteriocins purified from recombinant L. lactis NZ9000 hosts revealed that SakA purified from L. lactis (pNUS) had a lower molecular mass than SakA purified from Lb. sakei Lb706. The purified EntP/SakA showed a major fragment of a molecular mass of 4,410.5 Da, corresponding to its expected size, whereas the EntA/SakA showed a major fragment of a molecular mass of 3,805.8 Da (Fig. 1). Accordingly, since the theoretical molecular mass of SakA is 4,308.8 Da, that of EntP/SakA is 4,410.0 Da, and that of EntA/SakA is 4,769.3 Da, it seems that the SakA and the EntA/SakA produced by recombinant L. lactis NZ9000 are truncated bacteriocin fragments. Furthermore, all the analyzed bacteriocins manifest the presence of major bacteriocin fragments with presumably none, one (+16 Da), and two (+32 Da) methionine residues (Met30, Met41) oxidized to MetSO (Fig. 1). The oxidation of methionine residues during bacteriocin purification and recombinant production by LAB is common [9, 39]. Determination of the N-terminal amino acid sequence of SakA and EntA/SakA by Edman degradation revealed that both bacteriocins started with the amino acid sequence NGVY-, demonstrating that both are truncated forms of the bacteriocins cloned in L. lactis NZ9000. As far as we know, this is the first report of N-terminal truncated bacteriocins produced by recombinant LAB. However, the truncated SakA and EntA/SakA produced by L. lactis NZ9000 still maintain a lower, but measurable antimicrobial activity, perhaps owing to the nonspecific binding of the pediocin-like N-terminal sequences or their truncated forms to target cells through electrostatic interactions [12].
Transformation of Lb. sakei Lb706, natural producer of SakA, with pMG36c-derived vectors permitted a slightly higher production of SakA, EntP/SakA, and EntA/SakA by all transformants. Furthermore, Lb. sakei Lb706 (pGUS), producer of SakA, showed higher antimicrobial activity than supernatants of Lb. sakei Lb706 (Table 3), confirming the higher production and antimicrobial activity of bacteriocins produced by homologous LAB hosts [9]. In Lb. sakei Lb706 modification of the N-terminal sequence of SakA resulted in lower antimicrobial activity in the supernatants of Lb. sakei Lb706 (pGUPS) and Lb. sakei Lb706 (pGUAS), further supporting that different amino acid sequences for the signal and mature peptides may be required for optimal production and secretion depending on the bacterial host.
Supernatants of L. lactis NZ9000 (pNUPS), producers of EntP/SakA, and those of Lb. sakei Lb706 (pGUS), producers of SakA, showed up to a 5.4-fold higher antimicrobial activity against several Listeria spp. than any other recombinant LAB host (Table 4). These recombinant LAB strains as overproducers of SakA and EntP/SakA, with higher antimicrobial activity than the supernatants of Lb. sakei Lb706, may be considered as appropriate cellular factories and an alternative to Lb. sakei Lb706 for production and recovery of the SakA and EntP/SakA antilisterial bacteriocins. Moreover, the use as bacteriocin producers of Lactococcus spp. and Lactobacillus spp. strains, generally recognized as safe (GRAS) and with a qualified presumption of safety (QPS), may also provide means by which the potential benefits of antimicrobial compounds can be exploited in food.
Although heterologous expression systems for production of bacteriocins are being developed in bacteria, yeasts have not been yet fully exploited as alternative hosts for their production [10]. However, a number of yeast platforms have been developed for the successful heterologous production of peptides and proteins [6, 28]. In this work, linearized pPICZαA and pKLAC2 protein expression vectors [15, 17] containing SakA and SakA-derived chimeras have permitted the production of SakA and EntP/SakA by P. pastoris X-33SA and P. pastoris X-33PSA and the production and expression of these peptides by K. lactis GG799SA and K. lactis GG799PSA, although both expressed peptides are significantly less active than the SakA and EntP/SakA produced by Lb. sakei Lb706. All recombinant yeasts secreted SakA and EntP/SakA, although production of SakA was lower than that of EntP/SakA. Whereas positively charged amino acids immediately following the SP cleavage site do not seem to interfere with the yeasts secretion machinery, displacement of the arginine (R) residue from position +2 to position +3 of the bacteriocin improves EntP/SakA secretion in P. pastoris X-33PSA and K. lactis GG799PSA. It should be noted that growth of P. pastoris X-33SA was severely impaired as compared to that of P. pastoris X-33PSA (Table 5). Since one of the main bottlenecks in recombinant protein production is the inability of foreign peptides to reach their native conformation in heterologous yeast hosts, it may happen that incorrectly folded SakA is accumulated in the endoplasmic reticulum (ER), activating the unfolded protein response (UPR) and the ER-associated degradation with generation of reactive oxygen species (ROS), leading to persistent ER stress conditions causing apoptosis and yeast death [26]. In this context, recent studies show that synthetic antimicrobial peptides induce the accumulation of ROS and hydroxyl radicals known to be important regulators of apoptosis and cell death in Candida albicans [34].
The SakA and the EntP/SakA produced by the recombinant P. pastoris and K. lactis hosts showed no activity or a much lower antimicrobial activity than that deduced from their production (Table 5). This observation was not unexpected because bacteriocins cloned into S. cerevisiae [3, 47, 50], P. pastoris [4, 5, 10, 29, 45], and K. lactis, H. polymorpha, and A. adeninivorans [10] have been produced with variable success regarding their secretion and functional expression. In this work, MALDI–TOF MS analysis of the purified bacteriocins produced by the recombinant yeasts showed that, besides the presence of tentative oxidized forms of SakA and EntP/SakA, other fragments of high molecular mass were present (Fig. 2). In this respect, these peptides could be associated with unknown biological compounds, as suggested for the inactive pediocin PA-1 (PedPA-1) produced by P. pastoris [5]. However, more likely both bacteriocins may have been subjected to post-translational modifications (PTMs) (Fig. 2). Some common PTMs events in peptides and proteins are phosphorylation, acetylation, methylation, oxidation, formylation, disulfide bond formation, and N-linked and O-linked glycosylation [51].
The presence of cysteine and methionine residues in SakA and EntP/SakA (Table 2) may lead to the formation of correct disulfide bridges but also to oxidation of these residues and the covalent attachment of different compounds to cysteine. Cysteine is susceptible to chemical modifications such as glutathionylation and cysteinylation. Similarly, the oxidation of methionine residues to MetSO is common during production of bacteriocins by yeasts [4, 10]. Glycosylation is also a common PTM in eukaryotes involving linkage via the Asn-X-Ser/Thr sequence (N-glycosylation) or the side chain of serine and threonine (O-glycosylation). The absence in SakA and EntP/SakA of attachment sites for N-linkages precludes their N-glycosylation, but the presence of three serines and two threonines makes them suitable for O-glycosylation. However, the correct processing, secretion, and functional expression of the bacteriocins EntP [29], HirJM79 [45], and EntA [10] produced by recombinant yeasts contrast with the low biological activity of the SakA and EntP/SakA produced by the P. pastoris and K. lactis producers. Misfolding of SakA and EntP/SakA and induction of the yeasts’ UPR may be responsible for apoptosis in P. pastoris X-33SA and for extensive PTMs in P. pastoris X-33PSA, K. lactis G799SA, and K. lactis G799PSA.
The use of synthetic hybrid bacteriocins and the synthesis of bacteriocins containing modified amino acid sequences by site-directed mutagenesis, error-prone PCR, and gene shuffling techniques have permitted the design of more active bacteriocins [31]. The production of bacteriocins in heterologous LAB and yeasts has also permitted the use of safer microbial hosts, to increase their production, antimicrobial activity, and specific antimicrobial activity as compared to that of the natural producers, to provide antimicrobial capabilities to LAB that may be useful as starters or protective cultures, or to design potential cell factories for production and delivery of bacteriocins of interest in food, medical, veterinary, and animal production applications [10]. However, further efforts should be made to clarify those critical factors involved in the production and functional expression of different bacteriocins or their chimeras by recombinant LAB and yeasts.
References
Axelsson L, Holck A (1995) The genes involved in production of and immunity to sakacin A, a bacteriocin from Lactobacillus sake Lb706. J Bacteriol 177:2125–2137
Aymerich T, Holo H, Håverstein LS, Hugas M, Garriga M, Nes IF (1996) Biochemical and genetic characterization of enterocin A from Enterococcus faecium, a new antilisterial bacteriocin in the pediocin-family of bacteriocins. Appl Environ Microbiol 62:1676–1682
Basanta A, Herranz C, Gutiérrez J, Criado R, Hernández PE, Cintas LM (2009) Development of bacteriocinogenic strains of Saccharomyces cerevisiae heterologously expressing and secreting the leaderless enterocin L50 peptides L50A and L50B from Enterococcus faecium L50. Appl Environ Microbiol 75:2382–2392
Basanta A, Gómez-Sala B, Sánchez J, Dzung BD, Herranz C, Hernández PE, Cintas LM (2010) Use of the yeast Pichia pastoris as an expression host for secretion of enterocin L50, a leaderless two-peptide (L50A and L50B) bacteriocin from Enterococcus faecium L50. Appl Environ Microbiol 76:3314–3324
Beaulieu L, Groleau D, Míguez CB, Jetté JF, Aomari H, Subirade M (2005) Production of pediocin PA-1 in the methylotrophic yeast Pichia pastoris reveals unexpected inhibition of its biological activity due to the presence of collagen-like material. Prot Expr Purif 43:111–125
Böer E, Steinborn G, Kunze G, Gellissen G (2007) Yeast expression platforms. Appl Microbiol Biotechnol 77:513–523
Bøhle LA, Riaz T, Egge-Jacobsen W, Skaugen M, Busk ØL, Eijsink VGH, Mathiesen G (2011) Identification of surface proteins in Enterococcus faecalis V583. BMC Genomics 12:135
Borrero J, Jiménez JJ, Gútiez L, Herranz C, Cintas LM, Hernández PE (2011) Use of the usp45 lactococcal secretion sequence signal sequence to drive the secretion and functional expression of enterococcal bacteriocins in Lactococcus lactis. Appl Microbiol Biotechnol 89:131–143
Borrero J, Jiménez JJ, Gútiez L, Herranz C, Cintas LM, Hernández PE (2011) Protein expression vector and secretion signal peptide optimization to drive the production, secretion, and functional expression of the bacteriocin enterocin A in lactic acid bacteria. J Biotechnol 156:76–86
Borrero J, Kunze G, Jiménez JJ, Böer E, Gútiez L, Herranz C, Cintas LM, Hernández PE (2012) Cloning, production and functional expression of the bacteriocin enterocin A, produced by Enterococcus faecium T136, by the yeasts Pichia pastoris, Kluyveromyces lactis, Hansenula polymorpha and Arxula adeninivorans. Appl Environ Microbiol 78:5956–5961
Casaus P, Nilsen T, Cintas LM, Nes IF, Hernández PE, Holo H (1997) Enterocin B, a new bacteriocin from Enterococcus faecium T136 which can act synergistically with enterocin A. Microbiology 143:2287–2294
Chen Y, Ludescher RD, Montville TJ (1997) Electrostaic interactions but not the YGNGV consensus motif, govern the binding of pediocin PA-1 and its fragments to phospholipid vesicles. Appl Environ Microbiol 63:4770–4777
Cintas LM (1995) PhD thesis. Universidad Autónoma de Madrid, Spain
Cintas LM, Casaus P, Håvarstein LS, Hernández PE, Nes IF (1997) Biochemical and genetic characterization of enterocin P, a novel sec-dependent bacteriocin from Enterococcus faecium P13 with a broad antimicrobial spectrum. Appl Environ Microbiol 63:4321–4330
Colussi PA, Taron CH (2005) Kluyveromyces lactis LAC4 promoter variants that lack function in bacteria but retain full function in K. lactis. Appl Environ Microbiol 71:7092–7098
Cotter PD, Hill C, Ross RP (2005) Bacteriocins: developing innate immunity for food. Nat Rev Microbiol 3:777–788
Cregg JM, Cereghino JL, Shi J, Higins DR (2000) Recombinant protein expression in Pichia pastoris. Mol Biotechnol 16:23–52
Darmon E, Noone D, Masson A, Bron S, Kuipers OP, Devine KM, van Dijl JM (2002) A novel class of heat and secretion stress-responsive genes is controlled by the autoregulated CssRS two-component system of Bacillus subtilis. J Bacteriol 184:5661–5671
De Ruyter PGGA, Kuipers OP, de Vos WM (1996) Controlled gene expression systems for Lactococcus lactis with the food-grade inducer nisin. Appl Environ Microbiol 62:3662–3667
Diep DB, Axelsson L, Grefsli C, Nes IF (2000) The synthesis of the bacteriocin sakacin A is a temperature-sensitive process regulated by a pheromone peptide through a three-component regulatory system. Microbiology 146:2155–2160
Diep DB, Skaugen M, Salehian Z, Holo H, Nes IF (2007) Common mechanisms of target cell recognition and immunity for class II bacteriocins. Proc Natl Acad Sci U S A 104:2384–2389
Diep DB, Mathiesen G, Eijsink VGH, Nes IF (2009) Use of lactobacilli and their pheromone-based regulatory mechanism in gene expression and drug delivery. Curr Pharm Biotechnol 10:62–73
Dobson A, Cotter PD, Ross RP, Hill C (2012) Bacteriocin production: a probiotic trait? Appl Environ Microbiol 78:1–6
Driessen AJM, Nouwen N (2007) Protein translocation across the bacterial cytoplasmic membrane. Annu Rev Biochem 77:643–667
Freitas DA, Leclerc S, Miyoshi A, Oliveira SC, Sommer PSM, Rodrigues L, Correa A, Gautier M, Langella P, Azevedo VA, Le Loir Y (2005) Secretion of Streptomyces tendae antifungal protein 1 by Lactococcus lactis. Braz J Med Biol Res 38:1585–1592
Gasser B, Saloheimo M, Rinas U, Dragosits M, Rodríguez Carmona E, Baumann K, Giuliani M, Parrilli E, Branduardi P, Lang C, Porro D, Ferrer P, Tutino ML, Mattanovich D, Villaverde A (2008) Protein folding and conformational stress in microbial cells producing recombinant proteins: a host comparative overview. Microb Cell Fact 7:11. doi:10.1186/1475-2859-7-11
Gasson MJ (1983) Plasmid complements of Streptococcus lactis NCDO 712 and other lactic streptococci after protoplast-induced curing. J Bacteriol 154:1–9
Gellissen G, Kunze G, Gaillardin C, Cregg JM, Berardi E, Veenhuis M, van der Klei I (2005) New yeast expression platforms based on methylotrophic Hansenula polymorpha and Pichia pastoris and on dimorphic Arxula adeninivorans and Yarrowia lipolytica. A comparison. FEMS Yeast Res 5:1079–1096
Gutiérrez J, Criado R, Martín M, Herranz C, Cintas LM, Hernández PE (2005) Production of enterocin P, an antilisterial pediocin-like bacteriocin from Enterococcus faecium P13, in Pichia pastoris. Antimicrob Agents Chemother 49:3004–3008
Gutiérrez J, Larsen R, Cintas LM, Kok J, Hernández PE (2006) High-level heterologous production and functional expression of the sec-dependent enterocin P from Enterococcus faecium P13 in Lactococcus lactis. Appl Microbiol Biotechnol 72:41–51
Haugen GS, Fimland G, Nissen-Meyer J (2011) Mutational analysis of residues in the helical region of the class IIa bacteriocin pediocin PA-1. Appl Environ Microbiol 77:1966–1972
Håvarstein LS, Diep DB, Nes IF (1995) A family of bacteriocin ABC transporters carry out proteolytic processing of their substrates concomitantly with export. Mol Microbiol 16:229–240
Holck A, Axelsson L, Birkeland SE, Aukrust T, Blom H (1992) Purification and amino acid sequence of sakacin A, a bacteriocin from Lactobacillus sake Lb706. J Gen Microbiol 138:2715–2720
Hwang B, Hwang JS, Lee J, Lee DG (2011) The antimicrobial peptide psacotheasin induces reactive oxygen species and triggers apoptosis in Candida albicans. Biochem Biophys Res Commun 405:267–271
Kjos M, Borrero J, Opsata M, Birri DJ, Holo H, Cintas LM, Snipen L, Hernández PE, Nes IF, Diep DB (2011) Target recognition, resistance, inmunity and genome mining of class II bacteriocins from Gram-positive bacteria. Microbiology 157:3256–3267
Kuipers OP, de Ruyter PGGA, Kleerebezem M, de Vos WM (1998) Quorum sensing-controlled gene expression in lactic acid bacteria. J Biotechnol 64:15–21
Le Loir Y, Nouaille S, Commissaire J, Bretigny L, Gruss A, Langella P (2001) Signal peptide and propeptide optimization for heterologous protein secretion in Lactococcus lactis. Appl Environ Microbiol 67:4119–4127
Martín M, Gutiérrez J, Criado R, Herranz C, Cintas LM, Hernández PE (2007) Chimeras of mature pediocin PA-1 fused to the signal peptide of enterocin P permits the cloning, production, and expression of pediocin PA-1 in Lactococcus lactis. J Food Prot 70:2792–2798
Martín M, Gutiérrez J, Criado R, Herranz C, Cintas LM, Hernández PE (2007) Cloning, production and expression of the bacteriocin enterocin A produced by Enterococcus faecium PLBC21 in Lactococcus lactis. Appl Microbiol Biotechnol 76:667–675
Mathiesen G, Svee A, Brurberg MB, Fredriksen L, Axelsson L, Eijsink VGH (2009) Genome-wide analysis of signal peptide functionality in Lactobacillus plantarum WCFS1. BMC Genomics 10:425. doi:10.1186/1471-2164-10-425
Montalbán-López M, Sánchez-Hidalgo M, Valdivia E, Martínez-Bueno M, Maqueda M (2011) Are bacteriocins underexploited? Novel applications for old antimicrobials. Curr Pharm Biotechnol 12:1205–1220
Natale P, Brüsser T, Driessen AJM (2008) Sec- and Tat-mediated protein secretion across the bacterial cytoplasmic membrane: distinct translocases and mechanisms. Biochim Biophys Acta 1778:1735–1756
Nes IF, Yoon SS, Diep DB (2007) Ribosomally synthesized antimicrobial peptides (bacteriocins) in lactic acid bacteria: a review. Food Sci Biotechnol 16:675–690
Sánchez J, Diep DB, Herranz C, Nes IF, Cintas LM, Hernández PE (2007) Amino acid and nucleotide sequence, adjacent genes, and heterologous expression of hiracin JM79, a Sec-dependent bacteriocin produced by Enterococcus hirae DCH5, isolated from Mallard ducks (Anas platyrhynchos). FEMS Microbiol Lett 270:227–236
Sánchez J, Borrero J, Gómez-Sala B, Basanta A, Herranz C, Cintas LM, Hernández PE (2008) Cloning and heterologous production of hiracin JM79, a sec-dependent bacteriocin produced by Enterococcus hirae, in lactic acid bacteria and Pichia pastoris. Appl Environ Microbiol 74:2471–2479
Sarvas M, Harwood CR, Bron S, van Dijl JM (2004) Post-translocational folding of secretory proteins in Gram-positive bacteria. Biochim Biophys Acta 1694:311–327
Schoeman H, Vivier MA, du Toit M, Dicks LMT, Pretorius IS (1999) The development of bactericidal yeast strains by expressing the Pediococcus acidilactici pediocin gene (pedA) in Saccharomyces cerevisiae. Yeast 15:647–656
van Asseldonk M, Rutten G, Oteman M, Siezen RJ, de Vos WM, Simons G (1990) Cloning of usp45, a gene encoding a secreted protein from Lactococcus lactis subsp. lactis MG1363. Gene 95:155–160
van de Guchte M, van der Vossen JMBM, Kok J, Vemena G (1989) Construction of a lactococcal expression vector: expression of hen egg white lysozyme in Lactococcus lactis subsp. lactis. Appl Environ Microbiol 55:224–228
van Reenen CA, Chikindas ML, van Zyl WH, Dicks LMT (2002) Characterization and heterologous expression of a class IIa bacteriocin, plantaricin 423 from Lactobacillus plantarum 423, in Saccharomyces cerevisiae. Int J Food Microbiol 81:29–40
Zhao Y, Jensen ON (2009) Modification-specific proteomics: strategies for characterization of post-translational modifications using enrichment techniques. Proteomics 9:4362–4641
Acknowledgments
The authors express their gratitude to Prof. J. Kok (Department of Genetics, University of Groningen, the Netherlands) for supplying plasmid pMG36c. This work was partially supported by grants AGL2012-34829 from the Ministerio de Economía y Competitividad (MINECO) and AGL2009-08348 from the Ministerio de Ciencia e Innovación (MICINN), by grant GR35-10A from the BSCH-UCM, and by grant S2009/AGR-1489 from the Comunidad de Madrid (CAM). J.J. Jiménez is recipient of a fellowship (FPI) from the Ministerio de Ciencia e Innovación (MICINN), J. Borrero held a research contract from the CAM, and L. Gútiez holds a fellowship (FPU) from the Ministerio de Educación y Ciencia (MEC), Spain.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jiménez, J.J., Borrero, J., Diep, D.B. et al. Cloning, production, and functional expression of the bacteriocin sakacin A (SakA) and two SakA-derived chimeras in lactic acid bacteria (LAB) and the yeasts Pichia pastoris and Kluyveromyces lactis . J Ind Microbiol Biotechnol 40, 977–993 (2013). https://doi.org/10.1007/s10295-013-1302-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-013-1302-6